Abstract
Nucleic acids derived from pathogens induce potent innate immune responses1,2,3,4,5,6. Cyclic GMP–AMP synthase (cGAS) is a double-stranded DNA sensor that catalyses the synthesis of the cyclic dinucleotide cyclic GMP–AMP, which mediates the induction of type I interferons through the STING–TBK1–IRF3 signalling axis7,8,9,10,11. cGAS was previously thought to not react with self DNA owing to its cytosolic localization2,12,13; however, recent studies have shown that cGAS is localized mostly in the nucleus and has low activity as a result of tight nuclear tethering14,15,16,17,18. Here we show that cGAS binds to nucleosomes with nanomolar affinity and that nucleosome binding potently inhibits its catalytic activity. To elucidate the molecular basis of cGAS inactivation by nuclear tethering, we determined the structure of mouse cGAS bound to human nucleosome by cryo-electron microscopy. The structure shows that cGAS binds to a negatively charged acidic patch formed by histones H2A and H2B via its second DNA-binding site19. High-affinity nucleosome binding blocks double-stranded DNA binding and maintains cGAS in an inactive conformation. Mutations of cGAS that disrupt nucleosome binding alter cGAS-mediated signalling in cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The three-dimensional cryo-EM density maps are deposited into the Electron Microscopy Data Bank (EMDB) under accession numbers EMD-22046, EMD-22206 and EMD-22047. The coordinates were deposited in the Protein Data Bank (PDB) with accession numbers 6X59, 6XJD and 6X5A. Source data are provided with this paper.
References
Wu, J. & Chen, Z. J. Innate immune sensing and signaling of cytosolic nucleic acids. Annu. Rev. Immunol. 32, 461–488 (2014).
Hopfner, K. P. & Hornung, V. Molecular mechanisms and cellular functions of cGAS–STING signalling. Nat. Rev. Mol. Cell Biol. 21, 501–521 (2020).
Roers, A., Hiller, B. & Hornung, V. Recognition of endogenous nucleic acids by the innate immune system. Immunity 44, 739–754 (2016).
Kato, H., Takahasi, K. & Fujita, T. RIG-I-like receptors: cytoplasmic sensors for non-self RNA. Immunol. Rev. 243, 91–98 (2011).
Paludan, S. R. & Bowie, A. G. Immune sensing of DNA. Immunity 38, 870–880 (2013).
Ablasser, A. & Chen, Z. J. cGAS in action: Expanding roles in immunity and inflammation. Science 363, eaat8657 (2019).
Burdette, D. L. & Vance, R. E. STING and the innate immune response to nucleic acids in the cytosol. Nat. Immunol. 14, 19–26 (2013).
Barber, G. N. Innate immune DNA sensing pathways: STING, AIMII and the regulation of interferon production and inflammatory responses. Curr. Opin. Immunol. 23, 10–20 (2011).
Ablasser, A. et al. cGAS produces a 2′-5′-linked cyclic dinucleotide second messenger that activates STING. Nature 498, 380–384 (2013).
Gao, P. et al. Cyclic [G(2′,5′)pA(3′,5′)p] is the metazoan second messenger produced by DNA-activated cyclic GMP-AMP synthase. Cell 153, 1094–1107 (2013).
Zhao, B. et al. A conserved PLPLRT/SD motif of STING mediates the recruitment and activation of TBK1. Nature 569, 718–722 (2019).
Stetson, D. B. & Medzhitov, R. Recognition of cytosolic DNA activates an IRF3-dependent innate immune response. Immunity 24, 93–103 (2006).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Volkman, H. E., Cambier, S., Gray, E. E. & Stetson, D. B. Tight nuclear tethering of cGAS is essential for preventing autoreactivity. eLife 8, e47491 (2019).
Zierhut, C. et al. The cytoplasmic DNA sensor cGAS promotes mitotic cell death. Cell 178, 302–315 (2019).
Jiang, H. et al. Chromatin-bound cGAS is an inhibitor of DNA repair and hence accelerates genome destabilization and cell death. EMBO J. 38, e102718 (2019).
Gentili, M. et al. The N-terminal domain of cGAS determines preferential association with centromeric DNA and innate immune activation in the nucleus. Cell Rep. 26, 2377–2393 (2019).
Liu, H. et al. Nuclear cGAS suppresses DNA repair and promotes tumorigenesis. Nature 563, 131–136 (2018).
Li, X. et al. Cyclic GMP-AMP synthase is activated by double-stranded DNA-induced oligomerization. Immunity 39, 1019–1031 (2013).
Wang, W. W., Zeng, Y., Wu, B., Deiters, A. & Liu, W. R. A chemical biology approach to reveal Sirt6-targeted histone H3 sites in nucleosomes. ACS Chem. Biol. 11, 1973–1981 (2016).
Tachiwana, H. et al. Structural basis of instability of the nucleosome containing a testis-specific histone variant, human H3T. Proc. Natl Acad. Sci. USA 107, 10454–10459 (2010).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Acknowledgements
We acknowledge University of Texas, McGovern Medical School in Houston for providing access to their cryo-EM facility for data collection. We thank V. Mallampalli, G. Fan and I. Serysheva for assistance with cryo-EM data collection and Z. Cui at Texas A&M University for assistance with cryo-EM data processing. This research was supported in part by the Welch Foundation (Grants A-1931-20170325 to P.L. and A-1715 to W.R.L.) and National Institutes of Health (Grants R01 AI145287 to P.L., R01 GM121584 and R01 GM127575 to W.R.L.). A.P.W. and Y.L. were supported by NIH grant R01 HL148153 and award W81XWH-17-1-0052 through the Office of the Assistant Secretary of Defense for Health Affairs, Peer Reviewed Medical Research Programs.
Author information
Authors and Affiliations
Contributions
P.L. conceived the study. B.Z. and P.X. expressed proteins, conducted binding studies and determined the structures. C.M.R. prepared the reconstituted nucleosome. T.J. conducted the binding studies and generated and purified the cGAS mutants. O.S. purified the cGAS mutants. Y.L. and A.P.W. studied the nuclear localization of cGAS. B.Z., P.X., W.R.L. and P.L. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Yuan He, Jae U Jung and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 cGAS binds tightly with nucleosome.
a, Gel-filtration chromatography and SDS–PAGE analyses of nucleosomes purified from HEK293T cells. b, Gel-filtration chromatography and SDS–PAGE analyses of the in vitro reconstituted nucleosomes. c, Gel-filtration chromatography showing that mouse cGAS catalytic domain binds to nucleosomes. d, SDS–PAGE analysis of nucleosome and mouse cGAS catalytic domain complex purified by gel-filtration chromatography. e, Polyacrylamide gel EMSA showing the interactions between mouse cGAS catalytic domain with reconstituted nucleosomes and nucleosomes purified from HEK293T cells. cGAS is mixed with nucleosomes at a molar ratio of 6:1. f, Agarose gel electrophoresis of the dsDNA from purified mononucleosomes and oligonucleosomes. The histones have been digested with proteinase K. g, SDS–PAGE analyses of purified mononucleosomes and oligonucleosomes. h, Polyacrylamide gel EMSA showing that mouse cGAS catalytic domain binds to mononucleosomes and oligonucleosomes (left panel). In the mixtures, the molar ratio of cGAS/mononucleosome is 3:1 and cGAS/oligonucleosome is 6:1. SDS–PAGE analyses of the input samples for EMSA were shown in the right panel. i, SDS–PAGE analyses of biotin-Avi-His6-SUMO fusion of human and mouse cGAS full length and catalytic domain proteins used for nucleosome binding studies.
Extended Data Fig. 2 cGAS-nucleosome binding studies and activity assays of cGAS-nucleosome complex.
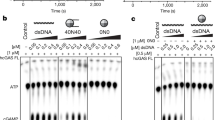
a, SPR binding studies show that full-length human cGAS binds to nucleosomes purified from HEK293T cells with nanomolar affinity. b, SPR binding studies show that human cGAS catalytic domain binds to reconstituted nucleosomes with nanomolar affinity. c, d, Bio-layer interferometry binding studies of full-length human cGAS and its catalytic domain with nucleosomes (HEK293T). e, f, SPR binding studies show that full-length mouse cGAS and its catalytic domain bind nucleosomes (HEK293T) with nanomolar affinities. g, h, Bio-layer interferometry binding studies of full-length mouse cGAS and its catalytic domain with nucleosomes (HEK293T). i, Gel-filtration chromatography (top) and SDS–PAGE (bottom) analyses of 45-bp ISD dsDNA, nucleosome and cGAS mixture show that the nucleosome efficiently competes with dsDNA to bind cGAS. j, Agarose gel electrophoresis of the salmon sperm DNA used in cGAS activity assays. k, cGAS activity assays by ion exchange chromatography show that oligonucleosome binding potently inhibits the activity of cGAS and ligand-free cGAS can be activated by the cGAS-oligonucleosome complex.
Extended Data Fig. 3 Cryo-EM analysis of mouse cGAS catalytic domain in complex with the reconstituted nucleosome.
a, Purification and SDS–PAGE analysis of nucleosomes from HEK293T cells in complex with mouse cGAS catalytic domain. b, Purification and SDS–PAGE analysis of reconstituted nucleosomes in complex with mouse cGAS catalytic domain. c, Representative micrograph of mouse cGAS -nucleosome complex in vitrified ice. Scale bar, 50 nm. d, 2D class averages of mouse cGAS-nucleosome complex particles. Scale bar, 10 nm. e, Flowchart of data processing; see Methods for details. f, Angular distribution of the mouse cGAS-nucleosome (1:1) particles included in the final reconstruction. g, Angular distribution of the mouse cGAS-nucleosome (2:1) particles included in the final reconstruction. h, Final 3D reconstruction of the mouse cGAS-nucleosome (1:1) complex, coloured according to the local resolution. i, Final 3D reconstruction of the mouse cGAS-nucleosome (2:1) complex, coloured according to the local resolution. j, Corrected Gold-standard Fourier shell correlation curves of the mouse cGAS-nucleosome (1:1) complex for the 3D electron microscopy reconstruction. k, Corrected Gold-standard Fourier shell correlation curves of the mouse cGAS-nucleosome (2:1) complex for the 3D electron microscopy reconstruction. l, Polyacrylamide gel shift assay (left) showing that one nucleosome can bind to two molecules of mouse cGAS catalytic domain. Nucleosome is mixed with mouse cGAS at molar ratio of 1:1, 1:2 and 1:3. SDS–PAGE analysis of the samples used for the gel shift assays was shown on right panel. m, Density map (contoured at 3σ) and structural model of mouse cGAS-nucleosome (2:1) complex at 6.8 Å resolution.
Extended Data Fig. 4 Density maps and structural models of cGAS-nucleosome (reconstituted, 1:1) complex.
a–f, The density maps (grey mesh) of histones H2A, H2B, H3, H4, part of mouse cGAS catalytic domain, and the Widom 601 nucleosome positioning sequence DNA contoured at 3σ. The protein and DNA structures fitted into the density map are shown by the stick models.
Extended Data Fig. 5 Cryo-EM analysis of mouse cGAS domain in complex with nucleosome purified from HEK293T cells.
a, Representative micrograph of mouse cGAS-nucleosome complex in vitrified ice. Scale bar, 50 nm. b, 2D class averages of mouse cGAS-nucleosome complex particles. Scale bar, 10 nm. c, Flowchart of data processing; see Methods for details. d, Angular distribution of particles included in the final reconstruction. e, Final 3D reconstruction, coloured according to the local resolution. f, Corrected Gold-standard Fourier shell correlation curves for the 3D electron microscopy reconstruction. g, Density map (contoured at 3σ) and structural model of cGAS-nucleosome (1:1) complex.
Extended Data Fig. 6 Mutations in the acidic patch of the nucleosome abolished cGAS binding.
a, Superposition of structures of ligand-free mouse cGAS (PDB, 4K8V), mouse cGAS in complex with dsDNA (PDB, 4LEY), and mouse cGAS bound to the nucleosome. b, Sequence alignment of human and mouse cGAS around the nucleosome binding site. The conserved basic residues around the nucleosome binding site are coloured red. Residues that abolish nucleosome binding when mutated are highlighted yellow. c, Superposition for the structures of ligand-free human cGAS (PDB, 4LEV), ligand-free mouse cGAS (PDB, 4K8V) and mouse cGAS bound to the nucleosome. d, Ni-NTA agarose pull-down assays of mouse cGAS catalytic domain by His-tagged H2A-H2B dimer. The 6A dimer contains mutations E61A, E64A, D90A, E91A, E92A of H2A and D51A of H2B. The 6K dimer contains mutations E61K, E64K, D90K, E91K, E92K of H2A and D51K of H2B. e, Sequence alignment of wild-type (WT) human H2A and human H2A.Bbd. The acidic patch residues of WT H2A are coloured red. f, Polyacrylamide gel shift assay (left) showing that the recombinant nucleosome variant (H2A.Bbd) does not bind to mouse cGAS catalytic domain. In this assay, mouse cGAS was mixed with nucleosomes at a molar ratio of 3:1. SDS–PAGE analysis of the samples for gel shift assay were shown on the right panel.
Extended Data Fig. 7 Characterization of mouse cGAS catalytic domain mutants and oligonucleosome binding studies.
a, SDS–PAGE analysis of mouse cGAS catalytic domain mutants used for the gel shift assays and enzyme activity assays. b, SDS–PAGE analysis of mouse cGAS and nucleosome mixture samples used for the gel shift assay. c, Polyacrylamide gel EMSA shows that mutations at the cGAS-nucleosome interface affect oligonucleosome binding by cGAS. In these samples, mouse cGAS was mixed with oligonucleosomes at a molar ratio of 6:1. The samples used for the binding studies were analysed by SDS–PAGE (right panel). d, Circular dichroism of mouse cGAS catalytic domain and its mutants used for gel shift assays and enzyme activity assays. cGAS mutants that have strong binding to nucleosomes are shown by the spectra on the left. cGAS mutants that have weak or no binding to nucleosomes are shown by the spectra on the right.
Extended Data Fig. 8 Mutations at the cGAS-nucleosome interface affect nucleosome binding by human cGAS.
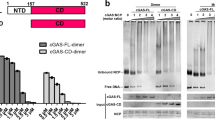
a, SDS–PAGE analysis of biotin-labelled full-length human cGAS mutants used for the SPR binding studies. b–n, SPR binding studies of full-length human cGAS mutants with reconstituted nucleosomes.
Extended Data Fig. 9 Mutations at the cGAS-nucleosome interface affect dsDNA binding, cGAS activity and cGAS-mediated signalling.
a, Agarose gel shift assay shows that mutations at the nucleosome binding surface of human cGAS affect the binding of a 45-bp dsDNA. b, Enzyme activity assays of mouse cGAS catalytic domain mutants by ion exchange chromatography. In this assay, 2.5 μM mouse cGAS was incubated with 0.2 mg ml−1 salmon sperm DNA. Wild-type mouse cGAS and negative control without DNA are coloured in green and blue. Mutations that abolish nucleosome binding are coloured red. The negative control mutation K382E is coloured orange. c, Enzyme activity assays of full-length human cGAS mutants by ion exchange chromatography. In this assay, 2.5 μM human cGAS was incubated with 0.2 mg ml−1 salmon sperm DNA. WT human cGAS and negative control without DNA are coloured in green and blue. Mutations that abolish nucleosome binding are coloured red. The negative control mutation K394E is coloured orange. d, IFN-β luciferase reporter assays show that mutations of human cGAS affect signalling in HEK293T cells. Luciferase reporter signals from the cells transfected with 0.025 ng cGAS and 0.4 ng STING are indicated by the orange bars, from cells transfected with 0.00625 ng cGAS and 0.4 ng STING by the cyan bars, from cells transfected with 0.025 ng cGAS, 0.4 ng STING, or the vector control by the green, brown and purple bars, respectively. The data (mean ± s.e.m.) are representative of three independent experiments. Each dot represents a biological replicate (n = 3). Two-tailed Student’s t-test: *P < 0.05, **P < 0.01, ***P < 0.001; NS, not significant. e, Western blot shows that WT mouse cGAS and its mutants have similar expression level in the transfected HEK293T cells. f, Western blot shows WT human cGAS and its mutants have similar expression level in the transfected HEK293T cells.
Supplementary information
Supplementary Figure 1
This file contains uncropped images with molecular mass markers.
Rights and permissions
About this article
Cite this article
Zhao, B., Xu, P., Rowlett, C.M. et al. The molecular basis of tight nuclear tethering and inactivation of cGAS. Nature 587, 673–677 (2020). https://doi.org/10.1038/s41586-020-2749-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2749-z
This article is cited by
-
The CRL5–SPSB3 ubiquitin ligase targets nuclear cGAS for degradation
Nature (2024)
-
TXNRD1 drives the innate immune response in senescent cells with implications for age-associated inflammation
Nature Aging (2024)
-
When DNA-damage responses meet innate and adaptive immunity
Cellular and Molecular Life Sciences (2024)
-
MRE11 liberates cGAS from nucleosome sequestration during tumorigenesis
Nature (2024)
-
ALDH2 mitigates LPS-induced cardiac dysfunction, inflammation, and apoptosis through the cGAS/STING pathway
Molecular Medicine (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.