Abstract
Typical 2-Cys Peroxiredoxins (2-Cys Prxs) reduce hydroperoxides with extraordinary rates due to an active site composed of a catalytic triad, containing a peroxidatic cysteine (CP), an Arg, and a Thr (or Ser). 2-Cys Prx are involved in processes such as cancer; neurodegeneration and host-pathogen interactions. During catalysis, 2-Cys Prxs switch between decamers and dimers. Analysis of 2-Cys Prx structures in the fully folded (but not locally unfolded) form revealed a highly conserved, non-conventional hydrogen bond (CH-π) between the catalytic triad Thr of a dimer with an aromatic residue of an adjacent dimer. In contrast, structures of 2-Cys Prxs with a Ser in place of the Thr do not display this CH-π bond. Chromatographic and structural data indicate that the Thr (but not Ser) destabilizes the decamer structure in the oxidized state probably through steric hindrance. As a general trend, mutations in a yeast 2-Cys Prx (Tsa1) favoring the dimeric state also displayed a decreased catalytic activity. Remarkably, yeast naturally contains Thr-Ser variants (Tsa1 and Tsa2, respectively) with distinct oligomeric stabilities in their disulfide states.
Similar content being viewed by others
Introduction
Typical 2-Cys peroxiredoxins (2-Cys Prx) from the Prx1/AhpC sub-group comprise a large class of thiol-specific antioxidant enzymes capable of reducing hydroperoxides with high efficiency (k = 106–108 M−1 s−1) and specificity1,2,3,4,5. The reactivity of 2-Cys Prx toward peroxides is considered extraordinary as they react one to ten million times faster than free Cys6. Due to their catalytic properties these enzymes are considered H2O2 sensors, which is probably related with the association of 2-Cys Prx with processes such as tumor suppression, neuronal differentiation and cardiovascular disease (reviewed in refs 7,8). 2-Cys Prxs make use of a fully conserved Cys residue, known as the peroxidatic cysteine (CP), for the reduction of hydroperoxides, which results in the formation of a sulfenic acid (CP-SOH) (Fig. 1a,b). The basic unit of 2-Cys Prxs is a homodimer (α2) and during the catalytic cycle a second Cys residue, the so-called resolving cysteine (CR) reacts with CP-SOH of the adjacent monomer (Fig. 1b,c) to form an intermolecular disulfide bond (reviewed in refs 8 and 9). A partial unfolding of the α-helix containing CP is required for the intermolecular disulfide formation and thioredoxin (Trx) is commonly the electron donor to reduce the disulfide7,8,9 (Fig. 1a–c). Therefore, during catalysis, 2-Cys Prx enzymes assume two structural states: fully folded (FF) and locally unfolded (LU). Alternatively, CP-SOH can further react with additional hydroperoxide molecules giving rise to hyperoxidized states such as sulfinic (CP-SO2H−) and sulfonic (CP-SO3H−) acids (Fig. 1b–d). These hyperoxidized forms cannot be reduced by Trx, however in eukaryotic organisms CP-SO2H can be reduced back to CP-SOH by an ATP dependent sulfiredoxin8.
Proposed mechanism for hydroperoxide reduction by 2-CysPrx (Ahpc-Prx1).
(a) CP in the fully folded (FF) form (S−p) displays high reactivity towards hydroperoxides and stabilizes the decamer state. This quaternary structure corresponds to a strong decamer. (b) The oxidation product (sulfenic acid - SpOH), also a strong decamer, and can have two outcomes: condensation or hyperoxidation. (c) The intermolecular disulfide is formed after a local unfolding process, because the two Cys residues are far away in the FF state, which destabilizes the decamer. In this work, we show that the extra Cγ in the side chain of catalytic Thr (Thr 44 in Tsa1) contributes to decamer destabilization. The reduced form of typical 2-Cys Prx, which favors the strong decamer, is regenerated by Trx. The disulfide form of 2-Cys Prx can switch from a weak metastable decamer (c) to dimers (e). Alternatively, the sulfenic acid of 2-Cys Prx (b) can be hyperoxidized (d) by further oxidation of CP to SpO2H by a second molecule of hydroperoxide. This process can be reversed by Srx, with ATP consumption.
In the FF state, CP is located in the first turn of an α-helix, surrounded by the Thr (or Ser, in some cases) and the Arg9,10. These three residues compose the catalytic triad, which has been implicated in the reactivity and specificity of Prx towards hydroperoxides8,9,10. Furthermore, transient interactions in the active site of Prxs are thought to play a crucial role in the stabilization of the transition state formed during the SN2 reaction11,12. There is evidence that the Arg residue is indispensable for Prx activity, since the substitution of the residue profoundly affects CP reactivity13,14,15,16,17. In contrast, studies involving the substitution of the conserved Thr (or Ser) residue are scarcer and show less drastic effects on catalysis13,17.
Another intriguing feature of 2-Cys Prx enzymes is their ability to switch among distinct quaternary structures, which is affected by redox state, among other factors8,9,18,19,20,21. These enzymes are α2 homodimers that can associate in (α2)5 decamers, i.e. pentamers of dimers. The factors governing the stabilities and the possible significance of these distinct quaternary structures are not completely understood, yet the formation of an intermolecular disulfide bond is known to destabilize the decamer (Fig. 1b–e). In the case of a 2-Cys Prx from Salmonella typhimurium (StAhpC), decamer stability was associated with an increase in the peroxidase activity4. Like the reduced enzymes, hyperoxidized 2-Cys Prxs appear preferentially as decamers (or even higher molecular weight complexes), whereas the presence of intermolecular disulfides destabilizes decamers into dimers4,20,21,22,23,24,25 (Fig. 1). Besides redox state, several post-translational modifications also affect the stabilities of 2-Cys Prx quaternary structures, which impact their activities26,27,28.
2-Cys Prxs play central roles in redox signaling but mechanisms governing their structural transitions are poorly understood. In this work, all analyzed 2-Cys Prx structures in the FF state (but not LU state) showed that the catalytic Thr (Thr44 in Tsa1, a 2-Cys Prx from S. cerevisiae) is located at the interface between two dimers, making a non-conventional hydrogen bond (H bond) with the π system of a Tyr/Phe residue (Tyr77 in Tsa1) of the adjacent dimer. In contrast, 2-Cys Prxs with Ser in place of catalytic Thr (Ser45 in Tsa2 from S. cerevisiae) always appear as decamers and do not present any CH-π bond. Chromatographic and structural data indicate that the Thr (but not Ser) destabilizes the decamer structure during disulfide formation (FF to LU transition) probably through steric hindrance, which affect the ability of 2-Cys Prx to reduce hydroperoxides.
Results
Structural characterization of typical 2-Cys Prx enzymes
The reversible transition between the dimeric and decameric states is a remarkable feature of 2-Cys Prxs4,18,19,20,21,22,23,24,25,26,27,28,29,30. Since the interactions between dimers are relevant for decamer stabilization, we analyzed the structures of 2-Cys Prx enzymes available in Protein Data Bank. The analysis of the yeast Tsa1 crystallographic structure31 in the reduced form revealed that (α2)5 decamer is stabilized by several non-covalent interactions at the dimer-dimer interface (Fig. 2a–c). Among these interactions, the Cγ atom of Thr44 acts as H-bond donor to the π-system of Tyr77 in the adjacent dimer (Fig. 2d). This CH-π bond is a non-conventional H bond that plays important contributions in protein structure and function (reviewed in refs 32,33). In the Tsa1 decameric structure, all five dimer-dimer interfaces have two CH-π H bonds between Thr44 and Tyr77 residues (Fig. 2e), and involves residues that are part of the so-called region I (Thr44) and region II (Tyr77), which are complementary at the dimer-dimer interface21.
Structural features at the dimer-dimer interface.
(a–d) Representations of Tsa1C47S structure. (a) Tsa1 homodimer represented in cartoon with a chain colored in green and the other in red. (b) Tsa1 decamer as a pentamer of homodimers (pdb code = 3SBC). (c) Amino acid residues involved in the interactions between dimers are represented by spheres. (d) Thr44 Cγ perform a CH-π bond with the Tyr77 of the adjacent dimer, while the Oγ of Thr44 side chain is involved in polar interactions with CP Sγ, which, in turn, is stabilized by the guanidine group of the Arg123. For (c,d), the carbon atoms are represented by colors similar to the polypeptide chain to which they belong, whereas N = blue, O = red and S = orange. For (e,f), the crystallographic structures of human Prx4 in the FF state ((e) 3TKP) and in the LU state ((f) 3TJB). The residues of catalytic triad and the conserved Tyr/Phe are represented by spheres and atoms are colored as follows: C = yellow, gray or light blue, according to the cartoon representation of the chain; N = blue; O = red; S = gold. (g) Wire representations of Thr/Ser region of yeast Tsa1 and Tsa2 (pdb codes 3SBC and 5DVB, respectively). The normalized B-factor was obtained by dividing the B-factor value of each Cα residue by the B-factor average value of all the Cα residues (normalized B-factor = B-factor of the residue/average B-factor of all residues). The wire thickness and color (blue = low to red = high) denotes the higher B factors for Tsa2 as compared to Tsa1. (h) Graphical detail of B-factor variation by amino acid in the portion containing the Thr or Ser from Tsa1 (blue curves) or Tsa2 (red curves), respectively. In the crystallographic structure of Tsa1, all chains are in the FF state while in the Tsa2 chains a mixture of FF, LU and intermediate states is found. Therefore, Tsa2 chains in FF state (C, D and G) and the respective chains of the Tsa1 were compared.
Remarkably, the CH-π H bonds between the catalytic Thr and the Tyr of the adjacent dimer are fully conserved among all analyzed structures of 2-Cys Prx (Prx1/AhpC) in the FF state (See Supplementary Fig. 1 and Fig. 2d and Supplementary Table 1). In some cases, the Tyr is substituted by a Phe residue, which can also accept H bonds through electrons of its π-system (See Supplementary Fig. 1 and Supplementary Fig. 2a,c,d).
In the LU state, the orientation between Thr44 and Tyr77 residues changes, and consequently the CH-π H bond is disrupted (Fig. 2e,f; See Supplementary Fig. 2b). As the microenvironment around the active site Thr is highly crowded (See Supplementary Fig. 3a,b,d–g), structural rearrangements of the neighboring residues during FF → LU transition are likely required to allow Thr44 to swing (Fig. 2e,f), probably provoking disruption of other non-covalent interactions at the dimer-dimer interface.
In contrast, the microenvironment around the active site Ser (equivalent to Thr44 in Tsa1) in AhpC from Amphibacillus xylanus (See Supplementary Fig. 3c) and in Tsa2 from Saccharomyces cerevisiae (See Supplementary Fig. 3h–k) is less crowded. Ser45 in the decameric structure of Tsa2 (which shares 86% of identity with Tsa1) appears as distinct rotamers in distinct chains, indicating high flexibility of this residue34. Remarkably, the CH-π hydrogen bond was not observed between Ser45 and Tyr78 (Tyr77 in Tsa1) in any of the Ser45 rotamers in Tsa2. Additionally, some of the chains from the Tsa2 decamer are in the FF state, while others are in the LU state. In fact, analysis of B factors in the α-helix containing the CP of the decameric Tsa1 and Tsa2 structures revealed that the thermal variation of Ser45 in Tsa2 is remarkably higher than the corresponding value for Thr44 in Tsa1 (Fig. 2g,h).
Therefore, it is possible that the conserved Thr/Ser not only participates in catalysis10,12, but is also involved in modulating the oligomeric state of the enzyme. For instance, while Thr44 Cγ atom of Tsa1 interacts with the neighboring dimer, the Oγ atom has a polar interaction with the Sγ atom of CP, which is further stabilized by polar contacts with the guanidine group of Arg123 (Fig. 2d,e; See Supplementary Fig. 2d). Furthermore, the catalytic triad of 2-Cys Prx is located near the dimer-dimer interface delimited by Tyr77 (Fig. 2d). Taken together, Thr44 provides a physical interaction between the catalytic center and the dimer-dimer interface, and indicates that the presence of Thr or Ser in the catalytic triad can influence the equilibrium among 2-Cys Prx quaternary structures.
Structural characterization of Tsa1 Thr44 mutants
Based on the structural considerations described above, we hypothesized that substitution of Thr44 in Tsa1 by other residues would affect decamer stability. Since the theoretical molecular weight of Tsa1 homodimers (~48 kDa) in solution is easily distinguishable from Tsa1 decamers (~240 kDa), we employed size exclusion chromatography (SEC) to evaluate the effects of Thr44 substitutions on Tsa1 quaternary structure. Under reducing conditions, Tsa1WT, Tsa1T44A, and Tsa1T44S eluted as decamers (Fig. 3a–c, solid lines), whereas Tsa1T44V eluted as a mixture of dimers and decamers, with the dimeric forms being clearly more predominant (Fig. 3d, solid line). As expected, the oxidized (disulfide) samples favored dissociation of decamers21 as compared with the reduced forms (Fig. 3, dashed lines). The exception was Tsa1T44S, which migrated exclusively as decamers in both the oxidized and reduced forms (Fig. 3c). In contrast, oxidized Tsa1T44V migrated mainly as dimer (Fig. 3d, dashed line). With intermediate behavior, comparable amounts of dimers and decamers of Tsa1WT and Tsa1T44A were detected in oxidizing conditions (Fig. 3a,b, dashed lines). In summary, two of the Thr mutations provoked dramatic effects on the quaternary structure, with Tsa1T44S favoring decamer stabilization and Tsa1T44V favoring dimer stabilization.
Size exclusion chromatography (SEC) profiles of Tsa1WT and mutants in reduced and oxidized states.
Samples were reduced with tris(2-carboxyethyl)phosphine (TCEP 5 mM; solid lines) or previously reduced with TCEP (5 mM) and then oxidized with H2O2 (1.2 molar equivalents; dashed lines) for 30 min at 25 °C. Tsa1WT (a), Tsa1T44A (b), Tsa1T44S (c) and Tsa1T44V (d). Enzyme (50 μM) was injected onto a HPLC system equipped with Phenomenex BioSep-SEC-S3000 column and resolved at a flow rate of 1 mL/min. The SEC column was calibrated with the following molecular weight standards: bovine thyroglobulin (670 kDa), bovine gamma globulin (158 kDa), ovalbumin (44 kDa), myoglobin (17 kDa), vitamin B12 (1.35 kDa) which are assigned at the top of the figure. The 100% value on the y-axis corresponds to the maximum absorbance detected in each chromatogram.
Kinetic characterization of Tsa1 Thr44 mutants
Next, we examined the effects of the mutations on Tsa1 reactivity towards hydroperoxides using the Trx system coupled assay. Tsa1WT and Tsa1T44S presented catalytic efficiencies towards H2O2 in the 104 M−1 s−1 range (See Supplementary Fig. 5a,c), similar to the previously obtained value35. On the other hand, Tsa1T44A presented a slight decrease in activity (See Supplementary Fig. 5b), whereas the catalytic efficiency of Tsa1T44V for H2O2 decreased by two orders of magnitude and the KM value (520 μM) increased by two orders of magnitude (See Supplementary Fig. 5d). All enzymatic parameters are summarized in Supplementary Table 2. The results for Tsa1T44V are consistent with a previous study, which showed that the catalytic efficiency of human PrxV, with a Thr for Val mutation, also decreased the catalytic efficiency by a factor of one thousand13. All kinetic parameters were determined under conditions where Trx (at 2 μM) was not limiting the catalysis (See Supplementary Fig. 5e).
In the case of cumene hydroperoxide (CHP) reduction, Tsa1WT and Tsa1T44S also presented similar enzymatic parameters (See Supplementary Fig. 5a–c, insets). In contrast to H2O2 reduction, Tsa1T44A presented a very high KM (300 μM) for CHP and consequently a four-fold decrease in the catalytic efficiency (See Supplementary Fig. 5b, inset). In the case of the Tsa1T44V mutant, an even more drastic drop in the reduction of CHP was observed (See Supplementary Fig. 5d, inset), precluding the determination of the kinetic parameters (Supplementary Table 2). Similarly, a 2-Cys Prx from L. donovani (TXNPx) also displayed negligible peroxidase activity towards organic peroxides when the homologous Thr residue was mutated to Val (TXNPxT49V)13. These results indicate that Val mutations are not well tolerated at this position, and that eliminating the hydroxyl group from the side chain impairs the catalytic activity of 2-Cys Prxs. The factors underlying why organic peroxide reduction is more inhibited than H2O2 reduction are not clear, and may be related to distinct effects of amino acid side chains on stabilization of the respective transition states with H2O2 and CHP; and/or with the release of the corresponding leaving groups.
We also evaluated the effects of the Thr mutations by non-reducing SDS-PAGE. Reduced proteins were oxidized with three equivalents of H2O2 or CHP and the disappearance of monomers was monitored. Among the three mutants, only Tsa1T44V presented a significant delay in disulfide formation for both H2O2 and CHP treatments (See Supplementary Fig. 6). For oxidations of 2-Cys Prx by H2O2, a similar trend in the reactivities as those observed in the Trx system coupled assay: Tsa1WT ~ Tsa1T44S > Tsa1T44A ≫ Tsa1T44V, taking into account that the SDS PAGE assay has a low resolution in time, being unable to discriminate moderate from very fast rates. For oxidation by CHP, similar tendency was also observed, but Tsa1T44V was oxidized in seconds (See Supplementary Fig. 6), whereas in the Trx system coupled assay, no NADPH consumption was detected (See Supplementary Fig. 5d). These apparent inconsistences are possibly related with differences in the principles underlying these two assays.
In the case of the Trx system coupled assay, a complete turnover of the enzyme is required for NADPH consumption. Mutations affecting any of the three reactions (hydroperoxide reduction; condensation; disulfide reduction) or structural transitions (described in Fig. 1) might impair or even block the turnover of the enzyme. In contrast, only the oxidation and condensation reactions are measured in the non-reducing SDS-PAGE assay.
Determination of apparent second order rate constants for the reaction between Tsa1 proteins and hydroperoxides
The interpretation of the results obtained by the Trx system - coupled assay is not always straightforward. Among other reasons this is complicated by the facts that the overall assay requires three different enzymes, involving various consecutive thiol-disulfide exchange reactions (among 2-Cys Prx, Trx and Trx reductase 1) and that catalytic cycle of 2-Cys Prx enzymes itself is complex, comprising three reactions (mentioned above) and conformational changes (LU-FF transitions; dimer–decamer switches)9. Therefore, direct rate constants between hydroperoxides and reduced Tsa1 proteins were determined by taking advantage of the changes in their intrinsic fluorescence upon oxidation (Fig. 4a). Under pseudo first-order conditions (excess peroxide), a two phase kinetic profile was observed for Tsa1WT (Fig. 4a,b). The first rapid phase is accompanied by a drop in fluorescence (first 5 ms), after an apparent 2 ms lag time, which coincides with the dead time of the instrument. The second phase is slower and is accompanied by a fluorescence increase.
Hydroperoxide reduction by Tsa1 monitored by changes in intrinsic fluorescence.
(a) Representative time course of Tsa1WT (1 μM) oxidation by H2O2 or CHP (20 μM). (b) Oxidation of Tsa1WT (1 μM) by increasing concentrations of H2O2 (4, 10 or 20 μM). Enzymes were previously reduced with 20 mM DTT for 30 minutes at room temperature, and excess DTT was removed by gel filtration. Total intrinsic fluorescence decays upon oxidation by hydroperoxides were followed (λex = 280 nm; λem > 310 nm using a cutoff filter) at 25 °C. The assays were performed in 40 mM phosphate buffer (pH 7.4). (c) The kobs values were obtained by fitting the stopped-flow data to single exponentials; second order rate constant was calculated from the slope of kobs plot versus oxidant concentration.
Recently, a similar kinetic profile was described for a bacterial 2-Cys Prx (StAhpC)36. In order to fit the experimental data, the authors proposed a three phase kinetic model. In the first step, H2O2 would reversibly bind to the StAhpC catalytic site, forming an enzyme - substrate complex, followed by CP peroxidation (which was associated to the fast fluorescence decay) and resolution (which was associated to second fluorescence increase phase). However, in the case of Tsa1 and under our experimental conditions the rise in fluorescence (second slower phase) could not be satisfactorily fitted and therefore was not further analyzed here. Therefore, Tsa1 and StAhpC display different fluorimetric properties, which might be due to distinct structural features of Trp residues (See Supplementary Fig. 7) or due to the fact that Tsa1 is sensitive to hyperoxidation, whereas StAhpC is resistant37.
Nevertheless, the rate of the initial rapid fluorescence decay increased linearly with peroxide concentrations (Fig. 4b,c), indicating that this phase is associated with CP oxidation. To test this assumption, we determined what would be the rate constant of CP oxidation under pseudo first-order conditions, fitting the fast decay in the fluorescence to a single first-order exponential equation or the whole biphasic time course to a two-exponential function (two consecutive first-order processes) for Tsa1WT. Using either approach, the rate constant for the Tsa1WT oxidation by H2O2 was close to 5 × 107 M−1 s−1 (Fig. 4c). Overall, this fluorimetric approach yielded second order rate constants for the Tsa1WT oxidation by H2O2 and CHP in the 106–107 M−1 s−1 range (Table 1), which agrees well with values previously determined by independent methods3. These convergences validated the fluorimetric method and made us confident to apply the same approach to the remaining Tsa1 mutants. Tsa1T44S mutant was also oxidized by H2O2 and CHP with extraordinary rates but the fluorimetric changes during the reaction was distinct from Tsa1WT, showing only a fast fluorescence decay associated with CP oxidation (See Supplementary Fig. 8). A decrease in reactivity was observed for both Tsa1T44A and Tsa1T44V enzymes (See Supplementary Fig. 8). Once again the Tsa1T44V mutant was far less reactive (Table 1), while the Tsa1T44A mutant was still quite active, one thousand fold faster than the free amino acid cysteine6, although at the same time, one thousand fold slower than the Tsa1WT. On the other hand, the corresponding rate constant for the oxidation of Tsa1T44V could not be determined due to its very low reactivity (Supplementary Fig. 8). The fluorimetric traces for Tsa1T44V correlated quite well with the non-reducing SDS-PAGE assay (See Supplementary Fig. 6), but not with the Trx system coupled assay possibly by reasons related to the distinct principles of these three assays, as discussed above.
It is worth mentioning that the values obtained for catalytic efficiencies in steady state kinetics (kcat/KM) were consistently in the range of 103–104 M−1 s−1, thus typically a thousand fold lower, than that obtained using fluorescence spectroscopy. This is probably due to other reactions and/or structural rearrangements limiting the overall turnover under steady-state conditions as mentioned above. It is worth mentioning that reduction of yeast TrxR1 by Trx1 and Trx2 are in order of 107 M−1s−1 38, reinforcing the hypothesis that structural rearrangements limit the reaction velocity. Nevertheless, the same trend was observed regardless of the approach (Trx coupled; non-reducing SDS-PAGE or fluorimetric assay) employed. In the case of Tsa1T44V, this mutation decreased the ability of Tsa1 to reduce hydroperoxides by several orders of magnitude, whereas the Tsa1T44A mutation induced a less pronounced effect. These results indicate the importance of the polar hydroxyl group in the catalytic mechanism.
Relationship between quaternary structures and catalytic activity of 2-Cys Prx
The results presented so far indicated that distinct residues present at position 44 of Tsa1 play critical roles in both decamer stabilization and peroxidase activity. Yeast Tsa2 (86% of amino acid identity with Tsa1) was also analyzed, since this enzyme naturally presents a Ser in the position equivalent to Thr44 in Tsa13. Similar to the Tsa1T44S mutant (Fig. 3c), Tsa2 migrated exclusively as decamers independent of the enzyme redox state (Fig. 5a), further supporting the hypothesis that Ser in place of Thr stabilizes the decameric state of 2-Cys Prxs. In addition, Tsa2 reduced H2O2 and CHP with extraordinary rates (Fig. 5b, Table 1; see Supplementary Fig. 9a,b).
Structural and kinetic characterization of Tsa2, Tsa1Y77A and Tsa1S78D.
SEC elution profiles of reduced (solid) and oxidized (dashed) Tsa2 (a), Tsa1Y77A (c) and Tsa1S78D (e) at 50 μM concentration, as described in Fig. 3. (b) Oxidation of Tsa2 (1 μM) by H2O2 or CHP (20 μM) performed in 40 mM phosphate buffer (pH 7.4). Tsa2 was previously reduced with 20 mM DTT for 30 minutes at room temperature, and the excess DTT was removed by gel filtration. (d,f) Trx system coupled assays: the reaction mixtures contained 2-Cys Prx (1 μM), Trx1 (2 μM), TrxR1 (0.3 μM), NADPH (150 μM), in HEPES-NaOH 50 mM (pH 7.4) and sodium azide (1 mM) and DTPA (0.1 mM). The reactions were performed at 30 °C. Tsa1Y77A (d) and Tsa1S78D (f) are represented by (▲), whereas negative control without Prx and positive control with Tsa1WT are represented by (●) and by (■), respectively.
As mentioned previously, the aromatic residue, Tyr77, also participates in the non-conventional CH-π H bond with Thr44. Therefore, the Tsa1Y77A mutant was also produced, overexpressed, and purified. This mutant protein always eluted as dimers (Fig. 5c) and the peroxidase activity of Tsa1Y77A was greatly reduced (Fig. 5d; see Supplementary Fig. 9c,e). Tyr77 performs multiple interactions besides the non-conventional CH-π hydrogen bond with Thr44 in the dimer-dimer interface (See Supplementary Fig. 3d–g) and its replacement by alanine may disrupt various forces that stabilize the decamer. Substitution of Tyr77 residue that plays no role on transition state stabilization11,12, but that dramatically affected the ability of Tsa1 to reduce peroxides suggested that the decameric quaternary structure is important to activity.
Furthermore, Tsa1S78D mutant was also generated based on the information that in StAhpC and in HsPrx4, similar substitutions provoked decamer destabilization4,25. Thr77 in StAhpC (Ser78 in Tsa1) is located in the decameric interface of the enzyme but does not have direct interactions with any of the active site residues. Substitution of Thr77 by Asp in StAhpC precluded decamer formation4. As predicted, Tsa1S78D also eluted as dimers, independent of the redox state (Fig. 5e) and the peroxidase activity was only residual in the Trx system coupled assay (Fig. 5f; see Supplementary Fig. 9d,f). Actually, the Tsa1S78D mutation resulted in a more pronounced loss of peroxidase activity than what was reported for StAhpCT77D 4.
As an independent and confirmatory assay, non-reducing SDS PAGE revealed that rates of disulfide formation in Tsa1Y77A and Tsa1S78D were extremely slow (See Supplementary Fig. 9g,h), precluding the determination of the rate constants values by available methods (Table 1). In contrast, Tsa2 displayed extraordinary second order rate constants for reduction of H2O2 and CHP (Table 1).
Since Thr44 provides a physical interaction between the catalytic center and the dimer-dimer interface it is intuitive a relationship between activity and oligomerization. However, the correlation between these two processes is not perfect, probably reflecting the fact that other factors interferes with the peroxidase activity and also with decamer stabilization. Nevertheless, if we analyze the more pronounced effects in the oligomeric state, such as Tsa2/Tsa1T44S (decamers with high activity) versus Tsa1T44V/Tsa1Y77A/Tsa1S78D (dimers with residual activity) the interplay between activity and oligomerization is clear (Fig. 5). These results further indicate a positive trend between decamer structure and peroxidase activity, although other factors such as pH and peroxiredoxin concentration also affect the enzymatic activity of 2-Cys Prx8,9,18,19,20,21,22,23,24,25,26,27,28,29,30.
Discussion
The equilibrium between dimers and decamers for 2-Cys Prx is dependent on CP redox state among other factors8,9,18,19,20,21,22,23,24,25,26,27,28,29,30, with oxidation to disulfides typically favoring dimers. Therefore, such a quaternary structure switch may be considered as part of the 2-Cys Prx catalytic cycle9 (Fig. 1). Here, we present evidences that the presence of a Thr or Ser in the catalytic triad (Thr44 in Tsa1; Ser45 in Tsa2) plays a critical structural role in the stabilization or destabilization of 2-Cys Prx decamers upon disulfide formation (Fig. 1; Supplementary Fig. 10). In proteins containing a Thr in the catalytic triad, oxidation leads to decamer dissociation into dimers9 (Fig. 1c–e), while with proteins containing a Ser, at the analogous position, such dissociation is not observed. This is an interesting structural finding by itself and may indicate different non-redundant or complementary enzyme functions.
Through a comparative structural analysis of 2-Cys Prx enzymes, it became clear that although most residues in the dimer-dimer interface retain similar conformation between the FF and LU states, the conserved Thr residue (Thr44 in Tsa1) swings considerably (Fig. 2e,f; See Supplementary Fig. 2a,b). This reorientation was observed in all of the analyzed decamer structures and is associated with the disruption of the Thr-Tyr (or Phe) CH-π bond in the LU state. Moreover, we observed that the decameric structure was prevalent (compare Ser44 versus Thr44 and Ala44 versus Val44 in Fig. 3) for protein containing residues with smaller side chains, consistent with a steric factor underlying the decamer stability. In fact, the microenvironment around the active site Thr is more crowded (See Supplementary Fig. 3) and significant structural rearrangements of neighboring residues are likely required to allow Thr44 to swing during the FF-LU transition. Possibly, the extra Cγ in the Thr side chain (in comparison with Ser side chain) may create steric effects with nearby residues, thus disfavoring the decameric state during the transition from the FF to LU states. In fact, a weakly associated decamer (in the LU state) has been proposed as an intermediate between the FF decamer and the LU dimer21 (Fig. 1c) and other intermediate oligomeric structures have also been observed14,19. On the other hand, it is possible that smaller side chains of residues at position 44 (for example, the active site Ser in AhpC from A. xylanus and Tsa2 from S. cerevisiae, Supplementary Fig. 3) might allow for smoother FF–LU transitions, by avoiding major structural perturbations at the dimer-dimer interface.
Other mutations in the dimer-dimer interface (Thr77 residue of StAhpC and Thr118 in Prx4) were reported to provoke major changes in the stability of decamers4,25. However, crystallographic structures of these proteins did not reveal major alterations in the dimer-dimer interface. In most of these structures, the 2-Cys Prx enzymes are in the LU state with an intermolecular disulfide bond and with the active site Thr (Thr43 in StAhpC) also swung in comparison to the FF state. Our structural analysis of StAhpC and Prx4 mutants indicated that subtle modifications in the dimer-dimer interface may induce major changes in the dimer-decamer transitions at lower protein concentrations.
The presence of a hydroxyl group in the side chain is also important for the stabilization of decamers (compare Ser44 versus Ala44 and Thr44 versus Val44 in Fig. 3). This is likely related to the fact that the O atom in the side chain of Thr residues performs distinct polar interactions in the FF and LU states, in both cases possibly these interactions help to stabilize 2-Cys Prx in the decameric form (See Supplementary Fig. 11). In contrast, the few available structures of 2-Cys Prx in the dimeric form show that the active site Thr of Prx1C83S from R. norvegicus possesses polar interactions distinct from the FF and LU decamers20. The hydroxyl group of the residue at position 44 is clearly important for the peroxidase activity (please compare the second order rate constants for Ser44 and Thr44 versus Ala44 and Val44, respectively in Table 1). Given the positive tendency between 2-Cys Prx decameric structure and activity, it is possible that Thr and certainly Ser residues stabilize the decamer, thereby allowing for optimal orientation of active site residues for peroxide reduction.
However, these associations between decamer and activity end after CP oxidation. For 2-Cys Prx containing Thr, this residue undergo a re-orientation during the FF-LU transition, which may disrupt the non-conventional CH-π hydrogen bond and perhaps various other neighboring interactions in the dimer-dimer interface, inducing decamer to dimer transition. In contrast, for 2-Cys Prx enzymes containing Ser, the FF-LU transition is smoother and the decameric oxidized structure is maintained throughout the catalytic cycle. It is tempting to speculate that the slow rise in fluorescence observed for the Tsa1WT is associated with this decamer dissociation (Fig. 4a,b), since it was not observed for the Tsa1T44S (See Supplementary Fig. 8) or for Tsa2 (Fig. 5b). If true, this would provide an opportunity to access the kinetics of decamer dissociation. However, at this moment, the mechanism behind of the slow rise in fluorescence during Tsa1 oxidation is still under investigation.
2-Cys Prx enzymes display complex structural and biochemical properties, involving distinct quaternary structures and oxidation states. Distinct post-translational modifications (hyperoxidation, glutathionylation and phosphorylation) in 2-Cys Prx also influence the quaternary structures they adopt and also their biological activities, indicating that mechanisms were positively selected for though evolution26,27,28. It is quite possible that these structural transitions among dimer, decamer and even higher molecular weight species relate to cellular functions, i.e. as peroxidase, chaperone, binding partner, enzyme activator and/or redox sensor (reviewed in ref. 18).
Our contribution here is to show that the active site Thr/Ser is also involved in modulating the stabilities of distinct 2-Cys Prx quaternary structures. The relationship between peroxidase activity and oligomerization is complex but disruption of dimer-dimer interactions in the reduced state is associated with a decreased catalytic efficiency with possible physiological significance as 2-Cys Prxs are considered the H2O2 sensors7,8,10.
Material and Methods
Site directed mutagenesis
The pET15b/Tsa1 plasmid was used as template to generate the individual Tsa1 mutants carrying Thr44 substitutions to Ala (Tsa1T44A), Ser (Tsa1T44S) and Val (Tsa1T44V); Tyr77 to Ala (Tsa1Y77A) and Ser78 to Asp (Tsa1S78D). The mutagenesis protocols were performed according manufacturer instructions, using Quick Change II Kit (Stratagene) and the following primers: Tsa1T44A_F (5′ TGGCCTTCGCTTTCGTCTGT 3′), Tsa1T44A_R (5′ ACAGACGAAAGCGAAGGCCA 3′); Tsa1T44S_F (5′ TTGGCCTTCAGTTTCGTCTGT 3′), Tsa1T44S_R (5′ ACAGACGAAACTGAAGGCCAA 3′); Tsa1T44V_F (5′ TTGGCCTTCGTTTTCGTCTG 3′), Tsa1T44V_R (5′ CAGACGAAAACGAAGGCCAA 3′); Tsa1S78D_F (5′ TCCGAATACGACCTTTTGGCA 3′), Tsa1S78D_R (5′ TGCCAAAAGGTCGTATTCGGA 3′), Tsa1Y77A_F (5′ GACTCCGAAGCCTCCCTTTTG 3′), Tsa1Y77A_R (5′ CAAAAGGGAGGCTTCGGAGTC 3′). The reaction products were treated with Dpn I to remove the parental methylated plasmids and E. coli XL1-Blue strain was used as host in the transformations. Single colonies were selected and their plasmids extracted and sequenced with BigDye Terminator v3.1 Cycle Sequencing Kit using the automatic sequencer ABI 3730 DNA Analyser (Thermo Scientific) to confirm codon substitutions. The plasmids harboring the mutations were transformed in E. coli BL21 (DE3) strain by electroporation.
Protein expression and purification
Single colonies of E. coli BL21 (DE3) strain containing the pET15b/tsa1, pET15b/tsa1T44A, pET15b/tsa1T44S, pET15b/tsa1T44V, pET15b/tsa1Y77A, pET15b/tsa1S78D or pPROEx/tsa2 plasmids were inoculated in LB medium (20 ml) containing 0.1 mg ampicillin/mL and grown overnight at 37 °C with shaking at 250 rpm. The following day the cultures were transferred to 1 L of fresh LB medium, and cultured further until OD600 reached 0.6–0.8. The overexpression of all the proteins was induced by the addition of 0.3 mM isopropyl-1-thio-β-D-galactopyranoside (IPTG) at 37 °C for 3hs with shaking at 250 rpm in an orbital shaker. Cells were harvested by centrifugation and cell pellets were resuspended in start buffer (50 mM sodium phosphate buffer, pH 7.4, 500 mM NaCl, 20 mM imidazole and 2 mM phenylmethyl sulfonyl fluoride - PMSF) and disrupted by sonication. The cell extracts were kept in ice during streptomycin sulfate (1%) treatment for 20 min. The supernatants were clarified by centrifugation, homogenized by filtration and purified by immobilized metal affinity chromatography (IMAC) using HisTrap column (GE Healthcare). Imidazole was removed by gel filtration using PD10 desalting column (GE Healthcare) and the purity of recombinant proteins was verified by SDS-PAGE.
Protein quantification
All of the purified enzymes were quantified using the molar extinction coefficient for reduced S. cerevisiae Tsa1WT obtained using the ProtParam tool (http://www.expasy.ch/tools/protparam.html) (ε280 = 23,950 M−1 cm−1), except for Tsa1Y77A for which molar extinction coefficient is different (ε280 = 22,460 M−1 cm−1).
Protein reduction
Wild type and mutant proteins were initially reduced with 5 mM TCEP in 50 mM sodium phosphate buffer (pH 7.4), 50 mM NaCl, 100 μM diethylenetriaminepentaacetic acid (DTPA) and 1 mM sodium azide, for 30 minutes, at room temperature. In some cases, TCEP was replaced by DTT (20 mM). After reduction, excess of TCEP or DTT was removed using a PD10 column (GE Healthcare).
Circular dichroism spectroscopy of Tsa1WT and mutants carrying Thr44 substitutions
The CD spectra of Tsa1WT and mutant proteins were obtained using a 0.1 cm path length cuvette containing 10 μM of protein sample in 10 mM Tris buffer (pH 7.4) and 100 mM NaF. The assays were carried out at 25 °C in a Jasco J-810 spectropolarimeter (Jasco Inc.). Spectra were presented as an average of eight scans recorded from 190 to 260 nm. The content of secondary structures in each protein was estimated using the CDNN 2.1 software39.
Size-exclusion chromatography
Size-exclusion chromatography was performed by analytical HPLC (Jasco LC-2000 Plus) equipped with a PU 2880 Plus injector and a PDA MD 2018 detector (Jasco). The samples (50 μM in 100 mM Tris-HCl at pH 7.4) were separated by a system containing a Phenomenex BioSep-SEC-S3000 column (7.8 × 300 mm, 5 μm, resolution range of 15 to 2000 kDa, Phenomenex, Inc., Torrance, California, USA) using a flow rate of 1.0 mL/min in 100 mM Tris-HCl buffer (pH 7.4) and 50 mM NaCl. The elution profile was monitored by absorbance (λ = 280 nm). Bovine thyroglobulin (670 kDa), bovine gamma globulin (158 kDa), ovalbumin (44 kDa), myoglobin (17 kDa) and vitamin B12 (1.35 kDa) were used as molecular weight standards (Bio-Rad). The chromatograms were analyzed using Jasco BORWIN, version 1.50, software (Jasco). The enzymes were reduced or oxidized by treatment with 5 mM TCEP or 1.2 molar excess of H2O2, respectively, for 30 min at 25 °C.
Determination of peroxidase activity using the Trx system coupled assay
Trx peroxidase activities were monitored by NADPH oxidation in a coupled assay35. S. cerevisiae Tsa1WT and mutants (1.0 μM) were incubated with Trx1 (2 μM), TrxR1 (0.3 μM) and NADPH (150 μM) in 50 mM HEPES buffer (pH 7.4) containing sodium azide (1 mM) and DTPA (100 μM), at 30 °C. Reactions were initiated by addition of H2O2 or cumene hydroperoxide (CHP). All kinetic data were analyzed by non-linear regression using Michaelis-Menten equation40 (GraphPad Prism 5 software, GraphPad Software, Inc., San Diego).
Kinetics of Tsa1 oxidation using the intrinsic fluorescence of the enzyme
Reduced Tsa1WT, Tsa1 mutants and Tsa2 enzymes (1 μM) in 40 mM sodium phosphate buffer (pH 7.4) were rapidly mixed with either H2O2 or CHP in excess in an Applied Photophysics SX-17MV stopped-flow spectrometer with a mixing time of approximately 2 ms. The reaction was followed by total fluorescence intensity decrease (λex = 280 nm; λem > 310 nm using a cutoff filter) of the enzymes upon oxidation. Observed rate constants (kobs) were determined by fitting the stopped-flow data to single exponential functions. The apparent second order rate constants were determined from the slope of kobs values plotted against hydroperoxide concentrations.
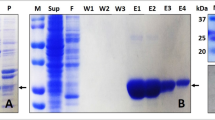
Non-reducing SDS PAGE assays
To study reduction of 2-Cys Prx by DTT and Trx1, wild-type Tsa1 and mutant proteins were previously reduced as described before and treated with H2O2 at 1.2 molar ratio in 50 mM sodium phosphate buffer (pH 7.4) containing 50 mM NaCl, 100 μM DTPA and 1 mM sodium azide. Next, the enzymes were reduced by increasing concentrations of DTT (10 μM to 10 mM) or Trx (2.5 to 25 μM) during 1 minute. The reaction was stopped with denaturing buffer (62.5 mM Tris-HCl, pH 6.8, containing 4% SDS and 10% glycerol) together with alkylation by 50 mM N-ethylmaleimide (NEM). Oxidized Tsa1WT (intermolecular disulfide) runs as a dimer in non-reducing 12% SDS-PAGE, whereas reduced Tsa1WT (dithiol) runs as a monomer in the same conditions. In order to evaluate the disulfide formation in the Tsa1WT and mutants, the proteins were initially reduced, as described above. The oxidation assays were performed using 10 μM of protein and 30 μM of H2O2, in 100 μL of final volume at room temperature, in 50 mM sodium phosphate buffer (pH 7.4) containing NaCl 50 mM, 100 μM DTPA and 1 mM sodium azide. The reactions were stopped with 10 μL of denaturing buffer and 50 mM NEM to alkylate the enzymes thiols, precluding formation unspecific disulfide bonds41 at the intervals of 0, 10, 30, 60, 90, 120, 300 and 600 seconds. Reactions were analyzed by non-reducing SDS-PAGE stained with Coomassie blue.
Analysis of Prx1/AhpC crystallographic structures
Analysis of available crystallographic structures and figures were performed using Pymol (http://www.pymol.org) and Discovery Studio 4.0. Distances and angles of non-conventional hydrogen bonds (CH-π interactions) in Prx structures were determined using Discovery Studio 4.0 software (http://accelrys.com/products/discovery-studio)32.
Additional Information
How to cite this article: Tairum, C. A. et al. Catalytic Thr or Ser Residue Modulates Structural Switches in 2-Cys Peroxiredoxin by Distinct Mechanisms. Sci. Rep. 6, 33133; doi: 10.1038/srep33133 (2016).
References
Cox, A. G., Peskin, A. V., Paton, L. N., Winterbourn, C. C. & Hampton, M. B. Redox potential and peroxide reactivity of human peroxiredoxin 3. Biochemistry 48, 6495–6501 (2009).
Horta, B. B., de Oliveira, M. A., Discola, K. F., Cussiol, J. R. R. & Netto, L. E. S. Structural and biochemical characterization of peroxiredoxin Qbeta from Xylella fastidiosa: catalytic mechanism and high reactivity. J. Biol. Chem. 285, 16051–16065 (2010).
Ogusucu, R., Rettori, D., Munhoz, D. C., Netto, L. E. S. & Augusto, O. Reactions of yeast thioredoxin peroxidases I and II with hydrogen peroxide and peroxynitrite: rate constants by competitive kinetics. Free Radic. Biol. Med. 42, 326–334 (2007).
Parsonage, D. et al. Analysis of the link between enzymatic activity and oligomeric state in AhpC, a bacterial peroxiredoxin. Biochemistry 44, 10583–10592 (2005).
Peskin, A. V. et al. The high reactivity of peroxiredoxin 2 with H2O2 is not reflected in its reaction with other oxidants and thiol reagents. J. Biol. Chem. 282, 11885–11892 (2007).
Winterbourn, C. C. & Metodiewa, D. Reactivity of biologically important thiol compounds with superoxide and hydrogen peroxide. Free Radic. Biol. Med. 27, 322–328 (1999).
Netto, L. E. S. & Antunes, F. The Roles of Peroxiredoxin and Thioredoxin in Hydrogen Peroxide Sensing and in Signal Transduction. Mol. Cells 39, 65–71 (2016).
Rhee, S. G., Woo, H. A., Kil, I. S. & Bae, S. H. Peroxiredoxin functions as a peroxidase and a regulator and sensor of local peroxides. J. Biol. Chem. 287, 4403–4410 (2012).
Hall, A., Nelson, K., Poole, L. B. & Karplus, P. A. Structure-based insights into the catalytic power and conformational dexterity of peroxiredoxins. Antioxid. Redox Signal. 15, 795–815 (2011).
Flohé, L., Toppo, S., Cozza, G. & Ursini, F. A comparison of thiol peroxidase mechanisms. Antioxid. Redox Signal. 15, 763–780 (2011).
Ferrer-Sueta, G. et al. Factors affecting protein thiol reactivity and specificity in peroxide reduction. Chem. Res. Toxicol. 24, 434–450 (2011).
Hall, A., Parsonage, D., Poole, L. B. & Karplus, P. A. Structural evidence that peroxiredoxin catalytic power is based on transition-state stabilization. J. Mol. Biol. 402, 194–209 (2010).
Flohé, L. et al. Tryparedoxin peroxidase of Leishmania donovani: molecular cloning, heterologous expression, specificity, and catalytic mechanism. Arch. Biochem. Biophys. 397, 324–335 (2002).
König, J. et al. Reaction mechanism of plant 2-Cys peroxiredoxin. Role of the C terminus and the quaternary structure. J. Biol. Chem. 278, 24409–24420 (2003).
Montemartini, M., Kalisz, H. M., Hecht, H. J., Steinert, P. & Flohé, L. Activation of active-site cysteine residues in the peroxiredoxin-type tryparedoxin peroxidase of Crithidia fasciculata. Eur. J. Biochem. FEBS 264, 516–524 (1999).
Nagy, P. et al. Model for the exceptional reactivity of peroxiredoxins 2 and 3 with hydrogen peroxide: a kinetic and computational study. J. Biol. Chem. 286, 18048–18055 (2011).
Portillo-Ledesma, S. et al. Deconstructing the catalytic efficiency of peroxiredoxin-5 peroxidatic cysteine. Biochemistry 53, 6113–6125 (2014).
Barranco-Medina, S., Lázaro, J.-J. & Dietz, K.-J. The oligomeric conformation of peroxiredoxins links redox state to function. FEBS Lett. 583, 1809–1816 (2009).
Jang, H. H. et al. Two enzymes in one; two yeast peroxiredoxins display oxidative stress-dependent switching from a peroxidase to a molecular chaperone function. Cell 117, 625–635 (2004).
Matsumura, T. et al. Dimer-oligomer interconversion of wild-type and mutant rat 2-Cys peroxiredoxin: disulfide formation at dimer-dimer interfaces is not essential for decamerization. J. Biol. Chem. 283, 284–293 (2008).
Wood, Z. A., Poole, L. B., Hantgan, R. R. & Karplus, P. A. Dimers to doughnuts: redox-sensitive oligomerization of 2-cysteine peroxiredoxins. Biochemistry 41, 5493–5504 (2002).
Demasi, A. P. D., Pereira, G. A. G. & Netto, L. E. S. Yeast oxidative stress response. Influences of cytosolic thioredoxin peroxidase I and of the mitochondrial functional state. FEBS J. 273, 805–816 (2006).
Manta, B. et al. The peroxidase and peroxynitrite reductase activity of human erythrocyte peroxiredoxin 2. Arch. Biochem. Biophys. 484, 146–154 (2009).
Saccoccia, F. et al. Moonlighting by different stressors: crystal structure of the chaperone species of a 2-Cys peroxiredoxin. Struct. Lond. Engl. 1993 20, 429–439 (2012).
Wang, X., Wang, L., Wang, X. e., Sun, F. & Wang, C. Structural insights into the peroxidase activity and inactivation of human peroxiredoxin 4. Biochem. J 441, 113–118 (2012).
Jang, H. H. et al. Phosphorylation and concomitant structural changes in human 2-Cys peroxiredoxin isotype I differentially regulate its peroxidase and molecular chaperone functions. FEBS Lett. 580, 351–355 (2006).
Park, J. W., Piszczek, G., Rhee, S. G. & Chock, P. B. Glutathionylation of peroxiredoxin I induces decamer to dimers dissociation with concomitant loss of chaperone activity. Biochemistry 50, 3204–3210 (2011).
Noichri, Y. et al. In vivo parameters influencing 2-Cys Prx oligomerization: The role of enzyme sulfinylation. Redox Biol. 6, 326–333 (2015).
Morais, M. A. B. et al. How pH modulates the dimer-decamer interconversion of 2-Cys peroxiredoxins from the Prx1 subfamily. J. Biol. Chem. 290, 8582–8590 (2015).
Kitano, K., Niimura, Y., Nishiyama, Y. & Miki, K. Stimulation of peroxidase activity by decamerization related to ionic strength: AhpC protein from Amphibacillus xylanus. J. Biochem. (Tokyo) 126, 313–319 (1999).
Tairum, C. A., de Oliveira, M. A., Horta, B. B., Zara, F. J. & Netto, L. E. S. Disulfide biochemistry in 2-cys peroxiredoxin: requirement of Glu50 and Arg146 for the reduction of yeast Tsa1 by thioredoxin. J. Mol. Biol. 424, 28–41 (2012).
Nishio, M., Umezawa, Y., Fantini, J., Weiss, M. S. & Chakrabarti, P. CH-π hydrogen bonds in biological macromolecules. Phys. Chem. Chem. Phys. PCCP 16, 12648–12683 (2014).
Weiss, M. S., Brandl, M., Sühnel, J., Pal, D. & Hilgenfeld, R. More hydrogen bonds for the (structural) biologist. Trends Biochem. Sci. 26, 521–523 (2001).
Nielsen, M. H., Kidmose, R. T. & Jenner, L. B. Structure of TSA2 reveals novel features of the active-site loop of peroxiredoxins. Acta Crystallogr. Sect. Struct. Biol. 72, 158–167 (2016).
Munhoz, D. C. & Netto, L. E. S. Cytosolic Thioredoxin Peroxidase I and II Are Important Defenses of Yeast against Organic Hydroperoxide Insult. 279, 35219–35227 (2004).
Parsonage, D. et al. Dissecting peroxiredoxin catalysis: separating binding, peroxidation, and resolution for a bacterial AhpC. Biochemistry 54, 1567–1575 (2015).
Wood, Z. A., Poole, L. B. & Karplus, P. A. Peroxiredoxin evolution and the regulation of hydrogen peroxide signaling. Science 300, 650–653 (2003).
Oliveira, M. A. et al. Insights into the specificity of thioredoxin reductase-thioredoxin interactions. A structural and functional investigation of the yeast thioredoxin system. Biochemistry 49, 3317–3326 (2010).
Böhm, G., Muhr, R. & Jaenicke, R. Quantitative analysis of protein far UV circular dichroism spectra by neural networks. Protein Eng. 5, 191–195 (1992).
Segel, I. H. Enzyme Kinetics: Behavior and Analysis of Rapid Equilibrium and Steady State Enzyme Systems (Wiley Inter- science, 1993).
Hansen, R. E. & Winther, J. R. An introduction to methods for analyzing thiols and disulfides: Reactions, reagents, and practical considerations. Anal. Biochem. 394, 147–158 (2009).
Acknowledgements
We acknowledge the financial support from the Fundação de Amparo à Pesquisa do Estado de São Paulo grants 07/50930-3 and 13/07937-8 (Redox Processes in Biomedicine -REDOXOMA).
Author information
Authors and Affiliations
Contributions
C.A.T. and M.C.S. performed point mutations, expressed and purified the proteins; C.A.T., M.C.S., C.A.B. and M.H.T. performed steady state kinetic analysis, SEC and CD; Spectrofluorometric approaches were performed by C.A.T., R.R.G., J.C.T.Jr., C.J.N., S.P.-L. and G.F.-S.; C.A.T., M.A.d.O. and L.E.S.N. executed the structural analysis. L.E.S.N., O.A. and M.A.d.O. provided overall project supervision and wrote the paper along with C.A.T. and M.C.S.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Tairum, C., Santos, M., Breyer, C. et al. Catalytic Thr or Ser Residue Modulates Structural Switches in 2-Cys Peroxiredoxin by Distinct Mechanisms. Sci Rep 6, 33133 (2016). https://doi.org/10.1038/srep33133
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep33133
This article is cited by
-
Basis for using thioredoxin as an electron donor by Schizosaccharomyces pombe Gpx1 and Tpx1
AMB Express (2022)
-
Relevance of peroxiredoxins in pathogenic microorganisms
Applied Microbiology and Biotechnology (2021)
-
Functional and structural evaluation of the antileukaemic enzyme l-asparaginase II expressed at low temperature by different Escherichia coli strains
Biotechnology Letters (2020)
-
Glutaredoxin-like protein (GLP)—a novel bacteria sulfurtransferase that protects cells against cyanide and oxidative stresses
Applied Microbiology and Biotechnology (2020)
-
Analyses of the three 1-Cys Peroxiredoxins from Aspergillus fumigatus reveal that cytosolic Prx1 is central to H2O2 metabolism and virulence
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.