Abstract
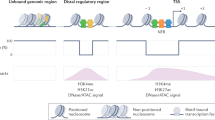
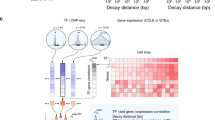
Dynamic access to genetic information is central to organismal development and environmental response. Consequently, genomic processes must be regulated by mechanisms that alter genome function relatively rapidly1,2,3,4. Conventional chromatin immunoprecipitation (ChIP) experiments measure transcription factor occupancy5, but give no indication of kinetics and are poor predictors of transcription factor function at a given locus. To measure transcription-factor-binding dynamics across the genome, we performed competition ChIP (refs 6, 7) with a sequence-specific Saccharomyces cerevisiae transcription factor, Rap1 (ref. 8). Rap1-binding dynamics and Rap1 occupancy were only weakly correlated (R2 = 0.14), but binding dynamics were more strongly linked to function than occupancy. Long Rap1 residence was coupled to transcriptional activation, whereas fast binding turnover, which we refer to as ‘treadmilling’, was linked to low transcriptional output. Thus, DNA-binding events that seem identical by conventional ChIP may have different underlying modes of interaction that lead to opposing functional outcomes. We propose that transcription factor binding turnover is a major point of regulation in determining the functional consequences of transcription factor binding, and is mediated mainly by control of competition between transcription factors and nucleosomes. Our model predicts a clutch-like mechanism that rapidly engages a treadmilling transcription factor into a stable binding state, or vice versa, to modulate transcription factor function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Data deposits
Data has been deposited in the Gene Expression Omnibus under accession numbers GSE32351 (ChIP-on-chip data), GPL14612 (ChIP platform), GSM677030–GSM677033 (RNA expression array data) and GPL4414 (expression platform).
References
Mueller, F., Wach, P. & McNally, J. G. Evidence for a common mode of transcription factor interaction with chromatin as revealed by improved quantitative fluorescence recovery after photobleaching. Biophys. J. 94, 3323–3339 (2008)
Yao, J., Munson, K. M., Webb, W. W. & Lis, J. T. Dynamics of heat shock factor association with native gene loci in living cells. Nature 442, 1050–1053 (2006)
Stavreva, D. A., Muller, W. G., Hager, G. L., Smith, C. L. & McNally, J. G. Rapid glucocorticoid receptor exchange at a promoter is coupled to transcription and regulated by chaperones and proteasomes. Mol. Cell. Biol. 24, 2682–2697 (2004)
Karpova, T. S. et al. Concurrent fast and slow cycling of a transcriptional activator at an endogenous promoter. Science 319, 466–469 (2008)
MacArthur, S. et al. Developmental roles of 21 Drosophila transcription factors are determined by quantitative differences in binding to an overlapping set of thousands of genomic regions. Genome Biol. 10, R80 (2009)
Dion, M. F. et al. Dynamics of replication-independent histone turnover in budding yeast. Science 315, 1405–1408 (2007)
van Werven, F. J., van Teeffelen, H. A., Holstege, F. C. & Timmers, H. T. Distinct promoter dynamics of the basal transcription factor TBP across the yeast genome. Nature Struct. Mol. Biol. 16, 1043–1048 (2009)
Lieb, J. D., Liu, X., Botstein, D. & Brown, P. O. Promoter-specific binding of Rap1 revealed by genome-wide maps of protein-DNA association. Nature Genet. 28, 327–334 (2001)
Piña, B., Fernández-Larrea, J., García-Reyero, N. & Idrissi, F. Z. The different (sur)faces of Rap1p. Mol. Genet. Genomics 268, 791–798 (2003)
Buck, M. J. & Lieb, J. D. A chromatin-mediated mechanism for specification of conditional transcription factor targets. Nature Genet. 38, 1446–1451 (2006)
Holstege, F. C. et al. Dissecting the regulatory circuitry of a eukaryotic genome. Cell 95, 717–728 (1998)
Layer, J. H., Miller, S. G. & Weil, P. A. Direct transactivator-transcription factor IID (TFIID) contacts drive yeast ribosomal protein gene transcription. J. Biol. Chem. 285, 15489–15499 (2010)
Pelechano, V., Chavez, S. & Perez-Ortin, J. E. A complete set of nascent transcription rates for yeast genes. PLoS ONE 5, e15442 (2010)
Polach, K. J. & Widom, J. Mechanism of protein access to specific DNA sequences in chromatin: a dynamic equilibrium model for gene regulation. J. Mol. Biol. 254, 130–149 (1995)
Segal, E. & Widom, J. From DNA sequence to transcriptional behaviour: a quantitative approach. Nature Rev. Genet. 10, 443–456 (2009)
Lam, F. H., Steger, D. J. & O'Shea, E. K. Chromatin decouples promoter threshold from dynamic range. Nature 453, 246–250 (2008)
Pokholok, D. K. et al. Genome-wide map of nucleosome acetylation and methylation in yeast. Cell 122, 517–527 (2005)
Anderson, J. D., Lowary, P. T. & Widom, J. Effects of histone acetylation on the equilibrium accessibility of nucleosomal DNA target sites. J. Mol. Biol. 307, 977–985 (2001)
John, S. et al. Chromatin accessibility pre-determines glucocorticoid receptor binding patterns. Nature Genet. 43, 264–268 (2011)
Zanton, S. J. & Pugh, B. F. Changes in genomewide occupancy of core transcriptional regulators during heat stress. Proc. Natl Acad. Sci. USA 101, 16843–16848 (2004)
Kaplan, N. et al. The DNA-encoded nucleosome organization of a eukaryotic genome. Nature 458, 362–366 (2009)
Koerber, R. T., Rhee, H. S., Jiang, C. & Pugh, B. F. Interaction of transcriptional regulators with specific nucleosomes across the Saccharomyces genome. Mol. Cell 35, 889–902 (2009)
Ganapathi, M. et al. Extensive role of the general regulatory factors, Abf1 and Rap1, in determining genome-wide chromatin structure in budding yeast. Nucleic Acids Res. 39, 2032–2044 (2011)
Mukherjee, S. et al. Rapid analysis of the DNA-binding specificities of transcription factors with DNA microarrays. Nature Genet. 36, 1331–1339 (2004)
Bosisio, D. et al. A hyper-dynamic equilibrium between promoter-bound and nucleoplasmic dimers controls NF-κB-dependent gene activity. EMBO J. 25, 798–810 (2006)
Voss, T. C. et al. Dynamic exchange at regulatory elements during chromatin remodeling underlies assisted loading mechanism. Cell 146, 544–554 (2011)
McNally, J. G., Muller, W. G., Walker, D., Wolford, R. & Hager, G. L. The glucocorticoid receptor: rapid exchange with regulatory sites in living cells. Science 287, 1262–1265 (2000)
Perlmann, T., Eriksson, P. & Wrange, O. Quantitative analysis of the glucocorticoid receptor-DNA interaction at the mouse mammary tumor virus glucocorticoid response element. J. Biol. Chem. 265, 17222–17229 (1990)
Gorski, S. A., Snyder, S. K., John, S., Grummt, I. & Misteli, T. Modulation of RNA polymerase assembly dynamics in transcriptional regulation. Mol. Cell 30, 486–497 (2008)
Rossetti, L. et al. Specific interactions of the telomeric protein Rap1p with nucleosomal binding sites. J. Mol. Biol. 306, 903–913 (2001)
Sikorski, R. S. & Hieter, P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122, 19–27 (1989)
Janke, C. et al. A versatile toolbox for PCR-based tagging of yeast genes: new fluorescent proteins, more markers and promoter substitution cassettes. Yeast 21, 947–962 (2004)
Gelbart, M. E., Rechsteiner, T., Richmond, T. J. & Tsukiyama, T. Interactions of Isw2 chromatin remodeling complex with nucleosomal arrays: analyses using recombinant yeast histones and immobilized templates. Mol. Cell. Biol. 21, 2098–2106 (2001)
Hoffman, C. S. in Current Protocols in Molecular Biology, Vol. 2 (eds Ausubel, F.M. et al.) 13.11.1–13.11.4 (John Wiley and Sons, 1997)
Cesaroni, M., Cittaro, D., Brofzi, A., Pelicci, P. G. & Luzi, L. CARPET: a web-based package for the analysis of ChIP-chip and expression tiling data. Bioinformatics 24, 2918–2920 (2008)
Liu, X., Brutlag, D. L. & Liu, J. S. BioProspector: discovering conserved DNA motifs in upstream regulatory regions of co-expressed genes. Pac. Symp. Biocomput. 6, 127–138 (2001)
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004)
Frith, M. C. et al. Detection of functional DNA motifs via statistical over-representation. Nucleic Acids Res. 32, 1372–1381 (2004)
Acknowledgements
We thank T. Kaplan and O. Rando for help with their turnover model, T. Palpant and S. Adar for help with time course experiments, and A. Leonardo Iniguez and H. Rosenbaum of Roche Nimblegen for pre-release custom HD4 12-plex microarrays. This work was supported by the US National Institutes of Health (NIH) Grant R01-GM072518 (to J.D.L.), and the intramural program of the NIH, National Cancer Institute, Center for Cancer Research (to J.G.M. and F.M.). F.M. was also supported in part by the Region Ile-de-France in the framework of C’Nano IdF, the nanoscience competence center of Paris Region.
Author information
Authors and Affiliations
Contributions
C.R.L., S.E.H. and J.D.L. designed the study. C.R.L. and S.E.H. performed the experiments. F.M. developed and implemented the binding dynamics model. C.R.L., F.M., J.G.M. and J.D.L. performed data analysis. C.R.L., F.M., J.G.M. and J.D.L. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Text, Supplementary Figures 1-12 and Supplementary References. (PDF 1885 kb)
Supplementary Table
This file contains Supplementary Table 1 which shows residence times and genomic coordinates for 439 Rap1 targets. (XLS 98 kb)
Rights and permissions
About this article
Cite this article
Lickwar, C., Mueller, F., Hanlon, S. et al. Genome-wide protein–DNA binding dynamics suggest a molecular clutch for transcription factor function. Nature 484, 251–255 (2012). https://doi.org/10.1038/nature10985
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10985
This article is cited by
-
Acetylation reprograms MITF target selectivity and residence time
Nature Communications (2023)
-
A spatio-temporally constrained gene regulatory network directed by PBX1/2 acquires limb patterning specificity via HAND2
Nature Communications (2023)
-
Molecular and evolutionary processes generating variation in gene expression
Nature Reviews Genetics (2021)
-
Single molecule studies reveal that p53 tetramers dynamically bind response elements containing one or two half sites
Scientific Reports (2020)
-
Network Walking charts transcriptional dynamics of nitrogen signaling by integrating validated and predicted genome-wide interactions
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.