Abstract
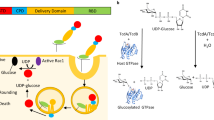
An internal cysteine protease domain (CPD) autoproteolytically regulates Clostridium difficile glucosylating toxins by releasing a cytotoxic effector domain into target cells. CPD activity is itself allosterically regulated by the eukaryote-specific molecule inositol hexakisphosphate (InsP6). Although allostery controls the function of most proteins, the molecular details underlying this regulatory mechanism are often difficult to characterize. Here we use chemical probes to show that apo-CPD is in dynamic equilibrium between active and inactive states. InsP6 markedly shifts this equilibrium toward an active conformer that is further restrained upon binding a suicide substrate. Structural analyses combined with systematic mutational and disulfide bond engineering studies show that residues within a β-hairpin region functionally couple the InsP6-binding site to the active site. Collectively, our results identify an allosteric circuit that allows bacterial virulence factors to sense and respond to the eukaryotic environment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Changeux, J.P. & Edelstein, S.J. Allosteric mechanisms of signal transduction. Science 308, 1424–1428 (2005).
del Sol, A., Tsai, C.J., Ma, B. & Nussinov, R. The origin of allosteric functional modulation: multiple pre-existing pathways. Structure 17, 1042–1050 (2009).
Goodey, N.M. & Benkovic, S.J. Allosteric regulation and catalysis emerge via a common route. Nat. Chem. Biol. 4, 474–482 (2008).
Tsai, C.J., Del Sol, A. & Nussinov, R. Protein allostery, signal transmission and dynamics: a classification scheme of allosteric mechanisms. Mol. Biosyst. 5, 207–216 (2009).
Egerer, M., Giesemann, T., Herrmann, C. & Aktories, K. Autocatalytic processing of Clostridium difficile toxin B. Binding of inositol hexakisphosphate. J. Biol. Chem. 284, 3389–3395 (2009).
Egerer, M. & Satchell, K.J. Inositol hexakisphosphate-induced autoprocessing of large bacterial protein toxins. PLoS Pathog. 6, e1000942 (2010).
Reineke, J. et al. Autocatalytic cleavage of Clostridium difficile toxin B. Nature 446, 415–419 (2007).
Shen, A. Allosteric regulation of protease activity by small molecules. Mol. Biosyst. 6, 1431–1443 (2010).
Michell, R.H. Inositol derivatives: evolution and functions. Nat. Rev. Mol. Cell Biol. 9, 151–161 (2008).
Egerer, M., Giesemann, T., Jank, T., Satchell, K.J. & Aktories, K. Auto-catalytic cleavage of Clostridium difficile toxins A and B depends on cysteine protease activity. J. Biol. Chem. 282, 25314–25321 (2007).
Prochazkova, K. et al. Structural and molecular mechanism for autoprocessing of MARTX toxin of Vibrio cholerae at multiple sites. J. Biol. Chem. 284, 26557–26568 (2009).
Rupnik, M. et al. Characterization of the cleavage site and function of resulting cleavage fragments after limited proteolysis of Clostridium difficile toxin B (TcdB) by host cells. Microbiology 151, 199–208 (2005).
Shen, A. et al. Mechanistic and structural insights into the proteolytic activation of Vibrio cholerae MARTX toxin. Nat. Chem. Biol. 5, 469–478 (2009).
Lupardus, P.J., Shen, A., Bogyo, M. & Garcia, K.C. Small molecule–induced allosteric activation of the Vibrio cholerae RTX cysteine protease domain. Science 322, 265–268 (2008).
Pruitt, R.N. et al. Structure-function analysis of inositol hexakisphosphate-induced autoprocessing in Clostridium difficile toxin A. J. Biol. Chem. 284, 21934–21940 (2009).
Pfeifer, G. et al. Cellular uptake of Clostridium difficile toxin B. Translocation of the N-terminal catalytic domain into the cytosol of eukaryotic cells. J. Biol. Chem. 278, 44535–44541 (2003).
Genth, H., Dreger, S.C., Huelsenbeck, J. & Just, I. Clostridium difficile toxins: more than mere inhibitors of Rho proteins. Int. J. Biochem. Cell Biol. 40, 592–597 (2008).
Jank, T. & Aktories, K. Structure and mode of action of clostridial glucosylating toxins: the ABCD model. Trends Microbiol. 16, 222–229 (2008).
Voth, D.E. & Ballard, J.D. Clostridium difficile toxins: mechanism of action and role in disease. Clin. Microbiol. Rev. 18, 247–263 (2005).
Kelly, C.P. & LaMont, J.T. Clostridium difficile—more difficult than ever. N. Engl. J. Med. 359, 1932–1940 (2008).
Rupnik, M., Wilcox, M.H. & Gerding, D.N. Clostridium difficile infection: new developments in epidemiology and pathogenesis. Nat. Rev. Microbiol. 7, 526–536 (2009).
Lyras, D. et al. Toxin B is essential for virulence of Clostridium difficile. Nature 458, 1176–1179 (2009).
Kuehne, S.A. et al. The role of toxin A and toxin B in Clostridium difficile infection. Nature 467, 711–713 (2010).
Aktories, K. Self-cutting to kill: new insights into the processing of Clostridium difficile toxins. ACS Chem. Biol. 2, 228–230 (2007).
Puri, A.W. et al. Rational design of inhibitors and activity-based probes targeting Clostridium difficile virulence factor TcdB. Chem. Biol. 17, 1201–1211 (2010).
Evans, M.J. & Cravatt, B.F. Mechanism-based profiling of enzyme families. Chem. Rev. 106, 3279–3301 (2006).
Greenbaum, D.C. et al. Small molecule affinity fingerprinting. A tool for enzyme family subclassification, target identification, and inhibitor design. Chem. Biol. 9, 1085–1094 (2002).
Berger, A.B. et al. Identification of early intermediates of caspase activation using selective inhibitors and activity-based probes. Mol. Cell 23, 509–521 (2006).
Kato, D. et al. Activity-based probes that target diverse cysteine protease families. Nat. Chem. Biol. 1, 33–38 (2005).
Powers, J.C., Asgian, J.L., Ekici, O.D. & James, K.E. Irreversible inhibitors of serine, cysteine, and threonine proteases. Chem. Rev. 102, 4639–4750 (2002).
Prochazkova, K. & Satchell, K.J. Structure-function analysis of inositol hexakisphosphate-induced autoprocessing of the Vibrio cholerae multifunctional autoprocessing RTX toxin. J. Biol. Chem. 283, 23656–23664 (2008).
Sheahan, K.L., Cordero, C.L. & Satchell, K.J. Autoprocessing of the Vibrio cholerae RTX toxin by the cysteine protease domain. EMBO J. 26, 2552–2561 (2007).
Chai, J. et al. Crystal structure of a procaspase-7 zymogen: mechanisms of activation and substrate binding. Cell 107, 399–407 (2001).
Romanowski, M.J., Scheer, J.M., O'Brien, T. & McDowell, R.S. Crystal structures of a ligand-free and malonate-bound human caspase-1: implications for the mechanism of substrate binding. Structure 12, 1361–1371 (2004).
Royer, C.A. Probing protein folding and conformational transitions with fluorescence. Chem. Rev. 106, 1769–1784 (2006).
Datta, D., Scheer, J.M., Romanowski, M.J. & Wells, J.A. An allosteric circuit in caspase-1. J. Mol. Biol. 381, 1157–1167 (2008).
Ferguson, A.D. et al. Signal transduction pathway of TonB-dependent transporters. Proc. Natl. Acad. Sci. USA 104, 513–518 (2007).
Süel, G.M., Lockless, S.W., Wall, M.A. & Ranganathan, R. Evolutionarily conserved networks of residues mediate allosteric communication in proteins. Nat. Struct. Biol. 10, 59–69 (2003).
Volkman, B.F., Lipson, D., Wemmer, D.E. & Kern, D. Two-state allosteric behavior in a single-domain signaling protein. Science 291, 2429–2433 (2001).
Fuentes-Prior, P. & Salvesen, G.S. The protein structures that shape caspase activity, specificity, activation and inhibition. Biochem. J. 384, 201–232 (2004).
Eisenmesser, E.Z. et al. Intrinsic dynamics of an enzyme underlies catalysis. Nature 438, 117–121 (2005).
Fraser, J.S. et al. Hidden alternative structures of proline isomerase essential for catalysis. Nature 462, 669–673 (2009).
Lechtenberg, B.C., Johnson, D.J., Freund, S.M. & Huntington, J.A. NMR resonance assignments of thrombin reveal the conformational and dynamic effects of ligation. Proc. Natl. Acad. Sci. USA 107, 14087–14092 (2010).
Lee, G.M. & Craik, C.S. Trapping moving targets with small molecules. Science 324, 213–215 (2009).
Shahian, T. et al. Inhibition of a viral enzyme by a small-molecule dimer disruptor. Nat. Chem. Biol. 5, 640–646 (2009).
Hardy, J.A., Lam, J., Nguyen, J.T., O'Brien, T. & Wells, J.A. Discovery of an allosteric site in the caspases. Proc. Natl. Acad. Sci. USA 101, 12461–12466 (2004).
Scheer, J.M., Romanowski, M.J. & Wells, J.A. A common allosteric site and mechanism in caspases. Proc. Natl. Acad. Sci. USA 103, 7595–7600 (2006).
Pruitt, R.N., Chambers, M.G., Ng, K.K., Ohi, M.D. & Lacy, D.B. Structural organization of the functional domains of Clostridium difficile toxins A and B. Proc. Natl. Acad. Sci. USA 107, 13467–13472 (2010).
Qa'Dan, M., Spyres, L.M. & Ballard, J.D. pH-induced conformational changes in Clostridium difficile toxin B. Infect. Immun. 68, 2470–2474 (2000).
Mittal, R., Peak-Chew, S.Y., Sade, R.S., Vallis, Y. & McMahon, H.T. The acetyltransferase activity of the bacterial toxin YopJ of Yersinia is activated by eukaryotic host cell inositol hexakisphosphate. J. Biol. Chem. 285, 19927–19934 (2010).
Hanakahi, L.A., Bartlet-Jones, M., Chappell, C., Pappin, D. & West, S.C. Binding of inositol phosphate to DNA-PK and stimulation of double-strand break repair. Cell 102, 721–729 (2000).
Macbeth, M.R. et al. Inositol hexakisphosphate is bound in the ADAR2 core and required for RNA editing. Science 309, 1534–1539 (2005).
Tan, X. et al. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446, 640–645 (2007).
Leslie, A.G. Recent changes to the MOSFLM package for processing film and image plate data. Joint CCP4 ESF-EAMCB Newslett. Protein Crystallogr. 26 (1992).
Potterton, E., Briggs, P., Turkenburg, M. & Dodson, E. A graphical user interface to the CCP4 program suite. Acta Crystallogr. D Biol. Crystallogr. 59, 1131–1137 (2003).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P.D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D Biol. Crystallogr. 58, 1948–1954 (2002).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D60, 2126–2132 (2004).
Davis, I.W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
DeLano, W.L. The PyMOL Molecular Graphics System, Version 1.3 (Schrödinger LLC 2002).
Acknowledgements
We thank J. Wells for helpful suggestions in the preparation of this manuscript. A.S. is supported by a US National Institutes of Health (NIH) National Institutes of General Medical Sciences grant (K99GM092934). P.J.L. is a Damon Runyon Fellow, supported by the Damon Runyon Cancer Research Foundation. A.W.P. is supported by a US National Science Foundation Graduate Research Fellowship. K.C.G. is supported by the Howard Hughes Medical Institute and NIH grant NIH-RO1AI48540. M.B. is supported by the Burroughs Wellcome Foundation and NIH grants R01EB005011 and R01AI078947.
Author information
Authors and Affiliations
Contributions
A.S. developed and executed autocleavage, probe labeling, limited proteolysis and tryptophan fluorescence assays; constructed, purified and expressed CPD variants; prepared CPD for crystallization studies; helped design the AWP19 probe; and wrote the manuscript. P.J.L. crystallized InsP6-bound TcdB CPD, collected the data, solved and analyzed the structure, generated Figure 4 and Table 1 and provided advice on the manuscript. M.M.G. synthesized AWP19, helped with initial characterization of TcdB CPD mutants using a fluorogenic substrate assay and the probe labeling assay and provided creative input. A.W.P. helped design and synthesize the AWP19 probe and performed initial characterization of probe labeling of TcdB CPD. V.E.A. helped design the AWP19 probe and provided advice on probe usage. K.C.G. provided advice on the manuscript and provided financial support. M.B. provided creative input and financial support, and helped write the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1 and 2, and Supplementary Methods (PDF 1277 kb)
Rights and permissions
About this article
Cite this article
Shen, A., Lupardus, P., Gersch, M. et al. Defining an allosteric circuit in the cysteine protease domain of Clostridium difficile toxins. Nat Struct Mol Biol 18, 364–371 (2011). https://doi.org/10.1038/nsmb.1990
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1990
This article is cited by
-
Clostridioides difficile toxins: mechanisms of action and antitoxin therapeutics
Nature Reviews Microbiology (2022)
-
Structure of the full-length Clostridium difficile toxin B
Nature Structural & Molecular Biology (2019)
-
Identification and characterization of a large family of superbinding bacterial SH2 domains
Nature Communications (2018)
-
Clostridium difficile infection
Nature Reviews Disease Primers (2016)
-
Crystal structure of Clostridium difficile toxin A
Nature Microbiology (2016)