Abstract
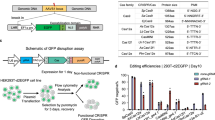
The utility of CRISPR-Cas9 and TALENs for genome editing may be compromised by their off-target activity. We show that integrase-defective lentiviral vectors (IDLVs) can detect such off-target cleavage with a frequency as low as 1%. In the case of Cas9, we find frequent off-target sites with a one-base bulge or up to 13 mismatches between the single guide RNA (sgRNA) and its genomic target, which refines sgRNA design.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Cong, L. et al. Science 339, 819–823 (2013).
Perez-Pinera, P. et al. Nat. Methods 10, 973–976 (2013).
Shalem, O. et al. Science 343, 84–87 (2014).
Mali, P. et al. Nat. Biotechnol. 31, 833–838 (2013).
Wang, T., Wei, J.J., Sabatini, D.M. & Lander, E.S. Science 343, 80–84 (2014).
Xiao, A. et al. Nucleic Acids Res. 41, e141 (2013).
Fu, Y. et al. Nat. Biotechnol. 31, 822–826 (2013).
Hsu, P.D. et al. Nat. Biotechnol. 31, 827–832 (2013).
Cho, S.W. et al. Genome Res. 24, 132–141 (2014).
Pattanayak, V., Ramirez, C.L., Joung, J.K. & Liu, D.R. Nat. Methods 8, 765–770 (2011).
Pattanayak, V. et al. Nat. Biotechnol. 31, 839–843 (2013).
Gabriel, R. et al. Nat. Biotechnol. 29, 816–823 (2011).
Osborn, M.J. et al. Mol. Ther. 21, 1151–1159 (2013).
Ran, F.A. et al. Cell 154, 1380–1389 (2013).
Lin, Y. et al. Nucleic Acids Res. 42, 7473–7485 (2014).
Mali, P. et al. Science 339, 823–826 (2013).
Doyle, E.L. et al. Nucleic Acids Res. 40, W117–W122 (2012).
Wang, X. et al. PLoS ONE 9, e93575 (2014).
Paruzynski, A. et al. Nat. Protoc. 5, 1379–1395 (2010).
Zhang, J., Kobert, K., Flouri, T.X. & Stamatakis, A. Bioinformatics 30, 614–620 (2014).
Acknowledgements
This work is supported by a grant RB3-02161 from California Institute of Regenerative Medicine (J.-K.Y.), grants from Nesvig Foundation and Beckman Research Institute Excellence Award (R.-J.L.). Y.B.W. is support by a grant from China Scholarship Council and Y.J.W. was supported by grants from National Natural Science Foundation of China, Zhejiang Provincial Natural Science Foundation of China, Zhejiang Provincial Major Science and Technology Project and China Scholarship Council.
Author information
Authors and Affiliations
Contributions
X.Wang, H.H., R.-J.L. and J.-K.Y. conceived of and designed experiments; X.Wang, Y.W., J.W., Z.Q. and T.C. performed experiments; X.Wang, X.Wu, R.-J.L. and J.-K.Y. data analysis and interpretation; X.Wang and J.-K.Y. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Indel formation by CRISPRs and TALENs targeting the WAS and TAT genes.
(a) Sequences of the CRISPRs and TALENs targeted regions in the WAS and TAT genes. Bases in grey indicate the PAM sequence. (b) Surveyor assay to detect CRISPR-mediated indel formation in the WAS and TAT genes (n=3). (c) Surveyor assay to detect TALEN-mediated indel formation in the WAS,TAT, ZRSR2 and miR-9-2 genes. HEK293T cells were transfected with the indicated CRISPR/Cas9 or TALEN pairs. Cells transfected with the GFP gene served as the negative control. Genomic DNA extracted 48 h later was assessed for target cleavage with the Surveyor assay (n=3). Results from duplicated transfection are shown. Efficiency of indel formation is shown below each lane. Arrows indicate the positions of the expected fragments.
Supplementary Figure 2 Genomic mapping of the IDLV-puro integration sites.
(a) Schematic display of mapping the IDLV-puro integration sites. Primer used for nrLAM-PCR is located in the U5 region of the lentiviral long terminal repeat (LTR) and is indicated with a half-arrowhead. The linear PCR products are ligated to a linker, followed by nested PCR to yield sufficient DNA for deep sequencing. (b) Representative photographs of IDLV-transduced colonies after puromycin selection. HEK293T cells were transfected with either CRISPR or TALEN, followed by tranduction twice with IDLV-puro. Puromycin selection was initiated two days after transduction. HEK293T cells transduced with IDLV-puro without CRISPR or TALEN served as the negative control. Expression of CRISPR or TALEN led to a 2-3 fold increase in the puromycin-resistant colonies (n=2).
Supplementary Figure 3 IDLV integration sites mapped within a ±60 bp window surrounding the on-target cleavage site of CRISPR or TALEN.
CRISPRs and TALENs were designed to target the same genomic region in the (a) WAS and (b)TAT genes. The expected cleavage sites are indicated by arrows. Black bars represent individual IDLV integration sites mapped by nrLAM-PCR.
Supplementary Figure 4 IDLV integration sites mapped within a ±60 bp window surrounding the off-target cleavage site of CRISPRs.
The off-target site was determined based on sequence similarity to the on-target site. Black bars represent individual IDLV integration sites mapped by nrLAM-PCR.
Supplementary Figure 5 Confirmation of IDLV mapped off-target sites with the Surveyor assay.
HEK293T cells were transfected with the expression plasmids for Cas9 and (a) WAS CR-4, (b) WAS CR-5, (c) TAT CR-1. Cells transfected with the expression plasmids for Cas9 and GFP genes served as the negative control. Genomic DNA isolated 48 h later was subjected to the Surveyor assay (n=3), and percent of indel formation was determined. Results from duplicated transfection are shown. Arrows indicate the positions of the expected fragments. The gene near the CRISPR/Cas9 cleavage site is indicated on top of each gel. UD, undetectable.
Supplementary Figure 6 Confirmation of the on- and off-target sites of CRISPR/Cas9 in HT1080 cells.
Human fibrosarcoma HT1080 cells were transfected with the expression plasmids for Cas9 and the indicated CRISPR. The genomic DNA extracted 48 h later was subjected to the Surveyor assay (n=3). (a) On-target cleavage by the indicated CRISPR/Cas9. (b) Off-target cleavage by the indicated CRISPR/Cas9. Cells transfected with the GFP/Cas9 genes served as the negative control. The gene near the off-target site is indicated on top of each gel. Arrows indicate the positions of the expected fragments. UD, undetectable.
Supplementary Figure 7 D10A Cas9 mediates efficient cleavage of the target in the WAS and TAT genes.
(a) Sequences of the human WAS and TAT genes targeted by CRISPR/D10A Cas9. Underlined bases indicate the CRISPR target and the PAM sequences. Bases labeled in grey indicate the PAM sequence; arrowheads indicate the positions of the nick created by D10A Cas9. Offset: the distance between the PAM-distal ends of a given pair of sgRNA. (b) Comparison of the on-target cleavage between individual CRISPR/Cas9 and CRISPR/D10A Cas9. HEK293T cells were transfected with the indicated CRISPR and Cas9 or D10A Cas9. The genomic DNA extracted 48 h later was analyzed with the Surveyor assay (n=3). Percent of indel is shown. Arrows indicate the positions of the expected fragments. UD, undetectable.
Supplementary Figure 8 Deep-sequencing analysis of CRISPR/Cas9-mediated cleavage of in silico predicted POT sites.
PCR primers were designed for each POT site with the expected cleavage site centrally located in the PCR product. The genomic sequence is shown in the first row and the red dash lines in this row indicate the space created to accommodate base insertion generated by NHEJ in this genomic region. The sequences of indel determined by deep sequencing are shown below each genomic sequence. Black dash lines indicate base deletion and red nucleotides indicate base insertion. Underlined bases indicate the off-target recognition sequence; bases labeled in grey denote the PAM sequence; arrows indicate the CRISPR/Cas9 cleavage site. The sequences with greater than 10 reads as determined by deep sequencing were calculated and only the top 20 reads were shown.
Supplementary Figure 9 Size of the colony input can modulate the sensitivity of the IDLV assay.
HEK293T cells were transfected with the WAS CR-4, followed by tranduction with IDLV-puro in three independent experiments. Approximately 500, 3000 or 15000 puromycin-resistant colonies were pooled for nrLAM-PCR and mapping of the IDLV integration site. With 500 colonies, only 5 of the 7 off-target sites detected in the 3000 colony pool could be identified. The 2 off-target sites, CD70 and HOXC13, with the lowest NHEJ frequency as determined by the Surveyor assay and deep sequencing failed to be detected in the 500 colony pool. From the 15000 colony pool, 4 additional off-target sites besides the 7 off-target sites present in the 3000 colony pool were identified.
Supplementary Figure 10 Base-skipping CRISPR mutants mediate efficient on-target cleavage.
HEK293T cells were transfected with the Cas9 expression plasmids with (a) WAS CR-4 and (b) WAS CR-5 as indicated, and the genomic DNA harvested 48 h later was assessed using the Surveyor assay (n=3). Efficient target cleavage was observed in some mutants with one-base skipping. Arrows indicate the positions of the expected fragments. UD, undetectable.
Supplementary Figure 11 Base-skipping CRISPR mutants mediated efficient cleavage with Cas9 and D10A Cas9.
HEK293T cells were transfected with the indicated plasmids and the genomic DNA harvested 48 h later was assessed using the Surveyor assay. The mutant name is as described in Fig. 2. wt: wild type; UD, undetectable.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 and Supplementary Tables 1–7 (PDF 10173 kb)
Rights and permissions
About this article
Cite this article
Wang, X., Wang, Y., Wu, X. et al. Unbiased detection of off-target cleavage by CRISPR-Cas9 and TALENs using integrase-defective lentiviral vectors. Nat Biotechnol 33, 175–178 (2015). https://doi.org/10.1038/nbt.3127
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3127
This article is cited by
-
Extru-seq: a method for predicting genome-wide Cas9 off-target sites with advantages of both cell-based and in vitro approaches
Genome Biology (2023)
-
Discovering CRISPR–Cas off-target breaks
Nature Methods (2023)
-
Deep sampling of gRNA in the human genome and deep-learning-informed prediction of gRNA activities
Cell Discovery (2023)
-
DNA as the main target in radiotherapy—a historical overview from first isolation to anti-tumour immune response
Strahlentherapie und Onkologie (2023)
-
Production of Duchenne muscular dystrophy cellular model using CRISPR-Cas9 exon deletion strategy
Molecular and Cellular Biochemistry (2023)