Abstract
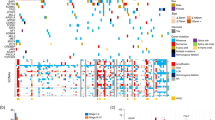
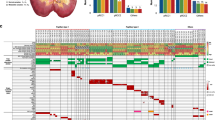
Clear cell renal cell carcinoma (ccRCC) is the most common form of adult kidney cancer, characterized by the presence of inactivating mutations in the VHL gene in most cases1,2, and by infrequent somatic mutations in known cancer genes. To determine further the genetics of ccRCC, we have sequenced 101 cases through 3,544 protein-coding genes. Here we report the identification of inactivating mutations in two genes encoding enzymes involved in histone modification—SETD2, a histone H3 lysine 36 methyltransferase, and JARID1C (also known as KDM5C), a histone H3 lysine 4 demethylase—as well as mutations in the histone H3 lysine 27 demethylase, UTX (KMD6A), that we recently reported3. The results highlight the role of mutations in components of the chromatin modification machinery in human cancer. Furthermore, NF2 mutations were found in non-VHL mutated ccRCC, and several other probable cancer genes were identified. These results indicate that substantial genetic heterogeneity exists in a cancer type dominated by mutations in a single gene, and that systematic screens will be key to fully determining the somatic genetic architecture of cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
Primary accessions
ArrayExpress
Gene Expression Omnibus
Data deposits
The patient and cell line expression data were deposited with the Gene Expression Omnibus and Array Express under accession numbers GSE17895 and E-TABM-770, respectively.
References
Eble, J., Epstein, J. & Sesterhann, I. Pathology and Genetics of Tumours of the Urinary System and Male Genital Organs (IARC Press, 2004)
Rini, B. I., Campbell, S. C. & Escudier, B. Renal cell carcinoma. Lancet 373, 1119–1132 (2009)
van Haaften, G. et al. Somatic mutations of the histone H3K27 demethylase gene UTX in human cancer. Nature Genet. 41, 521–523 (2009)
Banks, R. E. et al. Genetic and epigenetic analysis of von Hippel-Lindau (VHL) gene alterations and relationship with clinical variables in sporadic renal cancer. Cancer Res. 66, 2000–2011 (2006)
Bommi-Reddy, A. et al. Kinase requirements in human cells: III. Altered kinase requirements in VHL-/- cancer cells detected in a pilot synthetic lethal screen. Proc. Natl Acad. Sci. USA 105, 16484–16489 (2008)
Chi, J. T. et al. Gene expression programs in response to hypoxia: cell type specificity and prognostic significance in human cancers. PLoS Med. 3, e47 (2006)
Jones, S. et al. Core signaling pathways in human pancreatic cancers revealed by global genomic analyses. Science 321, 1801–1806 (2008)
Parsons, D. W. et al. An integrated genomic analysis of human glioblastoma multiforme. Science 321, 1807–1812 (2008)
Greenman, C. et al. Patterns of somatic mutation in human cancer genomes. Nature 446, 153–158 (2007)
Sun, X.-J. et al. Identification and characterization of a novel human histone H3 lysine 36-specific methyltransferase. J. Biol. Chem. 280, 35261–35271 (2005)
Iwase, S. et al. The X-linked mental retardation gene SMCX/JARID1C defines a family of histone H3 lysine 4 demethylases. Cell 128, 1077–1088 (2007)
Issaeva, I. et al. Knockdown of ALR (MLL2) reveals ALR target genes and leads to alterations in cell adhesion and growth. Mol. Cell. Biol. 27, 1889–1903 (2007)
Lee, M. G. et al. Demethylation of H3K27 regulates polycomb recruitment and H2A ubiquitination. Science 318, 447–450 (2007)
Ferner, R. E. Neurofibromatosis 1 and neurofibromatosis 2: a twenty first century perspective. Lancet Neurol. 6, 340–351 (2007)
Gordan, J. D., Bertout, J. A., Hu, C.-J., Diehl, J. A. & Simon, M. C. HIF-2α promotes hypoxic cell proliferation by enhancing c-Myc transcriptional activity. Cancer Cell 11, 335–347 (2007)
Zhang, H. et al. HIF-1 inhibits mitochondrial biogenesis and cellular respiration in VHL-deficient renal cell carcinoma by repression of C-MYC activity. Cancer Cell 11, 407–420 (2007)
Mandriota, S. J. et al. HIF activation identifies early lesions in VHL kidneys: Evidence for site-specific tumor suppressor function in the nephron. Cancer Cell 1, 459–468 (2002)
McKinnon, P. J. & Caldecott, K. W. DNA strand break repair and human genetic disease. Annu. Rev. Genomics Hum. Genet. 8, 37–55 (2007)
Kudlow, B. A., Kennedy, B. K. & Monnat, R. J. Werner and Hutchinson-Gilford progeria syndromes: mechanistic basis of human progeroid diseases. Nature Rev. Mol. Cell Biol. 8, 394–404 (2007)
Demuth, I. & Digweed, M. The clinical manifestation of a defective response to DNA double-strand breaks as exemplified by Nijmegen breakage syndrome. Oncogene 26, 7792–7798 (2007)
DeMasi, J., Huh, K.-W., Nakatani, Y., Münger, K. & Howley, P. M. Bovine papillomavirus E7 transformation function correlates with cellular p600 protein binding. Proc. Natl Acad. Sci. USA 102, 11486–11491 (2005)
Huh, K.-W. et al. Association of the human papillomavirus type 16 E7 oncoprotein with the 600-kDa retinoblastoma protein-associated factor, p600. Proc. Natl Acad. Sci. USA 102, 11492–11497 (2005)
Young, A. P. et al. VHL loss actuates a HIF-independent senescence programme mediated by Rb and p400. Nature Cell Biol. 10, 361–369 (2008)
Dicks, E. et al. AutoCSA, an algorithm for high throughput DNA sequence variant detection in cancer genomes. Bioinformatics 23, 1689–1691 (2007)
Greenman, C., Wooster, R., Futreal, P. A., Stratton, M. R. & Easton, D. F. Statistical analysis of pathogenicity of somatic mutations in cancer. Genetics 173, 2187–2198 (2006)
Fuhrman, S. A., Lasky, L. C. & Limas, C. Prognostic significance of morphologic parameters in renal cell carcinoma. Am. J. Surg. Pathol. 6, 655–663 (1982)
AJCC Cancer Staging Manual 7th edn. (eds Edge, S. B. et al.) (Springer, 2009)
Acknowledgements
We would like to acknowledge the Wellcome Trust for support under grant reference 077012/Z/05/Z and the Hauenstein and Gerber Foundations for support for the microarray expression work. We also thank S. Martin, W. McLaughlin and S. Noyes for administrative support and F. Brasseur for providing the matched-pair ccRCC cell lines.
Author Contributions G.L.D. directed the analytical aspects of the study. K.F., K.J.D. and L.C. performed the expression analyses. C.G. contributed statistical analyses. G.B., H.D., S.E., C.H., J.T., A.B., Je.A., S.B., D.B., G.B., P.J.C., S.F., M.J., D.J., H.K., C.Y.K., C.L., M.-L.L., D.J.M., M.M., S.M., K.M., A.M., T.M., Le.M., La.M., S.O., E.P., A.R., Re.S., Ra.S., L.S., P.S., G.T., P.S.T. and K.T. performed the sequencing, copy number, data analyses and provided comments on the manuscript. Jo.A., R.J.K., S.K.K., D.P., B.W. and B.T.T. contributed samples, data and comments on the manuscript. M.R.S. and P.A.F. conceived and directed the study and wrote the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary Data, Supplementary References and Supplementary Figures 1-3 with Legends. (PDF 1823 kb)
Supplementary Dataset 1
Germline variants. (PDF 386 kb)
Supplementary Dataset 2
Gene expression. (PDF 1109 kb)
Supplementary Table 1
This table contains the initial and follow up samples that were screened. (XLS 58 kb)
Supplementary Table 2
This table shows the ID of the genes sequenced. (XLS 333 kb)
Supplementary Table 3
In this table the initial screen somatic mutations are identified. (XLS 227 kb)
Supplementary Table 4
This table shows the mutation prevalence. (PDF 43 kb)
Supplementary Table 5
This table shows the genes sequenced in follow-up samples. (XLS 33 kb)
Supplementary Table 6
In this table the follow-up screen somatic mutations are identified. (XLS 128 kb)
Supplementary Table 7
This table contains the statistical analyses data. (XLS 567 kb)
Supplementary Table 8
This table shows the details of mutations in highlighted genes. (XLS 134 kb)
Supplementary Table 9
In this table the JARID1C, SETD2 cancer cell line screen variants are identified. (XLS 34 kb)
Supplementary Table 10
This table contains SETD2 and JARID1C expression analyses. (XLS 128 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Dalgliesh, G., Furge, K., Greenman, C. et al. Systematic sequencing of renal carcinoma reveals inactivation of histone modifying genes. Nature 463, 360–363 (2010). https://doi.org/10.1038/nature08672
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08672
This article is cited by
-
Lysine demethylase 5C inhibits transcription of prefoldin subunit 5 to activate c-Myc signal transduction and colorectal cancer progression
Molecular Medicine (2024)
-
The emerging roles of histone demethylases in cancers
Cancer and Metastasis Reviews (2024)
-
SETD2 variation correlates with tumor mutational burden and MSI along with improved response to immunotherapy
BMC Cancer (2023)
-
Identification and validation of a DNA methylation-driven gene-based prognostic model for clear cell renal cell carcinoma
BMC Genomics (2023)
-
The histone demethylase KDM5C functions as a tumor suppressor in AML by repression of bivalently marked immature genes
Leukemia (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.