Abstract
Extensins are plant cell wall glycoproteins that act as scaffolds for the deposition of the main wall carbohydrate polymers, which are interlocked into the supramolecular wall structure through intra- and inter-molecular iso-di-tyrosine crosslinks within the extensin backbone. In the conserved canonical extensin repeat, Ser-Hyp4, serine and the consecutive C4-hydroxyprolines (Hyps) are substituted with an α-galactose and 1–5 β- or α-linked arabinofuranoses (Arafs), respectively. These modifications are required for correct extended structure and function of the extensin network. Here, we identified a single Arabidopsis thaliana gene, At3g57630, in clade E of the inverting Glycosyltransferase family GT47 as a candidate for the transfer of Araf to Hyp-arabinofuranotriose (Hyp-β1,4Araf-β1,2Araf-β1,2Araf) side chains in an α-linkage, to yield Hyp-Araf4 which is exclusively found in extensins. T-DNA knock-out mutants of At3g57630 showed a truncated root hair phenotype, as seen for mutants of all hitherto characterized extensin glycosylation enzymes; both root hair and glycan phenotypes were restored upon reintroduction of At3g57630. At3g57630 was named Extensin Arabinose Deficient transferase, ExAD, accordingly. The occurrence of ExAD orthologs within the Viridiplantae along with its’ product, Hyp-Araf4, point to ExAD being an evolutionary hallmark of terrestrial plants and charophyte green algae.
Similar content being viewed by others
Introduction
Protein O-linked glycans in animal and plant cells surround the cell where they function as a surface protectant1,2,3,4. One group of such O-glycoproteins in plants is the extensins, which define a subgroup of the hydroxyproline rich glycoprotein (HRGP) superfamily and incorporate characteristic β-L-arabinofuranoside repetitive glycosylation motifs5,6,7. Whereas the extensin genes in their entirety are not conserved, the extensin glycosylation motifs and a core glycosylation machinery are conserved throughout the green plant lineage. In terrestrial plant cell walls, extensins constitute a minor component of both structural and functional importance during wall deposition and remodeling8. In contrast, in certain algae such as the chlorophyte Chlamydomonas reinhardtii, the wall is primarily made up of extensin-type glycoproteins9,10. The consecutive proline units in the canonical extensin repeat (SPPPP) (reviewed in Lamport et al.4), which is conserved in both chlorophytes and streptophytes [charophyacean green algae (CGA) and terrestrial plants], are hydroxylated by the glycosylation-defining prolyl-4-hydroxylases (P4Hs) to yield 4-Hydroxyproline (Hyp), which are subsequently O-glycosylated by the glycan initiating and elongating enzymes4. Extensins of chlorophyte green algae are often more diverse, with the two innermost β-linked arabinofuranoses (Arafs) shared with streptophytes9 implying that the extensin type O-glycosylation predates extensins of today. While the consecutive Hyps in C. reinhardtii are substituted with occasionally methylated l-Araf and d-Galactofuranose (Galf)9 (Fig. 1B), consecutive Hyps in CGAs and terrestrial plants are substituted with one to five Arafs with the linkages β-1,4, β-1,2, β-1,2 and α-1,311, with the linkage of the rare fifth Araf being currently unknown12. A sidechain of four Araf residues on Hyp (Hyp-Araf4) has not been found in C. reinhardtii and we have analyzed Ostreococcus tauri (a prasinophyte) and some taxa of the Pseudochloris wilhemii complex (chlorophytes) without detecting Hyp-Araf4 (unpublished observations). It thus appears that Hyp-Araf4 is a hallmark of CGA and terrestrial plant extensins. In addition, the serine in the glycosylation motif may be substituted with an α-1,3-linked galactose unit13. Surprisingly, Hyp-Araf3, but not Hyp-Araf1, −2 or −4, is found in a number of proteins that do not feature the contiguous Hyp-glycosylation motif. These include small peptide hormones (reviewed in Matsubayashi 201014), wall associated kinases (WAKs) (reviewed in Kohorn and Kohorn 200915) and proline-rich, extensin-like receptor kinases (PERKs) (reviewed in Nakhamchik et al.16). Furthermore, Hyp-Araf1–3 are detected in group 1 and 5 major pollen allergens17 which also do not comply with the contiguous Hyp definition of glycosylation sites. Elongation of Hyp-Araf3 with α-linked arabinose in CGAs and terrestrial plants seems to be confined to clustered proline arabinosylation of extensin substrates, and regulated accordingly.
(A) Canonical extensin repeat structures and extensin glycosylation enzymes. In the classic extensin repeat motif, SP3–59,10, serine is substituted with an α-1,3 linked galactose and the hydroxyprolines (Hyps) are substituted with β- and α-linked arabinofuranoses (Araf) of lengths from 1 to 5 in terrestrial plants and CGAs. Genes encoding GTs involved in building the structures are indicated. (B) The innermost two Arafs also exist in the chlorophyte alga C. reinhardtii, where they are extended by an α-1,2-galactofuranose (α-1,2Galf) and a β-1,5Araf, which may additionally be methylated at either the C-6 or C-3 position, respectively9. (C) Clade structure of family GT47. The tree was built from Arabidopsis thaliana (At), Physcomitrella patens (Pp), Selaginella moellendorffii (Sm), Oryza sativa (Os) and Klebsormidium flaccidum. The corresponding Newark tree file, GT47tree.txt, is provided as Supplementary Data. No known function has been demonstrated for members of the F-clade; GTs with known function, in the other clades are indicated (MUR361, ARAD62, XGD163, IRX7/IRX164,65).
The extensin glycosylation machinery is rather well characterized. A single gene encoding serine α-1,3-galactosyltransferase (SGT)18 of family GT96 was recently identified. Of the 13 glycosylation defining P4Hs present in A. thaliana19 AtP4H2, AtP4H5 and AtP4H13 have been implicated in hydroxylation of consecutive Pro residues of extensins20,21. Subsequent Hyp-linked arabinosylation is initiated by β-4-hydroxyproline arabinosyltransferases 1–3 (HPAT1-3)22 of family GT95. Three genes of family GT77, encoding the Reduced Residual Arabinose 1–3 (RRA1-3) that extend Hyp-Araf1 to Hyp-Araf2, were among the first β-arabinosyltransferases to be identified20,23. Shortly thereafter Xylo-EndoGlucanase 113 (XEG113), also of family GT77, was demonstrated to elongate Hyp-Araf2 to Hyp-Araf324. The extensin specific α-arabinosyltransferase that extends Hyp-Araf3 to Hyp-Araf4, however, has not yet been identified.
Here we report the identification of an extensin specific α-arabinosyltransferase, named ExAD, that extends Hyp-Araf3 to Hyp-Araf4 and map the occurrence within Viridiplantae of ExAD orthologs and its product, Hyp-Araf4, which appears to be hallmark of terrestrial plant and CGA cell wall extensins. The evolution of ExAD and its product Hyp-Araf4 is addressed and discussed.
Results
Candidate selection for the extensin specific α-Araf T activity
The RRA β-arabinosyltransferases in GT77 clade A were discovered by a comparative phylogenetic approach, which relied on the co-existence in this clade of terrestrial plant and C. reinhardtii genes, which we hypothesized to be involved in coat protein glycosylation. The co-existence of C. reinhardtii and plant glycosyltransferases in clade E of CAZy family GT47 led us to hypothesize that members of GT47-E were involved in extending Hyp-Araf3 to Hyp-Araf4 in extensins, as this particular linkage requires an inverting activity. The comparative phylogenetic approach used to identify the RRA genes relied on the fact that both Arabidopsis thaliana and C. reinhardtii feature Hyp-Araf2 in their cell walls and that both species are represented in the A-clade of GT7723. Direct application of this approach for identifying ExAD was not feasible since C. reinhardtii was reported to feature neither Hyp-Araf3 nor Hyp-Araf49 (Fig. 1).
However, our mass spectrometric analysis of C. reinhardtii extensin side chains additionally identified Hyp-Araf3 together with the earlier reported structures (see Supplementary Fig. S1). This observation is corroborated by analysis of the C. reinhardtii genome, which implies XP_001692787.1 and XP_001690605.1 to be orthologues of XEG113, the retaining β-arabinosyltransferase of family GT77 that catalyzes the formation of Hyp-Araf3. Nonetheless, Hyp-Araf4 was not detected. We reasoned that ExAD might be homologous to particular Hyp-Araf2, Hyp-Araf3, and/or Hyp-Araf2-Galf extending GTs from C. reinhardtii, and focused on the E-clade of family GT47 (Fig. 1C), which comprises a single A. thaliana gene, At3g57630, which we selected for further analysis.
Mutant knock out and molecular complementation links At3g57630 to arabinosylation of extensins
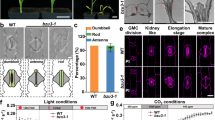
Two T-DNA knock-out lines of At3g57630 (Supplementary Fig. S2), designated exad1-1 and exad1-3, displayed a truncated root hair phenotype (Fig. 2A,B) similar to those previously reported for mutants of the extensin glycosylation-defining and elongating enzymes P4H2, P4H5, P4H13, RRA3, XEG11320,21. The root hairs were restored to WT lengths upon reintroduction of the At3g57630 cDNA under control of the CaMV 35S virus promotor in one of the mutant lines (Fig. 2A).
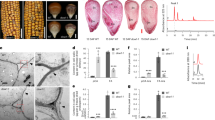
Root hairs of 10 days old light grown seedlings (N = 30) prepared and quantified as described in Material and Methods, NS, not significantly different, **denotes statistical significance at 1% level (student’s t-test). (A) Direct inlet ESI-MS of Ba(OH)2 hydrolysates of an extensin enriched fraction of 3 week old roots of WT and exad1-1 (B). LC-ESI-MS of Ba(OH)2 hydrolysates of an extensin enriched fraction of 3 week old rosette leaves of exad1-1, exad1-1 complemented with ExAD cDNA (exad1-1-35S::ExAD) and WT (extracted ion traces of [M + H+|Na+]+ 396 (Hyp-Araf2), 528/550 (Hyp-Araf3) and 660/682 (Hyp-Araf4) each eluting as twin peaks due to the C-4 R/S stereo chemistry of hydroxyproline (Hyp) (C). For MS2 spectrum of Hyp-Araf4 of exad1-1 -35S::ExAD see Supplementary Fig. S3. Hydroxyproline content (D), mono saccharide composition (E) and linkage analysis (F) of extensin enriched Ba(OH)2 extracts of 10 day old light grown seedlings of WT and the extensin glycosylation mutants (sgt1, rra3, xeg113-2, exad1-1).
Direct inlet Electrospray Ionization Mass Spectrometry (ESI-MS) analysis of an extensin-enriched fraction of roots of the exad1-1 T-DNA knock-out line subjected to Ba(OH)2 treatment (mediating peptide bond hydrolysis) showed the presence of Hyp-Araf1, Hyp-Araf2 and Hyp-Araf3, but the complete absence of Hyp-Araf4 (Fig. 2B,C). Again, introduction of the At3g57630 cDNA restored the Hyp-Araf4 species (Fig. 2C). Whereas α-arabinofuranosidase treatment trimmed the Hyp-Araf4 to Hyp-Araf3 in the WT and exad1-1 complemented line, Hyp-Araf1–3 of the T-DNA mutant remained resistant, strongly suggesting that the Hyp-Araf1–3 species of the mutant are the β-linked innermost WT Arafs as anticipated (Supplementary Fig. S3).
In 10 day old seedlings the Hyp content of an extensin enriched extract of the exad1-1, xeg113 and rra3 mutants is unaltered compared to WT (Fig. 2D). Corresponding monosaccharide composition analysis of Ba(OH)2 treated extensin-enriched extracts, however, showed significant reductions in arabinose; mutants of progressively earlier-acting extensin arabinosyltransferases had an increased arabinose reduction (xeg113, rra3), while the exad1-1 mutant had the smallest reduction (Fig. 2E). Galactose levels were only affected in the sgt1 mutant, which displayed a ca 30% reduction with respect to WT levels (Fig. 2E). This substitution pattern was substantiated by linkage analysis of the extensin extracts of xeg113, rra3 and exad1-1. Terminal arabinose was present at the same level in all mutants, indicating that the number of extensin arabinosyl-sidechains remained the same. The 3-linked arabinofuranosyl linkage was reduced in all three mutants to the same low level, whereas the 2-linked arabinofuranosyl linkage was not reduced in the exad mutant, but progressively in the xeg113 and rra3 mutants. These data are consistent with a role of ExAD in adding the last arabinosyl-residue at the 3 position, as shown in the model in Fig. 1A. In the sgt1 mutant 2- and 3-linked arabinose remained at WT levels (Fig. 2F) and only terminal galactose was reduced to ca. 35% of WT level. The relative (mol %) up-regulation of 5-, 2,5- and 3,5 linked arabinose and 3-linked galactose, which are diagnostic of arabinogalactan proteins, and 4-linked xylose from xylan can be attributed to compensation of the strong decrease of 2- and 3 and linked arabinose. Hence, extensin galactosylation and arabinosylation appear to be independent. In agreement with this, the Hyp-Araf1–4 distribution of an extensin-enriched extract of sgt1 was found to be similar to WT, as evidenced by LC-ESI-MS analysis (data not shown).
Taken together, we conclude that At3g57630 encodes an Araf T that adds the fourth α-1,3-linked Araf onto Hyp-Araf3 in classical extensin repeats. At3g57630 was named Extensin Arabinose Deficient (ExAD) after the mutant phenotype, accordingly.
In vitro assays of heterologous expressed ExAD
In order to assess the substrate specificity of ExAD we produced the soluble part of the At3g57630 protein by secreted expression in P. pastoris and baculovirus High Five cells. Assays for arabinosyltranferase activity were conducted using chemically synthesized UDP-β-L-Araf as donor substrate. Either a Hyp-Araf1–3 mixture [obtained as a Ba(OH)2 hydrolysate of the mutant exad1-1 (Fig. 2B)] or the human Mucin 1 peptide featuring Hyp-Araf3 [expressed in Tobacco Bright Yellow 2 BY2 suspension cells25], were used as acceptor substrates (Supplementary Fig. S4). Neither single Hyp nor Mucin 1 peptide substituted with Araf3 proved to be substrates. These negative results are in line with the reported extensin-type arabinosylation of endogenous and ectopically expressed substrates, where Hyp-Araf3 appear to be necessary but not sufficient for defining the ExAD acceptor substrate.
ExAD expression profiling
Geneinvestigator Affymetrix data (https://genevestigator.com/gv/plant.jsp) suggest that At3g57630 is ubiquitously expressed at medium to high levels in all major plant tissues (Supplementary Fig. S5). We analyzed extensin-enriched fractions from 7 day old etiolated seedlings as well as the major tissues (roots, rosette leaves, stems, flowerbuds and siliques) of 6 week old bolting plants of WT and exad1-1; we found that Hyp-Araf4 were present in all tissues in WT but absent in the mutant, thus unambiguously linking the single ExAD gene to the addition of the fourth extensin specific Araf of extensins in A. thaliana (Fig. 3).
In order to access the global developmental and organotypic consequence of knocking out ExAD we prepared extensin enriched fractions from (etiolated) 7 day old seedlings (See Fig. 2) and the major tissues (roots, rosette leaves, stems, flower buds and siliques) of 6- week old bolting plants of WT and exad1-1 and subjected these to LC-ESI-MS analysis and the extracted ion traces of [M + H+|Na+]+ (m/z 264 (Hyp-Araf1), 396 (Hyp-Araf2), 528/550 (Hyp-Araf3) and 660/682 (Hyp-Araf4) were semi quantified as described in the Material & Methods section and presented as the relative occurrence of each Hyp-Araf specie. In all tissues investigated Hyps substituted with Arafs exceeding the length of 3 were only found in WT, thus globally linking the single ExAD gene to the addition of the fourth extensin specific Araf of extensins in A. thaliana.
This suggests that ExAD represents a ubiquitously expressed housekeeping gene. The exad1-1 and exad1-3 root hair mutant phenotype (Fig. 2A) is in accordance with high ExAD expression in root hair cells and with similar high expression profiles of previously characterized extensin Araf Ts (HPAT322, RRA320, XEG11320) (Fig. 4) with strikingly similar expression to extensin specific Ser-GalT18 (SGT).
Expression of ExAD in root epidermis and root hair cells is similar to the expression pattern of the previously characterized extensin Araf Ts HPAT322, RRA320, XEG11320) and the Ser-GalT18 (SGT). Expression patterns were obtained using eFP-Browser66 (http://www.bar.utoronto.ca/) and the dataset of Brady et al.51 as described in the Material and Methods section.
Lamport and Miller5 observed a higher proportion of Hyp-Araf4 in suspension cultures of sycamore and other dicot species suggesting that Hyp-Araf3 arabinosylation is subject to regulation. To test whether this observation is ubiquitously valid, we analyzed extensin-type glycosylation in young expanding leaves from A. thaliana, on an AGP fraction and on an extensin fraction from sycamore. We found Hyp-Araf4/Hyp-Araf3 ratios of ca 0.25 and ca. 0.7 & 1.4, respectively (Supplementary Fig. S6). This shows substantial variation in comparable tissues between species, yet the higher ratios in sycamore leaves are still substantially lower than the ratio of 4.4 observed in sycamore suspension cultures by Lamport and Miller5. Suspension culture cells might preferentially slough off a higher amount of soluble wall proteins to the medium and for compensation purposes up-regulate their synthesis, but the ratio was lower in this fraction when extracted from leaves. Thus, up-regulation of Hyp-Araf3 arabinosylation in suspension cultures cannot be ruled out.
ExAD is ubiquitously expressed and there are thus limits to how strong co-expression patterns may be expected. Still, a co-expression network comprising several genes involved in extensin biosynthesis and post translational modifications suggests a tighter co-regulation of the extensin/extensin-like glycosylation enzyme machinery core (HPAT3, XEG113, SGT1, RRA3, P4H5 & -2) and ExAD being more moderately co-expressed with the core as expected. AtEXT3 and with it several other genes encoding extensin backbones, form a separate co-expression network (Supplementary Fig. S7).
ExAD is located in the cis to medial Golgi apparatus
The full-length genomic sequence of ExAD (At3g57630) C-terminally fused to mTurquoise3 was expressed under control of the CaMV 35S promoter in leaves of Nicotiana benthamiana to elucidate the subcellular localization of the At3g57630 protein (Fig. 5). N-Glycan N-acetylglucosaminyltransferase I fused to monomeric red fluorescent protein (GnTI-mRFP)26, N-glycan α-mannosidase II fused to monomeric RFP (GMII-mRFP)27 and sialyltransferase fused to yellow fluorescent protein (ST-YFP)28 were used as markers for cis-, medial- and trans-Golgi cisternae, respectively. Fluorescence of ExAD-mTurquoise3 co-localized with medial-Golgi-targeted mannosidase II (GMII) and partly with cis-Golgi localized GlcNAc transferase I (GnTI), both under control of the CaMV 35S promoter (Supplementary Material and Methods Fig. S11). This localization in the medial part of the Golgi membrane system is in accordance with the reported location of other extensin glycosylation-defining and elongating enzymes20,21.
The full-length genomic sequence of ExAD C-terminally fused to mTurquoise3 under control of the CaMV 35S virus promoter was expressed transiently in leaves of N. benthamiana using agrobacterium mediated infiltration. Fluorescence of EXAD-mTurquoise3 (green) partly co-localized (white) with co-expressed known cis Golgi localized GlcNAc transferase I (GnTI) (A–C), fully with co-expressed medial Golgi localized mannosidase II (Man II) (D–F), and not with trans Golgi localized co-expressed Sialyl-Transferase (ST) (G–I) all C-terminally fused to either mRFP or YFP (Magenta) and under control of the CaMV 35S promoter. Co-localization is evident (white) in section F, which, due to the nature of the experiment (transient expression), also contains cells that solely express the marker GMII-mRFP (upper left in the section). Scale bar = 10 μm.
ExAD protein structure
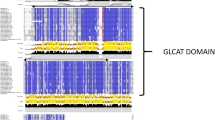
ExAD (At3g57630) encodes a typical Type-II membrane anchored protein of 793 amino acids, making it the largest of the 39 A. thaliana genes in family GT47. Our observation that a human MUC1 epitope expressed in A. thaliana was consistently arabinosylated to Hyp-Araf3 and never to Hyp-Araf425 just like the peptide hormones and that two Hyp-Araf3-containing acceptor substrates failed to work in in vitro ExAD assays lead us to hypothesize that the presence of Hyp-Araf3 does not fully define ExAD’s acceptor specificity. The additional structural requirements may be probed by ExAD through protein-protein or protein-carbohydrate interactions. Examination of the longer N-terminal domain reveals a highly structured polypeptide featuring two epidermal growth factor (EGF) like domains. These are characterized by cysteine knots, which are substrates for O-fucosylation in mammalian Notch signaling29. Figure 6A shows a partial alignment of ExAD with human Delta1, a Notch signaling component with the cysteine knot of the second EGF-like domain indicated. The first EGF-like domain also resembles a highly conserved domain in Lectin C (GlcNAc binding lectin30) albeit with an invariant Gly residue missing.
(A) ExAD (At3g57630) is a type-II membrane anchored GT with a short (11 aa) cytoplasmic tail preceding the transmembrane domain (TMD). The catalytic domain adopts a GT-B fold and is related to the mammalian type exostosin domain found in heparan sulfate synthases67. The N-terminal sequence between the TMD and the catalytic domain features two putative epidermal growth factor (EGF) like domains followed by what appears to be a flexible linker. (B) The domains that display sequence similarity with EGF motifs align with similar domains in human delta-like 1. Alignment of the first domain in ExAD and ExAD-like sequences is shown for selected Viridiplantae and haptophyte sequences. A cysteine knot, potential substrate for fucosylation, is indicated in the second domain. (C) Two curated alignments of part of the ExAD N-terminal with a partial sequence of pokeweed lectin-C. This lectin binds chitin and the region critical for binding is marked by a horizontal bar and with thicker boxes indicating fully conserved residues30.
We have also scanned full length ExAD homologous sequences, used in Fig. 1C, for domains in an unsupervised manner using SALAD31 (Supplementary Fig. S8) which is a comparative plant genomics motif-based database of protein annotations where similarity clustering is based on distribution patterns of such motifs in the query sequences. Interestingly both clades of sequences, including the two available full-length prasinophyte ExADs, contain EGF-like domains (domain 4 and 9) along with C-terminal exostosin-specific domains. The haptophyte sequences included in the tree feature one EGF-like domain; only in the rhodophyte Chondrus crispus sequence (which is a outgroup and not bona fide member of clade E) are the EGF-like domains completely missing. Beside the EGF-like domains in the N-terminal part, exostosin-specific domains are conserved. Obviously, the long N-terminal of the ExAD sequence is not merely an extended, flexible linker inserted between membrane anchor and catalytic domain. Rather it features highly structured domains that could recognize carbohydrate epitopes on the extensin polypeptide or serve the purpose of interacting with other protein(s) essential for its function.
Evolution of ExAD and ExAD-like genes
The substantial number of C. reinhardtii genes in clade E of GT47, combined with the absence of Hyp-Araf4 in C. reinhardtii, raises the question of when in evolution a GT47 clade E protein was first recruited to transfer an Araf to Hyp-Araf3 to form Hyp-Araf4. Likewise, what sets these GTs apart from the other clade E GTs with different catalytic activities. We propose to name the Hyp-Araf4 forming GTs ExAD and the other clade E members ExAD-like.
Figure 7 shows an expansion of the E-clade with prasinophyte, chlorophyte and charophyte sequences. Red algae were selected as an outgroup for rooting the tree, which comprises one charophyte sensu stricto (CGA), one streptophyte senso lato and three chlorophyte sub-clades for which naming are proposed. All clades feature ancestral prasinophyte sequences. Close examination (see Supplementary Fig. S9) reveals that while some charophytes are present in both the streptophyte and charophyte clades, others are present in only one of the two. Embryophytes were only found in sub-clade E1. The simplest assumption that accounts for the structure of the whole E-clade is that the ancestral organism had two ExAD-like genes, each giving rise to the clades on either side of the red algal clade. Embryophytes appear to have lost one of them and Klebsormidium flaccidum, the only CGA that has had its genome sequenced32, is only present in the opposite CGA-clade, subclade E2. It is noteworthy that none of the chlorophyte sequences were predicted to be membrane-anchored, while those of the terrestrial plants and K. flaccidum all appear to be type-II membrane-anchored proteins. All other CGA-sequences are transcripts from the 1 Kp-project (www.onekp.com) and none of these have complete N-terminals.
Rhodophyte homologs are used as outgroup. Three chlorophyte, one pure CGA and one streptophyte sub-clade, are proposed. Subtending prasinophyte sequences are not considered included in the sub-clades. All CGAs except Klebsormidium flaccidum are transcripts, i.e. partial. Chlamydomonas reinhardtii has 19 clade-E sequences. Only one is indicated, the one with the highest similarity to AtExAD. The complete tree with all taxa is provided as Supplementary Fig. S9.
To ascertain the presence of Hyp-Araf4 we subjected a Ba(OH)2 hydrolysate of K. flaccidum Alcohol Insoluble Residue (AIR) to LC-ESI-MS analysis.
The MS/MS spectrum in Fig. 8 demonstrates the presence of Hyp-Araf4. Taken together with the presence of Hyp-Araf2 and Hyp-Araf3 as evidenced by MS/MS analysis (Supplementary Fig. S10), it appears that K. flaccidum has an embryophyte-type extensin arabinosylation, which is thus not confined to species with an ExAD sequence belonging in sub-clade E1. K. flaccidum is an early diverging CGA amongst those with a polysaccharide-based cell wall, suggesting that Hyp-Araf4 is a general characteristic of streptophytes and thus that both the charophyte and embryophytes harbor true ExAD sequences.
Discussion
The study of Lamport and Miller5 documented the occurrence of Embryophyte-type Hyp-arabinosylation as far back as mosses and liverworts, albeit with overall shorter side-chains than in higher plants. A recent bioinformatics study33 pointed out that only genes of vascular plants encode extensins in the strict sense. The genome of the moss Physcomitrella patens does not encode repeated SP3–4 (SP3–4SP3–4) motifs, as do the genomes of A. thaliana, the lycopod Selaginella moellendorfii, the charophycean green alga K. flaccidum, the chlorophyte C. reinhardtii, the prasinophytes Micromonas pusilla and O. tauri. P. patens and K. flaccidum lack the cross-linking motif SP3–5X2–3YXY that is present in the proteomes of the other species, yet wall integrity in the moss was found to rely on Hyp-arabinosylation in a recent study of knock-outs of HPAT34. These observations lead us to conclude that the O-glycosylation machinery and the glycosylation sites preceded the evolution of extensins sensu strictu, which should be regarded a vascular plant invention, in accordance with the study by Liu et al.33. While this would make attempts at tracking the phylogeny of extensin genes to the origin of Viridiplantae uncertain at best, this does not hold true for the arabinosylating GTs, as evidenced by the discovery of the RRAs of family GT77 and ExAD of family GT47 (present study). The C. reinhardtii genome comprises putative orthologs of RRAs and XEG11335 and it encodes several GTs belonging to family GT47, all belonging to the E-clade. It seemed reasonable that these might be involved in synthesis of the elaborate extensin-like side-chains found in its coat proteins. The extensive gene duplication would suggest functional diversification in C. reinhardtii so that the absence of Hyp-Araf4 would not preclude that embryophyte orthologs could encode the Hyp-Araf3 elongating α-arabinosyltransferase. Hence, At3g57630 was selected for characterization.
ExAD is a single copy gene in A. thaliana making mutant analysis promising; indeed we observed both the expected root hair phenotype, and the biochemical phenotype in terms of presence of Hyp-Araf1–3 but not Hyp-Araf4 in the mutant. The root hair phenotype does not imply a role of Hyp-Araf4 confined to root hairs. Hyp-Araf4 is present and ExAD is expressed throughout the plant. Root hairs are exposed cells in which loss of cell wall integrity readily leads to an observable phenotype.
Extensin from the Arabidopsis ExAD mutant was insensitive to α-arabinofuranosidase treatment, suggesting that it comprised exclusively β-arabinofuranosides. Complementation did not fully restore the WT phenotype, but qualitatively the expected re-emergence of Hyp-Araf4 was observed, which could be trimmed down to Hyp-Araf3, diagnostic of the exad mutant, by α–arabinofuranosidase treatment.
Our attempts to demonstrate activity of heterologously expressed ExAD in vitro were not successful, even though two expression hosts combined with two a priori relevant acceptor substrates were evaluated and UDP-Araf was used as a donor substrate, thus bypassing the complexities of GT75 involvement and membrane transport. The protein structure of ExAD points to protein-protein or protein-carbohydrate interactions and precise substrate recognition is required for ExAD to distinguish between e.g. peptide hormones and true extensins. We thus assume that heterologously expressed ExAD was not presented with the correct acceptor or not accompanied by a required protein partner. ExAD is Golgi localized as expected and the sub-compartmental localization that we observed corroborates the notion of an assembly line for arabinosylation of extensins in the secretory pathway.
The ExAD gene product is nearly twice in size as a typical single domain GTs of family 47. The longer N-terminal extension was examined for sites for post-translational modification e.g. for regulatory purposes, as well as for domains suggestive of either protein-protein or protein-carbohydrate interaction. GTs with domains for carbohydrate recognition are well-known from starch biosynthesis36 and at least one interesting example is known from a mycobacterial arabinosyltransferase featuring a C-terminal CBM37. N-terminal lectin domains are observed in several plant GTs of family GT31, including the hydroxyproline-β-O-galactosyltransferase AtGALT238.
The observation discussed above that endogenous peptide hormones and human mucin sequence are glycosylated with tri-arabinofuranose glycans but are not acceptor substrates for ExAD could lead to the proposition that recognition of the single galactosyl sidechain on serine would allow ExAD to discriminate extensins from other β-arabinosylated proteins. Serine galactosylation was recently shown to require C4 hydroxylated proline (O) in the ‘KSOOOO’ peptide substrate in vitro18. This suggests that galactosylation depends on hydroxylation but not on arabinosylation of the KSPPPP motif and leaves the possibility that ExAD could probe the peptide for the galactosyl residue. Knock-out of SGT1 did not lead to an altered arabinosylation pattern, including an unaltered Hyp-Araf4 level, speaking against this. Another possibility is recognition of β-arabinofuranosides in the vicinity of the Hyp-Araf3 is required for elongation. The ordering of extensin side-chains with respect to each other is unknown, making this question difficult to address. A domain with similarity to a conserved region in lectin C is present in the N-terminal of ExAD. The pokeweed orthologue of lectin C binds chitin, with the ‘CCS’ sequence preceding a residue that is directly involved in carbohydrate binding; this motif is strictly conserved and also found in other α-D-GlcNAc-binding proteins, such as wheat germ agglutinin and nettle agglutinin isolectin 630. The ExAD sequence does not comprise this precise motif, but is anyway not expected to bind GlcNAc. However, sequences from C. reinhardtii, rice and S. moellendorffii produce similar alignments, but ExAD-like sequences from P. patens and micromonas do not (data not shown). Since Hyp-Araf4 is known from both grasses and mosses we conclude that this domain is unlikely to represent a lectin domain.
Another striking feature is the occurrence of two epidermal growth factor (EGF) like domains of which the first is overlapping with the lectin C similar sequence. EGF-domains are, for example, featured in several proteins of the NOTCH signaling pathway in non-plant eukaryotes, where it is involved in cell-cell interactions39. EGF-domains are e.g. present in the apoplastic part of Wall-Associated Kinases (WAKs) involved in cell wall sensing, which bind pectin or short pectic oligogalacturonic acid fragments15. However, there is no homolog of the NOTCH receptor itself in A. thaliana clearly indicating that plants do not have a NOTCH signaling pathway analogous to that of animals. EGF features so-called cysteine knots that are fucosylated by members of CAZy-family 65 or 6829. The A. thaliana genome comprises genes classified to GT68 but not to GT65. A glucosyltransferase from family GT90 glycosylates EGF domains in Notch signaling40 and A. thaliana also has genes classified to family GT90. Sequence similarity does not prove formation of the –S-S-bridges that make up cysteine knots. Putative cysteine knots are also found in ExAD or ExAD-like sequences from C. reinhardtii, rice and S. moellendorffii. EGF-domains usually occur in many copies in apoplastic plasma membrane bound proteins among which the WAKs are typical examples. ExAD is Golgi localized and the domain occurs only twice suggesting that these highly structured protein domains have been recruited to serve a new function in ExAD. The two EGF-like domains are present in all ExAD and ExAD-like sequences from the green plant lineage, one in haptophyte sequences and none in the red algal sequences. It is noteworthy that red algae are not known to have prolyl hydroxylases and hence cannot make HRGPs. The red algal sequences homologous to ExAD thus cannot fulfill the same biological function. The observation that the two haptophytes included in the analysis have a GT47 sequence which features an EGF-like domain and which is more similar to A. thaliana EXAD than to any other A. thaliana sequence, suggests that ExAD predates the green plant lineage. Late acquisition of a metazoan EGF by horizontal gene transfer may thus be ruled out. The K. flaccidum proteins are minimalistic ExADs in the sense that the SALAD analysis only detected domains that are conserved in all full length ExAD sequences analyzed. The included prasinophytes are also only sharing a limited numbers of domains with core chlorophytes and plants. The differences observed in the N-terminus of ExAD and ExAD-like are distinct, but hard to link to biochemical activity, as we have only identified the activity of AtExAD and do not know if the ExAD-like GTs from C. reinhardtii are responsible for the Hyp-Araf2-Galf-Araf series of extensin-like side-chains for example.
Only Chlorokybus atmophyticus among extant CGA species represents an earlier evolutionary state than K. flaccidum yet features a polysaccharide-based cell wall41 (Mesostigma viride is also charophyacean but it’s wall is made of elaborate proteinaceous scales42). Our observation that extensin side-chains in A. thaliana and K. flaccidum are indistinguishable may be viewed as being at odds with the conclusion above that true extensins are a vascular plant invention. The extensin repetitive sequence structure, however, prevents a precise mapping of the origin of land plant extensins and all four extensin genes of K. flaccidum have been classified as ‘other chimeric EXTs’39. The important observation is that the Hyp-Araf4 structure must be older than the classical extensins and it may be inferred that glycosylation machinery did not evolve in step with the extensins. Prolyl hydroxylation is a shared eukaryotic phenomenon, with Hyp-Araf3 common to the green plant lineage. Hyp-Araf4 is characteristic of streptophytes at least as far back as K. flaccidum, and classical extensins with their crosslinking motifs are an embryophyte feature. The finding that the K. flaccidum ExADs are type-II membrane anchored proteins like those of embryophytes, yet belong to a clade E2 that comprises no vascular plants raises the question whether membrane anchoring evolved twice in the gene family. More sequence information and extensin structures from CGA families are required before is it possible to determine whether ExAD-like sequences were recruited to become true ExADs more than once and whether this recruitment is correlated with membrane anchoring.
Identification of the acceptor substrate recognized by ExAD and determining the biochemical function of the EGF-like domains are important challenges still unresolved.
Material and Methods
Plant Material, growth conditions and transformation of A. thaliana
T-DNA insertional mutants in At3g57630 (exad1-1 (SAIL_843_G12), exad-2 (SALK_206288C), and exad1-3 (SALK_204414C) were obtained through Syngenta (SALK-204414C) and the Salk Institute (SAIL lines), respectively.
Dark grown etiolated seedlings were grown for 7 days on Murashige-Skoog-medium 0 (MS0) plates in a controlled-environment growth chamber (Percival AR-60 I, Boone, IA, USA), 20 °C and 70% relative humidity. Bolting plants of exad1-1 &-3 and WT A. thaliana ecotype Columbia (Col-0) were grown for 4–6 weeks in soil in a controlled-environment growth chamber (Percival AR-60 I, Boone, IA, USA) at a photosynthetic flux of 100–120 mol photons m–2 s–1 at 8 h light/16 h dark cycle, 20 °C and 70% relative humidity. Growth conditions for soil grown plants were essentially as described23.
Agrobacterium tumefaciens strain pGV3850 was used for agrobacterium-mediated transfomation of A. thaliana (ecotype Col-0) and mutant T-DNA mutant exad1-1 (SAIL_843_G12), which was performed essentially as described by Clough and Bent (1998). In brief, Agrobacterium was transformed by electroporation and selected with appropriate antibiotics. Agrobacterium cultures were grown overnight in Luria-Bertani (LB) medium, harvested by centrifugation, and re-suspended in a 1 l buffer containing 2.3 g MS-salt (Sigma no. M-5524), 3.2 g Gamborg-5 media (Sigma no. G-2519), 50 g sucrose and 300 μl silwet-L77, Adjusted to pH 5.7 with 1 M KOH. The aerial part of bolting A. thaliana plants were submerged for 5 min in the bacteria suspension and allowed to dry. Scoring of primary transformants: Seeds were sterilized with 95% ethanol and 5% hypochlorite (Clough and Bent 1998) and plated on Murashige and Skoog plates (1 l H2O, containing 4.3 g Murashige and Skoog-salt, 10 g sucrose and 8 g agar (Scharlau no. 07004), pH 5.8) containing appropriate antibiotics stored in the dark for 2 days at 4 °C for synchronization of germination and transformants selected after 10–14 days and transferred on to soil.
Bioinformatics
The general clade structure of family GT47 was determined as in Harholt et al.43 augmented with sequences from Klebsormidium flaccidum so that the clade structure now also embraces an early diverging charophycean green alga. E-clade sequences from this analysis were included in the analysis of the expanded GT47 clade E and were supplemented with sequences from full-length sequences from C. reinhardtii, Volvox carteri, Micromonas sp. RCC299, Bathycoccus prasinos, Chondrus crispus, Emiliania huxleyi, Chrysochromulina tobin and a large number of charophyte, chlorophyte and rhodophyte partial sequences forthcoming from the 1 Kp project44,45,46,47. The 1 Kp sequences were retrieved by BLAST as described43 using a cut off of 1E-25 in E value due to the shorter reads of the 1 Kp sequences. Trees were constructed on the basis of curated Muscle alignments48 followed by PhyML as described35. Identification of EGF-like domains was carried out using Interproscan49 and Phobius50 as a component of Interproscan for prediction of signal peptides and membrane anchors.
Organotypic and developmental expression profiles were obtained by use of Geneinvestigator (https://genevestigator.com/gv/plant.jsp).
The electronic Fluorescent Pictograph (eFP) – Browser available at http://www.bar.utoronto.ca/ was used to map the expression of ExAD and related genes in the root based on data published51.
Co-expression data and trees were obtained from ATTED (http://atted.jp).
SALAD analyses were performed as described (http://salad.dna.affrc.go.jp/salad/en/) using standard settings.
Root hair phenotype analysis
Seedlings were germinated on agar plates in a Percival incubator at 22 °C in a growth room for 10 days at 140 μmol m−2 s−1 light intensity. For quantitative analysis of root hair phenotypes, 200 fully elongated root hairs were measured (no roots = 30) from seedlings grown on vertical plates for 10 days. Values are reported as the mean ± SD using the ImageJ 1.50 b software. Measurements on images were captured with an Olympus SZX7 Zoom microscope equipped with a Q-Colors digital camera.
Linkage analysis and monosaccharide composition analysis of extensin extracts
Extensin extraction from etiolated seedlings (7 day old), monosaccharide composition analysis and glycosidic linkage analysis of extensin enriched Ba(OH)2 extracts were performed as described24. In brief, alcohol- insoluble residue (AIR) of the seedlings was treated with aqueous buffer, pectinases and endoglucanases to remove the matrix wall polysaccharides. The remaining residue was treated with saturated barium hydroxide (0.22 M) to solubilize the extensins. The monosaccharide composition of the extensin fraction was determined by hydrolyzing cell wall material in trifluoroacetic acid (TFA) followed by alditol acetate derivatization and GC-MS analysis. Glycosidic linkage analysis on the Ba(OH)2 fraction was performed by GC-MS analysis of partially methylated alditol acetates as described52 with modifications53. The hydroxyproline (Hyp) content of was determined by the colorimetric assay as described54.
Subcellular localization of ExAD transiently expressed in leaves of N. benthamiana
DNA constructs comprising the first 77 N-terminal amino acids (CTS region) of N. tabacum β1,2-N-acetylglucosaminyltransferase I fused to mRFP (GnT1-mRFP)55 and the first 52 N-terminal amino acids of the A. thaliana α-mannosidase II fused to mRFP (GMII-mRFP)55 were kindly provided by Dr. Richard Strasser, University of Natural Resources and Life Sciences-BOKU, Austria.
Expression constructs were transformed into the Agrobacterium tumefaciens strain pGV3850 and Agrobacterium-mediated transient transformation of N. benthamiana leaves was carried out as described56. The Agrobacterium strains harbouring the appropriate construct (OD600 = 0.2) were co-infiltrated with the strain harbouring the p19 construct (OD600 = 0.1). Image acquisition was performed with a Leica SP5 confocal laser scanning microscope (Leica Microsystems GmbH) equipped with a 63× objective lens using CFP, YFP and RFP filter setting provided by Leica.
Complementation and Gfp fusion constructs
The coding region of full length ExAD cDNA Genbank accession: BT011693 was obtained from Eurofins, Europe (http://www.eurofinsdna.com/ (AT3G57630-pEX-A). The primerset: FP_EXAD: CCATGGTttctcaccagaaatggaagttc, RP_EXAD_STOP: ACTAGT tta atgatgatgatgatgatgcttgtagtcatcatcgtccttgtagtcggaggtcttatgcagacattcttgg where capital letters designate NcoI and SpeI restriction sites, ‘ tta ’ the stop codon, underlining His6 and Flag (DYKDDDDK) encoding tags, using At3G5760-pEX-A as template, were used to PCR-clone the full length coding region of EXAD incl. stopcodon into open source vector pCambia 1302D (GenBankTM accession number AF234298) under the control of the Cauliflower mosaic virus 35S (CaMV-35S) promoter and the nopaline synthase gene terminator (NOS) by use of NcoI and SpeI, yielding At3g57630-F-H-pCambia 1302D.
C-terminal flag tagged exon-intron containing genomic At3g57630 (gAt3g57630-F) (Genbank accession: BT011693) was generated by PCR amplification of A. thaliana gDNA using the primers 5′-GGACTCTTGACCATGTTTTCTCACCAGAAATGGAAG-3′ and 5′-CTTCTCCTTTACTAGTCATTTATCGTCATCATCTTTGTAGTCGCTGGTTTTATGGAGACATTCTTG-3′ and inserted into pCambia1302D under control of the CaMV35 promotor and NOS terminator, using SpeI and NcoI and the In-Fusion cloning kit (Clontech), yielding At3g57630-F-pCambia1302D.
gAt3g57630-RFP-pCambia1302D was generated by PCR amplification of gAt3g57630-F and RFP55 followed by insertion into pCambia1302D using NcoI and BstEII and the In-Fusion cloning kit (Clontech). Primers for generation of gAt3g57630-RFP: 5′-GACTCTTGACCATGTTTTCTCACCAGAAATGGAAG-3′ and 5′-GCTGGTTTTATGGAGACATTCTTG-3′, using gAt3g57630-F-pCambia1302D as template, and 5′-CTCCATAAAACCAGCCCTAGGATGGCCTCCTCCGAGGAC-3′ and 5′-ATTCGAGCTGGTCACTTATCTAGAGGCGCCGGTGGAGTG-3′ using RFP55 as template.
gAt3g57630-mTurquoise-pCambia1302D, gAt3g57630-YFP-pCambia1302D and gAt3g57630-CFP-pCambia1302D were made by PCR amplification of mTurquoise, CFP and YFP followed by replacement of RFP in gAt3g57630-RFP-pCambia1302D using AvrII and XbaI. Primers for generation of mTurquoise: 5′-TAAAACCAGCCCTAGGATGGTGTCGAAGGGCGAG-3′ and 5′-TGGTCACttATCTAGTTACTTGTACAGTTCGTCCATGC-3′ using mTurquoise2 (obtained from Genscript) as template. Primers for generation of YFP & CFP: 5′-TAAAACCAGCCCTAGGGTGAGCAAGGGC-3′ and 5′-TGGTCACttATCTAGTTAAGCGTAATCTGGAACATCGTATG-3′ using pEarlygate10157 & mCerulean358 as template, respectively.
PCR was performed in 25 μl reaction volumes using the ClonAMP HiFi Master Mix system (Clontech) with the cycle parameters: 3 min at 98 °C followed by 30 cycles of 30 sec at 98 °C, 30 sec at 58 °C and 30 sec at 72 °C followed by 5 min at 72 °C.
Construct designs are visualized in Supplementary Material and Methods Fig. S11.
Isolation of Extensin enriched cell wall fractions
Alcohol insoluble residues (AIR) were isolated as described59 with few modifications. Briefly, plant material was harvested and frozen in liquid nitrogen. The tissue was then homogenized using a Retschmill machine (model MM200, Retsch, Haan, Germany) at 25 Hz for 1 min. The ground tissue was then suspended in 70% ethanol, vortexed, and pelleted by centrifugation at 10,000 × g for 10 min. The ethanol was decanted, and this procedure was then repeated using chloroform: methanol (1:1, vol/vol), until all chlorophyll was removed. The pellet was then washed twice in acetone. The remaining pellet was dried under vacuum for 5 min and either processed directly or stored until further use. AIR was subjected to sequential enzymatic treatment using pectin methylesterase and endo-polygalacturonanase followed by endo-cellulase (EGII; Megazyme) as described20,23, resulting in the EXT enriched cell wall fraction.
Barium hydroxyide mediated peptide bond hydrolysis
EXT enriched cell wall fractions or pre and post assay arabinosylated Gf(Muc1-2.2TR)p were subjected to Ba(OH)2 mediated hydrolysis, by dissolving it in 100 μl 0.22 M Ba(OH)2, incubated at 108 °C overnight, spun 10 min at 12000 × g and the supernatant transferred to a new Eppendorf tube and carefully neutralized using H2SO4. The precipitate was pelleted by centrifugation 12000 × g, 10 min. The Supernatant was used as acceptor substrate and subjected to (LC)-ESI-MS analysis.
Expanding leaves of more (Acer pseudoplatanus) were processed similarly albeit in larger scale and with a step introduced for the isolation of loosely bound cell wall proteins presumed to include AGPs. In brief: 10 g of AIR prepared as above was stirred for 1 h in 100 mL 0.2 M CaCl2. The solution was dialysed against water, freezedried and then hydrolysed in saturated Ba(OH)2 at 108 °C overnight. The solids were washed, resuspended in 150 mL 40 mM Na-oxalate pH = 4.8 and treated with endo-PG1 as described60 and destarched following the protocol of the same paper. The washed and freezedried residue was subjected to Ba(OH)2 hydrolysis as above.
Direct inlet and Liquid Chromatography (LC) Electrospray ionization mass spectrometry (ESI-MS)
LC-ESI-MS (3 μl supernatant injected) was carried out using an Agilent 1100 Series LC (www.agilent.com) coupled to a Bruker HCT Ultra ion trap mass spectrometer (www.bruker.com). The column was a Phenomenex Luna C8(2) (3 microM, 100A, 150 × 2.0 mm; www.phenomenex.com) preceded by a Phenomenex Gemini C18 SecurityGuard (4 × 2 mm). The oven temperature was maintained at 35 °C. The mobile phases were A, water; B, acetonitrile, both with 0.1% (v/v) HCOOH, and the flow rate was 0.2 mL min−1. The gradient was: 0 to 2 min, isocratic 1% B; 2 to 8.5 min, linear gradient 1 to 3% B; 8.6 to 9.6 isocratic 99% B; 9.7 to 17 min, isocratic 1% B. The mass spectrometer was run in positive electrospray mode.
Selected ion traces were semi-quantified in accordance the instructions of and using the software Bruker HCT Compass Data analysis 4.0 software.
Additional Information
How to cite this article: Moeller, S. R. et al. Identification and evolution of a plant cell wall specific glycoprotein glycosyl transferase, ExAD. Sci. Rep. 7, 45341; doi: 10.1038/srep45341 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Change history
23 May 2017
A correction has been published and is appended to both the HTML and PDF versions of this paper. The error has been fixed in the paper.
23 May 2017
Scientific Reports 7: Article number: 45341; published online: 30 March 2017; updated: 23 May 2017. The original version of this Article contained an error in the spelling of the author Zhang Yang, which was incorrectly given as Yang Zhang. The Author Contributions Statement, S.R.M., X.Y., S.M.V., S.G.
References
Breton, C., Heissigerova, H., Jeanneau, C., Moravcova, J. & Imberty, A. Comparative aspects of glycosyltransferases. Biochem Soc Symp 23–32 (2002).
Coutinho, P. M., Stam, M., Blanc, E. & Henrissat, B. Why are there so many carbohydrate-active enzyme-related genes in plants? Trends in plant science 8, 563–565, doi: 10.1016/j.tplants.2003.10.002 (2003).
Ellis, M., Egelund, J., Schultz, C. J. & Bacic, A. Arabinogalactan-proteins: key regulators at the cell surface? Plant Physiol 153, 403–419, doi: 10.1104/pp.110.156000 (2010).
Lamport, D. T., Kieliszewski, M. J., Chen, Y. & Cannon, M. C. Role of the extensin superfamily in primary cell wall architecture. Plant Physiol 156, 11–19, doi: 10.1104/pp.110.169011 (2011).
Lamport, D. T. & Miller, D. H. Hydroxyproline arabinosides in the plant kingdom. Plant Physiol 48, 454–456 (1971).
Tan, L., Leykam, J. F. & Kieliszewski, M. J. Glycosylation Motifs That Direct Arabinogalactan Addition to Arabinogalactan-Proteins. Plant Physiology 132, 1362–1369, doi: 10.1104/pp.103.021766 (2003).
Xu, J., Tan, L., Lamport, D. T., Showalter, A. M. & Kieliszewski, M. J. The O-Hyp glycosylation code in tobacco and Arabidopsis and a proposed role of Hyp-glycans in secretion. Phytochemistry 69, 1631–1640, doi: 10.1016/j.phytochem.2008.02.006 (2008).
Chakraborty, S. et al. YeATS - a tool suite for analyzing RNA-seq derived transcriptome identifies a highly transcribed putative extensin in heartwood/sapwood transition zone in black walnut. F1000 Res 4, 155, doi: 10.12688/f1000research.6617.2 (2015).
Bollig, K. et al. Structural analysis of linear hydroxyproline-bound O-glycans of Chlamydomonas reinhardtii–conservation of the inner core in Chlamydomonas and land plants. Carbohydr Res 342, 2557–2566, doi: 10.1016/j.carres.2007.08.008 (2007).
Miller, D. H., Lamport, D. T. & Miller, M. Hydroxyproline heterooligosaccharides in Chlamydomonas. Science 176, 918–920 (1972).
Akiyama, Y., Mori, M. & Katō, K. 13C-NMR Analysis of Hydroxy-proline Arabinosides from Nicotiana tabacum. Agricultural and Biological Chemistry 44, 2487–2489, doi: 10.1080/00021369.1980.10864347 (1980).
Campargue, C., Lafitte, C., Esquerre-Tugaye, M. T. & Mazau, D. Analysis of hydroxyproline and hydroxyproline-arabinosides of plant origin by high-performance anion-exchange chromatography-pulsed amperometric detection. Anal Biochem 257, 20–25, doi: 10.1006/abio.1997.2526 (1998).
Lamport, D. T. A., Katona, L. & Roerig, S. Galactosylserine in extensin. Biochemical Journal 133, 125–132, doi: 10.1042/bj1330125 (1973).
Matsubayashi, Y. Post-translational modifications in secreted peptide hormones in plants. Plant Cell Physiol 52, 5–13, doi: 10.1093/pcp/pcq169 (2011).
Kohorn, B. D. & Kohorn, S. L. The cell wall-associated kinases, WAKs, as pectin receptors. Front Plant Sci 3, 88, doi: 10.3389/fpls.2012.00088 (2012).
Nakhamchik, A. et al. A comprehensive expression analysis of the Arabidopsis proline-rich extensin-like receptor kinase gene family using bioinformatic and experimental approaches. Plant Cell Physiol 45, 1875–1881, doi: 10.1093/pcp/pch206 (2004).
Halim, A. et al. Glycoproteomic Analysis of Seven Major Allergenic Proteins Reveals Novel Post-translational Modifications. Molecular & Cellular Proteomics 14, 191–204, doi: 10.1074/mcp.M114.042614 (2015).
Saito, F. et al. Identification of Novel Peptidyl Serine alpha-Galactosyltransferase Gene Family in Plants. The Journal of biological chemistry 289, 20405–20420, doi: 10.1074/jbc.M114.553933 (2014).
Showalter, A. M., Keppler, B., Lichtenberg, J., Gu, D. & Welch, L. R. A bioinformatics approach to the identification, classification, and analysis of hydroxyproline-rich glycoproteins. Plant Physiol 153, 485–513, doi: 10.1104/pp.110.156554 (2010).
Velasquez, S. M. et al. O-glycosylated cell wall proteins are essential in root hair growth. Science 332, 1401–1403, doi: 10.1126/science.1206657 (2011).
Velasquez, S. M. et al. Complex regulation of prolyl-4-hydroxylases impacts root hair expansion. Mol Plant 8, 734–746, doi: 10.1016/j.molp.2014.11.017 (2015).
Ogawa-Ohnishi, M., Matsushita, W. & Matsubayashi, Y. Identification of three hydroxyproline O-arabinosyltransferases in Arabidopsis thaliana. Nat Chem Biol 9, 726–730, doi: 10.1038/nchembio.1351 (2013).
Egelund, J. et al. Molecular characterization of two Arabidopsis thaliana glycosyltransferase mutants, rra1 and rra2, which have a reduced residual arabinose content in a polymer tightly associated with the cellulosic wall residue. Plant Mol Biol 64, 439–451, doi: 10.1007/s11103-007-9162-y (2007).
Gille, S., Hansel, U., Ziemann, M. & Pauly, M. Identification of plant cell wall mutants by means of a forward chemical genetic approach using hydrolases. Proc Natl Acad Sci USA 106, 14699–14704, doi: 10.1073/pnas.0905434106 (2009).
Yang, Z. et al. Toward Stable Genetic Engineering of Human O-Glycosylation in Plants. Plant Physiology 160, 450–463, doi: 10.1104/pp.112.198200 (2012).
Schoberer, J. et al. Arginine/lysine residues in the cytoplasmic tail promote ER export of plant glycosylation enzymes. Traffic 10, 101–115, doi: 10.1111/j.1600-0854.2008.00841.x (2009).
Strasser, R. et al. Molecular cloning and characterization of Arabidopsis thaliana Golgi α-mannosidase II, a key enzyme in the formation of complex N-glycans in plants. The Plant Journal 45, 789–803, doi: 10.1111/j.1365-313X.2005.02648.x (2006).
Boevink, P. et al. Stacks on tracks: the plant Golgi apparatus traffics on an actin/ER network. Plant J 15, 441–447 (1998).
Ma, B., Simala-Grant, J. L. & Taylor, D. E. Fucosylation in prokaryotes and eukaryotes. Glycobiology 16, 158R–184R, doi: 10.1093/glycob/cwl040 (2006).
Hayashida, M., Fujii, T., Hamasu, M., Ishiguro, M. & Hata, Y. Similarity between protein-protein and protein-carbohydrate interactions, revealed by two crystal structures of lectins from the roots of pokeweed. J Mol Biol 334, 551–565 (2003).
Mihara, M., Itoh, T. & Izawa, T. SALAD database: a motif-based database of protein annotations for plant comparative genomics. Nucleic Acids Research 38, D835–D842, doi: 10.1093/nar/gkp831 (2010).
Hori, K. et al. Klebsormidium flaccidum genome reveals primary factors for plant terrestrial adaptation. Nat Commun 5, 3978, doi: 10.1038/ncomms4978 (2014).
Liu, X. et al. Bioinformatic Identification and Analysis of Extensins in the Plant Kingdom. PLoS One 11, e0150177, doi: 10.1371/journal.pone.0150177 (2016).
MacAlister, C. A., Ortiz-Ramirez, C., Becker, J. D., Feijo, J. A. & Lippman, Z. B. Hydroxyproline O-arabinosyltransferase mutants oppositely alter tip growth in Arabidopsis thaliana and Physcomitrella patens. Plant J 85, 193–208, doi: 10.1111/tpj.13079 (2016).
Ulvskov, P., Paiva, D. S., Domozych, D. & Harholt, J. Classification, naming and evolutionary history of glycosyltransferases from sequenced green and red algal genomes. PLoS One 8, e76511, doi: 10.1371/journal.pone.0076511 (2013).
Christiansen, C. et al. The carbohydrate-binding module family 20–diversity, structure, and function. FEBS J 276, 5006–5029, doi: 10.1111/j.1742-4658.2009.07221.x (2009).
Alderwick, L. J. et al. The C-terminal domain of the Arabinosyltransferase Mycobacterium tuberculosis EmbC is a lectin-like carbohydrate binding module. PLoS Pathog 7, e1001299, doi: 10.1371/journal.ppat.1001299 (2011).
Basu, D. et al. Functional identification of a hydroxyproline-o-galactosyltransferase specific for arabinogalactan protein biosynthesis in Arabidopsis. J Biol Chem 288, 10132–10143, doi: 10.1074/jbc.M112.432609 (2013).
Gazave, E. et al. Origin and evolution of the Notch signalling pathway: an overview from eukaryotic genomes. BMC Evol Biol 9, 249, doi: 10.1186/1471-2148-9-249 (2009).
Acar, M. et al. Rumi is a CAP10 domain glycosyltransferase that modifies Notch and is required for Notch signaling. Cell 132, 247–258, doi: 10.1016/j.cell.2007.12.016 (2008).
Sorensen, I. et al. The charophycean green algae provide insights into the early origins of plant cell walls. Plant J 68, 201–211, doi: 10.1111/j.1365-313X.2011.04686.x (2011).
Domozych, S., Wells, D. B. & Shaw, J. P. Basket Scales of the Green Alga, Mesostigma Viride: Chemistry and Ultrastructure. Journal of Cell Science 100, 397 (1991).
Harholt, J. et al. The glycosyltransferase repertoire of the spikemoss Selaginella moellendorffii and a comparative study of its cell wall. Plos One 7, e35846, doi: 10.1371/journal.pone.0035846 (2012).
Wickett, N. J. et al. Phylotranscriptomic analysis of the origin and early diversification of land plants. Proc Natl Acad Sci USA 111, E4859–4868, doi: 10.1073/pnas.1323926111 (2014).
Matasci, N. et al. Data access for the 1,000 Plants (1KP) project. Gigascience 3, 17, doi: 10.1186/2047-217X-3-17 (2014).
Xie, Y. et al. SOAPdenovo-Trans: de novo transcriptome assembly with short RNA-Seq reads. Bioinformatics 30, 1660–1666, doi: 10.1093/bioinformatics/btu077 (2014).
Johnson, M. T. et al. Evaluating methods for isolating total RNA and predicting the success of sequencing phylogenetically diverse plant transcriptomes. PLoS One 7, e50226, doi: 10.1371/journal.pone.0050226 (2012).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32, 1792–1797, doi: 10.1093/nar/gkh340 (2004).
Mitchell, A. et al. The InterPro protein families database: the classification resource after 15 years. Nucleic Acids Res 43, D213–221, doi: 10.1093/nar/gku1243 (2015).
Kall, L., Krogh, A. & Sonnhammer, E. L. A combined transmembrane topology and signal peptide prediction method. J Mol Biol 338, 1027–1036, doi: 10.1016/j.jmb.2004.03.016 (2004).
Brady, S. M. et al. A high-resolution root spatiotemporal map reveals dominant expression patterns. Science 318, 801–806, doi: 10.1126/science.1146265 (2007).
Ciucanu, I. & Kerek, F. A simple and rapid method for the permethylation of carbohydrates. Carbohydrate Research 131, 209–217, doi: http://dx.doi.org/10.1016/0008-6215(84)85242-8 (1984).
Ciucanu, I. Per-O-methylation reaction for structural analysis of carbohydrates by mass spectrometry. Anal Chim Acta 576, 147–155, doi: 10.1016/j.aca.2006.06.009 (2006).
Kivirikko, K. I. & Liesmaa, M. A Colorimetric Method for Determination of Hydroxyproline in Tissue Hydrolysates. Scand. J. Clin. Lab. Invest. 11, 128–133, doi: 10.3109/00365515909060420 (1959).
Schoberer, J. et al. Sequential depletion and acquisition of proteins during Golgi stack disassembly and reformation. Traffic 11, 1429–1444, doi: 10.1111/j.1600-0854.2010.01106.x (2010).
Knoch, E. et al. A beta-glucuronosyltransferase from Arabidopsis thaliana involved in biosynthesis of type II arabinogalactan has a role in cell elongation during seedling growth. Plant J 76, 1016–1029, doi: 10.1111/tpj.12353 (2013).
Earley, K. W. et al. Gateway-compatible vectors for plant functional genomics and proteomics. Plant J 45, 616–629, doi: 10.1111/j.1365-313X.2005.02617.x (2006).
Markwardt, M. L. et al. An improved cerulean fluorescent protein with enhanced brightness and reduced reversible photoswitching. Plos One 6, e17896, doi: 10.1371/journal.pone.0017896 (2011).
Abdulrazzak, N. et al. A coumaroyl-ester-3-hydroxylase insertion mutant reveals the existence of nonredundant meta-hydroxylation pathways and essential roles for phenolic precursors in cell expansion and plant growth. Plant Physiol 140, 30–48, doi: 10.1104/pp.105.069690 (2006).
Byg, I. et al. Large-scale extraction of rhamnogalacturonan I from industrial potato waste. Food Chemistry 131, 1207–1216, doi: http://dx.doi.org/10.1016/j.foodchem.2011.09.106 (2012).
Madson, M. et al. The MUR3 gene of Arabidopsis encodes a xyloglucan galactosyltransferase that is evolutionarily related to animal exostosins. Plant Cell 15, 1662–1670 (2003).
Harholt, J. et al. Arabinan Deficient 1 Is a Putative Arabinosyltransferase Involved in Biosynthesis of Pectic Arabinan in Arabidopsis. Plant Physiology 140, 49–58, doi: 10.1104/pp.105.072744 (2006).
Jensen, J. K. et al. Identification of a xylogalacturonan xylosyltransferase involved in pectin biosynthesis in Arabidopsis. Plant Cell 20, 1289–1302, doi: 10.1105/tpc.107.050906 (2008).
Brown, D. M., Zhang, Z., Stephens, E., Dupree, P. & Turner, S. R. Characterization of IRX10 and IRX10-like reveals an essential role in glucuronoxylan biosynthesis in Arabidopsis. Plant J 57, 732–746, doi: 10.1111/j.1365-313X.2008.03729.x (2009).
Zhong, R. et al. Arabidopsis fragile fiber8, which encodes a putative glucuronyltransferase, is essential for normal secondary wall synthesis. Plant Cell 17, 3390–3408, doi: 10.1105/tpc.105.035501 (2005).
Winter, D. et al. An “Electronic Fluorescent Pictograph” browser for exploring and analyzing large-scale biological data sets. PLoS One 2, e718, doi: 10.1371/journal.pone.0000718 (2007).
Lind, T., Tufaro, F., McCormick, C., Lindahl, U. & Lidholt, K. The putative tumor suppressors EXT1 and EXT2 are glycosyltransferases required for the biosynthesis of heparan sulfate. The Journal of biological chemistry 273, 26265–26268 (1998).
Acknowledgements
This work was supported by The Danish Councils for Strategic and Independent Research (12-125709, 12-131859), The Danish National Research Foundation (DNRF107), The Innovation Fund Denmark (contract 5152-0000), The Carlsberg Foundation DK, The Villum Foundation’s Young Investigator Program DK, The Copenhagen University Excellence Program for Interdisciplinary Research (CDO2016), The EU Marie Skłodowska-Curie program (ExHoMo, PIEF-GA-2011-301401), The Deutsche Forschungsgemeinschaft grant (EXC 1028), The UK BBSRC Institute Strategic Programme (MET, BB/J004561/1), The UK John Innes Foundation and The ANPCyT-Argentina (PICT2013-003, PICT2014-0504) and ICGEB (CRP/ARG16-03) programmes.
Author information
Authors and Affiliations
Contributions
S.R.M., X.Y., S.M.V., S.G., P.L.M.H., C.P.P., C.E.O., M.R., H.P., Z.Y., J.H., P.U., B.L.P. conducted the experiments. H.H.W., H.C., R.A.F. assisted with data analysis. M.P., J.M.E., J.H., P.U., B.L.P. drafted the manuscript. All authors reviewed the manuscript. Svenning Rune Møller (S.R.M.), Xueying Yi (X.Y.), Silvia Melina Velásquez (S.M.V.), Sascha Gille (S.G.), Pernille Louise Munke Hansen (P.L.M.H.), Christian P. Poulsen (C.P.P.), Carl Erik Olsen (C.E.O.), Martin Rejzek (M.R.), Harriet Parsons (H.P.), Zhang Yang (Z.Y.), Hans H. Wandall (H.H.W.), Henrik Clausen (H.C.), Robert A. Field (R.A.F.), Markus Pauly (M.P.), Jose M Estevez (J.M.E.), Jesper Harholt (J.H.), Peter Ulvskov (P.U.), Bent Larsen Petersen (B.L.P.).
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Møller, S., Yi, X., Velásquez, S. et al. Identification and evolution of a plant cell wall specific glycoprotein glycosyl transferase, ExAD. Sci Rep 7, 45341 (2017). https://doi.org/10.1038/srep45341
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep45341
This article is cited by
-
The hydroxyproline O-arabinosyltransferase FIN4 is required for tomato pollen intine development
Plant Reproduction (2023)
-
Functional characterization of chitin-binding lectin from Solanum integrifolium containing anti-fungal and insecticidal activities
BMC Plant Biology (2018)
-
Cell walls have a new family
Nature Plants (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.