Abstract
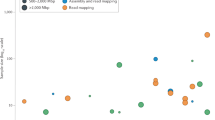
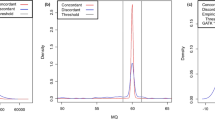
Whole-genome sequencing is becoming commonplace, but the accuracy and completeness of variant calling by the most widely used platforms from Illumina and Complete Genomics have not been reported. Here we sequenced the genome of an individual with both technologies to a high average coverage of ∼76×, and compared their performance with respect to sequence coverage and calling of single-nucleotide variants (SNVs), insertions and deletions (indels). Although 88.1% of the ∼3.7 million unique SNVs were concordant between platforms, there were tens of thousands of platform-specific calls located in genes and other genomic regions. In contrast, 26.5% of indels were concordant between platforms. Target enrichment validated 92.7% of the concordant SNVs, whereas validation by genotyping array revealed a sensitivity of 99.3%. The validation experiments also suggested that >60% of the platform-specific variants were indeed present in the genome. Our results have important implications for understanding the accuracy and completeness of the genome sequencing platforms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Change history
07 June 2012
In the version of this article initially published, the accession code to obtain raw sequence data was given as SRA045736.2; the correct code is SRA045736. The error has been corrected in the HTML and PDF versions of the article.
References
Ajay, S.S., Parker, S.C., Ozel Abaan, H., Fuentes Fajardo, K.V. & Margulies, E.H. Accurate and comprehensive sequencing of personal genomes. Genome Research 21, 1498–1505 (2011).
Ashley, E.A. et al. Clinical assessment incorporating a personal genome. Lancet 375, 1525–1535 (2010).
Wheeler, D.A. et al. The complete genome of an individual by massively parallel DNA sequencing. Nature 452, 872–876 (2008).
McKernan, K.J. et al. Sequence and structural variation in a human genome uncovered by short-read, massively parallel ligation sequencing using two-base encoding. Genome Res. 19, 1527–1541 (2009).
Roach, J.C. et al. Analysis of genetic inheritance in a family quartet by whole-genome sequencing. Science 328, 636–639 (2010).
Pushkarev, D., Neff, N. & Quake, S. Single-molecule sequencing of an individual human genome. Nat. Biotechnol. 27, 847–852 (2009).
Korbel, J.O. et al. Paired-end mapping reveals extensive structural variation in the human genome. Science 318, 420–426 (2007).
Snyder, M., Du, J. & Gerstein, M. Personal genome sequencing: current approaches and challenges. Genes Dev. 24, 423–431 (2010).
Rios, J., Stein, E., Shendure, J., Hobbs, H.H. & Cohen, J.C. Identification by whole-genome resequencing of gene defect responsible for severe hypercholesterolemia. Hum. Mol. Genet. 19, 4313–4318 (2010).
Lee, W. et al. The mutation spectrum revealed by paired genome sequences from a lung cancer patient. Nature 465, 473–477 (2010).
The 1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Lander, E.S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Sherry, S.T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Chen, R., Davydov, E.V., Sirota, M. & Butte, A.J. Non-synonymous and synonymous coding SNPs show similar likelihood and effect size of human disease association. PLoS ONE 5, e13574 (2010).
Kaur, I. et al. Variants in the 10q26 gene cluster (LOC387715 and HTRA1) exhibit enhanced risk of age-related macular degeneration along with CFH in Indian patients. Invest. Ophthalmol. Vis. Sci. 49, 1771–1776 (2008).
Tam, P.O. et al. HTRA1 variants in exudative age-related macular degeneration and interactions with smoking and CFH. Invest. Ophthalmol. Vis. Sci. 49, 2357–2365 (2008).
Yamaguchi, H. et al. Mutations in TERT, the gene for telomerase reverse transcriptase, in aplastic anemia. N. Engl. J. Med. 352, 1413–1424 (2005).
Albers, C.A. et al. Dindel: Accurate indel calls from short-read data. Genome Res. 21, 961–973 (2011).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Clark, M.J. et al. Performance comparison of exome DNA sequencing technologies. Nat. Biotechnol. 29, 908–914 (2011).
Acknowledgements
This work is supported by the Stanford Department of Genetics and the US National Institutes of Health.
Author information
Authors and Affiliations
Contributions
H.Y.K.L. and M.J.C. did the analysis. G.N. and L.H. assisted in the analysis. Rui C. did DNA sequencing. Rong C. did the disease-association study. Rui C. and M.O'H. did the validation experiments. H.Y.K.L., F.E.D., E.A.A., M.B.G., A.J.B., H.P.J. and M.S. coordinated the analysis and revised the manuscript. H.Y.K.L., M.J.C. and M.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.S. is a scientific advisory board member for Genapsys, Inc.; a scientific advisory board member and cofounder of Personalis, Inc.; and a consultant for Illumina.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2 (PDF 1129 kb)
Supplementary Table 1
Disease association of all platform-specific SNPs. (XLSX 40 kb)
Rights and permissions
About this article
Cite this article
Lam, H., Clark, M., Chen, R. et al. Performance comparison of whole-genome sequencing platforms. Nat Biotechnol 30, 78–82 (2012). https://doi.org/10.1038/nbt.2065
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2065
This article is cited by
-
MicrosatNavigator: exploring nonrandom distribution and lineage-specificity of microsatellite repeat motifs on vertebrate sex chromosomes across 186 whole genomes
Chromosome Research (2023)
-
Methods for exploring the faecal microbiome of premature infants: a review
Maternal Health, Neonatology and Perinatology (2021)
-
Estimating sequencing error rates using families
BioData Mining (2021)
-
New genetic variants associated with major adverse cardiovascular events in patients with acute coronary syndromes and treated with clopidogrel and aspirin
The Pharmacogenomics Journal (2021)
-
PacBio sequencing output increased through uniform and directional fivefold concatenation
Scientific Reports (2021)