Abstract
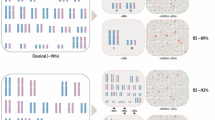
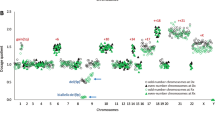
Hyperhaploid clones (24–34 chromosomes) were identified in 33 patients with multiple myeloma (MM), demonstrating a novel numerical cytogenetic subgroup. Strikingly, all hyperhaploid karyotypes were found to harbor monosomy 17p, the single most important risk stratification lesion in MM. A catastrophic loss of nearly a haploid set of chromosomes results in disomies of chromosomes 3, 5, 7, 9, 11, 15, 18, 19 and 21, the same basic set of odd-numbered chromosomes found in trisomy in hyperdiploid myeloma. All other autosomes are found in monosomy, resulting in additional clinically relevant monosomies of 1p, 6q, 13q and 16q. Hypotriploid subclones (58–68 chromosomes) were also identified in 11 of the 33 patients and represent a duplication of the hyperhaploid clone. Analysis of clones utilizing interphase fluorescence in situ hybridization (iFISH), metaphase FISH and spectral karyotyping identified either monosomy 17 or del17p in all patients. Amplification of 1q21 was identified in eight patients, demonstrating an additional high-risk marker. Importantly, our findings indicate that current iFISH strategies may be uninformative or ambiguous in the detection of these clones, suggesting this patient subgroup maybe underreported. Overall survival for patients with hyperhaploid clones was poor, with a 5-year survival rate of 23.1%. These findings identify a distinct numerical subgroup with cytogenetically defined high-risk disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Morgan GJ, Walker BA, Davies FE . The genetic architecture of multiple myeloma. Nat Rev Cancer 2012; 12: 335–348.
Smadja NV, Bastard C, Brigaudeau C, Leroux D, Fruchart C, Groupe Français de Cytogénétique Hématologique. Hypodiploidy is a major prognostic factor in multiple myeloma. Blood 2001; 98: 2229–2238.
Bergsagle PL, Chesi M . Molecular classification and risk stratification of myeloma. Hematol Oncol 2013; 31 (Suppl 1): 38–41.
Shaffer LG, McGowan-Jordan J, Schmid M (eds). ISCN (2013): An International System for Human Cytogenetic Nomenclature. S. Kager: Basel, 2013; pp 88–95.
Mandahl N, Johansson B, Mertens F, Mitelman F . Disease-associated patterns of disomic chromosomes in hyperhaploid neoplasms. Genes Chrom Cancer 2012; 51: 536–544.
Pantou D, Rizou H, Tsarouha H, Pouli A, Papanastasiou K, Stamatellou M et al. Cytogenetic manifestations of multiple myeloma heterogeneity. Genes Chromosomes Cancer 2005; 42: 44–57.
Mohamed AN, Bentley G, Bonnett ML, Zonder J, Al-Katib A . Chromosome aberrations in a series of 120 multiple myeloma cases with abnormal karyotypes. Am J Hematol 2007; 82: 1080–1087.
Gabrea A, Martelli ML, Qi Y, Roschke A, Barlogie B, Shaughnessy JD Jr et al. Secondary genomic rearrangements involving immunoglobulin or MYC loci show similar prevalences in hyperdiploid and nonhyperdiploid myeloma tumors. Genes Chromosomes Cancer 2008; 47: 573–590.
Sawyer JR, Tian E, Thomas E, Koller M, Stangeby C, Sammartino G et al. Evidence for a novel mechanism for gene amplification in multiple myeloma: 1q12 pericentromeric heterochromatin mediates breakage-fusion-bridge cycles of a 1q12-23 amplicon. Br J Haematol 2009; 147: 484–494.
Hoctor VT, Campbell LJ . Hyperhaploid plasma cell myeloma. Cancer Genet 2012; 205: 414–418.
Harrison C, Johansson B Acute lymphoblastic leukemia. In: Heim S, Mitelman F (eds). Cancer Cytogenetics, 3rd edn . Chapter 9. Wiley-Blackwell: Hoboken, NJ, 2009; pp 233–296.
Fonseca R, Bergsagel PL, Drach J, Shaughnessy J, Gutierrez N, Stewart AK et al. International Myeloma Working Group molecular classification of multiple myeloma: spotlight review. Leukemia 2009; 23: 2210–2221.
Avet-Loiseau H, Li C, Magrangeas F, Gouraud W, Charbonnel C, Harousseau JL et al. Prognostic significance of copy-number alterations in multiple myeloma. J Clin Oncol 2009; 27: 4585–4590.
Boyd KD, Ross FM, Chiecchio L, Dagrada GP, Konn ZJ, Tapper WJ et al. A novel prognostic model in myeloma based on co-segregating adverse FISH lesions and the ISS: analysis of patients treated in the MRC Myeloma IX trial. Leukemia 2012; 26: 349–355.
Mikhael JR, Dingli D, Roy V, Reeder CB, Buadi FK, Hayman SR et al. Management of newly diagnosed symptomatic multiple myeloma: updated Mayo Stratification of Myeloma and Risk-Adapted Therapy (mSMART) consensus guidelines 2013. Mayo Clin Proc 2013; 88: 360–376.
Avet-Loiseau H, Durie BG, Cavo M, Attal M, Gutierrez N, Haessler J et al. Combining fluorescent in situ hybridization data with ISS staging improves risk assessment in myeloma: an International Myeloma Working Group collaborative project. Leukemia 2013; 27: 711–717.
Chng WJ, Dispenzieri A, Chim CS, Fonseca R, Goldschmidt H, Lentzsch S et al. IMWG consensus on risk stratification in multiple myeloma. Leukemia 2014; 28: 269–277.
Tian E, Sawyer JR, Heuck CJ, Zhang Q, van Rhee F, Barlogie B et al. In multiple myeloma, 14q32 translocations are nonrandom chromosomal fusions driving high expression levels of the respective partner genes. Genes Chrom Cancer 2014; 53: 549–557.
Sawyer JR, Lukacs JL, Munshi N, Desikan KR, Singhal S, Mehta J et al. Identification of new nonrandom translocations in multiple myeloma with multicolor spectral karyotyping. Blood 1998; 92: 4269–4278.
Kaplan EL, Meier P . Nonparametric estimation from incomplete observations. J Am Stat Assoc 1958; 53: 457–481.
Barlogie B, Anaissie E, van Rhee F, Haessler J, Hollmig K, Pineda-Roman M et al. Incorporating bortezomib into upfront treatment for multiple myeloma: early results of total therapy 3. Br J Haematol 2007; 138: 176–185.
Barlogie B, Mitchell A, van Rhee F, Epstein J, Morgan GJ, Crowley J . Curing myeloma at last: defining criteria and providing the evidence. Blood 2014; 124: 3043–3051.
Shaughnessy JD, Zhan F, Burington BE, Huang Y, Colla S, Hanamura I et al. A validated gene expression model of high-risk multiple myeloma is defined by deregulated expression of genes mapping to chromosome 1. Blood 2007; 109: 2276–2284.
van Buuren S, GroothuIs-Oudshoorn K . Mice: multivariate imputation by chained equations in R. J Stat Softw 2011; 45: 1–67.
Ross FM, Avet-Loiseau H, Ameye G, Gutiérrez NC, Liebisch P, O'Connor S et al. European Myeloma Network. Report from the European Myeloma Network on interphase FISH in multiple myeloma and related disorders. Haematologica 2012; 97: 1272–1277.
An G, Li Z, Tai YT, Acharya C, Li Q, Qin X et al. The impact of clone size on the prognostic value of chromosome aberrations by fluorescence in situ hybridization in multiple myeloma. Clin Cancer Res 2015; 21: 2148–2156.
Gordon DJ, Resio B, Pellman D . Causes and consequences of aneuploidy in cancer. Nat Rev Genet 2012; 13: 189–203.
Krem MM, Press OW, Horwitz MS, Tidwell T . Mechanisms and clinical applications of chromosomal instability in lymphoid malignancy. Br J Haematol 2015; 171: 13–28.
Chng WJ, Ahmann GJ, Henderson K, Santana-Davila R, Greipp PR, Gertz MA et al. Clinical implication of centrosome amplification in plasma cell neoplasm. Blood 2006; 107: 3669–3675.
Safavi S, Forestier E, Golovleva I, Barbany G, Nord KH, Moorman AV et al. Loss of chromosomes is the primary event in near-haploid and low-hypodiploid acute lymphoblastic leukemia. Leukemia 2013; 27: 248–250.
Paulsson K, Morse H, Fioretos T, Behrendtz M, Strombeck B, Johansson B . Evidence for a single-step mechanism in the origin of hyperdiploid childhood acute lymphoblastic leukemia. Genes Chrom Cancer 2005; 44: 113–122.
Magrangeas F, Avet-Loiseau H, Gouraud W, Lodé L, Decaux O, Godmer P et al. Minor clone provides a reservoir for relapse in multiple myeloma. Leukemia 2013; 27: 473–481.
Corre J, Munshi N, Avet-Loiseau H . Genetics of multiple myeloma: another heterogeneity level? Blood 2015; 12: 1870–1876.
Avet-Loiseau H, Soulier J, Fermand JP, Yakoub-Agha I, Attal M, Hulin C et al. Impact of high-risk cytogenetics and prior therapy on outcomes in patients with advanced relapsed or refractory multiple myeloma treated with lenalidomide plus dexaméthasone. Leukemia 2010; 24: 623–628.
Avet-Loiseau H, Attal M, Campion L, Caillot D, Hulin C, Marit G et al. Long-term analysis of the IFM 99 trials for myeloma: cytogenetic abnormalities [t(4;14), del(17p), 1q gains] play a major role in defining long-term survival. J Clin Oncol 2012; 30: 1949–1952.
Stark B, Jeison M, Gobuzov R, Krug H, Glaser-Gabay L, Luria D et al. Near haploid childhood acute lymphoblastic leukemia masked by hyperdiploid line: detection by fluorescence in situ hybridization. Cancer Genet Cytogenet 2001; 128: 108–113.
Chang H, Jiang A, Qi C, Trieu Y, Chen C, Reece D . Impact of genomic aberrations including chromosome 1 abnormalities on the outcome of patients with relapsed or refractory multiple myeloma treated with lenalidomide and dexamethasone. Leuk Lymphoma 2010; 51: 2084–2091.
Boyd KD, Ross FM, Walker BA, Wardell CP, Tapper WJ, Chiecchio L et al. Mapping of chromosome 1p deletions in myeloma identifies FAM46C at 1p12 and CDKN2C at 1p32.3 as being genes in regions associated with adverse survival. Clin Cancer Res 2011; 17: 7776–7784.
Hebraud B, Magrangeas F, Cleynen A, Lauwers-Cances V, Chretien ML, Hulin C et al. Role of additional chromosomal changes in the prognostic value of t(4;14) and del(17p) in multiple myeloma: the IFM experience. Blood 2015; 125: 2095–2100.
Sawyer JR, Tricot G, Mattox S, Jagannath S, Barlogie B . Jumping translocations of chromosome 1q in multiple myeloma: evidence for a mechanism involving decondensation of pericentromeric heterochromatin. Blood 1998; 91: 1732–1741.
Sawyer JR, Tian E, Heuck CJ, Epstein J, Johann DJ, Swanson CM et al. Jumping translocations of 1q12 in multiple myeloma: a novel mechanism for deletion of 17p in cytogenetically defined high-risk disease. Blood 2014; 123: 2504–2512.
Sawyer JR, Tian E, Heuck CJ, Johann DJ, Epstein J, Swanson CM et al. Evidence of an epigenetic origin for high-risk 1q21 copy number aberrations in multiple myeloma. Blood 2015; 125: 3756–3759.
Melchor L, Brioli A, Wardell CP, Murison A, Potter NE, Kaiser MF et al. Single-cell genetic analysis reveals the composition of initiating clones and phylogenetic patterns of branching and parallel evolution in myeloma. Leukemia 2014; 28: 1705–1715.
Moorman AV, Enshaei A, Schwab C, Wade R, Chilton L, Elliott A et al. A novel integrated cytogenetic and genomic classification refines risk stratification in pediatric acute lymphoblastic leukemia. Blood 2014; 124: 1434–1444.
Acknowledgements
We thank the patients and staff of the Myeloma Institute for Research and Therapy. This work was supported in part by PO1 Grant CA 0055819 from the National Cancer Institute.
Author contributions
JRS analyzed and interpreted data and wrote the manuscript. CMS, CS, CLH, LP, ML, GS and JDS analyzed data. ET performed iFISH and JLL performed mFISH and SKY studies. JE, CS and CB provided statistical analysis. MZ, FED, FvR and BB provided patient samples. BB and GJM wrote and reviewed the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
About this article
Cite this article
Sawyer, J., Tian, E., Shaughnessy Jr, J. et al. Hyperhaploidy is a novel high-risk cytogenetic subgroup in multiple myeloma. Leukemia 31, 637–644 (2017). https://doi.org/10.1038/leu.2016.253
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2016.253
This article is cited by
-
Homozygous BCMA gene deletion in response to anti-BCMA CAR T cells in a patient with multiple myeloma
Nature Medicine (2021)
-
Hyperhaploid plasma cell myeloma characterized by poor outcome and monosomy 17 with frequently co-occurring TP53 mutations
Blood Cancer Journal (2019)
-
An acquired high-risk chromosome instability phenotype in multiple myeloma: Jumping 1q Syndrome
Blood Cancer Journal (2019)
-
Evolutionary biology of high-risk multiple myeloma
Nature Reviews Cancer (2017)
-
Multiple myeloma
Nature Reviews Disease Primers (2017)