Abstract
The primary two-dose SARS-CoV-2 mRNA vaccine series are strongly immunogenic in humans, but the emergence of highly infectious variants necessitated additional doses and the development of vaccines aimed at the new variants1,2,3,4. SARS-CoV-2 booster immunizations in humans primarily recruit pre-existing memory B cells5,6,7,8,9. However, it remains unclear whether the additional doses induce germinal centre reactions whereby re-engaged B cells can further mature, and whether variant-derived vaccines can elicit responses to variant-specific epitopes. Here we show that boosting with an mRNA vaccine against the original monovalent SARS-CoV-2 mRNA vaccine or the bivalent B.1.351 and B.1.617.2 (Beta/Delta) mRNA vaccine induced robust spike-specific germinal centre B cell responses in humans. The germinal centre response persisted for at least eight weeks, leading to significantly more mutated antigen-specific bone marrow plasma cell and memory B cell compartments. Spike-binding monoclonal antibodies derived from memory B cells isolated from individuals boosted with either the original SARS-CoV-2 spike protein, bivalent Beta/Delta vaccine or a monovalent Omicron BA.1-based vaccine predominantly recognized the original SARS-CoV-2 spike protein. Nonetheless, using a more targeted sorting approach, we isolated monoclonal antibodies that recognized the BA.1 spike protein but not the original SARS-CoV-2 spike protein from individuals who received the mRNA-1273.529 booster; these antibodies were less mutated and recognized novel epitopes within the spike protein, suggesting that they originated from naive B cells. Thus, SARS-CoV-2 booster immunizations in humans induce robust germinal centre B cell responses and can generate de novo B cell responses targeting variant-specific epitopes.
Similar content being viewed by others
Main
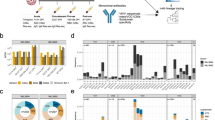
The emergence of SARS-CoV-2 variants with increasing numbers of mutations in the spike (S) protein has decreased the effectiveness of primary series vaccinations and led to a recommendation for booster immunizations in most populations10,11,12,13,14,15,16. Multiple reports have documented that third-dose booster immunizations based on the ancestral Washington strain (WA1/2020) enhanced antibody responses to the ancestral strain as well as emerging variants of concern and correspondingly increased protective effectiveness5,6,7,8,17,18,19,20,21. In addition, new vaccines based on circulating variants have been developed to enhance the ability of induced antibodies to combat such variants. Indeed, recent evidence indicates that a B.1.351 (Beta)-containing booster can generate higher titres of neutralizing antibodies against both B.1.351 and B.1.1.529 (Omicron) BA.1 strains of SARS-CoV-2 compared with a booster based on the original strain alone22. Additionally, bivalent boosters encoding the original strain and either the BA.1 or BA.5 subvariant induced broader neutralizing antibody responses than the constituent monovalent vaccines23. Whether re-exposure to mRNA vaccines encoding S protein from the original SARS-CoV-2 strain or from variants of concern induce robust germinal centre reactions that are critical for refining high-affinity and durable antibody responses has not been examined in humans. To address this question, we conducted an immunization study of 46 healthy adults with no history of SARS-CoV-2 infection, all of whom had completed a primary two-dose vaccination course with either the Pfizer–BioNTech (BNT162b2) or Moderna (mRNA-1273) SARS-CoV-2 mRNA vaccines, both of which encode a prefusion-stabilized full-length SARS-CoV-2 S protein based on the WA1/2020 strain24,25. Recruited individuals received a third booster dose of 50 μg mRNA-1273 or mRNA-1273.213, which contains a total of 100 μg of mRNA encoding B.1.351 (Beta; sequence conservation of 99.1% for S protein and 98.7% for the receptor binding domain (RBD) compared with WA1/2020) and B.1.617.2 (Delta; sequence conservation of 99.2% for S protein and 99.1% for the RBD compared with WA1/2020) SARS-CoV-2 S proteins (Extended Data Tables 1 and 2), as a sub-study of an ongoing clinical trial (Clinical NCT04927065).
Seven of the participants received a booster immunization of mRNA-1273, and 39 received mRNA-1273.213. Blood samples were collected at baseline and weeks 1, 2, 4, 8, 17 and 26 after vaccination. Five participants from the mRNA-1273 cohort and twenty from the mRNA-1273.213 cohort consented to collection of fine needle aspirates (FNAs) of draining axillary lymph nodes at weeks 2 and 8. Three participants from the mRNA-1273 cohort and 11 from the mRNA-1273.213 cohort consented to collection of bone marrow aspirates 26 weeks after their third dose (Fig. 1a). Circulating S protein-specific antibody-producing plasmablasts were measured by enzyme-linked immune absorbent spot (ELISpot) assay. S protein-specific IgG- and IgA-producing plasmablasts were detected one week after the booster immunization from all participants in the mRNA-1273 cohort. Robust circulating IgG-producing plasmablast responses against the original WA1/2020 S protein and against the vaccine-encoded B.1.351 and B.1.617.2 S proteins were detected in all participants in the mRNA-1273.213 cohort 1 week after immunization, with lower IgA responses detectable in most participants (Fig. 1b and Extended Data Fig. 1a). Plasma IgG antibody titres against S protein from the WA1/2020, B.1.351, B.1.617.2 and BA.1 strains were measured by multiplex bead binding assay. In both cohorts, plasma antibody binding levels against all strains were increased 4 weeks after immunization and declined approximately 1.3- to 2-fold by 17 weeks (Extended Data Fig. 1b,c).
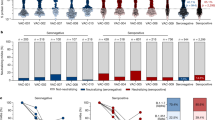
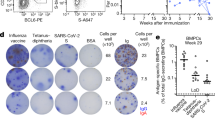
a, Seven healthy adults received an mRNA-1273 booster and thirty-nine healthy adults received a mRNA-1273.213 booster. Blood was collected at baseline and at 1, 2, 4, 8, 17 and 26 weeks after the booster. FNAs of ipsilateral axillary lymph nodes were collected 2 and 8 weeks after the booster from 5 participants from the mRNA-1273 cohort and from 20 participants from the mRNA-1273.213 cohort; bone marrow aspirates were collected 26 weeks after the booster from 3 participants from the mRNA-1273 cohort and 11 participants from the mRNA-1273.213 cohort. LN, lymph node. b, Frequencies of IgG plasmablasts (PBs) specific for the indicated antigens measured by ELISpot in week (wk) 1 PBMCs from participants who received mRNA-1273 (left; n = 7) and mRNA-1273.213 (right; n = 38). c,d, Representative flow cytometry plots of BCL6 and CD38 (c) or CD20 and CD38 (d) on IgDlo B cells in FNA (c) or PBMC (d) samples (left), pooled (WA1/2020, B.1.351, B.1.617.2 and BA.1) S protein probe (BV421 and BV650) staining on BCL6+CD38int germinal centre B cells (c) and CD20+CD38lo/int MBCs (d) (second from left), and frequencies of S+ germinal centre B cells from FNA (c) or S+ MBCs from PBMCs (d) from participants who received mRNA-1273 (second from right) and mRNA-1273.213 (right). c, mRNA-1273, n = 5; mRNA-1273.213, n = 20. d, mRNA-1273, n = 6; mRNA-1273.213, n = 28. e, Representative ELISpot wells coated with the indicated antigens, developed in blue (IgG) and red (IgA) after plating the indicated numbers of BMPCs. f, Frequencies of IgG BMPCs specific for the indicated antigens 26 weeks after participants received mRNA-1273 (left) or mRNA-1273.213 (right) boosters. Lines indicate medians. Symbols at each time point represent one sample. mRNA-1273, n = 3; mRNA-1273.213, n = 10. g,h, Binding of monoclonal antibodies from week 17 S+ MBCs from participants who received mRNA-1273 (left) and mRNA-1273.213 (right) to indicated antigens (g) and constituent domains of WA1/2020 S protein (h), measured by ELISA.
Ultrasonography was used to identify and guide FNA of accessible axillary lymph nodes on the same side as the site of immunization two and eight weeks after the booster. FNA samples were stained with pooled fluorescently labelled S protein probes from the WA1/2020, B.1.351, B.1.617.2 and BA.1 strains and then analysed by flow cytometry to detect S protein-specific B cells. We included BA.1 S protein probes in the pool to detect memory B cells (MBCs) that may have stochastically mutated to better recognize the variant antigen26. Spike-recognizing (S+) germinal centre B cells (CD19+CD3−IgDloBCL6+CD38int) and follicular helper T (TFH) cells (defined as CD3+CD19−CD4+CD8−CD14−CXCR5+PD-1+BCL6+FOXP3−) were detected in FNAs from all participants analysed at week 2. Frequencies of S+ germinal centre B cells and TFH cells correlated significantly and remained readily detectable in all but one participant from each cohort at week 8, consistent with the robust germinal centre response observed after primary vaccination with SARS-CoV-2 mRNA vaccines2,27,28 (Fig. 1c and Extended Data Fig. 1d–f). Circulating S+ MBCs in blood were identified as CD19+CD3−IgDloCD20+ cells that bound the pooled fluorescently labelled S protein probes. S+ MBCs were detected in all participants before the booster administration and at similar frequencies 17 weeks after the booster, with median frequencies of 0.52% and 0.55% of the total circulating B cells in the mRNA-1273 cohort, and 0.42% and 0.49% in the mRNA-1273.213 cohort (Fig. 1d). Consistent with sustained plasma antibody titres, bone marrow plasma cells (BMPCs) producing IgG that bound WA1/2020, B.1.351, B.1.617.2 and BA.1 S proteins were detected in all thirteen participants with enough BMPCs for ELISpot (3 from the mRNA-1273 and 10 from the mRNA-1273.213 cohort). Although the differences were not statistically significant, the frequencies of BMPCs that bound WA1/2020, B.1.351 and B.1.617.2 S proteins were approximately half of those of seasonal influenza vaccine-specific BMPCs, which accumulate over a lifetime of repeated antigen exposures. Similarly, the frequencies of BA.1-binding BMPCs were approximately one-quarter of those binding the other strains. S protein-binding IgA-producing BMPCs were considerably rarer or undetectable (Fig. 1e,f and Extended Data Fig. 1g,h).
To characterize the breadth of the MBC repertoire after boosting, we selected three participants from each cohort for whom we had characterized the B cell response to the primary vaccination series: 382-02, 382-07 and 382-08 from the mRNA-1273 cohort and 382-01, 382-13 and 382-15 from the mRNA-1273.213 cohort2,29. For both cohorts, we stained week 17 post-boost peripheral blood mononuclear cells (PBMCs) with pooled fluorescently labelled S protein probes from the WA1/2020, B.1.351, B.1.617.2 and BA.1 strains, enabling us to sort MBCs for monoclonal antibody generation independently of the S protein probe that they bound. We also bulk-sorted week 17 total MBCs for heavy chain immunoglobulin sequencing to broaden the clonal repertoire analyses (Extended Data Fig. 2a and Extended Data Table 3). We generated 81, 59, 73, 94, 90 and 59 clonally distinct monoclonal antibodies from participants 382-02, 382-07, 382-08, 382-01, 382-13 and 382-15, respectively. We next assessed the binding of these antibodies to S protein from the WA1/2020, 1.351, 1.617.2 and BA.1 strains by enzyme-linked immunosorbent assay (ELISA). In total, 205 out of 213 (96%) and 222 out of 243 (91%) monoclonal antibodies derived from the MBCs from the mRNA-1273 and mRNA-1273.213 cohorts, respectively, recognized the original WA1/2020, B.1.351 and B.1.617.2 S proteins, with 129 (61%) and 138 (57%), respectively, also recognizing BA.1 (Fig. 1g). Evaluation of the domains targeted by each monoclonal antibody revealed that 40%, 18% and 42% from the mRNA-1273 cohort and 43%, 16% and 39% from the mRNA-1273.213 cohort bound RBD, the N-terminal domain (NTD) and S2, respectively (Fig. 1h).
To track the clonal dynamics of the B cell response in the two cohorts, we performed single-cell RNA sequencing (scRNA-seq) on week 1 sorted plasmablasts and week 2 and week 8 FNA specimens from the same 6 participants and bulk-sorted germinal centre B cells and lymph node plasma cells (LNPCs) for heavy chain immunoglobulin sequencing (Fig. 2a,e, Extended Data Fig. 3a–g and Extended Data Tables 3–5). We then computationally inferred clonal relationships on the basis of common heavy chain V and J gene usage, CDR3 lengths and CDR3 similarity30,31 to link the B cell receptor sequences to known S-specific clones identified among monoclonal antibodies derived from MBCs isolated 17 weeks after the third vaccine dose or from the previously characterized response to the primary vaccination series2,29 (Fig. 2b,e, Extended Data Fig. 3h and Extended Data Table 6). The majority of S-specific plasmablasts identified by scRNA-seq after the booster dose were clonally related to MBCs, germinal centre B cells and/or plasma cells induced by primary vaccination (Fig. 2c). Multiple S-specific clones were detected in the plasmablast response after both the second dose of the primary vaccination series and the booster. Representatives of these clones participating in the booster plasmablast response had significantly higher somatic hypermutation (SHM) frequencies in their immunoglobulin heavy chain variable region (IGHV) genes than those from the primary response, consistent with their recall from affinity-matured MBCs (Fig. 2d). S-specific plasmablast clones identified one week after the booster dose were identified in germinal centre responses in all six analysed participants, although the peak frequencies of the S-specific germinal centre clonal repertoire occupied by plasmablast clones varied from 17% to 100% among participants (Extended Data Fig. 3i,j). SHM frequencies among S-specific MBCs isolated 17 weeks after the booster were significantly higher than their clonally related counterparts isolated 6 months after the primary vaccination, consistent with additional rounds of germinal centre-driven maturation (Fig. 2f). Similar trends were observed among paired S-specific BMPC clones analysed 6 or 9 months after the primary immunization and 6 months after the booster. However, SHM did not increase in all clonal families, consistent with durable populations of MBCs and BMPCs generated by the primary vaccine response persisting through the booster response (Fig. 2f,g). mRNA-1273 and mRNA-1273.213 both elicited robust germinal centre responses and maturation of the MBC and BMPC responses, but we did not isolate any antibodies specifically targeting S proteins from the variant strains encoded by the mRNA-1273.213 vaccine that did not cross-react to the original WA1/2020 S protein. Thus, the B cell response after boosting with the mRNA-1273.213 vaccine was imprinted by the primary vaccination series with mRNA-1273, which encodes the ancestral S protein.
a,b,e, Uniform manifold approximation and projection (UMAP) of scRNA-seq transcriptional clusters of B cells from sorted circulating plasmablasts 1 week after booster immunization, with log-normalized XBP1 gene expression (a) and S-specific clones (b) overlaid, or from FNAs of draining lymph nodes, with S-specific clones overlaid (e). Each dot represents a cell. c, Clonal overlap and percentages of S-specific plasmablast clones related to clones generated during the primary vaccine response among participants receiving mRNA-1273 (top) and mRNA-1273.213 (bottom). Peripheral arc lengths correspond to the number of B cell receptor sequences in the respective compartment and chord widths correspond to clone size. Purple chords connect clonally related sequences between plasmablast and pre-boost arcs. Percentages are of plasmablast clones related to pre-boost S-specific clones. The number of sequences is indicated below each plot. d,f,g, Paired median IGHV gene mutation frequencies of S-specific clones found among plasmablasts 1 week after the second dose of the primary vaccine series and 1 week after booster vaccination (d), MBCs identified 6 months after primary vaccination and 17 weeks after booster vaccination (f), and BMPCs identified 6 and/or 9 months after primary vaccination and 6 months after booster vaccination (g). Each symbol represents the median mutation frequency of a clone; horizontal lines indicate medians. P values from two-sided Wilcoxon test. d, mRNA-1273, n = 52; mRNA-1273.213, n = 104. f, mRNA-1273, n = 42; mRNA-1273.213, n = 39. g, mRNA-1273, n = 7; mRNA-1273.213, n = 15.
To determine whether a more antigenically divergent booster could generate a detectable response targeting novel epitopes, we recruited 8 participants who had received a 2-dose mRNA primary vaccination series and had no history of SARS-CoV-2 infection to receive 50 μg of mRNA-1273.529, which encodes BA.1 S protein (97.1% S protein and 93.3% RBD sequence conservation versus WA1/2020), as a first (third dose) or second (fourth dose at least 3 months after a third dose of mRNA-1273) booster (Extended Data Fig. 4a and Extended Data Tables 1 and 2). We analysed peripheral blood samples 1 and 17 weeks after the mRNA-1273.529 booster. All seven participants analysed one week after receiving the booster exhibited robust circulating IgG-producing plasmablast responses against the original WA1/2020 S protein as well as the vaccine-encoded BA.1 S protein, with lower IgA responses detectable in most participants (Fig. 3a and Extended Data Fig. 4b). To analyse the breadth of the MBC repertoire, we sorted MBCs from PBMCs isolated 17 weeks after the booster from participants 382-53, 382-54 and 382-55 (all of whom received mRNA-1273.529 as a third dose booster) using the same pooled fluorescent S protein probes from the WA1/2020, B.1.351, B.1.617.2 and BA.1 strains to detect MBCs regardless of their strain specificity and to maintain consistency with the previously generated monoclonal antibodies. Similar to the monoclonal antibodies from the mRNA-1273 and mRNA-1273.213 cohorts, 303 out of 310 monoclonal antibodies (98%) were cross-reactive, binding S protein from WA1/2020, B.1.351 and B.1.617.2, including 220 (71%) that also recognized the BA.1 S protein encoded by the mRNA-1273.529 vaccine (Fig. 3b). Evaluation of the domains targeted by each monoclonal antibody revealed that 35% bound the RBD, 15% bound the NTD and 48% bound S2 (Fig. 3c). To assess the neutralization capacity of the MBC-derived monoclonal antibodies, we first screened all 730 S-binding monoclonal antibodies from the 3 cohorts with a high-throughput assay using chimeric vesicular stomatitis virus (VSV) in which the native envelope glycoprotein was replaced with S protein from the WA1/2020 strain with the D614G substitution32. Thirty monoclonal antibodies from the mRNA-1273 cohort, 49 from the mRNA-1273.213 cohort and 52 from the mRNA-1273.529 cohort neutralized infection by at least 80% at 5 μg ml−1 (Extended Data Fig. 4c). We then evaluated the neutralizing capacity of these 131 monoclonal antibodies against a panel of authentic, infectious SARS-CoV-2 variants, including WA1/2020 D614G, B.1.351, B.1.617.2, BA.1 and BA.5, against which 67%, 47%, 67%, 10% and 10% from the mRNA-1273 cohort, 76%, 51%, 76%, 20% and 24% from the mRNA-1273.213 cohort, and 77%, 65%, 71%, 33% and 23% from the mRNA-1273.529 cohort, respectively, reduced infection by at least 90% at 5 μg ml−1 (Fig. 3d).
a, Frequencies of S-binding IgG-producing plasmablasts in PBMC 1 week after booster vaccination, measured by ELISpot in participants who received mRNA-1273.529. Horizontal lines indicate medians. Each symbol represents a sample; n = 7. b,c, Binding of monoclonal antibodies from S+ MBCs 17 weeks after mRNA-1273.529 booster immunization to S proteins from indicated SARS-CoV-2 strains (b) and constituent domains of WA1/2020 S protein (c), measured by ELISA. d, Neutralizing activity of monoclonal antibodies from week 17 S+ MBCs against indicated strains of authentic SARS-CoV-2 virus from participants who received indicated third-dose booster vaccines. Each symbol represents one monoclonal antibody. mRNA-1273, n = 30; mRNA-1273.213, n = 49; mRNA-1273.529, n = 52. Percentages indicate the proportion of monoclonal antibodies above the 90% infection reduction threshold. P values are from Chi-squared tests between the three vaccine cohorts.
Given the high frequency of cross-reactive monoclonal antibodies, we used a more targeted approach to sort MBCs specific for BA.1 S protein that did not bind WA1/2020 S protein from participants 382-53, 382-54 and 382-55 (Fig. 4a). Sixty-five out of the seventy-eight monoclonal antibodies (83%) isolated using this approach still cross-reacted with WA1/2020 S protein, indicating that the S-dependent sorting approach is not ideally sensitive for determining variant binding specificity. Out of the 78 monoclonal antibodies, 6 bound BA.1 S protein only, all of which were from participants 382-54 and 382-55 (Fig. 4b). Evaluation of the domains targeted by each monoclonal antibody revealed an increased proportion of NTD-targeting monoclonal antibodies at the expense of those specific for S2 compared with the monoclonal antibodies that were not sorted to exclude WA1/2020, with 31%, 53% and 10% binding RBD, NTD and S2, respectively (Fig. 4c). The SHM frequencies among the clonal families of the six monoclonal antibodies that bound BA.1 S protein but not WA1/2020 S protein were significantly lower than those from the same participants that cross-reacted with WA1/2020 S protein (Fig. 4d). We tested the neutralizing capacity of the six monoclonal antibodies that bound BA.1 but not WA1/2020 against the panel of SARS-CoV-2 variants. As expected, none of the six antibodies neutralized the ancestral D614G strain. Notably, all six neutralized BA.1, one also neutralized BA.5 (17%) and none neutralized B.1.351 or B.1.617.2 at 5 μg ml−1 (Fig. 4e). All six of the BA.1-neutralizing monoclonal antibodies targeted the RBD of S protein (Extended Data Fig. 4d). To identify the amino acid residues targeted by the six BA.1-specific monoclonal antibodies, we used VSV–SARS-CoV-2 S chimeric viruses (S from BA.1 strain) to select variants that escaped neutralization, as previously described32,33. We performed plaque assays on Vero E6 cells with the six neutralizing monoclonal antibodies in the overlay, purified the neutralization-resistant plaques, and sequenced the S genes (Fig. 4f and Extended Data Fig. 4e,f). Sequence analysis identified S protein escape mutations at R498 for monoclonal antibodies 382-54 3B02, 382-55 1A11 and 382-55 1C04; R498 and H505 for monoclonal antibody 382-55 1A05; L371, A372, F374, F375 and K378 for monoclonal antibody 382-55 2B01; and A484 and F490S for monoclonal antibody 382-55 3A07. Notably, many of the mutations were reversions to the ancestral strain of SARS-CoV-2. (Fig. 4g).
a, Gating strategy for sorting BA.1+ WA1/2020− MBCs from PBMCs 17 weeks post booster. b,c, Binding of monoclonal antibodies from BA.1+ WA1/2020− sorted MBCs to indicated strains of SARS-CoV-2 S protein (b), and constituent domains of S protein (c), measured by ELISA. d, IGHV mutation frequencies of clones related to monoclonal antibodies from participants 382-54 and 382-55 that bound WA1/2020 (left) and BA.1 but not WA1/2020 (right) S protein. Each symbol represents the median mutation frequency of a clone. WA1/2020+, n = 266; BA.1+ WA1/2020−, n = 6. Black lines indicate medians. P value from two-sided Mann–Whitney test. e, Neutralizing activity of BA.1+ WA1/2020− S-binding monoclonal antibodies against indicated strains of authentic SARS-CoV-2 virus. Numbers above each virus are of monoclonal antibodies above the 90% infection reduction threshold. f, Plaque assays on Vero E6 cells with indicated monoclonal antibody in the overlay to isolate escape mutants (red arrows). Images are representative of three experiments per monoclonal antibody. g, Structure of the RBD with the human ACE2 footprint highlighted in brown. BA.1 mutations are highlighted in blue and amino acids whose substitution confers resistance to indicated monoclonal antibodies in plaque assays are highlighted in red. Asterisks indicate substitutions that confer resistance to the indicated monoclonal antibody and are reversions from BA.1 to the WA1/2020 sequence.
Here we have evaluated antigen-specific B cell responses to SARS-CoV-2 mRNA-based booster immunizations in humans. We show that boosting with ancestral SARS-CoV-2 or bivalent B.1.351/B.1.617.2-matched vaccines induce robust S-specific germinal centre responses in draining axillary lymph nodes of all sampled participants that last until at least eight weeks after vaccination. S-specific plasmablasts and germinal centre B cells originate predominantly from pre-existing clonal lineages, consistent with most of the MBC-derived monoclonal antibodies that we isolated recognizing the original SARS-CoV-2 S protein. We also demonstrated that immunization with the monovalent B.1.1.529 BA.1-matched vaccine could induce rare de novo B cell responses against novel epitopes in the B.1.1.529 BA.1 S protein. These observations expand the large body of data showing improved potency and breadth of serum antibody responses after SARS-CoV-2 booster immunization in humans5,6,7,8,9,18.
Several critical insights relevant to cellular immunity to SARS-CoV-2 and recall responses to vaccination can be drawn from this study. B cell clones comprising the plasmablast compartment induced after the booster immunization were significantly more mutated than the same clones detected one week after the completion of the primary vaccination series, a clear indication of the robust maturation process triggered by the primary vaccination2,29. Consistent with our previous work on B cell responses to seasonal influenza virus vaccination in humans34, the data presented here show that pre-existing MBCs can be efficiently re-engaged into recall germinal centre reactions. The frequencies of IgG-secreting BMPCs specific for the original SARS-CoV-2 S protein were several fold higher than those measured seven months after mild SARS-CoV-2 infections or six months after the primary mRNA vaccination series29,35. These increased frequencies are likely to be the result of the persistent germinal centre responses induced after the primary vaccination series and the new germinal centre reactions seeded by the booster immunization, highlighting the critical contribution of repeated antigen exposures to increasing antigen-specific BMPC frequency36,37.
A key unexpected finding in our study was the exceptionally high percentage of circulating MBCs that recognize the S protein from the original SARS-CoV-2 strain in the individuals boosted with variant S antigens, particularly as the booster immunizations did not encode S protein from the original strain. Of note, none of the participants from whom the monoclonal antibodies were derived had documented SARS-CoV-2 infection or seroconverted against the virus nucleocapsid protein during the course of the study. These data are consistent with MBCs generated by the primary vaccination series dominating the recall response induced by the booster and potentially out-competing clones specific for novel epitopes. It is possible that we were unable to isolate more memory B cell-derived monoclonal antibodies specific for novel epitopes on the B.1.351, B.1.617.2 or B.1.1.529 BA.1 S proteins because their affinity for the probes was below the limit of detection. The high antigenic similarity between the variant-derived and original SARS-CoV-2 S antigens in the case of the bivalent B.1.351/B.1.617.2 vaccine may have contributed to the low frequency of de novo clones recognizing the variants, and our data lend support for the decision to not adopt this vaccine for clinical use. Given the degree of cross-reactivity in the response we observed to purely variant-based vaccines, it is likely that the recently licensed bivalent boosters encoding WA1/2020 and either BA.1 or BA.4/5 will elicit similarly cross-reactive responses. We also speculate that an additional immunization with a variant-based vaccine may be needed to amplify the exclusively variant-specific B cell clones, similar to what has been observed upon H5N1 influenza virus immunization in humans38. We note that many of the mutations selected when we cultured recombinant vesicular stomatitis virus expressing SARS-CoV-2 Omicron (BA.1) S protein in the presence of the de novo monoclonal antibodies were reverted to the residues in the original strain. This suggests that newly escaped viruses are likely to be sensitive to potently neutralizing antibodies, including some that were clinically approved for therapeutic use, that were thought to be no longer useful because of the changes at the E484 residue, for example39,40,41,42,43.
The high prevalence of MBCs recognizing the original SARS-CoV-2 S protein is evidence of antigenic imprinting, in which B cell responses to previously encountered antigens remain dominant even after exposure to different but antigenically related antigens44,45. Here we provide evidence that immunization with a distinct antigen can engage rare naive B cells that target novel epitopes, but the overall response remains dominated by clones that are cross-reactive to the original strain.
Methods
Sample collection, preparation and storage
All studies were approved by the Institutional Review Board of Washington University in St Louis. Written consent was obtained from all participants. Fifty-four healthy volunteers were enroled, of whom 26 and 15 provided axillary lymph node and bone marrow aspirate samples, respectively (Extended Data Table 1). Blood samples were collected in ethylenediaminetetraacetic acid (EDTA) evacuated tubes (BD), and PBMCs were enriched by density gradient centrifugation over Lymphopure (BioLegend). The residual red blood cells were lysed with ammonium chloride lysis buffer, washed with PBS supplemented with 2% fetal bovine serum (FBS) and 2 mM EDTA (P2), and PBMC were immediately used or cryopreserved in 10% dimethylsulfoxide (DMSO) in FBS. Ultrasound-guided FNA of axillary lymph nodes was performed by a radiologist. Lymph node dimensions and cortical thickness were measured, and the presence and degree of cortical vascularity and location of the lymph node relative to the axillary vein were determined prior to each FNA. For each FNA sample, six passes were made under continuous real-time ultrasound guidance using 22- or 25-gauge needles, each of which was flushed with 3 ml of RPMI 1640 supplemented with 10% FBS and 100 U ml−1 penicillin/streptomycin, followed by three 1-ml rinses. Red blood cells were lysed with ammonium chloride buffer (Lonza), washed with P2, and immediately used or cryopreserved in 10% DMSO in FBS. Participants reported no adverse effects from phlebotomies or serial FNAs. Bone marrow aspirates of approximately 30 ml were collected in EDTA tubes from the iliac crest. Bone marrow mononuclear cells were enriched by density gradient centrifugation over Ficoll-Paque PLUS (Cytiva), and remaining red blood cells were lysed with ammonium chloride buffer (Lonza) and washed with P2. BMPCs were enriched from bone marrow mononuclear cells using the EasySep Human CD138 Positive Selection Kit II (StemCell Technologies) and immediately used for ELISpot or cryopreserved in 10% DMSO in FBS.
Cell lines
Expi293F cells were cultured in Expi293 Expression Medium (Gibco). Vero-TMPRSS2 cells46 (a gift from S. Ding) were cultured at 37 °C in Dulbecco’s modified Eagle medium (DMEM) supplemented with 10% FBS, 10 mM HEPES (pH 7.3), 1 mM sodium pyruvate, 1× nonessential amino acids, 100 U ml−1 of penicillin/streptomycin, and 5 μg ml−1 of blasticidin. Vero E6 cells were cultured at 37 °C in DMEM supplemented with 10% FBS, 10 mM HEPES (pH 7.3), 1 mM sodium pyruvate, 1× nonessential amino acids, and 100 U ml−1 of penicillin/streptomycin.
Antigens
Recombinant soluble S protein from WA1/2020, B.1.351, B.1.617.2 and B.1.1.529 (BA.1) strains of SARS-CoV-2 and their Avi-tagged counterparts were expressed as previously described29,47. In brief, mammalian cell codon-optimized nucleotide sequences coding for the soluble ectodomain of S (GenBank: MN908947.3, amino acids 1–1213) including a C-terminal thrombin cleavage site, T4 foldon trimerization domain, and hexahistidine tag were cloned into mammalian expression vector pCAGGS. The S sequences were modified to remove the polybasic cleavage site (RRAR to A), and two prefusion stabilizing proline mutations were introduced (K986P and V987P, wild type numbering). For expression of Avi-tagged variants, the CDS of pCAGGS vector containing the sequence for the relevant soluble S was modified to encode 3′ Avitag insert after the 6×His tag (5′-His tag-GGCTCCGGGCTGAACGACATCTTCGAAGCCCAGAAGATTGAGTGGCATGAG-Stop-3′; HHHHHHGSGLNDIFEAQKIEWHE-) using inverse PCR mutagenesis as previously described48. Recombinant proteins were produced in Expi293F cells (ThermoFisher) by transfection with purified DNA using the ExpiFectamine 293 Transfection Kit (ThermoFisher). Supernatants from transfected cells were collected 3 days post-transfection, and recombinant proteins were purified using Ni-NTA agarose (ThermoFisher), then buffer exchanged into PBS and concentrated using Amicon Ultracel centrifugal filters (EMD Millipore). To biotinylate Avi-tagged S variants, the S-Avitag substrates were diluted to 40 μM and incubated for 1 h at 30°C with 15 μg ml−1 BirA enzyme (Avidity) in 0.05 M bicine buffer at pH 8.3 supplemented with 10 mM ATP, 10 mM magnesium acetate and 50 μM biotin. The protein was then concentrated and buffer exchanged with PBS using a 100 kDa Amicon Ultra centrifugal filter (EMD Millipore).
To generate antigen probes for flow cytometry staining and sorting, trimeric BirA-biotinylated recombinant S from WA1/2020, B.1.351. B.1.617.2, or B.1.1.529 (BA.1) were incubated with a 1.04-fold molar excess of BV421-, BV650-, or PE-conjugated streptavidin (BioLegend), on ice, with three equal additions of S spaced every 15 min. Fifteen minutes after the third S addition, d-biotin was added in sixfold molar excess to streptavidin to block any unoccupied biotin binding sites. Streptavidin-PE-Cy5 and streptavidin-APC-Fire750 (BioLegend) were blocked with a 6-fold molar excess of d-biotin and used as background staining controls. Bovine serum albumin (BSA) was biotinylated using the EZ-Link Micro NHS-PEG4-Biotinylation Kit (Thermo Fisher); excess unreacted biotin was removed using 7-kDa Zeba desalting columns (Pierce).
ELISpot assay
Wells were coated with Flucelvax Quadrivalent 2019/2020 seasonal influenza virus vaccine (Sequiris), recombinant S from the WA1/2020, B.1.351, B.1.617.2, or BA.1 strains of SARS-CoV-2, or pooled anti-κ and anti-λ light chain antibodies (Cellular Technology Limited). Direct ex vivo ELISpot assays were performed with non-cryopreserved PBMCs to determine the number of total, influenza vaccine-binding, or recombinant S-binding IgG- and IgA-secreting cells present in PBMC and non-cryopreserved enriched BMPC samples using IgG/IgA double-colour ELISpot Kits (Cellular Technology Limited) according to the manufacturer’s instructions. ELISpot plates were analysed using an ELISpot counter (Cellular Technology Limited).
Fluorescent bead antigen binding assay
Recombinant biotinylated S from WA1/2020, B.1.351, B.1.617.2 and BA.1 strains of SARS-CoV-2 and biotinylated BSA were incubated for 30 min on ice with different fluorescence intensity peaks of the Streptavidin Coated Fluorescent Yellow Particle Kit (Spherotech) at 9.12 ng per μg beads. Beads were washed twice with 0.05% Tween 20 in PBS, resusupended in monoclonal antibodies diluted to 65 μg ml−1 or plasma samples diluted 1:80 in 0.05% Tween 20 in PBS, and incubated for 30 min on ice. Beads were washed twice with 0.05% Tween 20 in PBS, stained with IgG-APC-Fire750 (M1310G05, BioLegend, 1:100), incubated for 30 min on ice, washed twice with 0.05% Tween 20 in PBS, and resuspended in 2% FBS and 2 mM EDTA in PBS and acquired on an Aurora using SpectroFlo v2.2 (Cytek). Data were analysed using FlowJo v10 (Treestar). Background-subtracted median fluorescence intensities were calculated for each sample by subtracting its median fluorescence intensity plus two times robust standard deviation for BSA and the median fluorescence intensity of an influenza virus haemagglutinin-specific monoclonal antibody (for monoclonal antibody samples) or plasma collected prior to the SARS-CoV-2 pandemic (for plasma samples) for the respective spike variant (Extended Data Fig. 1b).
ELISA
Assays were performed in 96-well plates (MaxiSorp; Thermo) coated with 100 µl of recombinant S from WA1/2020, B.1.351, B.1.617.2 and BA.1 strains of SARS-CoV-2, NTD of WA1/2020 or BA.1, receptor binding domain of WA1/2020 or BA.1, or S2 domain of WA1/2020, or bovine serum albumin diluted to 1 μg ml−1 in PBS, and plates were incubated at 4 °C overnight. Plates then were blocked with 10% FBS and 0.05% Tween 20 in PBS. Purified monoclonal antibodies were serially diluted in blocking buffer and added to the plates. Plates were incubated for 90 min at room temperature and then washed 3 times with 0.05% Tween 20 in PBS. Goat anti-human IgG-HRP (goat polyclonal, Jackson ImmunoResearch, 1:2,500) was diluted in blocking buffer before adding to plates and incubating for 60 min at room temperature. Plates were washed 3 times with 0.05% Tween 20 in PBS and 3 times with PBS before the addition of O-phenylenediamine dihydrochloride peroxidase substrate (Sigma-Aldrich). Reactions were stopped by the addition of 1 M hydrochloric acid. Optical density measurements were taken at 490 nm.
VSV–SARS-CoV-2-SΔ21 eGFP-reduction assay
The S gene of SARS-CoV-2 isolate WA1/2020 (with D614G mutation) was synthesized and replaced the native envelope glycoprotein of an infectious molecular clone of VSV, and resulting chimeric viruses expressing S protein from SARS-CoV-2 D614G were used for GFP reduction neutralization tests as previously described32. In brief, 103 PFU of VSV–SARS-CoV-2-SΔ21 was incubated for 1 h at 37 °C with recombinant monoclonal antibodies diluted to 5 μg ml−1. Antibody–virus complexes were added to Vero E6 cells in 96-well plates and incubated at 37 °C for 7.5 h. Cells were subsequently fixed in 2% formaldehyde (Electron Microscopy Sciences) containing 10 μg ml−1 Hoechst 33342 nuclear stain (Invitrogen) for 45 min at room temperature, when fixative was replaced with PBS. Images were acquired with an InCell 2000 Analyzer (GE Healthcare) automated microscope using the DAPI and FITC channels to visualize nuclei and infected cells (that is, eGFP-positive cells), respectively (4× objective, 4 fields per well, covering the entire well). Images were analysed using the Multi Target Analysis Module of the InCell Analyzer 1000 Workstation Software (GE Healthcare). GFP-positive cells were identified in the FITC channel using the top-hat segmentation method and subsequently counted within the InCell Workstation software. The sensitivity and accuracy of GFP-positive cell number determinations were validated using a serial dilution of virus. The percent infection reduction was calculated from wells to which no antibody was added. A background number of GFP-positive cells was subtracted from each well using an average value determined from at least four uninfected wells.
Focus reduction neutralization test
Each monoclonal antibody was incubated at 5 μg ml−1 in DMEM supplemented with 2% FBS, 10 mM HEPES, and 100 U ml−1 penicillin/streptomycin with 102 focus-forming units of different of SARS-CoV-2 strains (WA1/2020 D614G, B.1.351, B.1.617.2, BA.1, and BA.5) for 1 h at 37 °C49. Antibody–virus complexes were added to Vero-TMPRSS2 cell monolayers in 96-well plates and incubated at 37 °C for 1 h. Subsequently, cells were overlaid with 1% (w/v) methylcellulose in MEM supplemented with 2% FBS. Plates were harvested 30 h (D614G, B.1.351, or B.1.617.2-infected) or 70 h (BA.1 or BA.5-infected) later by removing overlays and fixed with 4% PFA in PBS for 20 min at room temperature. Plates were washed and incubated with an oligoclonal pool of anti-S antibodies (SARS2-2, SARS2-11, SARS2-16, SARS2-31, SARS2-38, SARS2-57, and SARS2-71)33, and an additional oligoclonal pool of anti-S antibodies with extended reactivity (SARS2-08, SARS2-09, SARS2-10, SARS2-13, SARS2-14, SARS2-17, SARS2-20, SARS2-26, SARS2-27, SARS2-28, SARS2-31, SARS2-41, SARS2-42, SARS2-44, SARS2-49, SARS2-62, SARS2-64, SARS2-65 and SARS2-67)50 were included for staining BA.1- or BA.5-infected plates. Plates were subsequently incubated with HRP-conjugated goat anti-mouse IgG (Sigma 12–349) in PBS supplemented with 0.1% saponin and 0.1% bovine serum albumin. SARS-CoV-2-infected cell foci were visualized using TrueBlue peroxidase substrate (KPL) and quantitated on an ImmunoSpot microanalyzer (Cellular Technologies Limited).
Selection of monoclonal antibody escape mutants in SARS-CoV-2 S
We generated VSV–SARS-CoV-2-S (BA.1 variant) chimera by inserting the BA.1 spike gene into the VSV genome and using the resulting VSV–SARS-CoV-2 BA.1 chimeric virus to select for SARS-CoV-2 S variants that escape monoclonal antibody neutralization as described previously32,33. Antibody neutralization-resistant mutants were recovered by plaque isolation. In brief, plaque assays were performed to isolate the VSV–SARS-CoV-2 escape mutant on Vero cells with each tested monoclonal antibody in the overlay. The concentration of each monoclonal antibody in the overlay was determined by neutralization assays at a multiplicity of infection of 100. Escape clones were plaque-purified on Vero cells in the presence of monoclonal antibodies, and plaques in agarose plugs were amplified on MA104 cells with the monoclonal antibodies present in the medium. Viral supernatants were collected after three days upon extensive cytopathic effect and clarified of cell debris by centrifugation at 1,000g for 5 min. Aliquots were maintained at −80 °C. Viral RNA was extracted from VSV–SARS-CoV-2 mutant viruses using RNeasy Mini kit (Qiagen), and the S gene was amplified using OneStep RT–PCR Kit (Qiagen). To sequence the 3,747-bp BA.1 spike gene, four sets of PCR were set up to efficiently cover the whole BA.1 spike gene in the virus as follows: first set of fragment PCR utilizing sequencing primers 15M (ctggattctatcagccac) and 369 (caatagattctgttggttgg); second set of fragment PCR utilizing sequencing primers 368 (ctctctcagaaacaaagtgt) and 372 (cctgcacgtgtttgaaaaa); third set of fragment PCR utilizing sequencing primers 370 (aactgcacagaagtccct) and 374 (agaacattctgtgtaactcc); and fourth set of fragment PCR utilizing sequencing primers 373 (tacaatcacttctggttgga) and 23L (cattgaactcgtcggtctc) (Extended Data Fig. 4f). The mutations were identified by Sanger sequencing (Genewiz).
Cell sorting and flow cytometry
Staining for flow cytometry analysis and sorting was performed using freshly isolated or cryopreserved FNA or PBMC samples. For analysis, PBMC were incubated for 30 min on ice with purified CD16 (3G8, BioLegend, 1:100), CD32 (FUN-2, BioLegend, 1:100), CD64 (10.1, BioLegend, 1:100) and PD-1-BB515 (EH12.1, BD Horizon, 1:100) in 2% FBS and 2 mM EDTA in PBS (P2), washed twice, then were stained for 30 min on ice with WA1/2020, B.1.351, B.1.617.2 and BA.1 spike probes pre-conjugated to SA–BV650 and SA–PE, S167–180-PE-Cy7 tetramer, S816–830-APC tetramer51, biotin-saturated SA–PE-Cy5, ICOS-SB436 (ISA-3, Invitrogen, 1:50), IgG–BV480 (goat polyclonal, Jackson ImmunoResearch, 1:100), IgA–FITC (M24A, Millipore, 1:500), CD8a–A532 (RPA-T8, Thermo, 1:100), CD38–BB700 (HIT2, BD Horizon, 1:500), CD71-BV421 (CY1G4, 1:400), CD20–Pacific Blue (2H7, 1:400), CD4–Spark Violet 538 (SK3, 1:400), CD19–BV750 (HIB19, 1:100), IgD–BV785 (IA6-2, 1:200), CXCR5–PE-Dazzle 594 (J252D4, 1:50), CD14–PerCP (HCD14, 1:50), CD27–PE-Fire810 (O323, 1:200), CCR7-Spark 685 (G043H7, 1:100), IgM-A700 (MHM-88, 1:400), CD3–APC-Fire810 (SK7, 1:50), and Zombie NIR (all BioLegend) diluted in Brilliant Staining buffer (BD Horizon). For analysis, FNA samples were incubated for 30 min on ice with purified CD16 (3G8, BioLegend, 1:100), CD32 (FUN-2, BioLegend, 1:100), CD64 (10.1, BioLegend, 1:100) and PD-1-BB515 (EH12.1, BD Horizon, 1:100) in P2, washed twice, then stained for 30 min on ice with WA1/2020, B.1.351, B.1.617.2, and B.1.1.529 spike probes pre-conjugated to SA–BV421 and SA–BV650, S167–180–APC tetramer, biotin-saturated SA–PE–Cy5, IgG–BV480 (goat polyclonal, Jackson ImmunoResearch, 1:100), IgA–FITC (M24A, Millipore, 1:500), CD8a–A532 (RPA-T8, Thermo, 1:100), CD38–BB700 (HIT2, BD Horizon, 1:500), CD20–Pacific Blue (2H7, 1:400), CD4–Spark Violet 538 (SK3, 1:400), IgM–BV605 (MHM-88, 1:100), CD19–BV750 (HIB19, 1:100), IgD–BV785 (IA6-2, 1:200), CXCR5–PE–Dazzle 594 (J252D4, 1:50), CD14–PerCP (HCD14, 1:50), CD71–PE–Cy7 (CY1G4, 1:400), CD27–PE–Fire810 (O323, 1:200), CD3–APC–Fire810 (SK7, 1:50) and Zombie NIR (all BioLegend) diluted in Brilliant Staining buffer (BD Horizon). Cells were washed twice with P2, fixed for 1 h at 25 °C using the True Nuclear fixation kit (BioLegend), washed twice with True Nuclear Permeabilization/Wash buffer, stained with Ki-67-BV711 (Ki-67, BioLegend, 1:200), Blimp1–PE (646702, R&D, 1:100), FOXP3–Spark 685 (206D, BioLegend, 1:200), and Bcl6–R718 (K112-91, BD Horizon, 1:200) for 1 h at 25 °C, and washed twice with True Nuclear Permeabilization/Wash buffer. Samples were resuspended in P2 and acquired on an Aurora using SpectroFlo v2.2 (Cytek). Flow cytometry data were analysed using FlowJo v10 (Treestar).
For sorting plasmablasts, PBMCs collected 1 week post-boost were stained for 30 min on ice with CD20–Pacific Blue (2H7, 1:400), CD71–FITC (CY1G4, 1:200), IgD–PerCP–Cy5.5 (IA6-2, 1:200), CD19–PE (HIB19, 1:200), CXCR5–PE–Dazzle 594 (J252D4, 1:50), CD38–PE–Cy7 (HIT2, 1:200), CD4–A700 (SK3, 1:400) and Zombie Aqua (all BioLegend) diluted in P2. Cells were washed twice, and plasmablasts (live singlet CD4−CD19+IgDloCD20loCD38+CXCR5loCD71+) were sorted using a Bigfoot (Invitrogen) into PBS supplemented with 0.05% BSA and immediately processed for scRNA-seq. For bulk sorting germinal centre and LNPC, lymph node FNA samples collected 2 or 8 weeks post-boost were stained for 30 min on ice with purified CD16 (3G8, BioLegend, 1:100), CD32 (FUN-2, BioLegend, 1:100), CD64 (10.1, BioLegend, 1:100) and PD-1–BB515 (EH12.1, BD Horizon, 1:100) in P2, washed twice, then stained for 30 min on ice with CD20–Pacific Blue (2H7, 1:400), CD19–BV750 (HIB19, 1:100), IgD–PerCP–Cy5.5 (IA6-2, 1:200), CD71–PE (CY1G4, 1:400), CXCR5–PE–Dazzle 594 (J252D4, 1:50), CD38–PE–Cy7 (HIT2, 1:200), CD4–A700 (SK3, 1:400) and Zombie Aqua (all BioLegend) diluted in P2. Cells were washed twice, and total germinal centre B cells (live singlet CD4−CD19+IgDloCD20+CD38intCXCR5+CD71+) and LNPC (live singlet CD4−CD19+IgDloCD20loCD38+CXCR5loCD71+) were sorted using a Bigfoot (Invitrogen) into Buffer RLT Plus (Qiagen) supplemented with 143 mM β-mercaptoethanol (Sigma-Aldrich) and immediately frozen on dry ice. For sorting memory B cells, PBMC collected 17 weeks post-boost were incubated for 10 min on ice with purified CD16 (3G8, BioLegend, 1:100), CD32 (FUN-2, BioLegend, 1:100), and CD64 (10.1, BioLegend, 1:100) in P2. For sorting S+ memory B cells, WA1/2020, B.1.351, B.1.617.2 and BA.1 spike probes pre-conjugated to SA–BV650 and SA–PE, biotin-saturated SA–PE–Cy5, CD20–Pacific Blue (2H7, 1:400), CD19–BV605 (HIB19, 1:100), IgD–BV785 (IA6-2, 1:200), CD3–FITC (HIT3a, 1:200), CD27–A700 (M-T271, 1:200) and Zombie NIR (all BioLegend) diluted in Brilliant Staining buffer (BD Horizon) were added and stained for an additional 30 min on ice. For sorting BA.1+ WA1/2020− memory B cells, WA1/2020 probes pre-conjugated to SA–BV650 and SA–APC, BA.1 probes pre-conjugated to SA–BV421 and SA–PE, biotin-saturated SA–PE–Cy5, CD20–Pacific Blue (2H7, 1:400), IgD–BV785 (IA6-2, 1:200), CD19–FITC (HIB19, 1:100), CD27–PE–Fire810 (O323, 1:200), CD3–A700 (HIT3a, 1:100) and Zombie NIR (all BioLegend) diluted in Brilliant Staining buffer (BD Horizon) were added and stained for an additional 30 min on ice. Cells were washed twice, and pooled S-binding single memory B cells (live singlet CD3−CD19+IgDloSA–PE–Cy5− pooled spikes double positive) or BA.1+ WA1/2020− single memory B cells (live singlet CD3−CD19+IgDloSA–PE-Cy5− BA.1+ WA1/2020−) were sorted using a Bigfoot (Invitrogen) into 96-well plates containing 2 µl Lysis Buffer (Clontech) supplemented with 1 U μl−1 RNase inhibitor (NEB), or total IgDlo memory B cells were bulk sorted into Buffer RLT Plus (Qiagen) supplemented with 143 mM β-mercaptoethanol (Sigma-Aldrich) and immediately frozen on dry ice.
Monoclonal antibody generation
Antibodies were cloned as described previously52. In brief, VH, Vκ and Vλ genes were amplified by PCR with reverse transcription and nested PCR reactions from singly-sorted S+ memory B cells using primer combinations specific for IgG, IgM/A, Igκ and Igλ from previously described primer sets53 and then sequenced. To generate recombinant antibodies, restriction sites were incorporated via PCR with primers to the corresponding heavy and light chain V and J genes. The amplified VH, Vκ and Vλ genes were cloned into IgG1 and Igκ or Igλ expression vectors, respectively, as described previously53,54,55. Heavy and light chain plasmids were co-transfected into Expi293F cells (Gibco) for expression, and antibody was purified using protein A agarose chromatography (Goldbio). Sequences were obtained from PCR reaction products and annotated using the ImMunoGeneTics (IMGT)/V-QUEST database (http://www.imgt.org/IMGT_vquest/)56,57. Mutation frequency was calculated by counting the number of nonsynonymous nucleotide mismatches from the germline sequence in the heavy chain variable segment leading up to the CDR3, while excluding the 5′ primer sequences that could be error-prone.
scRNA-seq library preparation and sequencing
Sorted plasmablast and lymph node FNA samples were processed using the following 10x Genomics kits: Chromium Next GEM Single Cell 5′ Kit v2 (PN-1000263); Chromium Next GEM Chip K Single Cell Kit (PN-1000286); BCR Amplification Kit (PN-1000253); Dual Index Kit TT Set A (PN-1000215). Chromium Single Cell 5′ Gene Expression Dual Index libraries and Chromium Single Cell V(D)J Dual Index libraries were prepared according to manufacturer’s instructions. Both gene expression and V(D)J libraries were sequenced on a Novaseq S4 (Illumina), targeting a median sequencing depth of 50,000 and 5,000 read pairs per cell, respectively.
Bulk BCR library preparation and sequencing
RNA was purified from sorted plasmablasts and memory B cells from PBMC, germinal centre B cells and plasma cells from lymph node FNA (LNPC), and CD138-enriched BMPC using the RNeasy Plus Micro kit (Qiagen). Reverse transcription, unique molecular identifier (UMI) barcoding, cDNA amplification, and Illumina linker addition to B cell heavy chain transcripts were performed using the human NEBNext Immune Sequencing Kit (New England Biolabs) according to the manufacturer’s instructions. High-throughput 2× 300 bp paired-end sequencing was performed on the Illumina MiSeq platform with a 30% PhiX spike-in according to manufacturer’s recommendations, except for performing 325 cycles for read 1 and 275 cycles for read 2.
Preprocessing of bulk sequencing BCR reads
Preprocessing of demultiplexed pair-end reads was performed using pRESTO v.0.6.258 as previously described2, with the exception that sequencing errors were corrected using the UMIs as they were without additional clustering (Extended Data Table 3). Previously preprocessed unique consensus sequences from samples corresponding to participants in the current study and previously reported in ref. 29 were included without additional processing. Participants 382-01, 382-02, 382-07, 382-08, 382-13, and 382-15 correspond to previously reported 368-22, 368-20, 368-02a, 368-04, 368-01a and 368-10, respectively2,29. Previously preprocessed unique consensus sequences from samples corresponding to participants in the current study and reported in2 were subset to those with at least two contributing reads and included.
Preprocessing of 10x Genomics single-cell BCR reads
Demultiplexed pair-end FASTQ reads were preprocessed using Cell Ranger v.6.0.1 as previously described29 (Extended Data Table 4). Previously preprocessed single-cell BCR reads from samples corresponding to participants in the current study and reported in ref. 29 were included.
V(D)J gene annotation and genotyping
Initial germline V(D)J gene annotation was performed on the preprocessed BCRs using IgBLAST v.1.17.159 with the deduplicated version of IMGT/V-QUEST reference directory release 202113-256. Further sequence-level and cell-level quality controls were performed as previously described29. Exceptions for monoclonal antibody sequences triggering QC filters were handled on a case-by-case basis upon inspection as follows. Indels detected in 382-01 1B04 heavy chain and 1F01 light chain and 382-53 2G07 heavy chain were accepted. The CDR3 annotations from IMGT/V-QUEST for 382-08 1G04 heavy chain, 382-54 1D09 heavy chain and 382-55 4H10 light chain were used in lieu of those from IgBLAST as the former had nucleotide lengths that were a multiple of 3 whereas the latter did not. Individualized genotypes were inferred based on sequences that passed all quality controls using TIgGER v.1.0.060 and used to finalize V(D)J annotations. Sequences annotated as non-productively rearranged by IgBLAST were removed from further analysis.
Clonal lineage inference
B cell clonal lineages were inferred on a by-individual basis based on productively rearranged sequences as previously described29. In brief, heavy chain-based clonal inference31 was performed by partitioning the heavy chains of bulk and single-cell BCRs based on common V and J gene annotations and CDR3 lengths, and clustering the sequences within each partition hierarchically with single linkage based on their CDR3s30. Sequences whose CDR3 nucleotides were within 0.15 normalized Hamming distance from each other were clustered as clones. Following clonal inference, full-length clonal consensus germline sequences were reconstructed using Change-O v.1.0.261. Within each clone, duplicate IMGT-aligned V(D)J sequences from bulk sequencing were collapsed using Alakazam v1.1.061 except for duplicates derived from different lymph nodes, time points, tissues, B cell compartments, isotypes, or biological replicates.
BCR analysis
B cell compartment labels were treated as previously described29. In brief, gene expression-based cluster annotation was used for single-cell BCRs; FACS-based sorting and magnetic enrichment were used for bulk BCRs, except that plasmablast sorts from lymph node FNA were labelled LNPCs; post-second dose week 2 IgDlo enriched B cells from blood were labelled activated; and post-second dose week 4 and post-third dose (booster) week 17 IgDlo enriched B cells from blood were labelled memory. For analysis involving the memory compartment, the memory sequences were restricted to those from blood. A heavy chain-based B cell clone was considered S-specific if it contained any sequence corresponding to a S-binding monoclonal antibody that was previously reported2,29 or from the current study. Clonal overlap between B cell compartments was visualized using circlize v.0.4.1362. SHM frequency was calculated for each heavy chain sequence using SHazaM v.1.0.261 as previously described29 by counting the number of nucleotide mismatches from the germline sequence in the variable segment leading up to the CDR3, while excluding the first 18 positions that could be error-prone due to the primers used for generating the monoclonal antibody sequences.
Processing of 10x Genomics single-cell 5′ gene expression data
Demultiplexed pair-end FASTQ reads were first preprocessed on a by-sample basis and samples were subsequently subsampled to the same effective sequencing length and aggregated using Cell Ranger v.6.0.1 as previously described29. Quality control was performed on the aggregate gene expression matrix consisting of 336,960 cells and 36,601 features using SCANPY v.1.7.263. In brief, to remove presumably lysed cells, cells with mitochondrial content greater than 17.5% of all transcripts were removed. To remove likely doublets, cells with more than 8,000 features or 80,000 total UMIs were removed. To remove cells with no detectable expression of common endogenous genes, cells with no transcript for any of a list of 34 housekeeping genes29 were removed. The feature matrix was subset, based on their biotypes, to protein-coding, immunoglobulin, and T cell receptor genes that were expressed in at least 0.05% of the cells in any sample. The resultant feature matrix contained 15,751 genes. Finally, cells with detectable expression of fewer than 200 genes were removed. After quality control, there were a total of 312,242 cells from 39 single-cell samples (Extended Data Table 4).
Single-cell gene expression analysis
Transcriptomic data was analysed using SCANPY v.1.7.263 as previously described29 with minor adjustments suitable for the current data. In brief, overall clusters were first identified using Leiden graph-clustering with resolution 0.50 (Extended Data Fig. 3b, Extended Data Table 5). UMAPs were faceted by participant and inspected for convergence to assess whether there was a need for integration (Extended Data Fig. 3c). Cluster identities were assigned by examining the expression of a set of marker genes34 for different cell types (Extended Data Fig. 3d). To remove potential contamination by platelets, 205 cells with a log-normalized expression value of >2.5 for PPBP were removed. From a cluster consisting primarily of monocytes, 36 cells originating from lymph node FNA and with a log-normalized expression value of >0 for at least two of FDCSP, CXCL1464, and FCAMR65 were annotated FDCs. Cells from the overall B cell cluster were further clustered to identify B cell subsets using Leiden graph-clustering resolution 0.35 (Extended Data Fig. 3e and Extended Data Table 5). Cluster identities were assigned by examining the expression of a set of marker genes34 for different B cell subsets (Extended Data Fig. 3f) along with the availability of BCRs. Despite being clustered with B cells during overall clustering, one group tended to have both BCRs and relatively high expression levels of CD2 and CD3E; one group tended to have no BCRs and relatively high CD2 and CD3E; and two unassigned groups tended to have no BCRs. These were excluded from the final B cell clustering. Ten cells that were found in the germinal centre B cell cluster but came from blood were labelled ‘PB-like’34. Four-hundred and seven cells that were found in the PB cluster but came from lymph node FNA were re-assigned as LNPCs. One cell that was found in the LNPC cluster but came from blood was re-assigned as PB. Heavy chain SHM frequency and isotype usage of the B cell subsets were inspected for consistency with expected values to further confirm their assigned identities.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Antibody sequences have been deposited in GenBank under accession numbers OQ634308–OQ635939. Newly reported raw sequencing data and the transcriptomics count matrix are deposited at the Sequence Read Archive and Gene Expression Omnibus under BioProject PRJNA800176. Processed transcriptomics and BCR data are deposited at Zenodo (https://doi.org/10.5281/zenodo.7719030). Previously reported BCR data used in this study were deposited under BioProjects PRJNA731610, PRJNA741267 and PRJNA777934. The IMGT/V-QUEST database is accessible at http://www.imgt.org/IMGT_vquest/. Materials are available upon request through an interinstitutional materials transfer agreement.
References
Krause, P. R. et al. SARS-CoV-2 variants and vaccines. N. Engl. J. Med. 385, 179–186 (2021).
Turner, J. S. et al. SARS-CoV-2 mRNA vaccines induce persistent human germinal centre responses. Nature 596, 109–113 (2021).
Laidlaw, B. J. & Ellebedy, A. H. The germinal centre B cell response to SARS-CoV-2. Nat. Rev. Immunol. 22, 7–18 (2022).
Amanat, F. et al. SARS-CoV-2 mRNA vaccination induces functionally diverse antibodies to NTD, RBD, and S2. Cell 184, 3936–3948.e10 (2021).
Muecksch, F. et al. Increased memory B cell potency and breadth after a SARS-CoV-2 mRNA boost. Nature 607, 128–134 (2022).
Goel, R. R. et al. Efficient recall of Omicron-reactive B cell memory after a third dose of SARS-CoV-2 mRNA vaccine. Cell 185, 1875–1887.e8 (2022).
Rodda, L. B. et al. Imprinted SARS-CoV-2-specific memory lymphocytes define hybrid immunity. Cell 185, 1588–1601.e14 (2022).
Pérez-Then, E. et al. Neutralizing antibodies against the SARS-CoV-2 Delta and Omicron variants following heterologous CoronaVac plus BNT162b2 booster vaccination. Nat. Med. 28, 481–485 (2022).
Sette, A. & Crotty, S. Immunological memory to SARS‐CoV‐2 infection and COVID‐19 vaccines. Immunol. Rev. 310, 27–46 (2022).
Lucas, C. et al. Impact of circulating SARS-CoV-2 variants on mRNA vaccine-induced immunity. Nature 600, 523–529 (2021).
Wang, Z. et al. mRNA vaccine-elicited antibodies to SARS-CoV-2 and circulating variants. Nature 592, 616–622 (2021).
Andrews, N. et al. Covid-19 vaccine effectiveness against the Omicron (B.1.1.529) variant. N. Engl. J. Med. 386, 1532–1546 (2022).
Kuhlmann, C. et al. Breakthrough infections with SARS-CoV-2 omicron despite mRNA vaccine booster dose. Lancet 399, 625–626 (2022).
Schmidt, F. et al. Plasma neutralization of the SARS-CoV-2 omicron variant. N. Engl. J. Med. 386, 599–601 (2022).
Cameroni, E. et al. Broadly neutralizing antibodies overcome SARS-CoV-2 Omicron antigenic shift. Nature 602, 664–670 (2022).
Cele, S. et al. Omicron extensively but incompletely escapes Pfizer BNT162b2 neutralization. Nature 602, 654–656 (2022).
Falsey, A. R. et al. SARS-CoV-2 Neutralization with BNT162b2 vaccine dose 3. N. Engl. J. Med. 385, 1627–1629 (2021).
Bowen, J. E. et al. Omicron spike function and neutralizing activity elicited by a comprehensive panel of vaccines. Science 377, 890–894 (2022).
Muik, A. et al. Neutralization of SARS-CoV-2 Omicron by BNT162b2 mRNA vaccine–elicited human sera. Science 375, 678–680 (2022).
Moreira, E. D. et al. Safety and efficacy of a third dose of BNT162b2 Covid-19 vaccine. N. Engl. J. Med. 386, 1910–1921 (2022).
Barda, N. et al. Effectiveness of a third dose of the BNT162b2 mRNA COVID-19 vaccine for preventing severe outcomes in Israel: an observational study. Lancet 398, 2093–2100 (2021).
Chalkias, S. et al. Safety, immunogenicity and antibody persistence of a bivalent Beta-containing booster vaccine against COVID-19: a phase 2/3 trial. Nat. Med. 28, 2388–2397 (2022).
Scheaffer, S. M. et al. Bivalent SARS-CoV-2 mRNA vaccines increase breadth of neutralization and protect against the BA.5 Omicron variant in mice. Nat. Med. 29, 247–257 (2023).
Jackson, L. A. et al. An mRNA vaccine against SARS-CoV-2—preliminary report. N. Engl. J. Med. 383, 1920–1931 (2020).
Walsh, E. E. et al. Safety and immunogenicity of two RNA-based Covid-19 vaccine candidates. N. Engl. J. Med. 383, 2439–2450 (2020).
Purtha, W. E., Tedder, T. F., Johnson, S., Bhattacharya, D. & Diamond, M. S. Memory B cells, but not long-lived plasma cells, possess antigen specificities for viral escape mutants. J. Exp. Med. 208, 2599–2606 (2011).
Lederer, K. et al. Germinal center responses to SARS-CoV-2 mRNA vaccines in healthy and immunocompromised individuals. Cell 185, 1008–1024.e15 (2022).
Röltgen, K. et al. Immune imprinting, breadth of variant recognition, and germinal center response in human SARS-CoV-2 infection and vaccination. Cell 185, 1025–1040.e14 (2022).
Kim, W. et al. Germinal centre-driven maturation of B cell response to mRNA vaccination. Nature 604, 141–145 (2022).
Gupta, N. T. et al. Hierarchical clustering can identify B cell clones with high confidence in Ig repertoire sequencing data. J. Immunol. 198, 2489–2499 (2017).
Zhou, J. Q. & Kleinstein, S. H. Cutting edge: Ig H chains are sufficient to determine most B cell clonal relationships. J. Immunol. 203, 1687–1692 (2019).
Case, J. B. et al. Neutralizing antibody and soluble ACE2 inhibition of a replication-competent VSV-SARS-CoV-2 and a clinical isolate of SARS-CoV-2. Cell Host Microbe 28, 475–485.e5 (2020).
Liu, Z. et al. Identification of SARS-CoV-2 spike mutations that attenuate monoclonal and serum antibody neutralization. Cell Host Microbe 29, 477–488.e4 (2021).
Turner, J. S. et al. Human germinal centres engage memory and naive B cells after influenza vaccination. Nature 586, 127–132 (2020).
Turner, J. S. et al. SARS-CoV-2 infection induces long-lived bone marrow plasma cells in humans. Nature 595, 421–425 (2021).
Radbruch, A. et al. Competence and competition: the challenge of becoming a long-lived plasma cell. Nat. Rev. Immunol. 6, 741–750 (2006).
Weisel, F. J., Zuccarino-Catania, G. V., Chikina, M. & Shlomchik, M. J. A temporal switch in the germinal center determines differential output of memory B and plasma cells. Immunity 44, 116–130 (2016).
Ellebedy, A. H. et al. Adjuvanted H5N1 influenza vaccine enhances both cross-reactive memory B cell and strain-specific naive B cell responses in humans. Proc. Natl Acad. Sci. USA 117, 17957–17964 (2020).
Zost, S. J. et al. Potently neutralizing and protective human antibodies against SARS-CoV-2. Nature 584, 443–449 (2020).
Chen, P. et al. SARS-CoV-2 neutralizing antibody LY-CoV555 in outpatients with COVID-19. N. Engl. J. Med. 384, 229–237 (2021).
Hansen, J. et al. Studies in humanized mice and convalescent humans yield a SARS-CoV-2 antibody cocktail. Science 369, 1010–1014 (2020).
Alsoussi, W. B. et al. A potently neutralizing antibody protects mice against SARS-CoV-2 infection. J. Immunol. 205, 915–922 (2020).
Chen, R. E. et al. In vivo monoclonal antibody efficacy against SARS-CoV-2 variant strains. Nature 596, 103–108 (2021).
Francis, T. On the doctrine of original antigenic sin. Proc. Am. Philos. Soc. 104, 572–578 (1953).
Gostic, K. M., Ambrose, M., Worobey, M. & Lloyd-Smith, J. O. Potent protection against H5N1 and H7N9 influenza via childhood hemagglutinin imprinting. Science 354, 722–726 (2016).
Zang, R. et al. TMPRSS2 and TMPRSS4 promote SARS-CoV-2 infection of human small intestinal enterocytes. Sci. Immunol. 5, eabc3582 (2020).
Stadlbauer, D. et al. SARS-CoV-2 seroconversion in humans: a detailed protocol for a serological assay, antigen production, and test setup. Curr. Protoc. Microbiol. 57, e100 (2020).
Fairhead, M. & Howarth, M. Site-specific biotinylation of purified proteins using BirA. Methods Mol. Biol. 1266, 171–184 (2015).
Chen, R. E. et al. Resistance of SARS-CoV-2 variants to neutralization by monoclonal and serum-derived polyclonal antibodies. Nat. Med. 27, 717–726 (2021).
VanBlargan, L. A. et al. A potently neutralizing SARS-CoV-2 antibody inhibits variants of concern by utilizing unique binding residues in a highly conserved epitope. Immunity 54, 2399–2416.e6 (2021).
Mudd, P. A. et al. SARS-CoV-2 mRNA vaccination elicits a robust and persistent T follicular helper cell response in humans. Cell 185, 603–613.e15 (2022).
Wrammert, J. et al. Broadly cross-reactive antibodies dominate the human B cell response against 2009 pandemic H1N1 influenza virus infection. J. Exp. Med. 208, 181–193 (2011).
Smith, K. et al. Rapid generation of fully human monoclonal antibodies specific to a vaccinating antigen. Nat. Protoc. 4, 372–384 (2009).
Wrammert, J. et al. Rapid cloning of high-affinity human monoclonal antibodies against influenza virus. Nature 453, 667–671 (2008).
Nachbagauer, R. et al. Broadly reactive human monoclonal antibodies elicited following pandemic H1N1 influenza virus exposure protect mice against highly pathogenic H5N1 challenge. J. Virol. 92, 1–17 (2018).
Brochet, X., Lefranc, M.-P. & Giudicelli, V. IMGT/V-QUEST: the highly customized and integrated system for IG and TR standardized V-J and V-D-J sequence analysis. Nucleic Acids Res. 36, W503–W508 (2008).
Giudicelli, V., Brochet, X. & Lefranc, M.-P. IMGT/V-QUEST: IMGT standardized analysis of the immunoglobulin (IG) and T cell receptor (TR) nucleotide sequences. Cold Spring Harb. Protoc. 2011, pdb.prot5633 (2011).
Vander Heiden, J. A. et al. pRESTO: a toolkit for processing high-throughput sequencing raw reads of lymphocyte receptor repertoires. Bioinformatics 30, 1930–1932 (2014).
Ye, J., Ma, N., Madden, T. L. & Ostell, J. M. IgBLAST: an immunoglobulin variable domain sequence analysis tool. Nucleic Acids Res. 41, W34–W40 (2013).
Gadala-Maria, D., Yaari, G., Uduman, M. & Kleinstein, S. H. Automated analysis of high-throughput B-cell sequencing data reveals a high frequency of novel immunoglobulin V gene segment alleles. Proc. Natl Acad. Sci. USA 112, E862–E870 (2015).
Gupta, N. T. et al. Change-O: a toolkit for analyzing large-scale B cell immunoglobulin repertoire sequencing data. Bioinformatics 31, 3356–3358 (2015).
Gu, Z., Gu, L., Eils, R., Schlesner, M. & Brors, B. circlize implements and enhances circular visualization in R. Bioinformatics 30, 2811–2812 (2014).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Haebe, S. et al. Single-cell analysis can define distinct evolution of tumor sites in follicular lymphoma. Blood 137, 2869–2880 (2021).
Mourcin, F. et al. Follicular lymphoma triggers phenotypic and functional remodeling of the human lymphoid stromal cell landscape. Immunity 54, 1788–1806.e7 (2021).
Acknowledgements
The authors thank all the donors for generously providing precious specimens; L. Kessels and the Washington University School of Medicine 382 Study Team (study coordinators A. Haile, R. Thompson, D. Carani, K. Gray and C. Ayres; pharmacists M. Royal and J. Tran; and laboratory technicians L. Blair, A. Afghanzada and N. Schodl) for assistance with scheduling participants and sample collection; P. Woodard, B. Thomas, M. Harrod, R. Hamlin, M. Rohn; the staff of the Center for Clinical Research Imaging at Washington University School of Medicine for assistance with sample collection; and C. Dalton and B. Roemmich for performing the nucleocapsid binding assay. The WU382 study was reviewed and approved by the Washington University Institutional Review Board (approval no. 202109021). This work was supported in part with funding from the US National Institutes of Health (NIH) and Moderna. The Ellebedy laboratory was supported by NIH grants U01AI141990, 1U01AI150747, 5U01AI144616-02 and R01AI168178-01. The Diamond laboratory was supported by NIH grant R01 AI157155. The Whelan laboratory was supported by NIH grant R01 AI163019. The content of this manuscript is solely the responsibility of the authors and does not necessarily represent the official view of NIAID or NIH.
Author information
Authors and Affiliations
Contributions
A.H.E., J.A.O., R.M.P., R.P., B.N. and S.C. conceived and designed the study. J.A.O., M.K.K. and R.M.P. wrote and maintained the IRB protocol, recruited participants and coordinated sample collection. W.B.A., W.K., F.H. and J.S.T. processed specimens. W.B.A., S.K.M., S.F.M. and J.S.T. performed multiplex bead array and ELISA. W.B.A., F.H. and J.S.T. performed ELISpot. W.B.A., S.K.M. and S.M. performed VSV neutralization assays. W.B.A., S.K.M., W.K., A. J. Schmitz, T.L., S.C.H., A. J. Sturtz, K.M.M., B.E., I.F.F. and J.S.T. generated and characterized monoclonal antibodies. J.Q.Z. analysed scRNA-seq and BCR repertoire data. Z.L. rescued and produced the chimeric vesicular stomatitis viruses for neutralization assays and performed and analysed epitope mapping. B.Y. performed the SARS-CoV-2 virus neutralization assays. W.K. and A. J. Sturtz prepared libraries for scRNA-seq. A. J. Schmitz performed RNA extractions and library preparation for BCR bulk sequencing and expressed SARS-CoV-2 S and variant proteins. S.M.S. generated the authentic SARS-CoV-2 virus stocks. C.W.F. supervised the nucleocapsid binding assay. I.P. supervised bone marrow specimen collection. B.S.S. and W.D.M. performed FNA. B.S.S., W.D.M. and S.A.T. supervised lymph node evaluation prior to FNA and specimen collection and evaluated lymph node ultrasound data. J.S.T. sorted cells and collected and analysed the flow cytometry data. L.P.-R., R.P., R.N., J.S.T. and A.H.E. analysed data. A.H.E., M.S.D. and S.P.J.W. supervised experiments and obtained funding. J.S.T. and A.H.E. composed the manuscript. All authors reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The Ellebedy laboratory and Infectious Disease Clinical Research Unit has received funding under sponsored research agreements from Moderna related to the data presented in the current study. The Ellebedy laboratory received funding from Emergent BioSolutions and AbbVie that are unrelated to the data presented in the current study. A.H.E. has received consulting and speaking fees from InBios International, Fimbrion Therapeutics, RGAX, Mubadala Investment Company, Moderna, Pfizer, GSK, Danaher, Third Rock Ventures, Goldman Sachs and Morgan Stanley and is the founder of ImmuneBio Consulting. W.B.A., A. J. Schmitz, S.P.J.W., M.S.D., J.S.T. and A.H.E. are recipients of a licensing agreement with Abbvie that is unrelated to the data presented in the current study. M.S.D. is a consultant for Inbios, Vir Biotechnology, Senda Biosciences, Moderna, Sterne-Kessler and Immunome. The Diamond laboratory has received unrelated funding support in sponsored research agreements from Vir Biotechnology, Emergent BioSolutions and Moderna. S.P.J.W. is a consultant for Thylacine Bio. S.P.J.W. is a recipient of a licensing agreement with Vir Biotechnology and Merck. The Whelan laboratory has received unrelated funding support in sponsored research agreements from Vir Biotechnology. R.P., B.N., S.C. and R.N. are employees of and shareholders in Moderna. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Sophie Valkenburg and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Robust GC and Tfh responses to mRNA-1273 and mRNA-1273.213 boosters.
(a) Frequencies of IgA plasmablasts specific for the indicated antigens measured by ELISpot in week-1 PBMC from participants who received mRNA-1273 (left) and mRNA-1273.213 (right). (b) Flow cytometry plots of influenza B/HA-specific mAb 321-05 6F0534, cross-reactive mAb 382-08 1C11, and residue E484-specific mAb mu 2B0442 binding to indicated strains of SARS-CoV-2 S by multiplex bead binding array. (c) Plasma IgG binding to indicated strains of SARS-CoV-2 S measured by multiplex bead binding array in participants who received mRNA-1273 (left) and mRNA-1273.213 (right). (d) Gating strategy for analyzing S+ germinal centre B cells and T follicular helper cells (TFH) in FNA. (e) Frequencies of TFH from FNA of draining lymph nodes. (f) Correlation between frequencies of S+ germinal centre B cells and TFH. P and r values from two-sided Spearman test. (g) Representative ELISpot wells coated with BSA, and developed in blue (IgG) and red (IgA) after plating the indicated numbers of BMPCs. (h) Frequencies of IgA-secreting BMPCs specific for the indicated antigens 26 weeks post-boost. Black lines indicate medians. Symbols at each time point represent one sample. For mRNA-1273 and mRNA-1273.213 respectively, n = 7 and 38 (a), n = 6 and 28 (c), n = 5 and 20 (e), n = 3 and 10 (h).
Extended Data Fig. 2 Breadth of MBC-derived mAbs after mRNA-1273 and mRNA-1273.213 boosters.
(a) Gating strategy for sorting S+ MBC from PBMC.
Extended Data Fig. 3 Maturation of S-specific MBCs in response to mRNA-1273 or mRNA-1273.213 booster.
(a) Gating strategy for sorting plasmablasts from PBMC. (b,c,e,g) UMAPs showing scRNA-seq transcriptional clusters of total cells (b,c) or B cells (e,g) from all participants (b,e) or from each participant separately (c,g). (d,f) Dot plots for the marker genes used for identifying the annotated clusters in (b,c) (d) and in (e,g) (f). (h) SARS-CoV-2 S-specific clones visualized in red on UMAPs of B cells from each participant separately and faceted by time point. (i) Clonal overlap between S-binding plasmablasts and germinal centre B cells at indicated time points. Arc length corresponds to the number of BCR sequences and chord width corresponds to clone size. Purple chords correspond to clones spanning both compartments. Percentages are of germinal centre B cell clones related to plasmablasts at each time point. (j) Percentages of S-specific germinal centre clones related to week 1 plasmablasts. Symbols at each time point represent one sample, n = 3 for mRNA-1273 and n = 3 for mRNA-1273.213.
Extended Data Fig. 4 Characterization of BA.1-specific mAbs.
(a) Study design for mRNA-1273.529 vaccine cohort. Three and five healthy adult volunteers were enrolled and received an mRNA-1273.529 booster as a third or fourth dose, respectively. Blood was collected at baseline and at 1 and 17 weeks post-boost. (b) Frequencies of S-binding IgA-producing plasmablasts in PBMC 1 week post-boost measured by ELISpot in participants who received mRNA-1273.529. Black lines indicate medians. Symbols represent one sample; n = 7. (c) Neutralizing activity of mAbs from week 17 S+ MBCs against chimeric vesicular stomatitis virus in which the native envelope glycoprotein was replaced with S from WA1/2020 (with D614G mutation). Each symbol represents one mAb, n = 205 for mRNA-1273, n = 222 for mRNA-1273.213, and n = 303 for mRNA-1273.529. Percentages are of mAbs above the 80% infection reduction threshold. (d) Binding of mAbs to constituent domains of BA.1 S by ELISA performed in duplicate, presented as mean OD490 values. S2-specific mAb 368-22 1B08 was described previously2. (e) Titration of mAbs to determine neutralizing concentrations against chimeric vesicular stomatitis virus in which the native envelope glycoprotein was replaced with S from SARS-CoV-2 BA.1. Data are representative of two independent experiments. (f) Plasmid map for chimeric vesicular stomatitis virus in which the native envelope glycoprotein was replaced with S from BA.1 with primers indicated in purple.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Alsoussi, W.B., Malladi, S.K., Zhou, J.Q. et al. SARS-CoV-2 Omicron boosting induces de novo B cell response in humans. Nature 617, 592–598 (2023). https://doi.org/10.1038/s41586-023-06025-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06025-4
This article is cited by
-
Activation-based repertoire analysis for T cell clonal dynamics in hybrid COVID-19 immunity
Nature Immunology (2024)
-
Repeated Omicron exposures override ancestral SARS-CoV-2 immune imprinting
Nature (2024)
-
Immune imprinting and next-generation coronavirus vaccines
Nature Microbiology (2023)
-
Three-month antibody persistence of a bivalent Omicron-containing booster vaccine against COVID-19
Nature Communications (2023)
-
Evolution of antibody immunity following Omicron BA.1 breakthrough infection
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.