Abstract
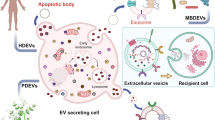
Extracellular vesicles (EV) are small membrane-bound structures that are secreted by various cell types, including tumor cells. Recent studies have shown that EVs are important for cell-to-cell communication, locally and distantly; horizontally transferring DNA, mRNA, microRNA (miRNA), proteins and lipids. In the context of cancer biology, tumor-derived EVs are capable of modifying the microenvironment, promoting tumor progression, immune evasion, angiogenesis and metastasis. miRNAs contained within EVs are functionally associated with cancer progression, metastasis and aggressive tumor phenotypes. These factors, along with their stability in bodily fluids, have led to extensive investigations on the potential role of circulating EV-derived miRNAs as tumor biomarkers. In this review, we summarize the current understanding of circulating EV miRNAs in human cancer, and discuss their clinical utility and challenges in functioning as biomarkers.
Similar content being viewed by others
Introduction
Extracellular vesicles (EV) are small membrane-bound vesicles that are secreted by various cells. EVs have been isolated from numerous cell types, including immune,1, 2, 3 stem,4, 5 nervous system,6, 7 epithelial,8 fibroblast,9 keratinocyte10 and a variety of tumor11 cells. EVs were originally described as vesicles that were released into the extracellular space from multi-vesicular bodies (MVBs) of reticulocytes.12, 13 They were initially considered to be a mechanism by which cells discard unnecessary molecules into the extracellular space.14, 15, 16 However, more recent studies have shown that EVs are an important mechanism by which cells communicate both locally and distantly, by transferring proteins, lipids and nucleic acids including DNA, mRNA and microRNA (miRNA).16 In the context of tumor biology, accumulating evidence suggests that EVs play an important role in communication between tumors and the microenvironment; by transferring EV cargo, tumor cells are able to alter the function of both local and distant normal cells, thereby promoting tumor growth and metastasis.17
miRNAs are small RNA molecules that regulate the expression of protein-coding genes by directly binding to target mRNAs in a sequence-specific manner.18 Bioinformatics algorithms predict that miRNAs regulate >60% of the protein-coding genes in the human genome.19 After the discovery of miRNA transfer between cells via EVs,20, 21, 22 EV miRNAs garnered increasing attention from the biomedical research community. Due to the stability of EV miRNAs in body fluids, and their functional association with tumor progression, circulating EV miRNAs are now extensively investigated for their potential use as cancer biomarkers. In this review, we briefly describe the functional significance of EV miRNAs in cancer biology, summarize circulating EV miRNA biomarker studies for human malignancies, and discuss their utility and limitations as circulating biomarkers.
EV biogenesis
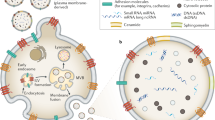
EVs are classified as either exosomes or microvesicles, as a function of their biogenesis. Exosomes are released on MVB fusion with the plasma membrane; whereas microvesicles are released by direct budding from the plasma membrane (Figure 1).16, 23, 24 Exosomes are slightly smaller in diameter (~30–150 nm) and have ‘cup-shaped’ morphology when observed under transmission electron microscopy; on the other hand, microvesicles are more heterogeneous in both size (~100–1000 nm) and shape.24
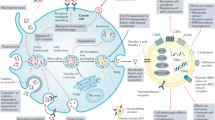
A schematic illustration of exosome and microvesicle biogenesis. Exosome biogenesis begins with the formation of MVBs via early-endosomal membrane invagination to create ILVs. Both ESCRT-dependent and -independent mechanisms are implicated in ILV/MVB formation. Although the majority of MVBs are degraded by fusion with lysosomes, some populations of MVBs fuse with the plasma membrane, releasing exosomes into the extracellular space. Exosome secretion involves RAB GTPases (RAB11, RAB27A/B and RAB35). Microvesicles are released by direct budding from the plasma membrane, and involve ARF1, ARF6 and TSG101. ARF, ADP-ribosylation factor, ESCRT, endosomal-sorting complex required for transport, ILV, intraluminal vesicle, MVB, multi-vesicular body, TSG101, tumor susceptibility gene 101.
Exosome biogenesis begins during the early-endosome maturation process (Figure 1). Early-endosomal membranes invaginate to create intraluminal vesicles (ILVs), thereby forming MVBs. There are several described mechanisms that mediate ILV/MVB formation (reviewed in23). The endosomal-sorting complex required for transport (ESCRT) is the best characterized mechanism, although other ESCRT-independent mechanisms also exist, which require the tetraspanin CD63 or lipid metabolism enzymes, such as neutral sphingomyelinase (nSMase)25 and phospholipase D2 (PLD2).26 The majority of MVBs are degraded by fusion with lysosomes; however, some populations of MVBs migrate towards and fuse with the plasma membrane, releasing their ILVs—now called exosomes—into the extracellular space. Although the mechanisms of exosome secretion have not yet been fully elucidated, RAB GTPases are known to be intricately involved. Savina et al.27 demonstrated that disruption of RAB11 via overexpression of its dominant-negative mutant resulted in inhibition of exosome release from the K562 erythroleukemia cell line. Ostrowski et al.28 demonstrated that knockdown of RAB27A or RAB27B resulted in effective reduction in exosome secretion by performing a shRNA screen for 59 members of the Rab GTPase family in HeLa cells. In another report, inhibition of RAB35 function led to intracellular accumulation of endosomes and impaired exosome secretion.29 Importantly, although RAB11, RAB27A/B and RAB35 play important roles in exosome secretion, the inhibition of just one of these RAB GTPases is insufficient for complete inhibition of exosome secretion.27, 28, 29 This therefore suggests the existence of distinct populations of ILVs and MVBs within the cell, requiring unique machinery for exosome secretion. Indeed, RAB27A inhibition in the mouse mammary carcinoma cell line 4T1 resulted in a decrease in exosome secretion for certain populations (CD63, TSG101, Alix and HSC70), while not affecting the secretion of others (CD9 and MFGE8).30
The biogenesis of microvesicles is less well-characterized (Figure 1). Muralidharan-Chari et al.31 found that the ADP-ribosylation factor 6 (ARF6) GTP/GDP cycle regulated the release of protease-loaded microvesicles from the LOX melanoma cell line. Similarly, ARF1 was found to be involved in the secretion of microvesicles in the MDA-MB-231 breast cancer cell line.32 Nabhan et al.33 demonstrated that TSG101, a component of ESCRT, regulated microvesicle secretion. These authors also showed that interaction of TSG101 with arrestin domain-containing protein 1 (ARRDC1) resulted in relocation of endosomal TSG101 to the plasma membrane, mediating release of microvesicles. Hence, insofar as some mechanisms are emerging in the differential biogenesis of exosomes vs microvesicles this is merely the beginning of an exciting field; future studies will be necessary to reveal further insight into the mechanism of microvesicular biogenesis.
EV isolation methods
A number of EV isolation methods have been developed for the purification of exosomes and microvesicles from body fluids and/or cell culture media. The most commonly used procedure employs a series of differential centrifugation steps.34 Following removal of cells and cell debris using low-speed centrifugations (300g and 2000g), the supernatant is spun at 10 000g to pellet microvesicles. Exosomes are subsequently pelleted from the supernatant via ultracentrifugation (100 000g).34 Although this protocol is straightforward and frequently used, contaminants (for example, protein aggregates) can also be co-precipitated. To overcome this problem, an additional density gradient centrifugation purification step is often combined with the differential centrifugation. Exosomes are then isolated from a buoyant density of 1.08–1.22 g ml−1 on sucrose or iodoxanol (OptiPrep, St Louis, MO, USA) gradients.1
An alternate EV purification method is immunoaffinity isolation using antibody-coated magnetic beads, which allows for the selection of a more restricted population of exosome. This method has been used for the isolation of HER2-positive exosomes from the culture media of breast cancer cell lines, exosomes from the ascites of a patient with advanced ovarian cancer35 and EpCAM-positive exosomes from the serum of patients with lung or ovarian cancer.36, 37 A common challenge with immunoaffinity isolation is the identification of exosome-specific markers. Recently, polymer-based exosome precipitation solutions have been developed (for example, ExoQuick, System Biosciences, Palo Alto, CA, USA) and widely used as a simple and rapid exosome isolation method, resulting in high yields of proteins and RNAs.38 It is unclear however, whether these polymer-based methods are capable of isolating pure exosomes.
To date, there are no reliable methods for purifying and discriminating between exosomes or microvesicles. Therefore, careful consideration of EV isolation methods must be taken into account when interpreting study results.
Composition of EVs
EVs contain many biomolecules, including proteins, lipids, DNA, mRNAs and non-coding RNAs, such as miRNAs.39 Among these, proteins are the best characterized EV cargo. The membrane protein composition of microvesicles resembles the parental cell more closely than that of exosomes.16 Exosomes, on the other hand, are enriched with components of endosomes and proteins found in the parental cell that are involved in MVB formation, such as tetraspanins (CD9, CD63 and CD81), Alix, flotillin, TSG101 and RAB GTPases.23 Mass spectrometry studies have shown that exosomes are enriched with proteins from the cytosol and plasma membrane, but lack proteins from the nucleus, endoplasmic reticulum, Golgi and mitochondria.40, 41 Some of the protein contents from EVs overlap between exosomes and microvesicles, thus limiting the distinction between EV types based on an individual protein.23, 42 Recent studies have used a combination of several protein markers to distinguish exosomes from microvesicles. The International Society for Extracellular Vesicles (ISEV) suggested that to claim the successful isolation of exosomes, a combination of at least three protein markers must be used,43 including: (a) the presence of transmembrane or lipid-bound extracellular proteins (CD9, CD63, CD81, cell adhesion molecules, growth factor receptors, heterotrimeric G proteins, integrins or MFGE8); (b) the presence of cytosolic proteins (TSG101, annexins, RAB GTPases or syntenin); and (c) the absence of intracellular proteins (from endoplasmic reticulum, Golgi, mitochondria, nucleus or Argonaute/RNA-induced silencing complex (RISC)).43
RNA is another biomolecule that is carried within EVs. Valadi et al.44 first identified the presence of mRNAs and miRNAs within the exosomes from human and mouse mast cells; these exosomes were markedly stable following treatment with RNase or trypsin. Notably, the authors also found that the mRNA contained within the mouse exosomes could be internalized into human mast cells, resulting in the production of protein from these mouse mRNAs in the recipient cells, suggesting that exosome mRNA remains functional and can be translated in other cells. This was the first identification of gene-based communication between mammalian cells. Subsequently, bioanalyzer analyses showed that exosomes and microvesicles have distinct RNA profiles.45, 46 Exosomal RNAs are generally enriched for small RNAs, including miRNAs, and lack ribosomal RNA. Interestingly, the RNA expression profiles are distinct between exosomes and their producer cells. Ohshima et al.47 found high exosomal let-7 family expression in the metastatic gastric cancer cell line AZ-P7a, but lower exosomal expression in several cell lines with higher intracellular let-7a expression. Lunavat et al.45 compared the RNA profiles between cells and different EV populations in the melanoma cell line MML-1 using next generation RNA sequencing. The authors found that out of 252 miRNAs that were detected, 113 miRNAs were shared between the EV sub-types and parental cells, whereas 23 miRNAs were detected exclusively in the exosomes. These observations suggest that cells have sorting mechanisms allowing for preferential secretion of specific miRNAs into the exosomes. Several mechanisms of miRNA sorting into exosomes have been reported (reviewed in48) such as nSMase2-dependent pathway,20 sumoylated heterogeneous nuclear ribonucleoprotein (hnRNP)-dependent,49 3′-end of the miRNA sequence-dependent,50 as well as the RISC-related pathways.51
EV miRNAs in cancer
EVs play important roles in physiological processes, such as immune system regulation, blood coagulation, stem cell and nervous system maintenance, as well as in pathologic states such as cancer.16, 17, 23 In the context of cancer biology, numerous studies have shown that tumor-derived EVs are capable of modifying the microenvironment to facilitate tumor progression, angiogenesis and metastasis.
Angiogenesis is essential for tumor development and metastasis, and tumor-derived EV miRNAs affect endothelial cells to promote this process.52, 53, 54 Exosomal miR-92a, derived from the leukemia cell line K562, was found to be transferred into endothelial cells, resulting in enhanced endothelial cell migration and tube formation.52 In addition, exosomal miR-135b, derived from hypoxia-resistant melanoma cells, enhanced endothelial tube formation under hypoxia by targeting factor-inhibiting hypoxia-inducible factor 1 (FIH-1).53 Zhuang et al.54 showed that SK23 melanoma cell-derived miRNAs, including miR-9, were transferred to endothelial cells largely via EVs, which they demonstrated using a transwell tumor–endothelial cell co-culture system. Functional studies showed that SK23-derived EVs led to enhanced migration in endothelial cells; this effect was attenuated by anti-miR-9 transfection, suggesting that EV-mediated miR-9 transfer was important for angiogenesis in this context.
Tumor-derived EV miRNAs have been shown to enhance metastasis by modifying the tumor microenvironment and promoting mesenchymal-to-epithelial transition (MET). Fong et al.55 identified that breast cancer cell-secreted EV miR-122 facilitated tumor metastasis by modifying glucose metabolism in the pre-metastatic niche environment. Zhou et al.56 showed that breast cancer-secreted EV miR-105 induced vascular permeability in endothelial cells by targeting the cellular tight junctions. Mice developed more metastases when pretreated with EVs secreted by the highly metastatic MDA-MB-231 cells, which contained high levels of miR-105. In both studies, therapeutic administration of anti-miR-122 or anti-miR-105, respectively, with tumor-derived EV injection resulted in suppression of brain and lung metastasis. MET is important for the population of metastatic sites, and the miR-200 family is a well-known mediator of this process.57 Le et al.58 showed that EV miR-200 conferred the ability to colonize lung metastatic sites via promotion of MET, using a highly metastatic breast cancer model.
EV miRNAs as cancer biomarkers
Given the role of tumor-derived EV miRNAs in tumor progression and metastasis, it is logical to investigate the role of EV miRNAs as biomarkers. EVs are known to exist in many types of bodily fluids, including blood,59 urine,60 saliva,61 breast milk,62 amniotic,63 ascites64 and cerebrospinal fluids.65 EV miRNAs have been primarily examined for their presence in the plasma and serum of cancer patients for diagnostic or prognostic purposes. A summary of circulating EV miRNAs identified in various cancer types, detected using either miRNA profiling studies or individual miRNA expression studies are shown in Tables 1 and 2, respectively. Important caveats for these Tables and in making comparisons include: (a) the varying terms used to define EVs; (b) sample type from which the EVs were derived (that is, plasma or serum); and (c) methodological differences in EV isolation, miRNA profiling and expression normalization.
The ISEV suggests the use of plasma rather than serum as a source for EV RNA for the purpose of biomarker studies,66 as platelets release EVs in serum during clot formation, which may account for over 50% of EVs in serum.67 However, a review of the literature indicates that two thirds of all circulating EV studies used serum as the EV source (Tables 1 and 2). Serum may be the traditional choice for source material for EV studies, as a significantly higher yield of EVs is obtained from serum.68
As indicated previously, no perfect method exists for the identification and purification of exosomes or microvesicles, and current methods are likely to yield diverse populations of EVs. Interestingly, only one third of studies adopted the ‘gold standard’ method of differential centrifugation for EV isolation. Instead, polymer-based EV precipitation solutions, such as ExoQuick and Total Exosome Isolation Reagent (Invitrogen, Carlsbad, CA, USA), were widely used, accounting for almost two thirds of these studies. The use of these methods is likely due to their simpler, less time consuming procedures, which might render polymer-based EV isolation methods more feasible options in the clinical setting. However, the purity of the polymer-precipitated EVs is yet to be proven, and these preparations likely do not exclude co-precipitation of other circulating miRNA carriers, such as Ago2 proteins69 and high-density lipoproteins.70
Although only two studies (from the same group) described the use of the immunoaffinity isolation method with magnetic-activated cell sorting, the data produced from these studies are promising.36, 37 Taylor et al.37 isolated EpCAM-positive EVs from patient plasma, and observed that miR-21, miR-141, miR-200a, miR-200c, miR-200b, miR-203, miR-205 and miR-214 were all significantly elevated in ovarian cancer patients compared with patients with benign tumors. The authors also showed that the protein yield of EpCAM-positive exosomes was clearly correlated with tumor stage. In cases where tumor-specific EV markers are available, immunoaffinity isolation may be a useful method for isolation of tumor-derived EV miRNAs. For example, glypican-1 (GPC1) was recently identified as a tumor-specific EV surface marker in pancreatic cancer.71 In a colorectal cancer study, CD147 was found to be an optimal EV surface marker to detect tumor-derived EVs.72 These specific EV molecular studies yield promising data for the development of an optimal EV isolation method for tumor-specific EVs.
Following EV isolation, miRNA expression may be profiled via quantitative real-time PCR-based arrays, hybridization-based arrays or next generation sequencing (Table 1). Current normalization methods are varied, but can be generally categorized into three types: internal control, spike-in miRNA or global mean normalization. Internal control normalization is a standard method for miRNA expression normalization in tissues. However, it is challenging to identify a reliable internal miRNA/small RNA control for EV studies, due to the limited consensus regarding a consistent miRNA/small RNA in EVs. Cazzoli et al.73 used let-7a as an internal control to normalize lung adenocarcinoma plasma EV miRNA expression profiles, identified following an examination of the expression of five candidate miRNAs in a training cohort. In a hepatocellular carcinoma serum exosome miRNA study, it was shown that a combination of miR-221, miR-191, let-7a, miR-181a and miR-26a was optimal for liver-specific miRNA normalization.74 Although miR-16, miR-451, miR-484 and U6 are often used as internal controls due to their use in whole plasma or serum studies, it remains unclear whether these miRNAs are suitable for EV miRNA normalization. Externally spiked-in miRNA is frequently used for normalization by adding a set amount of a unique miRNA species (for example, cel-miR-39, cel-miR-54 or ath-miR159a) into the sample prior to RNA isolation. The miRNA expression data derived from samples normalized using spike-in miRNA produces an expression value that is absolute rather than relative; thus the resulting miRNA profile may reflect biological relevance more precisely. However, as this method does not normalize the processes prior to RNA isolation, variation during EV isolation cannot be normalized. The third option for normalization of EV miRNA expression is the global mean normalization method, which uses the mean expression value of all miRNAs as a control, with the assumption that mean expression levels of detectable miRNAs are consistent if input amounts remain constant between samples.75 The primary advantages of this method are that it controls for external factors that lead to variability in miRNA expression due to sample processing, and it does not rely on a specific miRNA for normalization. The limitation of this method is that it may only be applicable to genome-wide miRNA profiling studies; therefore alternative normalization methods are still required for validation studies.
As a result of the wide-ranging methods for EV isolation, miRNA profiling and normalization, the resulting circulating EV miRNA expression signatures are highly variable between studies, even within a single cancer type (Tables 1 and 2). However, specific miRNAs were consistently deregulated among many of these studies, including the miR-200 family in ovarian cancer,37, 76 and miR-375 in prostate cancer.77, 78 Whole serum (not EV-specific) studies demonstrated that the miR-200 family of miRNAs were upregulated in the serum from ovarian cancer patients, which in turn correlated with increased tumor aggressiveness.79 Upregulation of miR-375 in the whole serum (not EV-specific) of prostate cancer patients was also frequently reported.80, 81, 82 These studies potentially indicate that the miR-200 family and miR-375 are promising EV-derived biomarkers, warranting further functional validation.
Our review of the literature revealed that miR-21 was upregulated in EVs from different kinds of cancers, including colorectal,83 lung,36 melanoma,84 ovarian,37 esophageal,85 hepatocellular carcinoma,86 laryngeal,87 pancreatic88 and prostate77 cancers (Tables 1 and 2). miR-21 is known to function as an onco-miR, downregulating several tumor suppressor genes such as phosphatase and tensin homolog (PTEN),89 programmed cell death 4 (PDCD4)90 and tropomyosin 1 (TPM1).91 Upregulation of miR-21 in tumor tissue has been shown to be a poor prognostic factor in numerous cancers.92 Although miR-21 is not cancer-type specific, due to its frequent association with malignancies, this miRNA could be a candidate prognostic biomarker in circulating EVs for certain cancer types.
Conclusions
EVs are secreted from numerous cell types, including tumor cells, and have been identified as an important mechanism by which cells communicate, via the transfer of DNA, mRNA, miRNA, proteins and lipids. Tumor-derived EVs impact the local and distal environment, aiding in tumor progression, angiogenesis and metastasis. EV-derived miRNAs, which are highly stable in bodily fluids, offer significant promise as circulating biomarkers for assessing tumor aggressiveness, and have been implicated in numerous human malignancies. However, current methods for examining tumor-derived EV miRNA biomarkers are highly variable. A standardized method of EV isolation and miRNA expression assessment and normalization would enable a more reliable inter-study validation of EV miRNAs as biomarkers. Nonetheless, the functional implication of EV-derived miRNAs in cancer, and the ability to detect tumor-derived EV miRNAs in plasma and serum, renders them as highly promising candidates for future application in the clinical setting.
References
Raposo, G., Nijman, H. W., Stoorvogel, W., Liejendekker, R., Harding, C. V., Melief, C. J. et al. B lymphocytes secrete antigen-presenting vesicles. J. Exp. Med. 183, 1161–1172 (1996).
Zitvogel, L., Regnault, A., Lozier, A., Wolfers, J., Flament, C., Tenza, D. et al. Eradication of established murine tumors using a novel cell-free vaccine: dendritic cell-derived exosomes. Nat. Med. 4, 594–600 (1998).
Chaput, N. & Thery, C. Exosomes: immune properties and potential clinical implementations. Semin. Immunopathol. 33, 419–440 (2011).
Ratajczak, J., Miekus, K., Kucia, M., Zhang, J., Reca, R., Dvorak, P. et al. Embryonic stem cell-derived microvesicles reprogram hematopoietic progenitors: evidence for horizontal transfer of mRNA and protein delivery. Leukemia 20, 847–856 (2006).
Lai, R. C., Yeo, R. W. & Lim, S. K. Mesenchymal stem cell exosomes. Semin. Cell Dev. Biol. 40, 82–88 (2015).
Faure, J., Lachenal, G., Court, M., Hirrlinger, J., Chatellard-Causse, C., Blot, B. et al. Exosomes are released by cultured cortical neurones. Mol. Cell. Neurosci. 31, 642–648 (2006).
Budnik, V., Ruiz-Canada, C. & Wendler, F. Extracellular vesicles round off communication in the nervous system. Nat. Rev. Neurosci. 17, 160–172 (2016).
Van Niel, G., Raposo, G., Candalh, C., Boussac, M., Hershberg, R., Cerf–Bensussan, N. et al. Intestinal epithelial cells secrete exosome–like vesicles. Gastroenterology 121, 337–349 (2001).
Lancaster, G. I. & Febbraio, M. A. Exosome-dependent trafficking of HSP70: a novel secretory pathway for cellular stress proteins. J. Biol. Chem. 280, 23349–23355 (2005).
Chavez-Munoz, C., Morse, J., Kilani, R. & Ghahary, A. Primary human keratinocytes externalize stratifin protein via exosomes. J. Cell. Biochem. 104, 2165–2173 (2008).
Mears, R., Craven, R. A., Hanrahan, S., Totty, N., Upton, C., Young, S. L. et al. Proteomic analysis of melanoma-derived exosomes by two-dimensional polyacrylamide gel electrophoresis and mass spectrometry. Proteomics 4, 4019–4031 (2004).
Pan, B. T. & Johnstone, R. M. Fate of the transferrin receptor during maturation of sheep reticulocytes in vitro: selective externalization of the receptor. Cell 33, 967–978 (1983).
Harding, C., Heuser, J. & Stahl, P. Endocytosis and intracellular processing of transferrin and colloidal gold-transferrin in rat reticulocytes: demonstration of a pathway for receptor shedding. Eur. J. Cell Biol. 35, 256–263 (1984).
Lerner, M. P., Lucid, S. W., Wen, G. J. & Nordquist, R. E. Selected area membrane shedding by tumor cells. Cancer Lett. 20, 125–130 (1983).
Ratajczak, M. Z. & Ratajczak, J. Horizontal transfer of RNA and proteins between cells by extracellular microvesicles: 14 years later. Clin. Transl. Med. 5, 7 (2016).
Andaloussi, S. E., Mager, I., Breakefield, X. O. & Wood, M. J. Extracellular vesicles: biology and emerging therapeutic opportunities. Nat. Rev. Drug Discov. 12, 347–357 (2013).
Tkach, M. & Thery, C. Communication by extracellular vesicles: where we are and where we need to go. Cell 164, 1226–1232 (2016).
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Friedman, R. C., Farh, K. K., Burge, C. B. & Bartel, D. P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 19, 92–105 (2009).
Kosaka, N., Iguchi, H., Yoshioka, Y., Takeshita, F., Matsuki, Y. & Ochiya, T. Secretory mechanisms and intercellular transfer of microRNAs in living cells. J. Biol. Chem. 285, 17442–17452 (2010).
Pegtel, D. M., Cosmopoulos, K., Thorley-Lawson, D. A., van Eijndhoven, M. A., Hopmans, E. S., Lindenberg, J. L. et al. Functional delivery of viral miRNAs via exosomes. Proc. Natl Acad. Sci. USA 107, 6328–6333 (2010).
Zhang, Y., Liu, D., Chen, X., Li, J., Li, L., Bian, Z. et al. Secreted monocytic miR-150 enhances targeted endothelial cell migration. Mol. Cell 39, 133–144 (2010).
Colombo, M., Raposo, G. & Thery, C. Biogenesis, secretion, and intercellular interactions of exosomes and other extracellular vesicles. Annu. Rev. Cell Dev. Biol. 30, 255–289 (2014).
Raposo, G. & Stoorvogel, W. Extracellular vesicles: exosomes, microvesicles, and friends. J. Cell Biol. 200, 373–383 (2013).
Trajkovic, K., Hsu, C., Chiantia, S., Rajendran, L., Wenzel, D., Wieland, F. et al. Ceramide triggers budding of exosome vesicles into multivesicular endosomes. Science 319, 1244–1247 (2008).
Ghossoub, R., Lembo, F., Rubio, A., Gaillard, C. B., Bouchet, J., Vitale, N. et al. Syntenin-ALIX exosome biogenesis and budding into multivesicular bodies are controlled by ARF6 and PLD2. Nat. Commun. 5, 3477 (2014).
Savina, A., Vidal, M. & Colombo, M. I. The exosome pathway in K562 cells is regulated by Rab11. J. Cell Sci. 115, 2505–2515 (2002).
Ostrowski, M., Carmo, N. B., Krumeich, S., Fanget, I., Raposo, G., Savina, A. et al. Rab27a and Rab27b control different steps of the exosome secretion pathway. Nat. Cell Biol. 12, 19–30 (2010).
Hsu, C., Morohashi, Y., Yoshimura, S., Manrique-Hoyos, N., Jung, S., Lauterbach, M. A. et al. Regulation of exosome secretion by Rab35 and its GTPase-activating proteins TBC1D10A-C. J. Cell Biol. 189, 223–232 (2010).
Bobrie, A., Colombo, M., Krumeich, S., Raposo, G. & Thery, C. Diverse subpopulations of vesicles secreted by different intracellular mechanisms are present in exosome preparations obtained by differential ultracentrifugation. J. Extracell. Vesicles 1, 18397 (2012).
Muralidharan-Chari, V., Clancy, J., Plou, C., Romao, M., Chavrier, P., Raposo, G. et al. ARF6-regulated shedding of tumor cell-derived plasma membrane microvesicles. Curr. Biol. 19, 1875–1885 (2009).
Schlienger, S., Campbell, S. & Claing, A. ARF1 regulates the Rho/MLC pathway to control EGF-dependent breast cancer cell invasion. Mol. Biol. Cell 25, 17–29 (2014).
Nabhan, J. F., Hu, R., Oh, R. S., Cohen, S. N. & Lu, Q. Formation and release of arrestin domain-containing protein 1-mediated microvesicles (ARMMs) at plasma membrane by recruitment of TSG101 protein. Proc. Natl Acad. Sci. USA 109, 4146–4151 (2012).
Thery, C., Amigorena, S., Raposo, G. & Clayton, A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr. Protoc. Cell Biol. Chapter 3 Unit 3.22 (2006).
Koga, K., Matsumoto, K., Akiyoshi, T., Kubo, M., Yamanaka, N., Tasaki, A. et al. Purification, characterization and biological significance of tumor-derived exosomes. Anticancer Res. 25, 3703–3707 (2005).
Rabinowits, G., Gercel-Taylor, C., Day, J. M., Taylor, D. D. & Kloecker, G. H. Exosomal microRNA: a diagnostic marker for lung cancer. Clin. Lung Cancer 10, 42–46 (2009).
Taylor, D. D. & Gercel-Taylor, C. MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol. Oncol. 110, 13–21 (2008).
Saenz-Cuesta, M., Arbelaiz, A., Oregi, A., Irizar, H., Osorio-Querejeta, I., Munoz-Culla, M. et al. Methods for extracellular vesicles isolation in a hospital setting. Front. Immunol. 6, 50 (2015).
Kalra, H., Simpson, R. J., Ji, H., Aikawa, E., Altevogt, P., Askenase, P. et al. Vesiclepedia: a compendium for extracellular vesicles with continuous community annotation. PLoS Biol. 10, e1001450 (2012).
Thery, C., Regnault, A., Garin, J., Wolfers, J., Zitvogel, L., Ricciardi-Castagnoli, P. et al. Molecular characterization of dendritic cell-derived exosomes. Selective accumulation of the heat shock protein hsc73. J. Cell Biol. 147, 599–610 (1999).
Thery, C., Boussac, M., Veron, P., Ricciardi-Castagnoli, P., Raposo, G., Garin, J. et al. Proteomic analysis of dendritic cell-derived exosomes: a secreted subcellular compartment distinct from apoptotic vesicles. J. Immunol. 166, 7309–7318 (2001).
Turiak, L., Misjak, P., Szabo, T. G., Aradi, B., Paloczi, K., Ozohanics, O. et al. Proteomic characterization of thymocyte-derived microvesicles and apoptotic bodies in BALB/c mice. J. Proteomics 74, 2025–2033 (2011).
Lotvall, J., Hill, A. F., Hochberg, F., Buzas, E. I., Di Vizio, D., Gardiner, C. et al. Minimal experimental requirements for definition of extracellular vesicles and their functions: a position statement from the International Society for Extracellular Vesicles. J. Extracell. Vesicles 3, 26913 (2014).
Valadi, H., Ekstrom, K., Bossios, A., Sjostrand, M., Lee, J. J. & Lotvall, J. O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 9, 654–659 (2007).
Lunavat, T. R., Cheng, L., Kim, D. K., Bhadury, J., Jang, S. C., Lasser, C. et al. Small RNA deep sequencing discriminates subsets of extracellular vesicles released by melanoma cells—Evidence of unique microRNA cargos. RNA Biol. 12, 810–823 (2015).
Crescitelli, R., Lasser, C., Szabo, T. G., Kittel, A., Eldh, M., Dianzani, I. et al. Distinct RNA profiles in subpopulations of extracellular vesicles: apoptotic bodies, microvesicles and exosomes. J. Extracell. Vesicles 2, 20677 (2013).
Ohshima, K., Inoue, K., Fujiwara, A., Hatakeyama, K., Kanto, K., Watanabe, Y. et al. Let-7 microRNA family is selectively secreted into the extracellular environment via exosomes in a metastatic gastric cancer cell line. PLoS ONE 5, e13247 (2010).
Zhang, J., Li, S., Li, L., Li, M., Guo, C., Yao, J. et al. Exosome and exosomal microRNA: trafficking, sorting, and function. Genomics Proteomics Bioinformatics 13, 17–24 (2015).
Villarroya-Beltri, C., Gutierrez-Vazquez, C., Sanchez-Cabo, F., Perez-Hernandez, D., Vazquez, J., Martin-Cofreces, N. et al. Sumoylated hnRNPA2B1 controls the sorting of miRNAs into exosomes through binding to specific motifs. Nat. Commun. 4, 2980 (2013).
Koppers-Lalic, D., Hackenberg, M., Bijnsdorp, I. V., van Eijndhoven, M. A., Sadek, P., Sie, D. et al. Nontemplated nucleotide additions distinguish the small RNA composition in cells from exosomes. Cell Rep 8, 1649–1658 (2014).
Guduric-Fuchs, J., O’Connor, A., Camp, B., O’Neill, C. L., Medina, R. J. & Simpson, D. A. Selective extracellular vesicle-mediated export of an overlapping set of microRNAs from multiple cell types. BMC Genomics 13, 357 (2012).
Umezu, T., Ohyashiki, K., Kuroda, M. & Ohyashiki, J. H. Leukemia cell to endothelial cell communication via exosomal miRNAs. Oncogene 32, 2747–2755 (2013).
Umezu, T., Tadokoro, H., Azuma, K., Yoshizawa, S., Ohyashiki, K. & Ohyashiki, J. H. Exosomal miR-135b shed from hypoxic multiple myeloma cells enhances angiogenesis by targeting factor-inhibiting HIF-1. Blood 124, 3748–3757 (2014).
Zhuang, G., Wu, X., Jiang, Z., Kasman, I., Yao, J., Guan, Y. et al. Tumour-secreted miR-9 promotes endothelial cell migration and angiogenesis by activating the JAK-STAT pathway. EMBO J. 31, 3513–3523 (2012).
Fong, M. Y., Zhou, W., Liu, L., Alontaga, A. Y., Chandra, M., Ashby, J. et al. Breast-cancer-secreted miR-122 reprograms glucose metabolism in premetastatic niche to promote metastasis. Nat. Cell Biol. 17, 183–194 (2015).
Zhou, W., Fong, M. Y., Min, Y., Somlo, G., Liu, L., Palomares, M. R. et al. Cancer-secreted miR-105 destroys vascular endothelial barriers to promote metastasis. Cancer Cell 25, 501–515 (2014).
Feng, X., Wang, Z., Fillmore, R. & Xi, Y. MiR-200, a new star miRNA in human cancer. Cancer Lett. 344, 166–173 (2014).
Le, M. T., Hamar, P., Guo, C., Basar, E., Perdigao-Henriques, R., Balaj, L. et al. miR-200-containing extracellular vesicles promote breast cancer cell metastasis. J. Clin. Invest. 124, 5109–5128 (2014).
Caby, M. P., Lankar, D., Vincendeau-Scherrer, C., Raposo, G. & Bonnerot, C. Exosomal-like vesicles are present in human blood plasma. Int. Immunol. 17, 879–887 (2005).
Pisitkun, T., Shen, R. F. & Knepper, M. A. Identification and proteomic profiling of exosomes in human urine. Proc. Natl Acad. Sci. USA 101, 13368–13373 (2004).
Ogawa, Y., Miura, Y., Harazono, A., Kanai-Azuma, M., Akimoto, Y., Kawakami, H. et al. Proteomic analysis of two types of exosomes in human whole saliva. Biol. Pharm. Bull. 34, 13–23 (2011).
Admyre, C., Johansson, S. M., Qazi, K. R., Filen, J. J., Lahesmaa, R., Norman, M. et al. Exosomes with immune modulatory features are present in human breast milk. J. Immunol. 179, 1969–1978 (2007).
Asea, A., Jean-Pierre, C., Kaur, P., Rao, P., Linhares, I. M., Skupski, D. et al. Heat shock protein-containing exosomes in mid-trimester amniotic fluids. J. Reprod. Immunol. 79, 12–17 (2008).
Andre, F., Schartz, N. E., Movassagh, M., Flament, C., Pautier, P., Morice, P. et al. Malignant effusions and immunogenic tumour-derived exosomes. Lancet 360, 295–305 (2002).
Vella, L. J., Sharples, R. A., Lawson, V. A., Masters, C. L., Cappai, R. & Hill, A. F. Packaging of prions into exosomes is associated with a novel pathway of PrP processing. J. Pathol. 211, 582–590 (2007).
Witwer, K. W., Buzas, E. I., Bemis, L. T., Bora, A., Lasser, C., Lotvall, J. et al. Standardization of sample collection, isolation and analysis methods in extracellular vesicle research. J. Extracell. Vesicles 2, 20360 (2013).
Gemmell, C. H., Sefton, M. V. & Yeo, E. L. Platelet-derived microparticle formation involves glycoprotein IIb-IIIa. Inhibition by RGDS and a Glanzmann’s thrombasthenia defect. J. Biol. Chem. 268, 14586–14589 (1993).
George, J. N., Thoi, L. L., McManus, L. M. & Reimann, T. A. Isolation of human platelet membrane microparticles from plasma and serum. Blood 60, 834–840 (1982).
Arroyo, J. D., Chevillet, J. R., Kroh, E. M., Ruf, I. K., Pritchard, C. C., Gibson, D. F. et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl Acad. Sci. USA 108, 5003–5008 (2011).
Vickers, K. C., Palmisano, B. T., Shoucri, B. M., Shamburek, R. D. & Remaley, A. T. MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat. Cell Biol. 13, 423–433 (2011).
Melo, S. A., Luecke, L. B., Kahlert, C., Fernandez, A. F., Gammon, S. T., Kaye, J. et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature 523, 177–182 (2015).
Yoshioka, Y., Kosaka, N., Konishi, Y., Ohta, H., Okamoto, H., Sonoda, H. et al. Ultra-sensitive liquid biopsy of circulating extracellular vesicles using ExoScreen. Nat. Commun. 5, 3591 (2014).
Cazzoli, R., Buttitta, F., Di Nicola, M., Malatesta, S., Marchetti, A., Rom, W. N. et al. microRNAs derived from circulating exosomes as noninvasive biomarkers for screening and diagnosing lung cancer. J. Thorac. Oncol. 8, 1156–1162 (2013).
Li, Y., Zhang, L., Liu, F., Xiang, G., Jiang, D. & Pu, X. Identification of endogenous controls for analyzing serum exosomal miRNA in patients with hepatitis B or hepatocellular carcinoma. Dis. Markers 2015, 893594 (2015).
Mestdagh, P., Van Vlierberghe, P., De Weer, A., Muth, D., Westermann, F., Speleman, F. et al. A novel and universal method for microRNA RT-qPCR data normalization. Genome Biol. 10, R64 (2009).
Meng, X., Muller, V., Milde-Langosch, K., Trillsch, F., Pantel, K. & Schwarzenbach, H. Diagnostic and prognostic relevance of circulating exosomal miR-373, miR-200a, miR-200b and miR-200c in patients with epithelial ovarian cancer. Oncotarget 7, 16923–16935 (2016).
Li, M., Rai, A. J., DeCastro, G. J., Zeringer, E., Barta, T., Magdaleno, S. et al. An optimized procedure for exosome isolation and analysis using serum samples: application to cancer biomarker discovery. Methods 87, 26–30 (2015).
Huang, X., Yuan, T., Liang, M., Du, M., Xia, S., Dittmar, R. et al. Exosomal miR-1290 and miR-375 as prognostic markers in castration-resistant prostate cancer. Eur. Urol. 67, 33–41 (2015).
Zuberi, M., Mir, R., Das, J., Ahmad, I., Javid, J., Yadav, P. et al. Expression of serum miR-200a, miR-200b, and miR-200c as candidate biomarkers in epithelial ovarian cancer and their association with clinicopathological features. Clin. Transl. Oncol. 17, 779–787 (2015).
Haldrup, C., Kosaka, N., Ochiya, T., Borre, M., Hoyer, S., Orntoft, T. F. et al. Profiling of circulating microRNAs for prostate cancer biomarker discovery. Drug Deliv. Transl. Res. 4, 19–30 (2014).
Nguyen, H. C., Xie, W., Yang, M., Hsieh, C. L., Drouin, S., Lee, G. S. et al. Expression differences of circulating microRNAs in metastatic castration resistant prostate cancer and low-risk, localized prostate cancer. Prostate 73, 346–354 (2013).
Wach, S., Al-Janabi, O., Weigelt, K., Fischer, K., Greither, T., Marcou, M. et al. The combined serum levels of miR-375 and urokinase plasminogen activator receptor are suggested as diagnostic and prognostic biomarkers in prostate cancer. Int. J. Cancer 137, 1406–1416 (2015).
Ogata-Kawata, H., Izumiya, M., Kurioka, D., Honma, Y., Yamada, Y., Furuta, K. et al. Circulating exosomal microRNAs as biomarkers of colon cancer. PLoS ONE 9, e92921 (2014).
Pfeffer, S. R., Grossmann, K. F., Cassidy, P. B., Yang, C. H., Fan, M., Kopelovich, L. et al. Detection of exosomal miRNAs in the plasma of melanoma patients. J. Clin. Med. 4, 2012–2027 (2015).
Tanaka, Y., Kamohara, H., Kinoshita, K., Kurashige, J., Ishimoto, T., Iwatsuki, M. et al. Clinical impact of serum exosomal microRNA-21 as a clinical biomarker in human esophageal squamous cell carcinoma. Cancer 119, 1159–1167 (2013).
Wang, H., Hou, L., Li, A., Duan, Y., Gao, H. & Song, X. Expression of serum exosomal microRNA-21 in human hepatocellular carcinoma. Biomed. Res. Int. 2014, 864894 (2014).
Wang, J., Zhou, Y., Lu, J., Sun, Y., Xiao, H., Liu, M. et al. Combined detection of serum exosomal miR-21 and HOTAIR as diagnostic and prognostic biomarkers for laryngeal squamous cell carcinoma. Med. Oncol. 31, 148 (2014).
Que, R., Ding, G., Chen, J. & Cao, L. Analysis of serum exosomal microRNAs and clinicopathologic features of patients with pancreatic adenocarcinoma. World J. Surg. Oncol. 11, 219 (2013).
Meng, F., Henson, R., Wehbe-Janek, H., Ghoshal, K., Jacob, S. T. & Patel, T. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology 133, 647–658 (2007).
Lu, Z., Liu, M., Stribinskis, V., Klinge, C. M., Ramos, K. S., Colburn, N. H. et al. MicroRNA-21 promotes cell transformation by targeting the programmed cell death 4 gene. Oncogene 27, 4373–4379 (2008).
Zhu, S., Si, M. L., Wu, H. & Mo, Y. Y. MicroRNA-21 targets the tumor suppressor gene tropomyosin 1 (TPM1). J. Biol. Chem. 282, 14328–14336 (2007).
Wang, W., Li, J., Zhu, W., Gao, C., Jiang, R., Li, W. et al. MicroRNA-21 and the clinical outcomes of various carcinomas: a systematic review and meta-analysis. BMC Cancer 14, 819 (2014).
Yeh, Y. Y., Ozer, H. G., Lehman, A. M., Maddocks, K., Yu, L., Johnson, A. J. et al. Characterization of CLL exosomes reveals a distinct microRNA signature and enhanced secretion by activation of BCR signaling. Blood 125, 3297–3305 (2015).
Matsumura, T., Sugimachi, K., Iinuma, H., Takahashi, Y., Kurashige, J., Sawada, G. et al. Exosomal microRNA in serum is a novel biomarker of recurrence in human colorectal cancer. Br. J. Cancer 113, 275–281 (2015).
Chiam, K., Wang, T., Watson, D. I., Mayne, G. C., Irvine, T. S., Bright, T. et al. Circulating serum exosomal mirnas as potential biomarkers for esophageal adenocarcinoma. J. Gastrointest. Surg. 19, 1208–1215 (2015).
Manterola, L., Guruceaga, E., Gallego Perez-Larraya, J., Gonzalez-Huarriz, M., Jauregui, P., Tejada, S. et al. A small noncoding RNA signature found in exosomes of GBM patient serum as a diagnostic tool. Neuro Oncol. 16, 520–527 (2014).
Sugimachi, K., Matsumura, T., Hirata, H., Uchi, R., Ueda, M., Ueo, H. et al. Identification of a bona fide microRNA biomarker in serum exosomes that predicts hepatocellular carcinoma recurrence after liver transplantation. Br. J. Cancer 112, 532–538 (2015).
Rodriguez, M., Silva, J., Lopez-Alfonso, A., Lopez-Muniz, M. B., Pena, C., Dominguez, G. et al. Different exosome cargo from plasma/bronchoalveolar lavage in non-small-cell lung cancer. Genes Chromosomes Cancer 53, 713–724 (2014).
Madhavan, B., Yue, S., Galli, U., Rana, S., Gross, W., Muller, M. et al. Combined evaluation of a panel of protein and miRNA serum-exosome biomarkers for pancreatic cancer diagnosis increases sensitivity and specificity. Int. J. Cancer 136, 2616–2627 (2015).
Eichelser, C., Stuckrath, I., Muller, V., Milde-Langosch, K., Wikman, H., Pantel, K. et al. Increased serum levels of circulating exosomal microRNA-373 in receptor-negative breast cancer patients. Oncotarget 5, 9650–9663 (2014).
Zhou, X., Zhu, W., Li, H., Wen, W., Cheng, W., Wang, F. et al. Diagnostic value of a plasma microRNA signature in gastric cancer: a microRNA expression analysis. Sci. Rep. 5, 11251 (2015).
Shi, R., Wang, P. Y., Li, X. Y., Chen, J. X., Li, Y., Zhang, X. Z. et al. Exosomal levels of miRNA-21 from cerebrospinal fluids associated with poor prognosis and tumor recurrence of glioma patients. Oncotarget 6, 26971–26981 (2015).
Sohn, W., Kim, J., Kang, S. H., Yang, S. R., Cho, J. Y., Cho, H. C. et al. Serum exosomal microRNAs as novel biomarkers for hepatocellular carcinoma. Exp. Mol. Med. 47, e184 (2015).
Alegre, E., Sanmamed, M. F., Rodriguez, C., Carranza, O., Martin-Algarra, S. & Gonzalez, A. Study of circulating microRNA-125b levels in serum exosomes in advanced melanoma. Arch. Pathol. Lab. Med. 138, 828–832 (2014).
Li, Z., Ma, Y. Y., Wang, J., Zeng, X. F., Li, R., Kang, W. et al. Exosomal microRNA-141 is upregulated in the serum of prostate cancer patients. Onco Targets Ther. 9, 139–148 (2016).
Ragusa, M., Barbagallo, C., Statello, L., Caltabiano, R., Russo, A., Puzzo, L. et al. miRNA profiling in vitreous humor, vitreal exosomes and serum from uveal melanoma patients: Pathological and diagnostic implications. Cancer Biol. Ther. 16, 1387–1396 (2015).
Acknowledgements
TK was supported by the Uehara Memorial Foundation Research Fellowship, and the Mochida Memorial Foundation for Medical and Pharmaceutical Research Fellowship. This work was supported by funds from the Canadian Institutes of Health Research; the Dr Mariano Elia Chair in Head and Neck Cancer Research; the Campbell Family Institute for Cancer Research; and the Ministry of Health and Long-Term Care. We also gratefully acknowledge the support from the Princess Margaret Cancer Center Head & Neck Translational Program, with philanthropic funds from the Wharton Family, Joe’s Team and Gordon Tozer.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Kinoshita, T., Yip, K., Spence, T. et al. MicroRNAs in extracellular vesicles: potential cancer biomarkers. J Hum Genet 62, 67–74 (2017). https://doi.org/10.1038/jhg.2016.87
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2016.87
This article is cited by
-
Non-destructive and efficient method for obtaining miRNA information in cells by artificial extracellular vesicles
Scientific Reports (2023)
-
Stability of exosomes in the postmortem serum and preliminary study on exosomal miRNA expression profiling in serum from myocardial infarction cadavers
International Journal of Legal Medicine (2023)
-
Protein and miRNA profile of circulating extracellular vesicles in patients with primary sclerosing cholangitis
Scientific Reports (2022)
-
Urinary exosome microRNA signatures as a noninvasive prognostic biomarker for prostate cancer
npj Genomic Medicine (2021)
-
Modulations of obesity-related microRNAs after exercise intervention: a systematic review and bioinformatics analysis
Molecular Biology Reports (2021)