Abstract
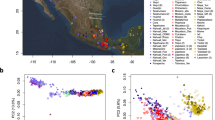
The possibility of voyaging contact between prehistoric Polynesian and Native American populations has long intrigued researchers. Proponents have pointed to the existence of New World crops, such as the sweet potato and bottle gourd, in the Polynesian archaeological record, but nowhere else outside the pre-Columbian Americas1,2,3,4,5,6, while critics have argued that these botanical dispersals need not have been human mediated7. The Norwegian explorer Thor Heyerdahl controversially suggested that prehistoric South American populations had an important role in the settlement of east Polynesia and particularly of Easter Island (Rapa Nui)2. Several limited molecular genetic studies have reached opposing conclusions, and the possibility continues to be as hotly contested today as it was when first suggested8,9,10,11,12. Here we analyse genome-wide variation in individuals from islands across Polynesia for signs of Native American admixture, analysing 807 individuals from 17 island populations and 15 Pacific coast Native American groups. We find conclusive evidence for prehistoric contact of Polynesian individuals with Native American individuals (around ad 1200) contemporaneous with the settlement of remote Oceania13,14,15. Our analyses suggest strongly that a single contact event occurred in eastern Polynesia, before the settlement of Rapa Nui, between Polynesian individuals and a Native American group most closely related to the indigenous inhabitants of present-day Colombia.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Individual-level genotypes for new data presented here are available through a data-access agreement to respect the privacy of the participants for the transfer of genetic data from the European Genome Archive (EGA; https://www.ebi.ac.uk/ega/home), under accession number EGAS00001004209.
Code availability
Details regarding the packages and versions used are included in the Methods. Codes for the matrix completions described in the paper are available at https://github.com/AlexIoannidis/completion.
References
Brown, J. M. The Riddle of the Pacific (T. F. Unwin, 1924).
Heyerdahl, T. American Indians in the Pacific (Allen & Unwin, 1952).
Yen, D. E. The Sweet Potato and Oceania (Bishop Museum Press, 1974).
Montenegro, A., Avis, C. & Weaver, A. Modeling the prehistoric arrival of the sweet potato in Polynesia. J. Archaeol. Sci. 35, 355–367 (2008).
Roullier, C., Benoit, L., McKey, D. B. & Lebot, V. Historical collections reveal patterns of diffusion of sweet potato in Oceania obscured by modern plant movements and recombination. Proc. Natl Acad. Sci. USA 110, 2205–2210 (2013).
Clarke, A. C., Burtenshaw, M. K., McLenachan, P. A., Erickson, D. L. & Penny, D. Reconstructing the origins and dispersal of the Polynesian bottle gourd (Lagenaria siceraria). Mol. Biol. Evol. 23, 893–900 (2006).
Muñoz-Rodríguez, P. et al. Reconciling conflicting phylogenies in the origin of sweet potato and dispersal to Polynesia. Curr. Biol. 28, 1246–1256 (2018).
Lie, B. A. et al. Molecular genetic studies of natives on Easter Island: evidence of an early European and Amerindian contribution to the Polynesian gene pool. Tissue Antigens 69, 10–18 (2007).
Thorsby, E. The Polynesian gene pool: an early contribution by Amerindians to Easter Island. Phil. Trans. R. Soc. Lond. B 367, 812–819 (2012).
Moreno-Mayar, J. V. et al. Genome-wide ancestry patterns in Rapanui suggest pre-European admixture with Native Americans. Curr. Biol. 24, 2518–2525 (2014).
Fehren-Schmitz, L. et al. Genetic ancestry of Rapanui before and after European contact. Curr. Biol. 27, 3209–3215 (2017).
Hagelberg, E., Quevedo, S., Turbon, D. & Clegg, J. B. DNA from ancient Easter Islanders. Nature 369, 25–26 (1994).
Wilmshurst, J. M., Hunt, T. L., Lipo, C. P. & Anderson, A. J. High-precision radiocarbon dating shows recent and rapid initial human colonization of East Polynesia. Proc. Natl Acad. Sci. USA 108, 1815–1820 (2011).
Hunt, T. L. & Lipo, C. P. Late colonization of Easter Island. Science 311, 1603–1606 (2006).
Mulrooney, M. A. An island-wide assessment of the chronology of settlement and land use on Rapa Nui (Easter Island) based on radiocarbon data. J. Archaeol. Sci. 40, 4377–4399 (2013).
Martinsson-Wallin, H., Wallin, P. & Anderson, A. Chronogeographic variation in initial East Polynesian construction of monumental ceremonial sites. J. Island Coastal Archaeol. 8, 405–421 (2013).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Maples, B. K., Gravel, S., Kenny, E. E. & Bustamante, C. D. RFMix: a discriminative modeling approach for rapid and robust local-ancestry inference. Am. J. Hum. Genet. 93, 278–288 (2013).
Liang, M. & Nielsen, R. The lengths of admixture tracts. Genetics 197, 953–967 (2014).
Gravel, S. et al. Demographic history and rare allele sharing among human populations. Proc. Natl Acad. Sci. USA 108, 11983–11988 (2011).
Fischer, S. R. Island at the End of the World (Reaktion Books, 2006).
Maude, H. E. Slavers in Paradise: The Peruvian Slave trade in Polynesia, 1862–1864 (Stanford Univ. Press, 1981).
Hurles, M. E. et al. Native American Y chromosomes in Polynesia: the genetic impact of the Polynesian slave trade. Am. J. Hum. Genet. 72, 1282–1287 (2003).
Seemann, B. Flora Vitiensis (L. Reeve, 1865).
Scaglion, R. in The Sweet Potato in Oceania: A Reappraisal (eds Ballard, C. et al.) 35–41 (Oceania Publications, 2005).
Hornell, J. Was there pre-Columbian Contact between the Peoples of Oceania and South America? J. Polynesian Soc. 54, 167–191 (1945).
Dewan, L. & Hosler, D. Ancient maritime trade on balsa rafts: an engineering analysis. J. Anthropol. Res. 64, 19–40 (2008).
Hosler, D. West Mexican metallurgy: revisited and revised. J. World Prehist. 22, 185–212 (2009).
Callaghan, R. T. Prehistoric trade between Ecuador and West Mexico: a computer simulation of coastal voyages. Antiquity 77, 796–804 (2003).
Irwin, G. J. Against, across and down the wind: a case for the systematic exploration of the remote Pacific Islands. J. Polynesian Soc. 98, 167–206 (1989).
Lewis, D. We, the Navigators (Univ. Hawaii Press, 1994).
Gravel, S. Population genetics models of local ancestry. Genetics 191, 607–619 (2012).
Porteous, J. D. The Modernization of Easter Island (Univ. Victoria, 1981).
Loh, P. R. et al. Inferring admixture histories of human populations using linkage disequilibrium. Genetics 193, 1233–1254 (2013).
Patterson, N. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012).
Walworth, M. Eastern Polynesian: the linguistic evidence revisited. Oceanic Linguistics 53, 256–272 (2014).
Kirch, P. V. & Green, R. C. Hawaiki, Ancestral Polynesia (Cambridge Univ. Press, 2001).
Hunt, T. & Lipo, C. The Statues That Walked (Free Press, 2011).
Heyerdahl, T. Fatu-Hiva: Back to Nature (Allen & Unwin, 1974).
Montenegro, Á., Callaghan, R. T. & Fitzpatrick, S. M. Using seafaring simulations and shortest-hop trajectories to model the prehistoric colonization of Remote Oceania. Proc. Natl Acad. Sci. USA 113, 12685–12690 (2016).
Di Piazza, A., Di Piazza, P. & Pearthree, E. Sailing virtual canoes across Oceania: revisiting island accessibility. J. Archaeol. Sci. 34, 1219–1225 (2007).
Kirch, P. V. On the Road of the Winds (Univ. California Press, 2017).
Durbin, R. et al. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010); corrigendum 473, 544 (2011).
de la Fuente, C. et al. Genomic insights into the origin and diversification of late maritime hunter-gatherers from the Chilean Patagonia. Proc. Natl Acad. Sci. USA 115, E4006–E4012 (2018).
Wojcik, G. L. et al. Genetic analyses of diverse populations improves discovery for complex traits. Nature 570, 514–518 (2019).
Bryc, K. et al. Genome-wide patterns of population structure and admixture among Hispanic/Latino populations. Proc. Natl Acad. Sci. USA 107 (Suppl 2), 8954–8961 (2010).
Posth, C. et al. Reconstructing the deep population history of Central and South America. Cell 175, 1185–1197 (2018).
Moreno-Mayar, J. V. et al. Early human dispersals within the Americas. Science 362, eaav2621 (2018).
Aitchison, J. The Statistical Analysis of Compositional Data (Chapman and Hall, 1986).
Novembre, J. et al. Genes mirror geography within Europe. Nature 456, 98–101 (2008).
Tyner, C. et al. The UCSC Genome Browser database: 2017 update. Nucleic Acids Res. 45 (D1), D626–D634 (2017).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Wickham, H. ggplot2 (Springer, 2016).
Holmes, S. & Huber, W. Modern Statistics for Modern Biology (Cambridge Univ. Press, 2019).
Lawson, D. J., van Dorp, L. & Falush, D. A tutorial on how not to over-interpret STRUCTURE and ADMIXTURE bar plots. Nat. Commun. 9, 3258 (2018).
van Dorp, L. et al. Evidence for a common origin of blacksmiths and cultivators in the Ethiopian Ari within the last 4500 years: lessons for clustering-based inference. PLoS Genet. 11, e1005397 (2015).
Alexander, D. H. & Lange, K. Enhancements to the ADMIXTURE algorithm for individual ancestry estimation. BMC Bioinformatics 12, 246 (2011).
Francis, R. M. pophelper: an R package and web app to analyse and visualize population structure. Mol. Ecol. Resour. 17, 27–32 (2017).
Pickrell, J. K. & Pritchard, J. K. Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet. 8, e1002967 (2012).
Meyer, B. S., Matschiner, M. & Salzburger, W. Disentangling incomplete lineage sorting and introgression to refine species-tree estimates for Lake Tanganyika cichlid fishes. Syst. Zool. 66, 531–550 (2017).
van den Boogaart, K. G. & Tolosana-Delgado, R. Analyzing Compositional Data with. (Springer Science & Business Media, 2013).
Homburger, J. R. et al. Genomic insights into the ancestry and demographic history of South America. PLoS Genet. 11, e1005602 (2015).
O’Connell, J. et al. A general approach for haplotype phasing across the full spectrum of relatedness. PLoS Genet. 10, e1004234 (2014).
Troyanskaya, O. et al. Missing value estimation methods for DNA microarrays. Bioinformatics 17, 520–525 (2001).
Browning, S. R. et al. Local ancestry inference in a large US-based Hispanic/Latino study: Hispanic community health study/study of Latinos (HCHS/SOL). G3 Genes Genom. Genet. 6, 1525–1534 (2016).
Makarenkov, V. & Lapointe, F.-J. A weighted least-squares approach for inferring phylogenies from incomplete distance matrices. Bioinformatics 20, 2113–2121 (2004).
Peter, B. M. Admixture, population structure, and F-statistics. Genetics 202, 1485–1501 (2016).
Hastie, T., Tibshirani, R. & Friedman, J. The Elements of Statistical Learning (Springer Science & Business Media, 2009).
Moreno-Estrada, A. et al. Reconstructing the population genetic history of the Caribbean. PLoS Genet. 9, e1003925 (2013).
Gusev, A. et al. Whole population, genome-wide mapping of hidden relatedness. Genome Res. 19, 318–326 (2009).
Moreno-Estrada, A. et al. The genetics of Mexico recapitulates Native American substructure and affects biomedical traits. Science 344, 1280–1285 (2014).
Longobardi, G. et al. Across language families: genome diversity mirrors linguistic variation within Europe. Am. J. Phys. Anthropol. 157, 630–640 (2015).
Han, E. et al. Clustering of 770,000 genomes reveals post-colonial population structure of North America. Nat. Commun. 8, 14238 (2017).
Epskamp, S., Cramer, A., Waldorp, L. J., Schmittmann, V. D. & Borsboom, D. qgraph: network visualizations of relationships in psychometric data. J. Stat. Softw. 48, 1–18 (2012).
Pickrell, J. K. et al. Ancient west Eurasian ancestry in southern and eastern Africa. Proc. Natl Acad. Sci. USA 111, 2632–2637 (2014).
Acknowledgements
We thank the participants and volunteers who donated DNA samples for this study and the many researchers who contributed to collecting samples, in particular J. Martinson, D. Weatherall and J. Clegg, as well as H.-W. Peng (Taiwan), T. Teariki (Cook Islands) and J. Roux (French Polynesia). We thank P. P. Edmunds Paoa, mayor of the Municipality of Easter Island, the Rapahango family, H. Huke, T. Hotu, O. Hey Riroroko, J. Emilio Estay and S. Fareea for providing local support during fieldwork and community engagement on Rapa Nui. We also thank the Rapa Nui Museum and the Office of Rapa Nui Patrimony for outreach support, and the people of Rapa Nui for making this study possible. We thank M. Stoneking for facilitating access to published data that enabled early stages of the analyses, as well as the ChileGenomico project consortium for providing access to reference genotype data from 16 Aymara individuals and 32 of Mapuche ancestry. We acknowledge the National Institutes of Health (NIH) genome-wide association study (GWAS) Data Repository for granting access to the POPRES data set. We also thank the support from the Core Staff at the UCSF Institute for Human Genetics for contributing with genotyping capacity, and the Stanford Center for Computational, Evolutionary and Human Genomics (CEHG) for supporting the initial stages of this project. We are grateful for genotyping and IT support from J. Cervantes, M. Torres and technicians from LANGEBIO’s Genomics Core Facility at CINVESTAV, Mexico. This work was supported by the George Rosenkranz Prize for Health Care Research in Developing Countries, Mexico’s CONACYT Basic Research Program (grant number CB-2015-01-251380), and the International Center for Genetic Engineering and Biotechnology (ICGEB, Italy) grant CRP/MEX15-04_EC (each awarded to A.M.-E.); the American Society of Engineering Education NDSEG Fellowship and the National Library of Medicine (NLM) training grant T15LM007033 (awarded to A.G.I.); the Chilean funding programs FONDEF, FONDECYT and CONICYT (grants D10I1007, 1130303 and USA2013-0015, respectively); and the National Institute for Health Research (NIHR) Oxford Biomedical Research Centre (BRC) and the Wellcome Trust Fellowship with reference 106289/Z/14/Z (to A.J.M.). Views expressed are those of the author(s) and not necessarily those of the NIHR, the NHS or the UK Department of Health.
Author information
Authors and Affiliations
Contributions
A.M.-E. and K.S. conceived the study. A.M.-E., A.G.I., A.J.M. and C.D.B. provided overall project supervision and management. A.M.-E., E.H., R.A.V., M.M., A.J.M., C.D.B. and C.R.G. contributed to study design. A.S., G.L.W., K.C.B., C.E., S.H., E.G.B., C.D.B. and A.M.-E. carried out genotyping experiments and quality control. A.G.I., J.B.-P., J.V.M.-M., J.E.R.-R., C.D.Q.-C., J.R.H. and A.M.-E. analysed the data. A.G.I. developed the analytical methods. A.M.-E., A.G.I. and J.B.-P. interpreted the results. A.M.-E., K.S., E.H., J.F.M.-P., K.A., T.P., K.R., A.V.S.H., M.C.A.-A., A.S., G.L.W., K.C.B., L.H., S.B., M.A., E.L., C.E., S.H., E.G.B., L.C., R.A.V., M.M., A.J.M. and C.D.B. contributed to acquisition of the data. A.G.I. wrote the manuscript, and A.G.I., A.M.-E., E.H., J.B.-P., J.V.M.-M. and M.M. edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
C.D.B. is a member of the scientific advisory boards for Liberty Biosecurity, Personalis, 23andMe Roots into the Future, Ancestry.com, IdentifyGenomics, Genomelink and Etalon and is a founder of CDB Consulting. C.R.G. owns stock in 23andMe and is a member of the scientific advisory board for Encompass Bioscience.
Additional information
Peer review information Nature thanks John Lindo, Paul Wallin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion, Supplementary References, Supplementary Tables 1-14 and Supplementary Figures 1-24.
Rights and permissions
About this article
Cite this article
Ioannidis, A.G., Blanco-Portillo, J., Sandoval, K. et al. Native American gene flow into Polynesia predating Easter Island settlement. Nature 583, 572–577 (2020). https://doi.org/10.1038/s41586-020-2487-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2487-2
This article is cited by
-
Rapa Nui’s population history rewritten using ancient DNA
Nature (2024)
-
Deep history of cultural and linguistic evolution among Central African hunter-gatherers
Nature Human Behaviour (2024)
-
Ancient Rapanui genomes reveal resilience and pre-European contact with the Americas
Nature (2024)
-
Ancient DNA uncovers past migrations in California
Nature (2023)
-
CINWA (database of terminology for cultivated plants in indigenous languages of northwestern South America): introducing a resource for research in ethnobiology, anthropology, historical linguistics, and interdisciplinary research on the neolithic transition in South America
Language Resources and Evaluation (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.