Abstract
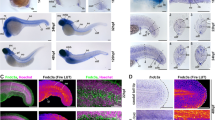
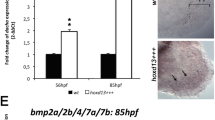
The early development of teleost paired fins is strikingly similar to that of tetrapod limb buds and is controlled by similar mechanisms1,2. One early morphological divergence between pectoral fins and limbs is in the fate of the apical ectodermal ridge (AER), the distal epidermis that rims the bud. Whereas the AER of tetrapods regresses after specification of the skeletal progenitors3, the AER of teleost fishes forms a fold that elongates4,5. Formation of the fin fold is accompanied by the synthesis of two rows of rigid, unmineralized fibrils called actinotrichia, which keep the fold straight6,7 and guide the migration of mesenchymal cells within the fold5,8. The actinotrichia are made of elastoidin, the components of which, apart from collagen, are unknown. Here we show that two zebrafish proteins, which we name actinodin 1 and 2 (And1 and And2), are essential structural components of elastoidin. The presence of actinodin sequences in several teleost fishes and in the elephant shark (Callorhinchus milii, which occupies a basal phylogenetic position), but not in tetrapods, suggests that these genes have been lost during tetrapod species evolution. Double gene knockdown of and1 and and2 in zebrafish embryos results in the absence of actinotrichia and impaired fin folds. Gene expression profiles in embryos lacking and1 and and2 function are consistent with pectoral fin truncation and may offer a potential explanation for the polydactyly observed in early tetrapod fossils. We propose that the loss of both actinodins and actinotrichia during evolution may have led to the loss of lepidotrichia and may have contributed to the fin-to-limb transition.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Grandel, H. & Schulte-Merker, S. The development of the paired fins in the zebrafish (Danio rerio). Mech. Dev. 79, 99–120 (1998)
Mercader, N. Early steps of paired fin development in zebrafish compared with tetrapod limb development. Dev. Growth Differ. 49, 421–437 (2007)
Pizette, S. & Niswander, L. BMPs negatively regulate structure and function of the limb apical ectodermal ridge. Development 126, 883–894 (1999)
Dane, P. J. & Tucker, J. B. Modulation of epidermal cell shaping and extracellular matrix during caudal fin morphogenesis in the zebrafish Brachydanio rerio. J. Embryol. Exp. Morphol. 87, 145–161 (1985)
Wood, A. Early pectoral fin development and morphogenesis of the apical ectodermal ridge in the killifish, Aphysosemion scheeli. Anat. Rec. 204, 349–356 (1982)
Bouvet, J. Différenciation et ultrastructure du squelette distal de la nageoire pectorale chez la truite indigène (Salmo Trutta Fario L.). I. Différenciation et ultrastructure des actinotriches. Arch. Anat. Microsc. Morphol. Exp. 63, 79–96 (1974)
Géraudie, J. Initiation of the actinotrichial development in the early fin bud of the fish, Salmo. J. Morphol. 151, 353–361 (1977)
Wood, A. & Thorogood, P. An analysis of in vitro cell migration during teleost fin morphogenesis. J. Cell Sci. 66, 205–222 (1984)
Garrault, H. Développement des fibres d’élastoidine (actinotrichia) chez les salmonides. Arch. Anat. Microsc. 130, 105–137 (1936)
Krukenberg, C. F. Ueber die chemische Beschaffenheit der sog. Hornfäden von Mustelus und über die Zusammensetzung der keratinösen Hüllen um den Eiern von Scyllium stellate. Mitt. Zool. Stat. Neapel 6, 286–296 (1885)
Damodaran, M., Sivaraman, C. & Dhavalikar, R. S. Amino acid composition of elastoidin. Biochem. J. 62, 621–625 (1956)
Kimura, S. & Kubota, M. Studies on elastoidin I. Some chemical and physical properties of elastoidin and its components. J. Biochem. 60, 615–621 (1966)
Kimura, S., Uematsu, Y. & Miyauchi, Y. Shark (Prionace glauca) elastoidin: characterization of its collagen as [alpha 1(E)]3 homotrimers. Comp. Biochem. Physiol. B 84, 305–308 (1986)
Padhi, B. K. et al. Screen for genes differentially expressed during regeneration of the zebrafish caudal fin. Dev. Dyn. 231, 527–541 (2004)
Seidah, N. G. & Chretien, M. Proprotein and prohormone convertases: a family of subtilases generating diverse bioactive polypeptides. Brain Res. 848, 45–62 (1999)
Andrade, M. A., Perez-Iratxeta, C. & Ponting, C. P. Protein repeats: structures, functions, and evolution. J. Struct. Biol. 134, 117–131 (2001)
Géraudie, J. in Biology of Invertebrate and Lower Vertebrate Collagens (eds Bairati, A. & Garrone, R.) 451–455 (Plenum, 1985)
Nishidate, M., Nakatani, Y., Kudo, A. & Kawakami, A. Identification of novel markers expressed during fin regeneration by microarray analysis in medaka fish. Dev. Dynamics 236, 2685–2693 (2007)
Venkatesh, B. et al. Survey sequencing and comparative analysis of the elephant shark (Callorhinchus milii). PLoS Biol. 5, e101 (2007)
Martin, G. R. The roles of FGFs in the early development of vertebrate limbs. Genes Dev. 12, 1571–1586 (1998)
Ahn, D. & Ho, R. K. Tri-phasic expression of posterior Hox genes during development of pectoral fins in zebrafish: implications for the evolution of vertebrate paired appendages. Dev. Biol. 322, 220–233 (2008)
Sordino, P., van der Hoeven, F. & Duboule, D. Hox gene expression in teleost fins and the origin of vertebrate digits. Nature 375, 678–681 (1995)
Mao, J., McGlinn, E., Huang, P., Tabin, C. J. & McMahon, A. P. Fgf-dependent Etv4/5 activity is required for posterior restriction of Sonic hedgehog and promoting outgrowth of the vertebrate limb. Dev. Cell 16, 600–606 (2009)
Zhang, Z., Verheyden, J. M., Hassel, J. A. & Sun, X. FGF-regulated Etv genes are essential for repressing Shh expression in mouse limb buds. Dev. Cell 16, 607–613 (2009)
Büscher, D., Bosse, B., Heymer, J. & Ruther, U. Evidence for genetic control of Sonic hedgehog by Gli3 in mouse limb development. Mech. Dev. 62, 175–182 (1997)
Zakany, J., Zacchetti, G. & Duboule, D. Interactions between HOXD and Gli3 genes control the limb apical ectodermal ridge via Fgf10. Dev. Biol. 306, 883–893 (2007)
Long, J. A. & Gordon, M. S. The greatest step in vertebrate history: a paleobiological review of the fish-tetrapod transition. Physiol. Biochem. Zool. 77, 700–719 (2004)
Hui, C. C. & Joyner, A. L. A mouse model of Greig cephalopolysyndactyly syndrome: the extra-toesJ mutation contains an intragenic deletion of the Gli3 gene. Nature Genet. 3, 241–246 (1993)
Smith, A. et al. Gene expression analysis on sections of zebrafish regenerating fins reveals limitations in the whole-mount in situ hybridization method. Dev. Dyn. 237, 417–425 (2008)
Choo, B. G. et al. Zebrafish transgenic enhancer TRAP line database (ZETRAP). BMC Dev. Biol. 6, 5 (2006)
Thisse, C. & Thisse, B. High-resolution in situ hybridization to whole-mount zebrafish embryos. Nature Protocols 3, 59–69 (2008)
Tyurina, O. V. et al. Zebrafish Gli3 functions as both an activator and a repressor in Hedgehog signaling. Dev. Biol. 277, 537–556 (2005)
Lang, C. et al. Molecular characterization and developmentally regulated expression of Xenopus lamina-associated polypeptide 2 (XLAP2). J. Cell Sci. 112, 749–759 (1999)
Thummel, R. et al. Inhibition of zebrafish fin regeneration using in vivo electroporation of morpholinos against fgfr1 and msxb. Dev. Dyn. 235, 336–346 (2006)
Avaron, F., Hoffman, L., Guay, D. & Akimenko, M. A. Characterization of two new zebrafish members of the hedgehog family: atypical expression of a zebrafish indian hedgehog gene in skeletal elements of both endochondral and dermal origins. Dev. Dyn. 235, 478–489 (2006)
Acknowledgements
The authors would like to thank I. Duran for technical advice, A. Maurya for assistance in the early stages of this work and M. Ekker, F. Avaron and M. Debiais for discussions and critical reading of the manuscript. L.S.-P. is supported by a European Modular Biology Organization Long-Term Fellowship (ALT 325-2008). This work was supported by grants to M.-A.A. from the Natural Science Engineering and Research Council of Canada and the Canadian Institutes of Health Research. M.A.A.-N. received funding from the Canada Research Chairs programme and from the Helmholtz Alliance on Systems Biology. V.K. received funding from the Agency for Science, Technology and Research of Singapore.
Author information
Authors and Affiliations
Contributions
J.Z., P.W., D.G. and B.K.P. performed the experiments; J.Z. collected and analysed the data; V.K. produced and provided the enhancer-trap transgenic line; M.A.A.-N. and L.S.-P. performed the analysis of the actinodin sequences; and M.-A.A. designed experiments, analysed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-11 with legends and References. (PDF 11241 kb)
Rights and permissions
About this article
Cite this article
Zhang, J., Wagh, P., Guay, D. et al. Loss of fish actinotrichia proteins and the fin-to-limb transition. Nature 466, 234–237 (2010). https://doi.org/10.1038/nature09137
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09137
This article is cited by
-
Gene expression changes during the evolution of the tetrapod limb
Biologia Futura (2022)
-
Spatial regulation by multiple Gremlin1 enhancers provides digit development with cis-regulatory robustness and evolutionary plasticity
Nature Communications (2021)
-
Hoxd13/Bmp2-mediated mechanism involved in zebrafish finfold design
Scientific Reports (2021)
-
ECM alterations in Fndc3a (Fibronectin Domain Containing Protein 3A) deficient zebrafish cause temporal fin development and regeneration defects
Scientific Reports (2019)
-
Expression analysis of And4 during fin regeneration in Misgurnus anguillicaudatus provides insights into its function
Fish Physiology and Biochemistry (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.