Abstract
During microbial reductive dechlorination of tetrachloroethene (PCE) and trichloroethene (TCE), trans-1, 2-dichloroethene (trans-DCE) has been observed to be produced predominantly by certain mixed and pure cultures. However, the reductive dehalogenase (RDase) genes involved in trans-DCE generation remain elusive. In this study, identification and transcriptional analysis of RDases were conducted on trans-DCE-producing Dehalococcoides sp. strain MB. Two pairs of degenerate primers targeting the conserved regions of RDases in known Dehalococcoides species were applied to amplify the putative RDase genes of strain MB. Cloning and restriction analysis revealed the presence of seven unique RDase gene fragments (dceA1 to dceA7) that possess sequence identity to known RDase genes. Gene expression analysis of the PCE-grown culture MB exhibited 10-fold regulation of the RDase gene dceA6 (designated mbrA gene), suggesting that it is involved in the production of trans-DCE. This is in agreement with the molecular size of the most abundant protein that is resolved on the denaturing protein gel. Complete sequence of the mbrA gene was obtained by chromosome walking, and the upstream of it is a regulator of transcription, indicating that the expression of this functional gene is tightly controlled in the microbe. The mbrA gene was subsequently found to be present in other trans-DCE-producing cultures containing Dehalococcoides sp. The new mbrA gene identified in this study may serve as an important biomarker for evaluating, predicting and elucidating the biological production of trans-DCE in the chloroethene-contaminated sites.
Similar content being viewed by others
Introduction
Tetrachloroethene (PCE) and trichloroethene (TCE) remain a critical environmental concern because of their poor disposal practices and accidental releases in various dry-cleaning and degreasing industries (Abelson, 1990; ATSDR, 1997a, 1997b). The released chloroethenes often contaminate groundwater, which in turn threatens drinking water supplies (Westrick et al., 1984). Under anaerobic conditions, PCE and TCE can be reductively dechlorinated to the less chlorinated ethenes cis-1, 2-dichloroethene (cis-DCE) and vinyl chloride (VC) by a variety of dechlorinators or to the harmless ethene by Dehalococcoides species (Holliger et al., 1998; Löffler et al., 2000; He et al., 2003; Luijten et al., 2003; Müller et al., 2004; Smidt and de Vos, 2004; He et al., 2005; Hiraishi, 2008). However, in PCE-/TCE-contaminated sites, dechlorination often terminates at DCEs and VC. Although cis-DCE is the major DCE isomer produced by the currently characterized dechlorinators, reductive dechlorination of PCE or TCE to trans-1, 2-dichloroethene (trans-DCE) has also been observed in enrichment cultures and the pure culture of Dehalococcoides sp. strain MB (Löffler et al., 1997; Griffin et al., 2004; Miller et al., 2005; Kittelmann and Friedrich, 2008; Cheng et al., 2010; Cheng and He, 2009). Although reports on the generation of trans-DCE by microbial dechlorination are relatively fewer than studies on cis-DCE production, the microbial generation of trans-DCE is equally important because of its recalcitrant nature. The accumulation of trans-DCE at contaminated sites poses a serious problem, as it is known to be persistent and to disperse at the subsurface. Therefore, the remediation of chloroethenes remains a challenging task.
A number of PCE-to-cis-DCE reductive dehalogenases (RDases) and their encoding gene sequences have been characterized to understand the generation of cis-DCE, such as RDases in Dehalospirullum multivorans (Neumann et al., 1996, 1998), Dehalobacter restrictus (Schumacher et al., 1997; Maillard et al., 2003), Desulfitobacterium sp. strain PCE-S (Miller et al., 1998), Clostridium bifermentans (Okeke et al., 2001) and Dehalococcoides ethenogenes strain 195 (Magnuson et al., 1998; Magnuson et al., 2000). Besides these PCE-RDases (PceA), Dehalococcoides species possess additional RDases that catalyze the dechlorination of DCEs to VC and ethene. For example, D. ethenogenes 195 contains an enzyme TceA, which is able to catalyze dechlorination of TCE and DCEs to VC and ethene (Magnuson et al., 2000). Two other VC dechlorinating RDases, VcrA and BvcA, distinguish Dehalococcoides sp. strains VS and BAV1 from other strains of the same genus (Krajmalnik-Brown et al., 2004; Müller et al., 2004). The uniqueness of dehalorespiring bacteria is that all the dehalogenation reactions are catalyzed by RDases. RDases are characterized to have a molecular weight between 45–65 kDa when subjected to SDS-polyacrylamide gel electrophoresis, and to contain a corrinoid factor and iron sulfur (Fe4S4) clusters (Maillard et al., 2003). Sequence similarity to other proteins is limited to the C-terminal part of the enzyme, which consists of the twin arginine translocation signal sequence and two Fe4S4 binding motifs with four conserved cysteines in each of the Fe4S4 clusters (Berks, 1996; Sticht and Rösch, 1998). As this class of RDases shares common features both at the biochemical and genetic levels, the sequence information of the known RDases is used for the design of degenerate primers to identify new RDase genes in this study.
To date, Dehalococcoides sp. strain MB is reported to be capable of reducing PCE/TCE to trans-DCE predominantly (Cheng et al., 2010; Cheng and He, 2009). Microarray analysis reveals that the genes of strains MB and 195 are highly conserved, reflected by the fact that the genomic DNA of MB hybridizes with 1389 out of the 1579 (88%) probe sets on the microarray targeting genes of strain 195 and 10 of them are RDase genes (Cheng and He, 2009). Little is known about the novel RDases and their transcription in the trans-DCE-producing culture MB. Therefore, the aim of this study is to investigate the genetic make-up of RDase genes and to provide insights on their protein subunits in the trans-DCE-producing culture MB. Expression analysis of multiple RDase genes of strain MB will help to elucidate how the microbes catalyze dechlorination of PCE/TCE to the predominant trans-DCE and will provide an enzymatic basis of trans-DCE production in contaminated sites.
Materials and methods
Chemicals
The chlorinated ethenes, universal vectorette system and other chemicals were purchased from Sigma-Aldrich (Saint Louis, MO, USA). H2 was obtained from a hydrogen generator (Schmidlin-DBS, AG, Neuheim, Switzerland). Lysozyme was purchased from Invitrogen (Carlsbad, CA, USA), and luciferase control RNA was obtained from Promega (Madison, WI, USA).
Culture and growth conditions
The trans-DCE-producing culture MB was grown in acetate (5 mM)-amended mineral salts medium spiked with 0.5 mM of PCE and a vitamin solution (Wolin et al., 1963) comprising 0.05 mg l−1 of vitamin B12 (He et al., 2007). H2 was added as the electron donor (5 00 000 p.p.m.v by volume). All bottles were inverted and incubated in the dark at 30 °C without shaking. Dechlorination activity was monitored by a gas chromatograph (GC-6890, Agilent, Wilmington, DE, USA) equipped with a flame ionizing detector as previously described (Cheng and He, 2009).
Genomic DNA extraction and construction of DNA clone library
Genomic DNA was extracted from the cell pellet of 1.0 ml trans-DCE-producing culture MB with Qiagen DNeasy Blood and Tissue Kit (QIAGEN GmbH, Hilden, Germany) as described in the manufacturer's instructions, except that 45 μl of proteinase K (25 mg ml−1), 20 μl of lysozyme (100 mg ml−1) and 10 μl of achromopeptidase (25 mg ml−1) were added to improve cell lysis. The extracted genomic DNA was then used as a template for the amplification of putative RDase genes with two pairs of degenerate primers (Table 1). In this study, primer pair 2 (RDH F1C and RDH R1C) was designed to cover ∼90% of the known RDases detected in Dehalococcoides spp. The subsequent construction of clone libraries and screening of major clone types were conducted as previously described (Hölscher et al., 2004).
RNA extraction and construction of cDNA clone library
Total RNA was extracted from the cell pellet of 1.5 ml pure culture MB by using the RNeasy extraction kit (QIAGEN GmbH) according to the manufacturer's instructions, except that the tubes holding the cells received 100 μl of 0.1 mm zirconia-silica beads (Biospec Products, Bartlesville, OK, USA) and were bead-beaten for 2 min to improve cell lysis.
Reverse transcription was carried out by using the two-step reverse transcription-PCR Sensiscript kit (QIAGEN GmbH). First, the corresponding cDNAs were synthesized by incubating 20–50 ng of extracted RNA in a 20 μl reaction mixture at 37 °C for 3 h. Then, the generated cDNAs were used as template for the amplification of RDase genes with degenerate primer pairs 1 and 2, respectively, as shown in Table 1. Two cDNA clone libraries were then established with the resulting amplicons using the TOPO-TA cloning kit (Invitrogen, Carlsbad, CA, USA). Screening and determination of the most abundant RDase genes were conducted by enzyme digestion using the HhaI and MspI enzymes (New England Biolabs, Ipswich, MA, USA), followed by visualization of digested products on 2.5% agarose gel (Metaphor agarose, FMC BioProducts, Rockland, ME, USA) after being stained with ethidium bromide.
Sequence analysis of RDase genes
The seven dominant RDase gene inserts in the cDNA clone library were sequenced with an ABI 3100 Sequencer (Applied Biosystems, Foster City, CA, USA) using primers M13F-20 (5′-GTAAAACGACGGCCAGT-3′) and M13R-24 (5′-GGAAACAGCTATGACCATG-3′) (Invitrogen). Five of the RDase genes (dceA1 to dceA5) required an additional internal primer to be designed (dceA1S to dceA5S) to obtain a complete sequence for each of the gene inserts (Table 1). Sequences were assembled and compared by using the BioEdit assembly software (http://www.mbio.ncsu.edu/BioEdit/bioedit.html) (Hall, 1999) and BLASTX search tool, respectively. The deduced amino-acid sequences of the identified putative RDase genes were obtained with the TRANSLATE program (http://us.expasy.org/tools/dna.html) (Geneva, Switzerland). Phylogenetic trees were constructed with the online MAFFT program (http://align.genome.jp/mafft/) (Katoh et al., 2002). FASTA program was used to compare the identity among the predicted amino-acid sequences of the identified putative RDases (http://fasta.bioch.virginia.edu/fasta_www2/fasta_www.cgi) (Pearson and Lipman, 1988).
Transcriptional analysis by quantitative real-time PCR
Gene expression analysis of the seven identified RDases in strain MB was conducted on an ABI 7500 fast quantitative real-time PCR (qPCR) (Applied Biosystems) by using SYBR green assays and their respective primers (primer pairs 3–9) (Table 1). Luciferase control RNA (Promega, Madison, WI, USA) was added as an internal reference transcript for mRNA losses during RNA isolation, reverse transcription and quantification (Johnson et al., 2005), and rpoB was included in this study as a housekeeping gene (a positive control) for all qPCR assays. The housekeeping gene, rpoB, was chosen to serve as an indicator of the basal metabolic activity of the bacterial cells because it is highly conserved among Dehalococcoides spp. The transcriptional levels of RDase genes were normalized against the housekeeping gene, rpoB, before comparing with their respective Dehalococcoides 16S rRNA gene copies. SYBR green dye bound to all amplified double-stranded DNA during qPCR reactions and the fluorescently tagged DNA in turn would be detected by the qPCR system. The specificity of such an assay was ensured by the use of the stringent HotStar Taq DNA polymerase (QIAGEN GmbH) as well as the inclusion of a melt curve analysis at the end of the entire amplification process. Reactions were performed in 20-μl volumes containing 10 μl of QuantiTect SYBR green PCR master mix (QIAGEN GmbH), 1.2 μl of each primer (5 pmol μl−1), 6.6 μl of distilled water, and 1 μl of template DNA or cDNA. The thermocycling program was as follows: an initial step of 15 min at 95 °C, followed by 40 cycles of 15 s at 94 °C and 30 s at 55–59 °C depending on the primer annealing temperature (Table 1), and then 30 s at 72 °C. Fluorescence data were collected after each elongation step. To create qPCR standard curves, PCR products of individual genes amplified with the primers specified in Table 1 were ligated into the pCR2.1 vector (Invitrogen) and transformed into Escherichia coli TOP10 chemically competent cells (Invitrogen). The plasmids were obtained by using the QIAprep Spin Miniprep kit (QIAGEN GmbH). A calibration curve was obtained by using serial dilutions of known plasmid DNA concentrations. The qPCR experiments were carried out in triplicate along with appropriate controls (reference luciferase mRNA and housekeeping rpoB controls).
Genome walking of the RDase genes
The upstream DNA sequences of the three most abundant genes (dceA1, dceA6 and dceA7) were mapped using the universal vectorette system (Sigma-Genosys, Saint Louis, MO, USA). A HindIII vectorette library was constructed using genomic DNA of strain MB according to the manufacturer's protocol. Gene-specific primers (VFs or VRs in Table 1) were designed to be used with the universal vectorette primers for the primary and secondary PCR. In addition to the above PCR reactions, three other reaction mixtures were set up as controls, which contained (1) the respective VF primer only, (2) the respective VR primer only and (3) the vectorette primer only. All amplified PCR products were then resolved on a 1% agarose gel (Invitrogen). Bands unique to the reaction that contained both vectorette primer and either VF or VR primer as compared with the above three controls were deemed as positive bands as either primer alone may prime/misprime at two sites generating a product. These positive bands were then gel purified with Nucleospin extract II kit (Macherey-Nagel, Düren, Germany) in accordance with the manufacturer's instructions. The gel-purified PCR products were then sequenced using the respective dceA gene-specific VF or VR primers and universal sequencing primers provided by the kit.
Gel electrophoresis, in-gel activity test and in-gel digestion for MALDI-TOF MS analysis
Cells from 300 ml of active culture MB were harvested by centrifugation at 23 000 × g for 20 min at 4 °C. The resulting cell pellet was resuspended in 3 ml of buffer 1 (Adrian et al., 2007) and sonicated for 5 s with a 15 s cooling interval between each pulse on a Vibra Cell Sonicator (Sonics and Materials, Danbury, CT, USA). Cell disruption by sonication was carried out for a total of 10 min and kept in ice-cold slurry. The cell lysate was centrifuged at 10 000 × g for 30 min at 4 °C. The supernatant-containing cell extract was then collected and resolved on a native polyacrylamide gel. The positions of the respective excised protein bands were determined by the method as described by Adrian et al. (2007). Gel fragments were used for in-gel activity assays and the dechlorination activity of PCE was conducted as previously described (Neumann et al., 2007). The protein band identified to show dechlorination activity was eluted and concentrated for a second gel electrophoresis step (Adrian et al., 2007).
Denaturing SDS-polyacrylamide gel with a 10% resolving and 4% stacking gel were prepared as described by Läemmli (1970). The denaturing protein gels were run at 200 V for 2 h under aerobic conditions. After electrophoresis, the gel was stained with BioSafe Coomassie Stain (Bio-Rad Laboratories, Hercules, CA, USA).
The desired gel band was then excised using a scalpel and ground using a clean spatula. Tryptic gel digestion and the subsequent purification of digested protein fragments were performed as previously described (http://www.dbs.nus.edu.sg/research/facilities/ppc/index.htm). The digested peptides were analyzed with a Micromass Q-TOF Tandem mass spectrometer (Micromass, Manchester, UK). The map of peptide mass fingerprint generated was compared with the primary sequence database, MASCOT (Matrix Science, Boston, MA, USA).
Nucleotide accession number
The coding sequences of the putative RDase genes and putative B gene fragments were deposited in GenBank under accession numbers EU625398-625402 (dceA1-5) and GU120392 (dceA7). GenBank accession number GU120391 was assigned to the complete sequence of the identified trans-DCE-producing RDase gene, dceA6 (designated the mbrA gene).
Results
Identification of RDase genes in culture MB
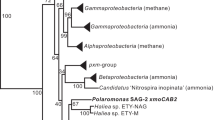
Two pairs of degenerate primers (RRF2–B1R, and RDH F1C–RDH R1C) were used to target the genomic DNA of strain MB. The primer pair 1 was designed on the basis of 17 RDase genes in the genome of strain 195 to target the conserved twin arginine motif (for RRF2) and WYEY motif (for B1R), which was used to identify the bvcA gene in strain BAV1 (Krajmalnik-Brown et al., 2004). However, the reverse primer B1R does not amplify tceA-like genes or other similar RDase genes, as it targets the conserved WYEY motif of the associated anchoring protein of nine RDase genes in strain 195 (Krajmalnik-Brown et al., 2004). Hence, to complement the RRF2-B1R primer pair, a second set of primers was designed according to the conserved twin arginine motif (for forward primer RDH F1C) and the conserved PIDD motif (for the reverse primer RDH R1C) in the RDase domain of Dehalococcoides species. When targeting the genomic DNA of strain MB, amplicons of expected size of 1.5–1.7 kb were obtained with primer pair 1 (RRF2 and B1R), whereas ∼1.2 kb gene fragments were amplified by primer pair 2 (RDH F1C–RDH R1C). The amplicons were used to establish two clone libraries. Restriction fragment length polymorphism analysis of 120 clones from each clone library revealed seven distinct clone types, designated dceA1 to dceA7, of which dceA1, dceA6 and dceA7 are the most abundant ones. The same RDase gene profiles were observed in all the DNA clone libraries established with DNA of enriched MB cultures and DNA of pure strain MB, indicating the high fidelity of the presence of RDases in these cultures. After sequencing the seven clone inserts, BLASTX analysis shows that strain MB shares three highly similar RDase genes (dceA3, 4 and 7) with those of strain 195 (95–100% identity), three (dceA1, 2 and 5) with those of strain VS (54–97% identity), and one (dceA6) with that of strain CBDB1 (98% identity). None of the seven identified RDase genes of MB are similar to the TCE RDase genes in Dehalococcoides (for example, strains 195 and FL2). Phylogenetic analysis of the RDase genes in strain MB and other Dehalococcoides using MAFFT software generated phylogenetic trees that are similar in topology to previous studies (Hölscher et al., 2004; Waller et al., 2005) (Figure 1).
A similarity analysis was carried out with the translated amino-acid sequences of the RDase genes in strain MB by using protein software (TRANSLATE). Alignment of the seven amino-acid sequences (DceA1 to 7) by ClustalW shows that they all share the twin arginine translocation signal peptide, RRXFXK. The seven RDases share an amino-acid identity ranging from 28.8% to 68.8% when compared with each other (Table 2). The two most similar proteins are DceA6 (from primer pair 1 clone library) and DceA7 (from primer pair 2 clone library), sharing 72.1% amino-acid similarity. However, the presence of an early termination codon in the RDase domain of DceA7 distinguishes it from the other six putative RDase genes, though it shares 100% homology with the RDase gene DET 0088 in strain 195.
According to the seven RDase gene sequences of strain MB (Table 1), primer pairs 3–9 were designed and specific bands were observed in the gel picture when targeted on the genomic DNA of culture MB (data not shown).
Transcriptional analysis of RDase genes in strain MB
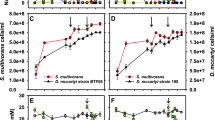
The seven identified RDase genes were detected at DNA level in strain MB; however, the transcription of DNA to copies of mRNA might not occur for all seven genes. Therefore, transcriptional analysis was conducted to find out the functional RDase gene(s) among the seven identified ones in strain MB on exposure to PCE. After starving the MB cells for 72 h, three parallel cultures were spiked with PCE (0.22 mM of each). Total RNA was extracted from cultures at different time points, followed by reverse transcription and subsequent qPCR with the primer pairs designed specifically for the RDase genes (primers 3–9 in Table 1). At time zero, negligible amounts of cDNAs were detected by qPCR. After 72 h, dceA6 and dceA7 genes expressed eight- and fivefold, whereas dceA6 gene reached its highest expression of 10-fold when compared with the 16S rRNA gene copy numbers at 120 h (Figure 2). The expressions of the other five identified RDase genes remained relatively low (less than twofold augmentation of gene copies) when compared with the 16S rRNA gene copies, suggesting little or no upregulation of the respective RDase genes. However, the gel picture of PCR showed positive bands when the genomic DNA of culture MB was targeted with the gene-specific primers, suggesting that these five genes were present in the genome of MB but were not expressing themselves (data not shown). Furthermore, the extracted RNA samples were not contaminated with genomic DNA, as confirmed by no visible amplicons from the amplification of RNA samples with the seven RDase gene-specific primers. Therefore, two (dceA6 and dceA7) out of the seven identified RDase genes are actively expressed in strain MB when fed with PCE.
Genome walking of dceA6 and dceA7 genes
Two new pairs of sequencing primers (Table 1) were designed for genome walking to obtain sequence information on the 5′ and 3′ flanking regions of the expressed RDase genes dceA6 and dceA7. Restriction enzymes, HindIII and RsaI, were chosen as unique base cutters for the extension of dceA6 and dceA7 genes, respectively. Amplification of the ligated DNA with the designed primer and the universal vectorette primer resulted in amplicon sizes of approximately 0.6 and 1.2 kb for the dceA6 gene and 0.8 and 2 kb for the dceA7 gene. Finally, complete sequences (2.4 and 1.9 kb) were obtained for the RDase genes, dceA6 and dceA7 (Figure 3), and deposited in GenBank with accession numbers GU120391and GU120392, respectively. The upstream region of dceA6 (∼300 bp in length) was revealed to share 100% identity with the DNA-binding response regulator protein of CBDB1, and the dceA6 gene is 98% similar to the putative RDase gene (cbdbA80) of Dehalococcoides sp. strain CBDB1. For the dceA7 gene, its upstream region shares 91% identity with the radical S-adenosylmethionine (SAM) domain-containing protein of strain 195, whereas the dceA7 RDase domain shares 100% identity with DET0088 of strain 195. Alignment of RDase gene sequences reveals that different groups of regulatory systems associate with the detected RDases. In addition to dceA6 and dceA7, the full-length sequence of the dceA1 (Cheng et al., 2010) gene was also obtained in this study. Comparison of the dceA1 and dceA6 genes shows that both RDases have an associated anchoring protein at the 3′ end (Figure 3).
Arrangements of the potential trans-1, 2-dichloroethene-producing reductive dehalogenase genes and their corresponding anchoring protein. Also shown are the conserved dehalogenase features including the signal peptide RRDFMK, and two iron-sulfur (Fe4S4) clusters near the C-terminal end for mbrA (dceA6).
Protein banding and in-gel dechlorination assay
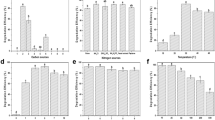
From the Swiss-Prot (Geneva, Switzerland) software prediction, the size of the DceA6 RDase protein subunit is ∼50 kDa. Protein extracts of culture MB fed with PCE were analyzed by native gel electrophoresis. A clear and reproducible gel pattern with one dominant band was observed on the Coomassie blue-stained native gel (Figure 4). Dechlorination activity of PCE to trans-DCE was detected after 24 h of incubating the corresponding unstained gel fragment. This protein band was subsequently eluted and concentrated for a second denaturing gel electrophoresis to verify the protein size. SDS-polyacrylamide gel electrophoresis analysis revealed that the size of the distinct band is ∼50 kDa, which is located in the 36–55 kDa range of the pre-stained protein marker. MALDI-TOF MS analysis of the predominant protein mass of this band was found to be most similar to cbdbA80, a putative RDase of 592 amino-acids (equivalent to 1778 bp of nucleic acids) in strain CBDB1. RDase gene cbdbA80 shows highest identity (98%) to the dceA6 gene of strain MB. Besides being the most abundant protein in strain MB, the dceA6 gene expressed 10-fold after exposure to PCE, suggesting that dceA6 is the RDase gene involved in trans-DCE production. The dceA6 gene is designated the mbrA gene based on the name of strain MB.
Detection of the mbrA (dceA6) gene in various trans-DCE-producing cultures
Gene-specific primers, mbrAF (dce6AF) and mbrAR (dce6AR), were designed to determine whether the highly expressed mbrA (dceA6) gene could be an appropriate biomarker for distinguishing trans-DCE-producing cultures from other dechlorinating cultures generating a negligible amount of trans-DCE. The presence of the mbrA gene was tested on genomic DNA extracted from non-trans-DCE-producing Dehalococcoides isolate ANAS1 (Cheng and He, 2009) (negative control) as well as various trans-DCE-producing mixed cultures. The presence of DNA was first verified by detecting 16S rRNA genes in all the cultures. Results show the presence of the mbrA gene in 5 out of 10 Dehalococcoides-containing cultures that generated trans-DCE from reductive dechlorination of PCE/TCE (Figure 5). However, the other five Dehalococcoides trans-DCE-generating cultures do not contain the mbrA gene, suggesting the presence of the other trans-DCE-producing RDase genes in those cultures. The absence of the mbrA gene in the non-Dehalococcoides trans-DCE-producing cultures suggests the correlation of this newly identified RDase gene with the presence of the Dehalococcoides species. Specificity of the primer pairs, mbrAF and mbrAR, was further confirmed by a negative signal when targeting the genomic DNA of non-trans-DCE-producing cultures (for example, ANAS1) (Figure 5).
(a) Genus-specific primers targeting the 16S rRNA genes of Dehalococcoides sp. (620 bp). (b) Detection of the trans-1, 2-dichloroethene (trans-DCE)-producing mbrA gene (427 bp) in various cultures. Lane 1: 100 bp DNA ladder (Promega); lanes 2–11: sediment-free trans-DCE-producing cultures established from soil sediments collected from various electrical waste landfill sites; lane 12: trans-DCE-producing enrichment culture MB; lane13: trans-DCE-producing culture from Sungei Buloh; lane14: pure culture MB; lane 15: non trans-DCE-producing culture Dehalococcoides sp. ANAS1; lane 16: water.
Discussion
Trans-DCE accumulation at chloroethene-contaminated sites has been an environmental concern because of their potential toxicity to humans (Mochida et al., 1995). However, molecular features of trans-DCE generation had been largely unknown. The mbrA RDase gene identified in this study is suggested to be involved in the reductive dechlorination of PCE/TCE to the major end product trans-DCE, corroborated by: (1) its elevated gene transcription level measured by qPCR; (2) protein expression observed on protein gels; and (3) in-gel PCE-to-trans-DCE dechlorination activity of the above most expressed protein. The increase in the mbrA gene expression leads to an augmented production of the functional protein, as the size of the mature MbrA (DceA6) RDase is estimated to be ∼50 kDa, corresponding to the calculated molecular mass of 55 kDa from the length of the mbrA RDase gene. The identified RDase gene, mbrA, is detected in 5 out of the 10 trans-DCE-producing cultures, which were derived from microcosms established with environmental soil or sediments. All these observations indicate that environmental microbes are capable of generating trans-DCE as the end products. Hence, biotic degradation of PCE/TCE may explain the presence of high levels of trans-DCE at contaminated sites (ATSDR, 2007; Nijenhuis et al., 2007). As for the dceA7 gene, its fold of expression level is lower than that of the mbrA gene and its predicted molecular mass is ∼17 kDa. Therefore, it is likely that the mRNA of dceA7 is either not translated into the functional RDase enzyme or the translated protein might not be directly involved in the generation of trans-DCE.
Strain MB contains RDase genes that share high identity to genes in strains 195, VS and CBDB1, suggesting that these RDases might be acquired through the process of horizontal gene transfer (Regeard et al., 2005). Although the identified mbrA gene consists of the same characteristics of typical cis-DCE-producing RDases (twin arginine translocation peptide signal and the Fe4S4 clusters), they differ functionally. Comparison of the amino-acid sequences of the newly identified mbrA gene and the known tceA gene reveals an identity of only 29%, thus indicating the uniqueness of the trans-DCE-producing mbrA gene. A previous microarray study also shows that strain MB shares high identity to 10 RDase genes of strain 195, among which three (dceA3, dceA4 and dceA7) were also found in this study, indicating the reproducibility of this and previous findings (Cheng and He, 2009). Therefore, 14 putative RDase genes (7 from this study and 10 from microarray analysis, whereas 3 are overlapping ones) could be present in the trans-DCE-producing culture MB. Observations in this study are consistent with the hypothesis of the existence of a pool of RDases in each different strain with different combinations. The amplicons generated for primer pair 2 are much smaller in size (∼1.2 kb) than those for primer pair 1, as the former targets the PIDD region of RDase genes. In this study, six intact coding sequences encode putative RDases (DceA1 to DceA6) with adjacent genes encoding for potential membrane-anchoring proteins. The dceA7 gene (100% identity to DET0088) is truncated, and the 3′ anchoring protein is replaced by a hypothetical protein. Nevertheless, recent studies have reported that the expression of DET0088 was upregulated during PCE dechlorination in Alameda Naval Air Station enrichments and mixed cultures containing strain 195 (Rahm et al., 2006; West et al., 2008). Also, the dceA7 gene was identified using the newly designed primer pairs, RDH F1C and RDH R1C, revealing inadequacies of the previous degenerate primer pair. In all, the presence of seven RDase genes provides an insight into the genetic makeup of strain MB for the first time, though the functions of the other non-expressing RDase genes identified in culture MB still remain to be elucidated.
The sequences of the two expressing RDase genes in strain MB possess two open reading frames, consisting of a dehalogenase-containing domain and an associated protein at the 3′ end (an anchoring protein and a hypothetical protein for dceA6 and dceA7, respectively). The upstream regions of the two expressing RDase genes differ from each other. The upstream region of the RDase gene mbrA (dceA6) is a DNA-binding response regulator, whereas the 5′ region of the dceA7 gene shares high identity with the SAM domain-containing protein as shown in strain 195. The transcription level of the mbrA gene is lower compared with some other known RDase genes such as the tceA gene, which is possibly due to the tight regulation of transcription by the upstream DNA-binding response regulator protein (Wagner et al., 2009; Kube et al., 2005). The upstream regions of the RDase genes have attracted much interest recently with current genome sequencing projects (Kube et al., 2005; Seshadri et al., 2005). Therefore, the DNA-binding response regulator upstream of the mbrA (dceA6) gene might be a part of the two-component regulatory system, whose function is to trigger downstream pathway cascades once it is activated by an extracellular signal (Fabret et al., 1999). This signal will trigger the phosphorylated DNA-binding response regulator to bind to a transcriptional factor that will affect the dehalogenation processes by altering the level of transcription RDase genes (Fabret et al., 1999). On the other hand, radical SAM proteins, such as the one detected in dceA7, are reported to catalyze unusual methylations, isomerization, sulfur insertion, ring formation, anaerobic oxidation and protein radical formation (Sofia et al., 2001). They also function in the biosynthesis of DNA precursors, vitamins, cofactors, antibiotics and herbicides, and in the biodegradation pathways (Sofia et al., 2001). Nonetheless, the function that the SAM radical protein has on its associated RDase gene dceA7 remains to be determined.
In our previous study, one of the abundant putative RDase genes detected in strain MB is the dceA1 gene (Cheng and He, 2009). Genome walking of the dceA1 gene reveals that upstream of this RDase gene shares high identity with the transcriptional regulator belonging to the MarR (multiple antibiotic resistance) superfamily (Figure 3). It is oriented in the direction opposite to that of the associated RDase gene and its associated anchoring protein—a typical RDase locus associated with the MarR regulator (Kube et al., 2005). MarR transcriptional regulators found in prokaryotes are often involved in the catabolism of aromatic compounds as well as the modulation of multiple responses to other toxic molecules such as antibiotics (Alekshun et al., 2001). The presence of the MarR transcriptional regulator and the DNA-binding response regulator nearby suggests that the expressions of the RDase genes are tightly regulated. Despite the different origins (for example, strains 195 and CBDB1) from which MB might have inherited the upstream regions (of dceA1, dceA6 and dceA7), these regions of the identified RDases should mainly function on the regulation of the downstream RDases.
In all, understanding the nature of the transcriptional regulators associated with the RDase genes is crucial to bioremediation applications as these regulatory elements often act as ‘switches’ for the microbial dechlorination activity. This study sheds light on the genetic pool of RDase genes present in the trans-DCE-producing cultures. The presence of the mbrA gene in other trans-DCE-producing cultures implicates the potential use of this gene as a functional biomarker, complementing the current inadequacies of the 16S rRNA gene approaches in assessing and monitoring in situ PCE/TCE dechlorination.
Accession codes
References
Adrian L, Rahnenführer J, Gobom J, Hölscher T . (2007). Identification of a chlorobenzene reductive dehalogenase in Dehalococcoides sp. strain CBDB1. Appl Environ Microbiol 73: 7717–7724.
Alekshun MN, Levy SB, Mealy TR, Seaton BA, Head JF . (2001). The crystal structure of MarR, a regulator of multiple antibiotic resistance, at 2.3 Å resolution. Nat Struct Biol 8: 710–714.
Abelson PH . (1990). Inefficient remediation of ground-water pollution. Science 250: 733.
ATSDR (1997a). Toxicological profile for tetrachloroethylene. Agency for Toxic Substance and Disease Registry, US Public Health Service, US Department of Health and Human Services: Atlanta, GA, USA.
ATSDR (1997b). Toxicological Profile for trichloroethylene. Agency for Toxic Substance and Disease Registry, US Public Health Service, US Department of Health and Human Services: Atlanta, GA, USA.
ATSDR (2007). Agency for Toxic Substance and Disease Registry, US Public Health Service, US Department of Health and Human Services: Atlanta, GA, USA.
Berks BC . (1996). A common export pathway for proteins binding complex redox cofactors? Mol Microbiol 22: 393–404.
Campanella JJ, Bitincks L, Smalley J . (2003). MatGAT: an application that generates similarity/identity matrices using protein or DNA sequences. BMC Bioinformatics 4: 29.
Cheng D, Chow WL, He J . (2010). A Dehalococcoides-containing coculture that dechlorinates tetrachloroethene to trans-1,2-dichloroethene. ISME J 4: 88–97.
Cheng D, He J . (2009). Isolation and characterization of “Dehalococcoides” sp. MB, which dechlorinates tetrachloroethene to trans-1,2-dichloroethene. Appl Environ Microbiol 75: 5910–5918.
Fabret C, Feher VA, Hoch JA . (1999). Two-component signal transduction in Bacillus subtilis: how one organism sees its world. J Bacteriol 181: 1975–1983.
Griffin BM, Tiedje J.M, Löffler FE . (2004). Anaerobic microbial reductive dechlorination of tetrachloroethene to predominately trans-1,2-dichloroethene. Environ Sci Technol 38: 4300–4303.
Hall TA . (1999). Bioedit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41: 95–98.
He J, Holmes VF, Lee PKH, Alvarez-Cohen L . (2007). Influence of vitamin B12 and cocultures on the growth of Dehalococcoides isolates in defined medium. Appl Environ Microbiol 73: 2847–2853.
He J, Ritalahti KM, Aiello MR, Löffler FE . (2003). Complete detoxification of vinyl chloride by an anaerobic enrichment culture and identification of the reductively dechlorinating population as a Dehalococcoides species. Appl Environ Microbiol 69: 996–1003.
He J, Sung Y, Krajmalnik-Brown R, Ritalahti KM, Löffler FE . (2005). Isolation and characterization of Dehalococcoides sp. strain FL2, a trichloroethene (TCE)- and 1,2-dichloroethene-respiring anaerobe. Environ Microbiol 7: 1442–1450.
Hiraishi A . (2008). Biodiversity of dehalorespiring bacteria with special emphasis on polychlorinated biphenyl/dioxin dechlorinators. Microbes Environ 23: 1–12.
Holliger C, Hahn D, Harmsen H, Ludwig W, Schumacher W, Tindall B et al. (1998). Dehalobacter restrictus gen. nov. and sp. nov., a strictly anaerobic bacterium that reductively dechlorinates tetra- and trichloroethene in an anaerobic respiration. Arch Microbiol 169: 313–321.
Hölscher T, Krajmalnik-Brown R, Ritalahti KM, von Wintzingerode F, Görisch H, Löffler FE et al. (2004). Multiple nonidentical reductive-dehalogenase-homologous genes are common in Dehalococcoides. Appl Environ Microbiol 70: 5290–5297.
Johnson DR, Lee PKH, Holmes VF, Alvarez-Cohen L . (2005). An internal reference technique for accurately quantifying specific mRNAs by real-time PCR with application to the tceA reductive dehalogenase gene. Appl Environ Microbiol 71: 3866–3871.
Katoh K, Misawa K, Kuma K, Miyata T . (2002). MAFTT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30: 3059–3066.
Kittelmann S, Friedrich MW . (2008). Novel uncultured Chloroflexi dechlorinate perchloroethene to trans-dichloroethene in tidal flat sediments. Environ Microbiol 10: 1557–1570.
Krajmalnik-Brown R, Hölscher T, Thomson IN, Saunders FM, Ritalahti KM, Löffler FE . (2004). Genetic identification of a putative vinyl chloride reductase in Dehalococcoides sp. strain BAV1. Appl Environ Microbiol 70: 6347–6351.
Kube M, Beck A, Zinder SH, Kuhl H, Reinhardt R, Adrian L . (2005). Genome sequence of the chlorinated compound-respiring bacterium Dehalococcoides species strain CBDB1. Nat Biotechnol 23: 1269–1273.
Läemmli UK . (1970). Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227: 680–685.
Löffler FE, Champine JE, Ritalahti KM, Sprague SJ, Tiedje JM . (1997). Complete reductive dechlorination of 1,2-dichloropane by anaerobic bacteria. Appl Environ Microbiol 63: 2870–2875.
Löffler FE, Sun Q, Li J, Tiedje JM . (2000). 16S rRNA gene-based detection of tetrachloroethene-dechlorinating Desulfuromonas and Dehalococcoides species. Appl Environ Microbiol 66: 1369–1374.
Luijten ML, de Weert J, Smidt H, Boschker HT, de Vos WM, Schraa G et al. (2003). Description of Sulfurospirillum halorespirans sp. nov., an anaerobic, tetrachloroethene-respiring bacterium, and transfer of Dehalospirillum multivorans to the genus Sulfurospirillum as Sulfurospirillum multivorans comb. nov. Int J Syst Evol Microbiol 53: 787–793.
Magnuson JK, Romine MF, Burris DR, Kingsley MT . (2000). Trichloroethene reductive dehalogenase from Dehalococcoides ethenogenes: sequence of tceA and substrate range characterization. Appl Environ Microbiol 66: 5141–5147.
Magnuson JK, Stern RV, Gossett JM, Zinder SH, Burris DR . (1998). Reductive dechlorination of tetrachloroethene to ethene by a two component enzyme pathway. Appl Environ Microbiol 64: 1270–1275.
Maillard J, Schumacher W, Vazquez F, Regeard C, Hagen WR, Holliger C . (2003). Characterization of the corrinoid iron-sulfur protein tetrachloroethene reductive dehalogenase of Dehalobacter restrictus. Appl Environ Microbiol 69: 4628–4638.
Miller E, Wohlfarth G, Diekert G . (1998). Purification and characterization of the tetrachloroethene reductive dehalogenase of strain PCE-S. Arch Microbiol 169: 497–502.
Miller GS, Milliken CE, Sowers KR, May HD . (2005). Reductive dechlorination of tetrachloroethene to trans-dichloroethene and cis-dichloroethene by PCB-dechlorinating bacterium DF-1. Environ Sci Technol 39: 2631–2635.
Mochida K, Gomyoda M, Fujita T . (1995). Toxicity of 1,1-dichloroethane and 1,2-dichloroethylene determined using cultured human KB cells. Bull Environ Contam Toxicol 55: 316–319.
Müller JA, Rosner BM, Von Abendroth G, Meshulam-Simon G, McCarty PL, Spormann AM . (2004). Molecular identification of the catabolic vinyl chloride reductase from Dehalococcoides sp. strain VS and its environmental distribution. Appl Environ Microbiol 70: 4880–4888.
Neumann A, Wohlfarth G, Diekert G . (1996). Purification and characterization of tetrachloroethene reductive dehalogenase from Dehalospirillum multivorans. J Biol Chem 271: 16515–16519.
Neumann A, Wohlfarth G, Diekert G . (1998). Tetrachloroethene dehalogenase from Dehalospirillum multivorans: cloning, sequencing of the encoding genes, and expression of the pceA gene in Escherichia coli. J Bacteriol 180: 4140–4145.
Neumann A, Siebert A, Trescher T, Reinhardt S, Wohlfart G, Diekert G . (2007). Tetrachloroethene reductive dehalogenase of Dehalospirillum multivorans: substrate specificity of the native enzyme and its corrinoid factor. Arch Microbiol 177: 420–426.
Nijenhuis I, Nikolausz M, Köth A, Felföld T, Weiss H, Drangmeister J et al. (2007). Assessment of the anaerobic natural attenuation of chlorinated ethenes in an anaerobic contaminated aquifer in the Bitterfield/Wolfen studies, and molecular biomarkers. Chemosphere 67: 300–311.
Okeke BC, Chang YC, Hatsu H, Suzuki T, Takamizawa K . (2001). Purification, cloning, and sequencing of an enzyme mediating the reductive dechlorination of tetrachloroethylene (PCE) from Clostridium bifermentans DPH-1. Can J Microbiol 47: 448–456.
Pearson WR, Lipman DJ . (1988). Improved tools for biological sequence comparison. Proc Natl Acad Sci USA 85: 2444–2448.
Rahm BG, Morris RM, Richardson RE . (2006). Temporal expression of respiratory genes in an enrichment culture containing Dehalococcoides ethenogenes. Appl Environ Microbiol 72: 5486–5491.
Regeard C, Maillard J, Dufraigne C, Deschavanne P, Holliger C . (2005). Indications for acquisition of reductive dehalogenase genes through horizontal gene transfer by Dehalococcoides ethenogenes strain 195. Appl Environ Microbiol 71: 2955–2961.
Schumacher W, Holliger C, Zehnder AJ, Hagen WR . (1997). Redox chemistry of cobalamin and iron-sulfur cofactors in the tetrachloroethene reductase of Dehalobacter restrictus. FEBS Lett 409: 421–425.
Seshadri R, Adrian L, Fouts DE, Eisen JA, Phillippy AM, Methe BA et al. (2005). Genome sequence of the PCE-dechlorinating bacterium Dehalococcoides ethenogenes. Science 307: 105–108.
Smidt H, de Vos WM . (2004). Anaerobic microbial dehalogenation. Annu Rev Microbiol 58: 43–73.
Sofia HJ, Chen G, Hetzler BG, Reyes-Spindola JF, Miller NE . (2001). Radical SAM, a novel protein superfamily linking unresolved steps in familiar biosynthetic pathways with radical mechanisms: functional characterization using new analysis and information visualization methods. Nucleic Acids Res 29: 1097–1106.
Sticht H, Rösch P . (1998). The structure of iron-sulfur proteins. Prog Biophys Mol Biol 70: 95–136.
Wagner A, Adrian L, Kleinsteuber S, Andreesen JR, Lechner U . (2009). Transcription analysis of genes encoding homologues of reductive dehalogenases in “Dehalococcoides” sp. strain CBDB1 by using terminal restriction fragment length polymorphism and quantitative PCR. Appl Environ Microbiol 75: 1876–1884.
Waller AS, Krajmalnik-Brown R, Löffler FE, Edwards EA . (2005). Multiple reductive-dehalogenase-homologous genes are simultaneously transcribed during dechlorination by Dehalococcoides-containing cultures. Appl Environ Microbiol 71: 8257–8264.
West KA, Johnson DR, Hu P, DeSantis TZ, Brodie EL, Lee PKH et al. (2008). Comparative genomics of “Dehalococcoides ethenogenes” 195 and an enrichment culture containing unsequenced “Dehalococcoides” strains. Appl Environ Microbiol 74: 3533–3540.
Westrick JJ, Mello JW, Thomas RF . (1984). The groundwater supply survey. J Am Water Works Assoc 76: 52–59.
Wolin EA, Wolin MJ, Wolfe RS . (1963). Formation of methane by bacterial extracts. J Biol Chem 238: 2882–2886.
Acknowledgements
This study was supported by the Academic Research Fund from the Singapore Ministry of Education under Project no. R-288-000-041-112 and the Singapore Agency for Science, Technology and Research (A*STAR) of the Science and Engineering Research Council under Project no. 062 101 0028.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chow, W., Cheng, D., Wang, S. et al. Identification and transcriptional analysis of trans-DCE-producing reductive dehalogenases in Dehalococcoides species. ISME J 4, 1020–1030 (2010). https://doi.org/10.1038/ismej.2010.27
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2010.27
Keywords
This article is cited by
-
Diversity of organohalide respiring bacteria and reductive dehalogenases that detoxify polybrominated diphenyl ethers in E-waste recycling sites
The ISME Journal (2022)
-
Microbial degradation of chloroethenes: a review
Environmental Science and Pollution Research (2017)
-
Spatial Abundance and Distribution of Potential Microbes and Functional Genes Associated with Anaerobic Mineralization of Pentachlorophenol in a Cylindrical Reactor
Scientific Reports (2016)
-
A comparative genomics and reductive dehalogenase gene transcription study of two chloroethene-respiring bacteria, Dehalococcoides mccartyi strains MB and 11a
Scientific Reports (2015)
-
Genome sequencing of a single cell of the widely distributed marine subsurface Dehalococcoidia, phylum Chloroflexi
The ISME Journal (2014)