Abstract
The core of the Vibrio Harveyi clade contains V. harveyi, V. campbellii, V. owensii, V. jasicida, and V. rotiferianus. They are well recognized aquatic animal pathogens, but misclassification has been common due to similarities in their rDNA sequences and phenotypes. To better understand their evolutionary relationships and functional features, we sequenced a shrimp pathogen strain V. harveyi 1114GL, reclassified it as V. campbellii and compared this and 47 other sequenced Vibrio genomes in the Harveryi clade. A phylogeny based on 1,775 genes revealed that both V. owensii and V. jasicida were closer to V. campbellii than to V. harveyi and that V. campbellii strains can be divided into two distinct groups. Species-specific genes such as intimin and iron acquisition genes were identified in V. campbellii. In particular, the 1114GL strain contains two bacterial immunoglobulin-like genes for cell adhesion with 22 Big_2 domains that have been extensively reshuffled and are by far the most expanded among all species surveyed in this study. The 1114GL strain differed from ATCC BAA-1116 by ~9% at the synonymous sites, indicating high diversity within V. campbellii. Our study revealed the characteristics of V. campbellii in the Harveyi clade and the genetic basis for their wide-spread pathogenicity.
Similar content being viewed by others
Introduction
The genus Vibrio is a bacterial group widely distributed in the marine environment. The core of the Vibrio Harveyi clade consist of V. harveyi and its closely related species V. campbellii, V. owensii, V. jasicida and V. rotiferianus1,2,3, all of which are well recognized aquatic animal pathogens4,5,6,7,8,9,10. Their members are commonly used as models to study bacterial luminescence11,12,13, quorum sensing14, biofilm formation, multi-chromosomal genome organization15,16, and recombination patterns2. Their genetic and phenotypic signatures are highly similar; conventional biochemical tests and 16 S rDNA sequencing frequently led to species misidentifications in this clade17,18. A well-known example is the misclassification of V. campbellii as V. harveyi – while the two species share only 61–74% DNA sequence similarity, the similarity in their 16 S rDNA sequences is over 97%1,17. Consequently, multi-locus sequence analysis (MLSA) was later used for species classification1,18,19,20. Given the sequence of two concatenated house-keeping gene sets (ftsZ, mreB and topA, or rpoD, rctB and toxR), the Harveyi clade ATCC BAA-1116 and HY01 were found to be V. campbellii rather than V. harveyi18.
The advent of next-generation sequencing has enabled multiple genome sequencing and assembly of Vibrio species. Such data have aided the study of phylogenomics2,3,21 and the identification of diagnostic features in Vibrio species22. A recent study of V. harveyi compared the genome of this species with those of three other related ones21. Nevertheless, a detailed genomic analysis of the frequently misidentified pathogen V. campbellii is still absent. High quality genomes of Vibrio species are difficult to obtain because of the high copy number rRNA operon present in the two chromosomes of Vibrio species (8–12). In the core Harveyi clade, to date, only the genomes of V. campbellii ATCC BAA-1116, V. harveyi ATCC 43516, and V. harveyi ATCC 33843 have been assembled into chromosomes. The other published genome assemblies remained fragmented, with 42 to 6,533 scaffolds (last assessed: 2015.09.17).
By applying genome-wide data, microbial taxonomy can be established via (1) average nucleotide identity (ANI) among sequences between two genomes23, and (2) the phylogeny of concatenated single copy orthologues, such as a core genome tree2,21. ANI ≥ 95% against type strains has become a commonly accepted definition of species3,24. Additionally, the combination of species assignment based on genomic distance and the monophyletic groupings allow clear species delineation2,3,25. Once the correct taxonomy is produced, shared genomic features and phenotypes of strains in a monophyletic group of interest can be established.
We had three goals in the current study. First, we sequenced the genome of V. harveyi 1114GL, which is a virulent strain isolated from a black tiger shrimp-culturing pond in Thailand in 200526,27. Second, we compared this new genome with 47 other published genome assemblies to identify species-specific differences in gene content and protein domains. Lastly, we established a reliable Harveyi clade phylogeny to reduce potential misclassification of species or strains in this clade.
Results and Discussion
Genome sequencing of V. harveyi 1114GL and re-classification of it as V. campbellii
V. harveyi 1114GL is the main cause of shrimp deaths in Thailand26,27,28, and thus, it attracted our interest to sequence its genome. The genome was sequenced using both Illumina and Roche 454 technologies (see Methods) and assembled into two scaffolds corresponding to chromosomes I (3,515,540 bp with 6 gaps) and II (2,118,471 bp with no gaps). The ANI between 1114GL and the type strain V. campbellii NBRC 15631 was 96.3%. Based on the threshold of 95–96% ANI for the same species, 1114GL was reclassified as V. campbellii. This is the most well assembled V. campbellii genome assembly after the genome assembly of ATCC BAA-1116. Among the 4,991 proteins predicted by PROKKA29, only 12 (0.24%) did not have significant matches in the nr database. There was no plasmid sequence found in the 1114GL assembly, an observation consistent with the negative result of plasmid extraction from 1114GL in a previous study27.
Phylogeny and genome content for a highly diverse set of Vibrio strains
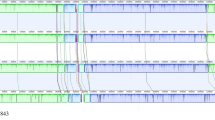
To infer an accurate phylogeny of available strains in the core Harveyi clade, we initially collected all 59 available genomes for the species in the core Harveyi clade and 6 additional genomes from V. parahaemolyticus as the outgroup (see Methods; Supplementary Table S1; Supplementary Table S2). However, only 362 single copy orthologous genes across all the species were identified, which is much lower than the 897 across 35 Vibrio strains of the Harveyi clade observed by Urbanczyk et al.2. This might be due to the lower annotation quality of the fragmented assemblies. Indeed, the number of single copy orthologous genes was found to decrease sharply when strains with poorer assemblies were included (Fig. 1). When assemblies with N90 < 10 k bp were filtered out, 1,775 single copy orthologous proteins (on average 34.8% of the proteome) were identified among 48 strains (12 from V. campbellii, 15 from V. harveyi, 7 from V. owensii, 5 from V. jasicida, 3 from V. rotiferianus, and 6 from V. parahaemolyticus) (see Supplementary Table S2). A concatenated alignment from these proteins was used to construct a maximum likelihood phylogeny tree with strong bootstrap clearly separating different species in the Harveyi clade and the V. parahaemolyticus outgroup (Fig. 2a).
The starting point includes 19 strains: V. campbellii 1114GL, ATCC BAA-1116, NBRC15631, UMTGB204, and HY01; V. owensii 47666-1, DY05, and 1DA3; V. jasicida BSW5 and BSW7; V. harveyi CAIM1792, VH5, and VH2; V. parahaemolyticus O3:K6 substr. RIMD 2210633, BB22OP, O1:Kuk str. FDA_R31, O1:K33 str. CDC_K4557, UCM-V493, and FORC_008. The N90 of these 19 genomes ranged from 13,457 to 2,195,939 bp. The number of single copy orthologous genes was found to decrease sharply when strains with N90 < 10 kb assemblies were included. To increase annotation quality, only the genomes with N90 > 10 kb were chosen for further analysis in this study. The final dataset consisted of 48 strains (12 from V. campbellii, 15 from V. harveyi, 7 from V. owensii, 5 from V. jasicida, 3 V. rotiferianus, and 6 from V. parahaemolyticus) and they shared 1,823 single copy orthologs.
(a) A total of 1,775 amino acid sequences from single copy orthologous genes were used to construct a maximum likelihood phylogeny tree, which separates the core clade into 5 groups. The strain names are indicated in each branch of the tree. The copy number of Big_2 domain, isolation source, N50 of the genome assembly (bar chart), and the copy numbers of rRNA (open circle) and tRNA (solid circle) are given on the right hand side of the figure. (b) The bootstrap values in the phylogenetic trees reconstructed using the proteins selected from the correlations between sequence similarities and branch length of the phylogenetic tree from (a). Node1 to Node5 of the trees correspond to the nodes of the phylogenetic tree in (a). The top ten proteins are: (1) hypothetical protein, (2) 1-deoxy-D-xylulose 5-phosphate reductoisomerase, (3) glutamate-cysteine ligase, (4) lipoprotein, (5) N-acetylglucosamine-6-phosphate deacetylase, (6) D-cysteine desulfhydrase, (7) NAD dependent epimerase/dehydratase family protein, (8) Peptidoglycan hydrolase FlgJ, (9) muropeptide transporter, and (10) Flagellar hook-associated protein 3. (c) The bootstrap values from the phylogentic trees reconstructed by concatenating phosphoethanolamine transferase (EptA) and the top one to top ten genes that were added serially. The tree reconstructed by concatenating EptA and the top four proteins correctly classified both the species and the two V. campbellii groups with strong bootstrap support.
The phylogenetic tree revealed inconsistencies in the species classification. Four strains in the phylogeny (1114GL, KC13.17.5, BSW7 and BSW5) were previously identified as V. harveyi30,31. Here we reclassified 1114GL and KC13.17.5 as V. campbellii strains, and BSW7 and BSW5 as V. jasicida strains. Their ANI values showed congruent results with this designation (see Supplementary Table S3): ANI was 96.3% between KC13.17.5 and the type strain V. campbellii NBRC 15631, 97.9% between BSW5 and its type strain V. jasicida LMG25398, and 97.9% between BSW7 and the type strain V. jasicida LMG25398. An interesting observation is that in the phylogeny V. owensii and V. jasicida were closer to V. campbellii than to V. harveyi, different from that of a previous study using only 138 genes3.
V. campbellii strains can be classified into two major groups, denoted as Vc-Gr1 and Vc-Gr2 (Fig. 2a). The separation of the two groups in V. campbellii had also been suggested by two previous studies, using concatenated sequences of 8972 or 1,61521 protein-coding genes.
We sought to identify a minimum number of key genes to reproduce the same classification based on calculations of correlations between phylogenetic topology from amino acid sequence and DNA sequence similarities of orthologous genes (see Methods). Briefly, we ranked the single copy orthologues by correlations and reconstructed the phylogeny (see Supplementary Table S4). The first gene (Pearson’s r = 0.99) was able to separate V. rotiferianus from the other species of the core Harveyi clade (Fig. 2b). Adding the second gene resulted in the correct classification of all the species. The first and second genes encode a hypothetical protein and 1-deoxy-D-xylulose 5-phosphate reductoisomerase, respectively. Adding more genes did not separate the two V. campbellii groups; hence we produced an alternative list which was ranked according to the topology of the V. campbellii subtree. Using the top gene from this new list (Pearson’s correlation coefficient = 0.960), the gene for a phosphoethanolamine transferase (EptA), and two more genes (for glutamate cysteine ligase and lipoprotein), which are from the old list, correctly classified both the species and the two V. campbellii groups with strong bootstrap support (Fig. 2b). The EptA is involved in the addition of phosphoethanolamine to lipid A and is required for polymyxin resistance32. It will be interesting to study the protein diversity in these species and the two groups of V. campbellii.
V. campbellii-specific features common to all strains
We sought to identify genes specific to V. campbellii. As some of the published genomes used in this study are fragmented, we focused on protein domains, which are modules of protein structure33, so that they are robust against potential misannotations such as gene fusion. Among the 313 enriched Pfam domains in V. campbellii by Wilcoxon rank-sum test (p ≤ 0.05) (see Supplementary Table S5a), 51 were present only in V. campbellii strains. Five of these domains in six proteins of ATCC BAA-1116 were found in all V. campbellii strains. The product descriptions and IDs of these genes in ATCC BAA-1116 and 1114GL are shown in Supplementary Table S5b. Three of these proteins are related to bacterial fitness: the rhizobactin siderophore biosynthesis protein with K_oxygenase domain which increases the ability of iron aquisition34, the putative neutral zinc metallopeptidase with Zn_peptidase domain known to increase fitness by cleaving proteins in Vibrio35, and the hypothetical protein with IAT_beta found in intimins and invasins, which are adhesin and virulence factors produced by gram-negative bacteria36. These V. campbellii -specific genes may be used as characteristics for distinguishing V. campbellii from the other members of the core Harveyi clade. The 313 expanded domains were enriched in GO terms related to bacterial flagellum formation including “bacterial-type flagellum-dependent cell motility”, “bacterial-type flagellum assembly”, and “bacterial-type flagellum organization”, as well as virulence-related GO terms such as “protein secretion by the type III secretion system”, “chitin catabolic process”, and “proteolysis”37,38 (see Supplementary Table S6).
However, V. campbellii also appears to have lost four domains that all other members of the core Harveyi clade and V. parahaemolyticus contained at least one copy of them. They are Glucodextran_C (C-terminal binding-module, SLH-like, of glucodextranase), Omp_AT (solitary outer membrane autotransporter beta-barrel domain), ydhR (putative mono-oxygenase ydhR), and YHS (found in copper transporting ATPases, some phenol hydroxylases and uncharacterized membrane proteins) (see Supplementary Table S5c).
Identification of determinants associated with V. campbellii Group 1 or 2
Our virulent isolate 1114GL was clustered in Group 1 (Vc-Gr1) with the known shrimp pathogen HY0139; half of the strains in this group were also associated with aquatic animals with strong bootstrap support (Fig. 2a). Group 2 (Vc-Gr2) included the published complete genome ATCC BAA-1116 and other strains isolated from seawater. There is no apparent ecological factor contributing to the separation of these two groups: only half of the Vc-Gr1 members were isolated from aquatic animals and all three Vc-Gr2 members were isolated from seawater. Next, we investigated the copy number differences in Pfam domains between these two groups (p ≤ 0.05; Wilcoxon rank-sum test). The phylogenetic position and the patterns of the expanded/lost domains of Vc-Gr1 were closer to the other four Vibrio species than to Vc-Gr2 (Fig. 2a, Supplementary Fig. S1, Table S5d, and Table S5e), suggesting that the domains enriched in Vc-Gr1 were likely present in their ancestors and were subsequently lost in Vc-Gr2.
A total of 386 Pfam domains were significantly expanded in Vc-Gr1 (see Supplementary Fig. S1a and Table S5d) and were enriched in the functions related to ‘antibiotic transport’ and ‘galactose metabolic process’ (Supplementary Table S7a). For example, there were on average 25.9 copies (20–29 copies) of the MatE domain (Multi antimicrobial extrusion protein) in Vc-Gr1, but only 18.7 copies in Vc-Gr2 (17–20 copies) (Fig. 3). In ATCC BAA-1116, there are ten proteins with two MatE domains (AGU93723.1, AGU93934.1, AGU94255.1, AGU95063.1, AGU95118.1, AGU95453.1, AGU95978.1, AGU97030.1, AGU97571.1, and AGU97766.1). In 1114GL, ten proteins (Vca1114GL_00245, Vca1114GL_00475, Vca1114GL_00731, Vca1114GL_01057, Vca1114GL_02103, Vca1114GL_02186, Vca1114GL_02598, Vca1114GL_03288, Vca1114GL_04301, and Vca1114GL_04662) were identified using OrthoMCL40 to be orthologous with the ten proteins in ATCC BAA-1116. In addition, there are three other proteins (Vca1114GL_03262, Vca1114GL_03492, and Vca1114GL_04269) each of which has two MatE domains. Because a protein with one or more MatE domains can function as a drug/sodium antiporter, variation in the copy number of these antiporter proteins among V. campbellii strains may cause variation in multidrug resistance strength. In addition, one adhesion-related autotransporter β-domain was present in every member of Vc-Gr1, but absent from Vc-Gr2 (Fig. 3, Supplementary Fig. S1a, and Table S5d). In contrast, 155 domains were significantly expanded in Vc-Gr2 (see Supplementary Fig. S1b and Table S5e) and were enriched in mobile genetic element terms (see Supplementary Table S7b). The galactose metabolic process was originally thought to be absent in V. campbellii and this phenotypic feature was one of those used in the traditional diagnosis to distinguish between V. campbellii and V. harveyi41,42. However, the genes responsible for D-galactose fermentation are all present in 1114GL, including 1114GL_02654 (Aldose 1-epimerase; EC 5.1.3.3), 1114GL_02653 (galactokinase; EC 2.7.1.6), 1114GL_02652 (UDP-glucose-hexose-1-phosphate uridylyltransferase; EC 2.7.7.12), 1114GL_02651 and 1114GL_04334 (both UTP-glucose 4-epimerase, EC 5.1.3.2), 1114GL_02322 (UTP-glucose-1-phosphate uridylyltransferase, EC 2.7.7.9), and 1114GL_01168 (phosphoglucomutase, EC 5.4.2.2). Additionally, a carbohydrate fermentation test of 1114GL showed acid production when using D-galactose as the sole carbon source (see Supplementary Fig. S2a), and 1114GL can grow better in basal medium (0.2% peptone and 0.1% yeast extract) with D-galactose than in basal medium without D-galactose (see Supplementary Fig. S2b). In V. campbellii strains, aldose 1-epimerase,galactokinase, and UDP-glucose-hexose-1-phosphate uridylyltransferase are absent only in NBRC15631 and DS40M4 (see Supplementary Fig. S3). This finding is consistent with a recent suggestion that the utilization of D-galactose is actually a variable feature among V. campbellii22 strains.
The number of squares displays the average number of domains in a genome from a cluster. The copy numbers for Vc-Gr1 include 1114GL. The numbers from each strain are shown in Supplementary Table S5.
In the Vc-Gr1 enriched domains, the Endotoxin_N domain was present in both KC13.17.5 and 151112 C, but absent in all other strains analyzed in this study. This led us to identify the two strains with a PirB, which is known as a cause for acute hepatopancreatic necrosis disease (AHPND) in shrimps by V. parahaemolyticus43. Compared with the PirB protein (accession number: AKC05670.1) from V. parahaemolyticus 3HP in which the virulence of this protein has been identified, the protein sequence identity of the PirB in KC13.17.5 (WP_025789543.1) is 100% and that in 151112 C (WP_045384430.1) is 70.3%. Although PirB was seen in KC13.17.5 in a previous study44, this strain was misidentified as V. harveyi. 151112 C containing PirB has never been studied with respect to whether it can cause AHPND or not. Here, we confirmed that the pirB gene is present only in Vc-Gr1.
Expanded domains, especially Big_2, found in V. campbellii 1114GL
To identify potential virulent factors of V. campbellii 1114GL to shrimps, expanded or reduced Pfam domains in 1114GL were found by comparing V. campbellii 1114GL to the 41 other strains (Wilcoxon rank-sum test; p < 0.05). We found 29 domain families significantly expanded in 1114GL (see Supplementary Table S5f), of which 15 displayed a 4-fold or higher average copy number. One interesting domain is Big_2 (22 versus 2.54 copies on average), which modulates bacterial cell-adhesion45. Another is the RebB domain (4 versus 0.15), which is responsible for the synthesis of R-body, a complex protein inclusion associated with toxic effects of Caedibacter cells on host paramecia46,47.
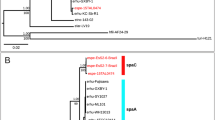
The Big_2 domain gene was found in tandem within bacterial immunoglobulin-like (Ig-like) genes, and was only observed in 6 Vc-Gr1 strains. Within the core of the Harveyi clade, Ig-like genes were found in ~47.6% of the strains, which on average had 4.6 copies of Big_2 domains. Strikingly, 1114GL contained two Ig-like genes with 9 and 13 Big_2 domains. In the light of the species phylogeny, Ig-like genes seemed to be ancestral to the core Harveyi clade and were subsequently lost, especially in the V. harveyi and V. rotiferianus groups (Fig. 2a). Interestingly, the phylogeny of V. campbellii Big_2 domains revealed strong clustering according to their physical location within Ig-like genes with the exception of those in 1114GL (Fig. 4). All of the 1114GL Big_2 domains were clustered in a clade, suggesting that strong intragenic or intergenic reshuffling among these domains has taken place. Little is known about the function of the Big_2 domain. The best studied case is in Escherichia coli: the Big_2 domain is found in the inimin protein, which is the ligand for Tir (Translocated intimin receptor)45. Currently, the function of Ig-like genes and the mechanism for the expansion of Big_2 domains in V. campbellii 1114GL remains elusive.
The Big_2 domains used were 47 Big_2 domains from 6 V. campbellii strains, 34 domains from 6 V. owensii strains, 44 domains from 7 V. jasicida strains, 1 domain from V. harveyi, and 21 domains from V. parahaemolyticus. The 9 Big_2 domains of gene 00953 and 13 of gene 00955 from 1114GL are highlighted by a1–a9 and b1–b13, respectively. The name of each unit is a domain from a gene (denoted as “gene_domain”). The genes are represented by the following numbers (in bold face): 1–7: V. campbellii 051011 F_04569, 051011G_03805, Vc1114GL_00953, 1114GL_00955, 151112C_02128, CCS02_04122, and HY01_EDL69217.1. 8: V. owensii 1DA3_EEZ87153.1. 9: V. harveyi AOD131_02851. 10–20: V. jasicida BSW5_KIP71511.1, BSW5_KIP75788.1, BSW7_ KIP71411.1, 090810c_00317, 090810c_00949, 200612G_01152, 200612G_02166, 201212A_03844, 201212A_05538, LMG25398_05354, and MWB21_01073. 21–28: V. owensii 47666-1_KIF47787.1, ATCC25919_00309, ATCC25919_01392, ATCC25919_03479, GRA50-12_04068, LMG25430_00078, LMG25430_00181, and OCN002_01444. Vp1–Vp7: V. parahaemolyticus BB22OP_AGB10101.1, CDCK4557_AGQ98222.1, FDAR31_AGQ91073.1, FDAR31_AGQ94143.1, FORC008_AKU54820.1, RIMD2210633_BAC60030.1, and UCMV493_AHI99806.1.
Mobile genetic elements and superintegrons are responsible for syntenic differences between V. campbellii 1114GL and ATCC BAA-1116
To reveal the syntenic differences between Vc-Gr1 and Vc-Gr2, we compared V. campbellii 1114GL, which belongs to Vc-Gr1, and ATCC BAA-1116, which belongs to Vc-Gr2. Most regions could be aligned between the two strains with an average nucleotide identity of 96% (see Supplementary Fig. S4). The relative numbers and positions of the nine rRNA operons were similar between the two strains, indicating that the assembly quality of 1114GL was comparable to the 454 and Sanger sequenced ATCC BAA-1116 (Accession numbers: NC_009777, NC_009783, and NC_009784). We further identified 95 (87.4% genome coverage) and 99 (85.4%) synteny blocks on ChrI and ChrII of 1114GL, and 96 (79.9%) and 98 (78.6%) blocks on ChrI and ChrII of ATCC BAA-1116 (see Supplementary Table S8). GO enrichment of genes in synteny break regions revealed abundant genetic mobile elements in both strains, especially in ATCC BAA-1116 (see Supplementary Table S9). The number of mobile elements correlated with the size of the break region between syntenies of ATCC BAA-1116 (see Supplementary Fig. S5), suggesting that transposition of these genes contributed to a lower synteny conservation in the genome.
It is known that mobile genetic elements can cause horizontal gene transfer and contribute to adaptation to varying environments and acquisition of virulent traits in Vibrio48. In an attempt to understand how these transposase genes evolved and led to the differences between ATCC BAA-1116 and 1114GL, a phylogenetic tree was constructed with the transposase sequences of GO:0006313 (transposition, DNA-mediated) from the ten biggest synteny breaks of ATCC BAA-1116 (see Supplementary Fig. S6). The phylogeny revealed two major groups with one having a much higher intra-group distance than the other (66.9% versus 1.3%). The distance between the two groups was 78.4%. We hypothesized that the longer syntenic break regions were a result of preferential transposition of mobile elements to already inserted regions49. Indeed, the phylogeny of these proteins showed no correlation in terms of amino acid distance between their relative orders inside syntenic break regions or their relative chromosome positions, suggesting that transpositions across the genome have been largely random but steadily accumulated in regions where syntenies were already broken up.
V. campbellii 1114GL harbors a 73 kb super integron (SI) on chromosome I (1114GL_00040 -1114GL_00134; see Supplementary Fig. S7a). This SI has several interesting features. First, its IntI4 is identical to the IntI4 from V. harveyi CAIM 1792 (accession number EMR38874) and has 65.3% sequence identity with the IntI4 from V. cholerae (accession number NP_232687.1). Second, we detected 39 conserved attC sites (see Supplementary Fig. S7b). These attC sites are 120 bp long and have 78.3–100% identity with the consensus sequence. Third, 76% of the genes are hypothetical proteins, similar to that in the SI of the other Vibrio genomes50. Although this SI has no syntenic relationship with V. campbellii ATCC BAA-1116, a remnant full-length IntI4 was identified on chromosome I.
Strong purifying selection on Vibrio campbellii genes
We next examined the degree of sequence divergence between V. campbellii strains and asked whether positive selection has occurred in any of the genes. dN (number of non-synonymous substitutions per site) and dS (the number of synonymous substitutions per site) were calculated for 4,239 one-to-one orthologous pairs in the syntenic regions between 1114GL and 051011F and for 3,672 pairs in the syntenic regions between 1114GL and ATCC BAA-1116 (Fig. 5). The mean dS value was 0.092 between 1114GL and 051011F and 0.130 between 1114GL and ATCC BAA-1116, while the mean dN values were 0.008 and 0.011, respectively. At a mean dS of 0.092, 051011F was the closest to 1114GL among the strains studied, indicating high genetic diversity of the V. campbellii species. The mean dN/dS value of gene pairs was 0.0981 between 1114GL and 051011F and 0.104 between 1114GL and ATCC BAA-1116 (Fig. 5). About 80% of the dN/dS values both between 1114GL and 051011F and between 1114GL and ATCC BAA-1116 were much smaller than 1, a result indicative of strong purifying selection on the majority of the genes in the genome. Moreover, there was no evidence of positive selection for any gene (Fig. 5). This predominant mode of purifying selection was also observed in the study of other prokaryotic genomes51.
Conclusions
In this study, we sequenced, assembled, and annotated the noncontiguous finished genome52 of a Vibrio isolate (V. campbellii 1114GL) that causes shrimp disease in Thailand. We compared this genome to 47 other sequenced Vibrio genomes in the core of the Harveyi clade. We showed that the species classification and phylogenetic relationships of the core species in the Harveyi clade could be correctly delineated based on a combination of average nucleotide identity and phylogenetic reconstruction of concatenated single copy orthologous genes. This enabled us to carry out an extensive comparative analysis of the Vibrio genomes to reveal how these organisms adapted to different environments. For example, the existence of iron acquisition genes in V. campbellii and the expansion of genes associated with transposition in Vc-Gr2 were discovered. This study demonstrated that a high quality genome assembly can provide more accurate gene annotation and taxonomic classification, and can enable detailed analyses including synteny analysis. We were thus able to describe the evolutionary dynamics at the genome level, such as strong purifying selection across the genome and numerous genomic rearrangements caused by transposition of mobile genetic elements. Areas worth further investigation have also been highlighted. First, the roles of Ig-like genes in different Vibrio species warrant functional studies. Second, it will be interesting to investigate whether the PirB protein in 151112 C can cause acute hepatopancreatic necrosis disease as in V. campbellii strain KC13.17.5. Finally, further comparative genomic analyses of the Harveyi clade members will provide deeper biological insights.
Materials and Methods
DNA preparation, sequencing and de novo assembly of the V. campbellii 1114GL genome
The V. campbellii 1114GL strain, provided by Timothy William Flegel, Centex Shrimp, Faculty of Science, Mahidol University, Bangkok, Thailand, was named VH1114GL in a previous study28. It was cultured from a glycerol stock in MHB + 3% ASW at 30 °C with shaking at 200 rpm for 16 hours. The bacterial cells were pelleted by centrifugation at 5,000 × g for 10 min. Genomic DNA was extracted using Qiagen Genomic-tip 100/G according to the manufacturer’s instructions. For genome sequencing, Illumina and Roche 454 platforms were used. Two paired-end libraries (insert size = ~320 bp) were constructed using the TruSeq DNA Preparation Kit with the standard protocol (Illumina) and sequenced by Illumina MiSeq to produce 250-bp paired end reads. Three mate-pair libraries of various jumping sizes (2 kb, 4 kb, and 6 kb) were constructed using the Nextera Mate Pair Sample Preparation Kit and sequenced by Illumina HiSeq2000 to produce 100-bp mate pair reads. The long single-end reads were produced by 454 GS FLX+.
Genomic data
Before the assembly of the 1114GL genome, adaptor and quality trimming (Q30 with minimum length = 70 bp) were conducted using Trimmomatic (version 0.32)53 leaving 4,968,772 paired-end sequences were retained. A total of 3,786,218 mate-pair sequence reads were retained after the detection and quality-trimming of TruSeq adaptor with at least 50 bp reserved in both reads using Nextclip54. The paired-end, mate-pair, and raw 454 reads (437,632 reads with mean length = 980 bp) of 1114GL were assembled de novo using the ALLPATH-LG assembler (ver. r48123)55. Subsequently, gap were closed using GapFiller (v1-10)56. Another 64 Vibrio genome assemblies were downloaded from NCBI (last retrieval date: 18th September 2015; see Supplementary Table S1).
For taxonomy confirmation, the Average Nucleotide Identity (ANI) for which pair-wise comparison of sequences between two strains or species was calculated using pyani (https://github.com/widdowquinn/pyani/tree/master/pyani) with BLAST method23.
Predicted proteins
The V. campbellii 1114GL assembly and the publicly available assemblies without annotation were annotated by the Prokaryotic Genome Annotation System (PROKKA) pipeline29. Functional annotation of the predicted proteins was obtained by Argot257. Protein domains of each gene were identified by pfam_scan.pl v1.5 by comparing against Pfam v27.058. The replication origin (oriC) regions of bacterial genomes were predicted by the online system Ori-Finder59 and DoriC60 tools.
Phylogenetic analysis
Single-copy orthologous genes in all 65 Vibrio strains of the core Harveyi members (21 from Vibrio campbellii, 24 from V. harveyi, 7 from V. owensii, 5 from V. jasicida, 3 V. rotiferianus, and 6 from V. parahaemolyticus) (Supplementary Table S1) were collected and identified based on the BLASTP results (E-value ≤ 10−5) using OrthoMCL40. To identify the relationship between the number of orthologous genes and assembly contiguity, the correlation between N90 of the genome assembly and number of orthologous genes were calculated (Fig. 1). N90 = 10 kb was chosen as the cutoff. The final dataset consisted of 48 strains (12 from V. campbellii (including 1114GL), 15 from V. harveyi, 7 from V. owensii, 5 from V. jasicida, 3 V. rotiferianus, and 6 from V. parahaemolyticus) (see Supplementary Fig. S1) with 1,729 single copy orthologs retained after removing alignments with more than 10% gaps of the 1,823 single copy orthologs. A phylogenetic tree was generated using RAxML (v8.1.17)61 with 500 bootstrap replicates from the alignment of these concatenated single copy orthologs by MAFFT (v7.123b, local option)62. The phylogeny was plotted using FigTree v1.4.2 (http://tree.bio.ed.ac.uk/software/figtree/).
For the phylogenetic tree of specific proteins such as Big_2 (Bacterial Ig-like, group 2) domains and transposase, the corresponding protein sequences were aligned by MAFFT (v7.123b, option localpair)57 and trimmed by TrimAl (with option strictplus)63. Maximum likelihood phylogenies were constructed by RAxML (v8.1.17)61 with 500 bootstraps.
Identification of genes for classification of the core Harveyi clade and V. campbellii groups
To identify the genes informative for taxonomic classification, we develop a method based on the calculation of correlations between the branch length of the Harveyi clade phylogenetic tree and sequence similarities of single copy orthologues. Our procedure includes three steps. First, for a given phylogenic tree with N nodes (species or strains), a distance vector (V) was constructed by  , where i and j were two species in the tree and d(i, j) was the summation of branch lengths between the two species. Second, for a set of orthologous genes (1-to-1 orthologous pairs among N species/strains), a vector of sequence dissimilarity (S) between the orthologous pairs was constructed by
, where i and j were two species in the tree and d(i, j) was the summation of branch lengths between the two species. Second, for a set of orthologous genes (1-to-1 orthologous pairs among N species/strains), a vector of sequence dissimilarity (S) between the orthologous pairs was constructed by  . For E(i, j), we took loge of the BLAST E-value between the orthologous pair in species i and j. If the E-value was zero, we set it to 0.1×(minimum non-zero E-value). The log-Evalue (e) was normalized by (e-Emin)/(Emax-Emin), where Emax and Emin were the maximum and minimum log-Evalue, respectively. For B(i, j), we normalized the bit score (bs) between the orthologous pair in species i and j by (bs-bs_max)/(bs_min-bs_max), where bs_max and bs_min were the maximum and minimum of the bit scores, respectively. We transformed the maximum bit score into zero to indicate the lowest sequence dissimilarity. Finally, we calculated Pearson Correlation Coefficient (PCC) between the distance vector (V) of the tree and the sequence vector (S) for each set of orthologous groups. The PCCs were then sorted in descending order. To obtain the granularity of taxon-specific genes in Vibrio species and V. campbellii strains, we used two phylogenies: (1) the tree of the Vibrio core Harveyi clade with V. parahaemolyticus as the outgroup and (2) its sub-tree in the V. campbellii species.
. For E(i, j), we took loge of the BLAST E-value between the orthologous pair in species i and j. If the E-value was zero, we set it to 0.1×(minimum non-zero E-value). The log-Evalue (e) was normalized by (e-Emin)/(Emax-Emin), where Emax and Emin were the maximum and minimum log-Evalue, respectively. For B(i, j), we normalized the bit score (bs) between the orthologous pair in species i and j by (bs-bs_max)/(bs_min-bs_max), where bs_max and bs_min were the maximum and minimum of the bit scores, respectively. We transformed the maximum bit score into zero to indicate the lowest sequence dissimilarity. Finally, we calculated Pearson Correlation Coefficient (PCC) between the distance vector (V) of the tree and the sequence vector (S) for each set of orthologous groups. The PCCs were then sorted in descending order. To obtain the granularity of taxon-specific genes in Vibrio species and V. campbellii strains, we used two phylogenies: (1) the tree of the Vibrio core Harveyi clade with V. parahaemolyticus as the outgroup and (2) its sub-tree in the V. campbellii species.
Genes identified by this method were aligned by MAFFT (v7.123b, option localpair) and the maximum likelihood phylogenetic tree was constructed by RAxML (v8.1.17) with 500 bootstraps.
Expansion and reduction of protein domain family sizes
Enrichment of Pfam domain number between two sets of interest was assessed by the Wilcoxon rank-sum test (p ≤ 0.05). We compared the Pfam copy number (1) between V. campbellii 1114GL and strains of all others species in the core Harveyi clade, (2) between groups 1 and 2 of V. campbellii strains, and (3) between V. campbellii and all other species in the core Harveyi clade. The expanded or reduced domains were assigned to the KEGG pathway by KAAS64. GO enrichments were identified for significant domain gains/losses using TopGO (version 2.10.0)65.
Synteny analysis
The Artemis Comparison Tool (ACT)66 was used to visualize whole-genome alignment between V. campbellii 1114GL and ATCC BAA-1116. Synteny block and orthologous gene pairs between 1114GL and ATCC BAA-1116 were defined using DAGCHAINER (-Z 12 -D 3 -g 1 -A 3)67, using the BLASTP output with an E-value < 1 × 10−10. The sequence regions not covered by synteny blocks were defined as synteny breaks. The genes located in synteny blocks or breaks were assessed by BEDTools68 and custom Python scripts.
Single orthologues between pairs of species in synteny blocks were obtained from DAGCHAINER62. Genes with one-to-many or many-to-many hits between two genomes were excluded in our sequence divergence analysis. There were 3,672 and 4,239 one-to-one orthologous pairs for 1114GL vs. ATCC BAA-1116 and for 1114GL vs. 051011 F, respectively. For each orthologous pair, dN (number of non-synonymous substitutions per site), dS (the number of synonymous substitutions per site), and dN/dS were computed using the Nei-Gojobori method69, and the likelihood values of dN/dS for gene pairs were calculated by PAML70. The likelihood ratio test was conducted to test if the dN/dS ratio was significantly different from 1.
Identification of integrons in V. cambellii 1114GL and ATCC BAA-1116
The complete proteome of V. campbellii 1114GL and ATCC BAA-1116 were searched against the IntI4 protein sequence of V. harveyi CAIM 1792 (Genbank accession EMR38874) and V. cholerae (Genabank accession NP_232687.1) using blastp. Conserved palindrome sequences were searched from 10 bp before the stop codon to 3′ intergenic sequences at 10 genes downstream of IntI4 using palindrome from the EMBOSS package71. A final conserved palindrome (TAACNNN[C/T]TGTTNAAG) was used to identify putative cassette-associated recombination (attC) site for all intergenic regions in the genomes of 1114GL and BAA-1116. Integron_finder72 was also used to check the accuracy of our approach. Multiple alignment of attC sites was manually checked using Jalview73.
D-galactose utilization test
1114GL was cultured from a glycerol stock in MHB + 3% ASW at 30 °C with shaking at 200 rpm for 16 hours. After the overnight culture, cells were inoculated, in three replicates, at a starting density of OD600 = 0.05 into flasks with 50 ml basal medium (0.2% of peptone and 0.1% of yeast extract broth) and into flasks with 50 ml basal medium plus 0.5% D-galactose. The cells were cultured at 30 °C with shaking at 200 rpm. Their growth was monitored by measuring the OD600 every hour.
For the D-galactose fermentation test, the overnight culture was inoculated into 4 ml basal media at a starting density of OD600 = 0.05 with and without 1.8 × 10−3% of phenol red. After cultures were grown at 30 °C for 10 hours, the culture color was assessed. A change in color from red to yellow indicates acid production.
Additional Information
How to cite this article: Ke, H. M. et al. Comparative genomics of Vibrio campbellii strains and core species of the Vibrio Harveyi clade. Sci. Rep. 7, 41394; doi: 10.1038/srep41394 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Sawabe, T., Kita-Tsukamoto, K. & Thompson, F. L. Inferring the evolutionary history of vibrios by means of multilocus sequence analysis. Journal of bacteriology 189, 7932–7936, doi: 10.1128/JB.00693-07 (2007).
Urbanczyk, H., Ogura, Y. & Hayashi, T. Contrasting Inter- and Intraspecies Recombination Patterns in the “Harveyi Clade” Vibrio Collected over Large Spatial and Temporal Scales. Genome biology and evolution 7, 71–80, doi: 10.1093/gbe/evu269 (2014).
Urbanczyk, H., Ogura, Y. & Hayashi, T. Taxonomic revision of Harveyi clade bacteria (family Vibrionaceae) based on analysis of whole genome sequences. Int J Syst Evol Microbiol 63, 2742–2751, doi: 10.1099/ijs.0.051110-0 (2013).
Cano-Gomez, A., Goulden, E. F., Owens, L. & Hoj, L. Vibrio owensii sp. nov., isolated from cultured crustaceans in Australia. FEMS microbiology letters 302, 175–181, doi: 10.1111/j.1574-6968.2009.01850.x (2010).
Ushijima, B., Smith, A., Aeby, G. S. & Callahan, S. M. Vibrio owensii induces the tissue loss disease Montipora white syndrome in the Hawaiian reef coral Montipora capitata. PloS one 7, e46717, doi: 10.1371/journal.pone.0046717 (2012).
Roy Chowdhury, P. et al. Genome sequence of Vibrio rotiferianus strain DAT722. Journal of bacteriology 193, 3381–3382, doi: 10.1128/JB.05089-11 (2011).
Yoshizawa, S. et al. Vibrio jasicida sp. nov., a member of the Harveyi clade, isolated from marine animals (packhorse lobster, abalone and Atlantic salmon). Int J Syst Evol Microbiol 62, 1864–1870, doi: 10.1099/ijs.0.025916-0 (2012).
Abraham, T. J., Palaniappan, R. & Dhevendaran, K. Simple taxonomic key for identifying marine luminous bacteria. Indian J Mar Sci 28, 35–38 (1999).
Karunasagar, I., Pai, R., Malathi, G. R. & Karunasagar, I. Mass Mortality of Penaeus-Monodon Larvae Due to Antibiotic-Resistant Vibrio-Harveyi Infection. Aquaculture 128, 203–209, doi: 10.1016/0044-8486(94)90309-3 (1994).
Diggles, B. K., Moss, G. A., Carson, J. & Anderson, C. D. Luminous vibriosis in rock lobster Jasus verreauxi (Decapoda: Palinuridae) phyllosoma larvae associated with infection by Vibrio harveyi. Diseases of aquatic organisms 43, 127–137, doi: 10.3354/dao043127 (2000).
Dunlap, P. V. In Encyclopedia of Microbiology (ed. M. Schaechter ) Ch. 45–61, (Elsevier, 2009).
Dunlap, P. Biochemistry and Genetics of Bacterial Bioluminescence. Adv Biochem Eng Biot 144, 37–64, doi: 10.1007/978-3-662-43385-0_2 (2014).
Dunlap, P. V. & Urbanczyk, H. in The Prokaryotes (eds E. Rosenberg et al.) Ch. 13, 495–528 (Springer, 2013).
Henke, J. M. & Bassler, B. L. Three parallel quorum-sensing systems regulate gene expression in Vibrio harveyi. Journal of bacteriology 186, 6902–6914, doi: 10.1128/JB.186.20.6902-6914.2004 (2004).
Kirkup, B. C. Jr., Chang, L., Chang, S., Gevers, D. & Polz, M. F. Vibrio chromosomes share common history. BMC microbiology 10, 137, doi: 10.1186/1471-2180-10-137 (2010).
Okada, K., Iida, T., Kita-Tsukamoto, K. & Honda, T. Vibrios commonly possess two chromosomes. Journal of bacteriology 187, 752–757, doi: 10.1128/JB.187.2.752-757.2005 (2005).
Gomez-Gil, B. et al. Molecular identification of Vibrio harveyi-related isolates associated with diseased aquatic organisms. Microbiology 150, 1769–1777, doi: 10.1099/mic.0.26797-0 (2004).
Lin, B. et al. Comparative genomic analyses identify the Vibrio harveyi genome sequenced strains BAA-1116 and HY01 as Vibrio campbellii. Environmental microbiology reports 2, 81–89, doi: 10.1111/j.1758-2229.2009.00100.x (2010).
Hoffmann, M., Monday, S. R., Fischer, M. & Brown, E. W. Genetic and phylogenetic evidence for misidentification of Vibrio species within the Harveyi clade. Letters in applied microbiology 54, 160–165, doi: 10.1111/j.1472-765X.2011.03183.x (2012).
Cano-Gomez, A., Hoj, L., Owens, L. & Andreakis, N. Multilocus sequence analysis provides basis for fast and reliable identification of Vibrio harveyi-related species and reveals previous misidentification of important marine pathogens. Systematic and applied microbiology 34, 561–565, doi: 10.1016/j.syapm.2011.09.001 (2011).
Espinoza-Valles, I. et al. Unique and conserved genome regions in Vibrio harveyi and related species in comparison with the shrimp pathogen Vibrio harveyi CAIM 1792. Microbiology, doi: 10.1099/mic.0.000141 (2015).
Amaral, G. R. S. et al. Genotype to phenotype: identification of diagnostic vibrio phenotypes using whole genome sequences. Int J Syst Evol Micr 64, 357–365, doi: 10.1099/Ijs.0.057927-0 (2014).
Goris, J. et al. DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Micr 57, 81–91, doi: 10.1099/ijs.0.64483-0 (2007).
Konstantinidis, K. T. & Tiedje, J. M. Genomic insights that advance the species definition for prokaryotes. Proceedings of the National Academy of Sciences of the United States of America 102, 2567–2572, doi: 10.1073/pnas.0409727102 (2005).
Chan, J. Z., Halachev, M. R., Loman, N. J., Constantinidou, C. & Pallen, M. J. Defining bacterial species in the genomic era: insights from the genus Acinetobacter. BMC microbiology 12, 302, doi: 10.1186/1471-2180-12-302 (2012).
Pasharawipas, T. et al. Partial characterization of a novel bacteriophage of Vibrio harveyi isolated from shrimp culture ponds in Thailand. Virus research 114, 63–69, doi: 10.1016/j.virusres.2005.05.012 (2005).
Khemayan, K. et al. Unstable lysogeny and pseudolysogeny in Vibrio harveyi siphovirus-like phage 1. Applied and environmental microbiology 72, 1355–1363, doi: 10.1128/AEM.72.2.1355-1363.2006 (2006).
Intaraprasong, A., Khemayan, K., Pasharawipas, T. & Flegel, T. W. Species-specific virulence of Vibrio harveyi for black tiger shrimp is associated with bacteriophage-mediated hemocyte agglutination. Aquaculture 296, 185–192, doi: 10.1016/j.aquaculture.2009.08.005 (2009).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069, doi: 10.1093/bioinformatics/btu153 (2014).
Kondo, H. et al. Draft Genome Sequences of Six Strains of Vibrio parahaemolyticus Isolated from Early Mortality Syndrome/Acute Hepatopancreatic Necrosis Disease Shrimp in Thailand. Genome announcements 2, doi: 10.1128/genomeA.00221-14 (2014).
Tatusova, T., Ciufo, S., Fedorov, B., O’Neill, K. & Tolstoy, I. RefSeq microbial genomes database: new representation and annotation strategy. Nucleic acids research 42, D553–559, doi: 10.1093/nar/gkt1274 (2014).
Needham, B. D. & Trent, M. S. Fortifying the barrier: the impact of lipid A remodelling on bacterial pathogenesis. Nature Reviews Microbiology 11, 467–481, doi: 10.1038/nrmicro3047 (2013).
Phillips, D. C. 3-Dimensional Structure of an Enzyme Molecule. Sci Am 215, 78-& (1966).
Krithika, R. et al. A genetic locus required for iron acquisition in Mycobacterium tuberculosis. Proceedings of the National Academy of Sciences of the United States of America 103, 2069–2074, doi: 10.1073/pnas.0507924103 (2006).
Miyoshi, S. & Shinoda, S. Microbial metalloproteases and pathogenesis. Microbes Infect 2, 91–98 (2000).
Leo, J. C., Oberhettinger, P., Schutz, M. & Linke, D. The inverse autotransporter family: Intimin, invasin and related proteins. International Journal of Medical Microbiology 305, 276–282, doi: 10.1016/j.ijmm.2014.12.011 (2015).
Aguirre-Guzman, G., Mejia Ruiz, H. & Ascencio, F. A review of extracellular virulence product of Vibrio species important in diseases of cultivated shrimp. Aquaculture Research 35, 1395–1404, doi: 10.1111/j.1365-2109.2004.01165.x (2004).
Henke, J. M. & Bassler, B. L. Quorum sensing regulates type III secretion in Vibrio harveyi and Vibrio parahaemolyticus. Journal of bacteriology 186, 3794–3805, doi: 10.1128/Jb.186.12.3794-3805.2004 (2004).
Rattanama, P. et al. Shrimp pathogenicity, hemolysis, and the presence of hemolysin and TTSS genes in Vibrio harveyi isolated from Thailand. Diseases of aquatic organisms 86, 113–122, doi: 10.3354/dao02119 (2009).
Li, L., Stoeckert, C. J., Jr. & Roos, D. S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome research 13, 2178–2189, doi: 10.1101/gr.1224503 (2003).
Alsina, M. & Blanch, A. R. A Set of Keys for Biochemical-Identification of Environmental Vibrio Species. J Appl Bacteriol 76, 79–85, doi: 10.1111/j.1365-2672.1994.tb04419.x (1994).
Farmer, J. J. I. & Hickman-Brenner, F. W. In The Prokaryotes: a handbook on the biology of bacteria: ecophysiology, isolation, identification, applications Vol. 3 (eds A. Balows et al.) 2952–3011 (Springer-Verlag, 1992).
Lee, C. T. et al. The opportunistic marine pathogen Vibrio parahaemolyticus becomes virulent by acquiring a plasmid that expresses a deadly toxin. Proceedings of the National Academy of Sciences of the United States of America 112, 10798–10803, doi: 10.1073/pnas.1503129112 (2015).
Kondo, H., Van, P. T., Dang, L. T. & Hirono, I. Draft Genome Sequence of Non-Vibrio parahaemolyticus Acute Hepatopancreatic Necrosis Disease Strain KC13.17.5, Isolated from Diseased Shrimp in Vietnam. Genome announcements 3, doi: 10.1128/genomeA.00978-15 (2015).
Kelly, G. et al. Structure of the cell-adhesion fragment of intimin from enteropathogenic Escherichia coli . Nature structural biology 6, 313–318, doi: 10.1038/7545 (1999).
Heruth, D. P., Pond, F. R., Dilts, J. A. & Quackenbush, R. L. Characterization of Genetic-Determinants for R-Body Synthesis and Assembly in Caedibacter-Taeniospiralis-47 and Caedibacter-Taeniospiralis-116. Journal of bacteriology 176, 3559–3567 (1994).
Jeblick, J. & Kusch, J. Sequence, transcription activity, and evolutionary origin of the R-body coding plasmid pKAP298 from the intracellular parasitic bacterium Caedibacter taeniospiralis. J Mol Evol 60, 164–173, doi: 10.1007/s00239-004-0002-2 (2005).
Hazen, T. H., Pan, L., Gu, J. D. & Sobecky, P. A. The contribution of mobile genetic elements to the evolution and ecology of Vibrios. FEMS microbiology ecology 74, 485–499, doi: 10.1111/j.1574-6941.2010.00937.x (2010).
Touchon, M. & Rocha, E. P. C. Causes of insertion sequences abundance in prokaryotic genomes. Molecular biology and evolution 24, 969–981, doi: 10.1093/molbev/msm014 (2007).
Boucher, Y. et al. Recovery and evolutionary analysis of complete integron gene cassette arrays from Vibrio. BMC evolutionary biology 6, 3, doi: 10.1186/1471-2148-6-3 (2006).
Novichkov, P. S., Wolf, Y. I., Dubchak, I. & Koonin, E. V. Trends in prokaryotic evolution revealed by comparison of closely related bacterial and archaeal genomes. Journal of bacteriology 191, 65–73, doi: 10.1128/JB.01237-08 (2009).
Chain, P. S. et al. Genome project standards in a new era of sequencing. Science 326, 236–237, doi: 10.1126/science.1180614 (2009).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120, doi: 10.1093/bioinformatics/btu170 (2014).
Leggett, R. M., Clavijo, B. J., Clissold, L., Clark, M. D. & Caccamo, M. NextClip: an analysis and read preparation tool for Nextera Long Mate Pair libraries. Bioinformatics 30, 566–568, doi: 10.1093/bioinformatics/btt702 (2014).
Ribeiro, F. J. et al. Finished bacterial genomes from shotgun sequence data. Genome research 22, 2270–2277, doi: 10.1101/gr.141515.112 (2012).
Nadalin, F., Vezzi, F. & Policriti, A. GapFiller: a de novo assembly approach to fill the gap within paired reads. BMC bioinformatics 13 Suppl 14, S8, doi: 10.1186/1471-2105-13-S14-S8 (2012).
Falda, M. et al. Argot2: a large scale function prediction tool relying on semantic similarity of weighted Gene Ontology terms. BMC bioinformatics 13 Suppl 4, S14, doi: 10.1186/1471-2105-13-S4-S14 (2012).
Punta, M. et al. The Pfam protein families database. Nucleic acids research 40, D290–301, doi: 10.1093/nar/gkr1065 (2012).
Gao, F. & Zhang, C. T. Ori-Finder: a web-based system for finding oriCs in unannotated bacterial genomes. BMC bioinformatics 9, 79, doi: 10.1186/1471-2105-9-79 (2008).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic acids research 25, 3389–3402 (1997).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313, doi: 10.1093/bioinformatics/btu033 (2014).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Molecular biology and evolution 30, 772–780, doi: 10.1093/molbev/mst010 (2013).
Capella-Gutierrez, S., Silla-Martinez, J. M. & Gabaldon, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973, doi: 10.1093/bioinformatics/btp348 (2009).
Moriya, Y., Itoh, M., Okuda, S., Yoshizawa, A. C. & Kanehisa, M. KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic acids research 35, W182–185, doi: 10.1093/nar/gkm321 (2007).
Alexa, A., Rahnenfuhrer, J. & Lengauer, T. Improved scoring of functional groups from gene expression data by decorrelating GO graph structure. Bioinformatics 22, 1600–1607, doi: 10.1093/bioinformatics/btl140 (2006).
Carver, T., Harris, S. R., Berriman, M., Parkhill, J. & McQuillan, J. A. Artemis: an integrated platform for visualization and analysis of high-throughput sequence-based experimental data. Bioinformatics 28, 464–469, doi: 10.1093/bioinformatics/btr703 (2012).
Haas, B. J., Delcher, A. L., Wortman, J. R. & Salzberg, S. L. DAGchainer: a tool for mining segmental genome duplications and synteny. Bioinformatics 20, 3643–3646, doi: 10.1093/bioinformatics/bth397 (2004).
Quinlan, A. R. BEDTools: The Swiss-Army Tool for Genome Feature Analysis. Curr Protoc Bioinformatics 47, 11 12 11–11 12 34, doi: 10.1002/0471250953.bi1112s47 (2014).
Nei, M. & Gojobori, T. Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Molecular biology and evolution 3, 418–426 (1986).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Molecular biology and evolution 24, 1586–1591, doi: 10.1093/molbev/msm088 (2007).
Rice, P., Longden, I. & Bleasby, A. EMBOSS: The European molecular biology open software suite. Trends Genet 16, 276–277, doi: 10.1016/S0168-9525(00)02024-2 (2000).
Cury, J., Jove, T., Touchon, M., Neron, B. & Rocha, E. P. C. Identification and analysis of integrons and cassette arrays in bacterial genomes. Nucleic acids research 44, 4539–4550, doi: 10.1093/nar/gkw319 (2016).
Waterhouse, A. M., Procter, J. B., Martin, D. M. A., Clamp, M. & Barton, G. J. Jalview Version 2-a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191, doi: 10.1093/bioinformatics/btp033 (2009).
Acknowledgements
The authors thank Timothy William Flegel for providing the 1114GL strain. This work was supported by Academia Sinica and Ministry of Science and Technology, Taiwan (MOST 104-2321-B-001-041). AP and SP acknowledged the support from Mahidol University and the National Center for Genetic Engineering and Biotechnology (BIOTEC) of the Thai National Science and Technology Development Agency (NSTDA). All the library construction and next-generation sequencing experiments were carried out by the High Throughput Genomics Core Facility of the Biodiversity Research Center in Academia Sinica, Taiwan, with help from YH Chen, KJ Yang, and MY Lee. Thanks also for the helpful comments by Drs Christine H-T Wang, John Wang and HsinHua C Cho.
Author information
Authors and Affiliations
Contributions
H.M.K., I.J.T., and W.-H.L. designed the research and wrote the paper. H.M.K., A.P, S.P., Y.-T. Y., K.-F.L., and C.-F.L. performed the experiment and confirmed the pathogenicity of 1114GL. M.-Y.J.L. designed the sequencing experiment. H.M.K., I.J.T., and C.-P.Y. analyzed the data. I.J.T., M.-C.L., and W.-H.L. supervised the experiments and manuscript preparation. All authors discussed the results and commented on the manuscripts.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ke, HM., Prachumwat, A., Yu, CP. et al. Comparative genomics of Vibrio campbellii strains and core species of the Vibrio Harveyi clade. Sci Rep 7, 41394 (2017). https://doi.org/10.1038/srep41394
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep41394
This article is cited by
-
Recovering high-quality bacterial genomes from cross-contaminated cultures: a case study of marine Vibrio campbellii
BMC Genomics (2024)
-
Genomic mining of Vibrio parahaemolyticus highlights prevalence of antimicrobial resistance genes and new genetic markers associated with AHPND and tdh + /trh + genotypes
BMC Genomics (2024)
-
Isolation and characterization of a Vibrio owensii phage phi50-12
Scientific Reports (2022)
-
Delineating virulence of Vibrio campbellii: a predominant luminescent bacterial pathogen in Indian shrimp hatcheries
Scientific Reports (2021)
-
A Novel Vibriophage vB_VcaS_HC Containing Lysogeny-Related Gene Has Strong Lytic Ability against Pathogenic Bacteria
Virologica Sinica (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.