Abstract
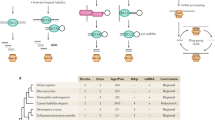
Heterochromatin comprises tightly compacted repetitive regions of eukaryotic chromosomes. The inheritance of heterochromatin through mitosis requires RNA interference (RNAi), which guides histone modification1 during the DNA replication phase of the cell cycle2. Here we show that the alternating arrangement of origins of replication and non-coding RNA in pericentromeric heterochromatin results in competition between transcription and replication in Schizosaccharomyces pombe. Co-transcriptional RNAi releases RNA polymerase II (Pol II), allowing completion of DNA replication by the leading strand DNA polymerase, and associated histone modifying enzymes3 that spread heterochromatin with the replication fork. In the absence of RNAi, stalled forks are repaired by homologous recombination without histone modification.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
GenBank/EMBL/DDBJ
Gene Expression Omnibus
Data deposits
Genomics data and analysis are available from the Gene Expression Omnibus accession number GSE30837. Individual cDNA sequences are available from GenBank with accession numbers JN388396–JN388565.
References
Volpe, T. A. et al. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297, 1833–1837 (2002)
Kloc, A., Zaratiegui, M., Nora, E. & Martienssen, R. RNA interference guides histone modification during the S phase of chromosomal replication. Curr. Biol. 18, 490–495 (2008)
Li, F., Martienssen, R. & Cande, W. Z. Coordination of DNA replication and histone modification by the Rik1–Dos2 complex. Nature 475, 244–248 (2011)
Verdel, A. et al. RNAi-mediated targeting of heterochromatin by the RITS complex. Science 303, 672–676 (2004)
Motamedi, M. R. et al. HP1 proteins form distinct complexes and mediate heterochromatic gene silencing by nonoverlapping mechanisms. Mol. Cell 32, 778–790 (2008)
Bayne, E. H. et al. Stc1: a critical link between RNAi and chromatin modification required for heterochromatin integrity. Cell 140, 666–677 (2010)
Irvine, D. V. et al. Argonaute slicing is required for heterochromatic silencing and spreading. Science 313, 1134–1137 (2006)
Svejstrup, J. Q. The interface between transcription and mechanisms maintaining genome integrity. Trends Biochem. Sci. 35, 333–338 (2010)
Cam, H. P. et al. Comprehensive analysis of heterochromatin- and RNAi-mediated epigenetic control of the fission yeast genome. Nature Genet. 37, 809–819 (2005)
Djupedal, I. et al. RNA Pol II subunit Rpb7 promotes centromeric transcription and RNAi-directed chromatin silencing. Genes Dev. 19, 2301–2306 (2005)
Bühler, M., Verdel, A. & Moazed, D. Tethering RITS to a nascent transcript initiates RNAi- and heterochromatin-dependent gene silencing. Cell 125, 873–886 (2006)
Rosonina, E., Kaneko, S. & Manley, J. L. Terminating the transcript: breaking up is hard to do. Genes Dev. 20, 1050–1056 (2006)
Djupedal, I. et al. Analysis of small RNA in fission yeast; centromeric siRNAs are potentially generated through a structured RNA. EMBO J. 28, 3832–3844 (2009)
Bühler, M., Haas, W., Gygi, S. P. & Moazed, D. RNAi-dependent and -independent RNA turnover mechanisms contribute to heterochromatic gene silencing. Cell 129, 707–721 (2007)
Murakami, H. et al. Ribonuclease activity of Dis3 is required for mitotic progression and provides a possible link between heterochromatin and kinetochore function. PLoS ONE 2, e317 (2007)
Smith, J. G. et al. Replication of centromere II of Schizosaccharomyces pombe . Mol. Cell. Biol. 15, 5165–5172 (1995)
Hayashi, M. T., Takahashi, T. S., Nakagawa, T., Nakayama, J. & Masukata, H. The heterochromatin protein Swi6/HP1 activates replication origins at the pericentromeric region and silent mating-type locus. Nature Cell Biol. 11, 357–362 (2009)
Lambert, S., Watson, A., Sheedy, D. M., Martin, B. & Carr, A. M. Gross chromosomal rearrangements and elevated recombination at an inducible site-specific replication fork barrier. Cell 121, 689–702 (2005)
Shimmoto, M. et al. Interactions between Swi1-Swi3, Mrc1 and S phase kinase, Hsk1 may regulate cellular responses to stalled replication forks in fission yeast. Genes Cells 14, 669–682 (2009)
Segurado, M., Gomez, M. & Antequera, F. Increased recombination intermediates and homologous integration hot spots at DNA replication origins. Mol. Cell 10, 907–916 (2002)
Minca, E. C. & Kowalski, D. Multiple Rad5 activities mediate sister chromatid recombination to bypass DNA damage at stalled replication forks. Mol. Cell 38, 649–661 (2010)
Deshpande, A. M. & Newlon, C. S. DNA replication fork pause sites dependent on transcription. Science 272, 1030–1033 (1996)
Scott, K. C., Merrett, S. L. & Willard, H. F. A heterochromatin barrier partitions the fission yeast centromere into discrete chromatin domains. Curr. Biol. 16, 119–129 (2006)
Lambert, S. et al. Homologous recombination restarts blocked replication forks at the expense of genome rearrangements by template exchange. Mol. Cell 39, 346–359 (2010)
Groth, A., Rocha, W., Verreault, A. & Almouzni, G. Chromatin challenges during DNA replication and repair. Cell 128, 721–733 (2007)
Pirngruber, J., Shchebet, A. & Johnsen, S. A. Insights into the function of the human P-TEFb component CDK9 in the regulation of chromatin modifications and co-transcriptional mRNA processing. Cell Cycle 8, 3636–3642 (2009)
Zhang, K. et al. Clr4/Suv39 and RNA quality control factors cooperate to trigger RNAi and suppress antisense RNA. Science 331, 1624–1627 (2011)
Ghazal, G. et al. Yeast RNase III triggers polyadenylation-independent transcription termination. Mol. Cell 36, 99–109 (2009)
Washburn, R. S. & Gottesman, M. E. Transcription termination maintains chromosome integrity. Proc. Natl Acad. Sci. USA 108, 792–797 (2011)
Pauler, F. M., Koerner, M. V. & Barlow, D. P. Silencing by imprinted noncoding RNAs: is transcription the answer? Trends Genet. 23, 284–292 (2007)
Moreno, S., Klar, A. & Nurse, P. Molecular genetic analysis of fission yeast Schizosaccharomyces pombe . Methods Enzymol. 194, 795–823 (1991)
Sallés, F. J., Richards, W. G. & Strickland, S. Assaying the polyadenylation state of mRNAs. Methods 17, 38–45 (1999)
Zaratiegui, M. et al. CENP-B preserves genome integrity at replication forks paused by retrotransposon LTR. Nature 469, 112–115 (2011)
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008)
Acknowledgements
We thank D. Roh and T. Volpe for isolating cDNA clones. D.V.I. was supported by a NHMRC CJ Martin Postdoctoral Research Fellowship. M.Z. was supported by a fellowship from the Spanish Ministry of Science. This work was supported by grants BFU2008-01919 and Consolider-Ingenio CSD2007-00015 from the Spanish Ministry of Science and Innovation to F.A., and NIH R01 GM076396 to W.Z.C. and R.A.M.
Author information
Authors and Affiliations
Contributions
S.C., D.V.I., A.K., J.R. contributed equally to this work and are listed in alphabetical order. M.Z., S.C., D.V.I., A.K., J.R., F.L., E.d.C., L.M., A.-Y.C. and D.G. performed experiments, and S.C. analysed the data. W.Z.C., F.A., B.A. and R.A.M. designed experiments and R.A.M. and M.Z. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Tables 1-4, Supplementary Figures 1-5 with legends and additional references. (PDF 5439 kb)
Rights and permissions
About this article
Cite this article
Zaratiegui, M., Castel, S., Irvine, D. et al. RNAi promotes heterochromatic silencing through replication-coupled release of RNA Pol II. Nature 479, 135–138 (2011). https://doi.org/10.1038/nature10501
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10501
This article is cited by
-
Dicer promotes genome stability via the bromodomain transcriptional co-activator BRD4
Nature Communications (2022)
-
Rbm10 facilitates heterochromatin assembly via the Clr6 HDAC complex
Epigenetics & Chromatin (2021)
-
Conserved chromosomal functions of RNA interference
Nature Reviews Genetics (2020)
-
The prevention and resolution of DNA replication–transcription conflicts in eukaryotic cells
Genome Instability & Disease (2020)
-
ETMR: a tumor entity in its infancy
Acta Neuropathologica (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.