Abstract
Horizontal gene transfer (HGT) often has strong benefits for fungi. In a study of samples from apple canker in Shaanxi Province, China, diverse microbes, along with the necrotrophic pathogen Valsa mali, were found to colonize the apple bark, thus providing ample opportunity for HGT to occur. In the present study, we identified 32 HGT events in V. mali by combining phyletic distribution-based methods with phylogenetic analyses. Most of these HGTs were from bacteria, whereas several others were from eukaryotes. Three HGTs putatively functioned in competition with actinomycetes, some of which showed a significant inhibitory effect on V. mali. Three HGTs that were probably involved in nitrogen uptake were also identified. Ten HGTs were thought to be involved in pathogenicity because they were related to known virulence factors, including cell wall-degrading enzymes and candidate effector proteins. HGT14, together with HGT32, was shown to contribute to bleomycin resistance of V. mali.These results suggest that HGT drives the adaptive evolution of V. mali. The HGTs identified here provide new clues for unveiling the adaptation mechanisms and virulence determinants of V. mali.
Similar content being viewed by others
Introduction
Horizontal gene transfer (HGT) is the stable transmission of genetic materials between species through any mechanism other than vertical inheritance1. HGT plays a significant role in the evolution of prokaryotic lineages, such as by providing novel genes involved in pathogenicity and contributing to adaptive traits2,3. In eukaryotes, compared with prokaryotes, there is less evidence for functional HGT, but the phenotypic consequences can also be significant in the adaptive evolution of eukaryotes4. Hundreds of fungal genomes are now available, and a growing body of data suggest that HGT has had a profound impact on the evolution of pathogenic traits in fungal pathogens5,6. When considering their functions in infection processes and ecological niches, HGTs in fungi can be divided into the following three categories: those competing with other microbes (antimicrobial genes7 and secondary metabolite toxins1), those utilizing nutrients (membrane transporter) and those interacting with hosts (reviewed in Soanes and Richards, 20146). Therefore, understanding of functional HGT in fungal pathogens should facilitate the discovery of novel genes involved in niche adaptation, particularly in pathogenicity.
Valsa canker caused by the ascomycetous Valsa mali, is one of the most destructive diseases affecting apple trees, causing significant yield losses in eastern Asia8,9. This necrotrophic pathogen infects trees mainly through wounds and results in severe maceration and necrosis of bark tissues10. The dead and dying tissues can be subsequently colonized by diverse saprophytes, thus implying that these microbes occupy the same environment as V. mali and therefore may contribute genes to V. mali via HGT, because shared habitat is a major factor driving transfers1,5,11. Thus, V. mali may have ample opportunity for HGT to occur. We have previously reported the V. mali genome sequence, which suggests a potential adaptation to colonize woody bark12. In the present study, phyletic distribution-based methods13,14 and phylogenetic analyses were used to identify potential HGTs in V. mali. The functions of two HGTs of interest were verified experimentally. The results will provide new clues for unveiling adaptation mechanisms and virulence determinants in V. mali.
Results and Discussion
Identification of HGT genes
Identifying HGT in eukaryotes is difficult because of their highly complex genome content. The gold standard for identifying HGT with confidence is phylogenetic incongruence15, although its throughput is much lower than that of surrogate methods especially when hundreds of genes or even a genome are being analysed. Thus, we used the effective surrogate tool HGT-Finer13 to identify HGT candidates and then verified the results gene-by-gene with phylogenetic analysis. In total, 345 candidates were identified by HGT-Finer, and 32 HGT genes were verified by phylogenetic analyses, manual checking and another phyletic distribution-based method, DarkHorse14 (Table 1, Supplementary Files).
Most of these HGTs were derived from bacterial sources (Table 1)16, probably because prokaryote-to-fungal HGT is more likely than eukaryote-to-fungal HGT. Consistently with their prokaryotic origin, 18 of the 25 bacterial HGTs had no introns. We also identified five HGTs that were transferred from ascomycetes into bacteria (HGT15, HGT21), basidiomycetes (HGT12, HGT15) and oomycetes (HGT18, HGT20). Compared with prokaryote-fungi transfers, fungi-fungi transfers, especially among closely related species, are difficult to identify because of independent gene loss17. Thus, most of the fungi-fungi HGT candidates identified by HGT-Finder were not well supported by phylogenetic analysis. Methods for specifically identifying HGTs among closely related eukaryotes are currently not available.
Functional annotations showed that the HGTs identified mainly affected enzymes (69%) from diverse metabolic pathways and genes (19%) with unknown functions (Table 2). The roles of HGTs in their recipient organisms are often unknown without further experiments18. Whether an HGT is expressed under specific conditions might provide clues regarding its adaptive value. Therefore, transcriptomes of V. mali during infection of apple bark, reported by us previously19, were used to identify HGTs potentially involved in pathogenicity. Six HGTs were strongly upregulated during infection (Table 2), thus suggesting a potential role in V. mali-apple interaction. Their specific adaptive values will be discussed in the following section.
Putative adaptive value of HGTs in V. mali
Competing with other microbes
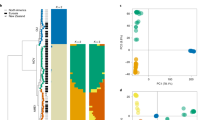
To protect themselves against competitors, microbes often produce secondary metabolite toxins that kill other microbes. The severe maceration and necrosis of apple bark caused by V. mali provide ample opportunity for other microbes to colonize. HGT14 and HGT32, bleomycin (Bm) resistance proteins, might enable V. mali to be resistant to bleomycin which is a family of antibiotics produced by actinomycetes that cause cell death in eukaryotes and prokaryotes20. To test this hypothesis, the capacity of Bm resistance of V. mali, Fusarium graminearum and Aspergillus nidulans was examined under three levels of Bm stress (10, 50 and 100 μg/ml). Compared wiht F. graminearum and A. nidulans, V. mali was significantly more resistant (Figs 1a and S2). RT-qPCR analysis showed that HGT32 but not HGT14 was induced at a high level (50 μg/ml) of Bm stress (Fig. 1b). However, a null mutant of HGT32 did not show reduced resistance (Fig. 1d), and HGT14 of the HGT32 null mutant was induced under Bm stress (50 μg/ml) at 36 hpi (Fig. 1c). Thus, we generated the double deletion mutant of HGT14 and HGT32, which showed a significantly reduced resistance compared with the resistance of the wild type and HGT32 null mutant (Fig. 1e). These results suggest that HGT14, together with HGT32, contributes to Bm-resistance.
HGT14, together with HGT32, contributes to bleomycin resistance.
(a) Inhibition ratios of radial growth of three fungi by bleomycin. (b) Relative expression of HGT14 and HGT32 genes under bleomycin stress (50 μg/ml). (c) Relative expression of HGT14 gene of the HGT32 null mutant under bleomycin stress (50 μg/ml). (d) Inhibition ratios of radial growth of the HGT32 null mutant by bleomycin (20 μg/ml). (e) Inhibition ratios of radial growth of the HGT14 HGT32 double deletion mutant by bleomycin (50 μg/ml). Bleomycin resistance assays and qRT-PCR analyses were repeated three and three times, respectively. Error bars represent the mean S.D. and asterisks (**) indicate significant differences (P < 0.01).
Another gene predicted to be associated with competition was HGT5, an O-methyltransferase involved in polyketide biosynthesis (Table 2). We reasoned that HGT5 probably functions in competition rather than in synthesizing toxic polyketides, because the genes involved in polyketide biosynthesis are often clustered in fungi and V. mali acquired HGT5 as a single gene from actinomycetes. HGT5 in V. mali is probably used as a protective device to modify or detoxify specific toxic polyketides produced by actinomycetes, as is the case for blmB in the bleomycin biosynthesis gene cluster. The blmB gene encodes a N-acetyltransferase, which modifies and inactivates bleomycin, conferring self-protection; moreover, other bacteria transformants carrying blmB may also gain bleomycin resistance21. In addition, three of the five fungi that acquired HGT5 were non-pathogens, including an endophytic Pestalotiopsis fici, an endomycorrhizal fungus Oidiodendron maius and the saprotrophic Penicillium roqueforti (Supplementary Files).
Intriguingly, these three HGTs are all involved in competition with actinomycetes. Indeed, an endophytic actinomycete strain Hhs.015 isolated from cucumber root22 significantly inhibits the conidial germination and mycelial growth of V. mali23. These findings suggest that HGT has driven the adaptive evolution of V. mali for competing with other microbes.
Nutrient uptake
The types of nutrients that can be utilized by fungi are determined largely by their membrane transporters, because these microbes feed exclusively by osmotrophy. Thus, acquiring a novel transporter gene might enable fungi to utilize a new source of nutrition or gain a competitive advantage against other microbes in their ecological niches. HGT11, a major facilitator superfamily (MFS) transporter of a basidiomycetous source, was found only in the genus Valsa (including the pear canker pathogen V. pyri) (Tables 1 and 2 and Supplementary Files). HGT11 has significant similarity to the Saccharomyces cerevisiae protein DAL5, an allantoate/dipeptide permease (family 2.A.1.14.4) in the Transporter Classification Database. DAL5 transports several substrates and is sensitive to nitrogen catabolite repression24,25,26,27,28. Thus, by gaining HGT11, V. mali might possibly utilize new sources of nitrogen.
Lysine is synthesized de novo by the α-aminoadipate (AAA) pathway in higher fungi, and by the diaminopimelate (DAP) pathway in bacteria and plants29. The two pathways evolved separately in organisms, and no organism is known to possess both pathways30. However, we found that some higher fungi acquired the DapE gene (HGT1) in the DAP pathway by horizontal gene transfer (Supplementary Files). HGT1 was transferred from Proteobacteria, which use the succinylase pathway (a variant of the DAP pathway) for lysine biosynthesis31. HGT1 was also found in many other fungi including Colletotrichum graminicola. The orthologous gene GLRG_10812 was horizontally transferred from bacteria32. Further analysis showed that the V. mali genome contained all of the essential genes of the AAA pathway and six of the nine genes in the succinylase pathway (Supplementary Table S1). However, in the case of selfish genetic elements33, the incomplete DAP pathway might still work through utilizing intermediates synthesized by symbiotic bacteria in the environment, thus providing a strong benefit to acquiring nitrogen from the nitrogen-limited apple bark.
Nitrogen is essential for growth, and disruption of nitrogen nutrition is often associated with the virulence of phytopathogens34,35. Nitrogen regulation is relatively well studied in yeast and filamentous fungi35,36. Glutamine and ammonium are preferentially used as nitrogen sources, and the NmrA protein can be activated to repress nitrogen catabolic genes when these sources are sufficient36. NmrA is highly conserved in filamentous ascomycetes and is also found in oomycetes which gain NmrA by HGT37. In this study, we found an HGT event of fungus-oomycete NmrA (HGT20), which appears to be a potential virulence factor, because both the donor and recipient groups are phytopathogens (Fig. 2). Indeed, the gene that encodes HGT20 was markedly upregulated (log2-fold change > 6) during V. mali infection (Table 2), thus suggesting that readily assimilated nitrogen sources were sufficient in the infected tissues and that NmrA might be important for regulating selective nitrogen utilization during infection. Consistently with this hypothesis, three of the five ammonia transporter genes have also been found to be significantly upregulated in planta12. Functions of HGT20 in nitrogen regulation and virulence in V. mali, and especially in oomycetes whose nitrogen regulation is poorly understood, are worth investigating.
Additional HGTs predicted to be associated with nutrient uptake include a phosphomannomutase (HGT6) and a uridine nucleosidase (HGT17), which putatively function in the mannose biosynthetic process and nucleotide metabolism, respectively. A putative fructosyl amino acid oxidase (HGT12), which putatively functions in catabolizing naturally occurring fructosyl amino acids38, was also identified. Intriguingly, most of these nutrient-related HGTs were predicted to be involved in obtaining nitrogen, whose content in apple bark is relatively low39. Therefore, these nitrogen-related HGTs of V. mali are possible drivers of adaptive evolution in nitrogen uptake.
Interacting with the host
After pathogen attack, many plants produce low molecular weight antimicrobial compounds. Likewise, several cytochrome P450 genes potentially involved in secondary metabolite biosynthesis in apples are upregulated after V. mali infection40. To counteract these toxic compounds, fungi possess membrane transporters that pump the toxins out of the cell or possess enzymes to detoxify them. Two enzymes with roles in detoxifying phytoalexins are HGT10 (dioxygenase) and HGT28 (thiosulphate sulphurtransferase). HGT10 putatively functions in the degradation of aromatic compounds via its LigB domain. HGT28 is found only in the genus Valsa and is a mitochondrial enzyme that detoxifies cyanide41. The production of poisonous hydrogen cyanide is known to be used by plants to protect against insects and other herbivorous animals42 and has been recently reported to be required for inducible pathogen defence in Arabidopsis43. To overcome plant cyanogenesis, arthropods have been found to detoxify plant-produced cyanide by acquiring a horizontally transferred gene involved in sulphur amino acid biosynthesis in bacteria44. In the current study, we found another HGT event of bacterial origin, which might enable fungi to detoxify cyanide through a different mechanism (HGT28). V. mali infects apple trees mainly through wounds10, where cyanide production may be induced. Thus, we speculate that the poisonous cyanide itself and its induced defence might possibly be inhibited or overcome by HGT28 if this protein is functional during the V. mali-apple interaction.
As a typical necrotrophic pathogen on apples, V. mali can degrade woody tissues through secreted cell wall-degrading enzymes (CWDEs)10,12,19. Diverse HGTs that putatively function in degrading plant cell wall have been found in eukaryotic plant pathogenic microbes6. In the genome of V. mali, at least six CWDEs involved in HGT events were identified in the current study (Table 2). Among these, only HGT2 and HGT4 contained the N-terminal signal peptide, thus suggesting that they are probably secreted proteins. However, most of the HGT2 proteins of the donor bacteria and the recipient fungi had no N-terminal signal peptide, except several proteins from two clades, including V. mali (Fig. 3). HGT2, a putative hydrolase and peptidase, was strongly activated during infection (Table 2), and null mutants of the HGT2 gene in V. mali show a significantly reduced virulence on apple trees (Feng et al., unpublished). These results suggest that HGT2 is a potentially important virulence factor in V. mali. Beyond their enzymatic functions, CWDEs can also act as a microbe-associated molecular pattern (MAMP) that activates plant defence responses, as reported for endopolygalacturonases (pectinase) of Botrytis cinerea45 and a glycoside hydrolase family 12 (GH12) protein of Phytophthora sojae46. Exploration of the specific functions of HGT2 during V. mali-apple interactions is in progress.
Salicylic acid (SA) is an important plant hormone involved in defence responses against biotrophic pathogens47. To counteract SA-induced defence responses, pathogens have several weapons that target and disturb SA biosynthesis and signalling, such as salicylate hydroxylase, which degrades SA48. Furthermore, genes involved in apple SA signalling are significantly upregulated and enriched after V. mali infection40. In addition, a putative salicylate hydroxylase (HGT9) transferred from bacteria was identified in V. mali (Supplementary Files). Nevertheless, only one of the three salicylate hydroxylases is active in the smut fungus Ustilago maydis49, and there are also three salicylate hydroxylase homologues in V. mali (data not shown). Another protein that probably functions in regulating plant immunity is HGT18, a small secreted protein with unknown function that was strongly activated during infection (Table 2). HGT18 seems to be a candidate effector protein with these characteristics, and it was transferred from fungi to phytopathogenic oomycetes (Phytophthora spp.) (Supplementary Files).
Conclusion
By combing phyletic distribution-based methods and phylogenetic analyses, we identified 32 HGT events in the apple canker pathogen V. mali. Most of these HGTs were of bacterial origin, and several were of eukaryotic origin. Three HGTs putatively functioned in competition with actinomycetes some of which showed a significant inhibitory effect on V. mali. Three HGTs that are likely to be involved in nitrogen uptake were identified in V. mali, which can effectively utilize nitrogen in apple bark. Ten HGTs were thought to be involved in pathogenicity, as they were related to known virulence factors. HGT14, together with HGT32, was shown to contribute to bleomycin resistance of V. mali. These results suggest that HGT drives the adaptive evolution of V. mali.
Methods
Identification of HGT candidates
The proteome of V. mali12 was searched using blastp (v2.2.30+) against GenBank NR database (data-version 20150706). The blastp output was then subjected to HGT-Finer (R threshold ranging from 0.2 to 0.9, Q value < 0.01)13 to identify HGT candidates. To confirm the results of HGT-Finder, another phyletic distribution-based method, DarkHorse14 was used to calculate LPI (lineage probability index) scores of HGTs.
Phylogenetic analysis
For phylogenetic analysis, the protein sequences of the top 100 blastp hits for each HGT candidate were retrieved from GenBank. Multiple sequence alignments were performed using MAFFT (v7.245)50, and poorly aligned regions were removed by trimAl (v1.4)51. Maximum likelihood phylogenetic trees were constructed using IQ-TREE (v 1.3.11)52 with the best-fit substitution model automatically selected, and branch supports were assessed with ultrafast bootstrap53 and SH-aLRT test (1000 replicates). Phylogenetic trees were viewed and produced by iTOL (v2, http://itol.embl.de/)54 and FigTree (v1.4.2, http://tree.bio.ed.ac.uk/software/figtree/).
Functional annotation of HGTs
The putative functions of HGTs were predicted with the Pfam55, NCBI CDD56 and KEGG57 databases. The N-terminal signal peptide was predicted with the SignalP 4.1 server58. The expression data were extracted from Yin et al.12.
Bleomycin resistance assays
The bleomycin resistance of three fungi (Valsa mali, Aspergillus nidulans and Fusarium graminearum) was evaluated under three levels of bleomycin stress (10, 50 and 100 μg/ml). Fungal isolates were grown on potato dextrose agar (PDA) media at 25 °C in the dark. Diameters of colonies were measured two days post inoculation. Each assay was repeated ten times. The data were analysed with ANOVA or t-test by using the online tool VassarStats (http://www.vassarstats.net/).
Functional studies on bleomycin resistance genes
For RT-qPCR analysis, fungi were grown in potato dextrose broth at 25 °C in the dark for two days. Total RNA was extracted using a Quick RNA isolation Kit (Huayueyang Biotechnology, Beijing, China) according to the manufacturer’s instructions. First strand cDNA synthesis and qPCR were performed using a RevertAid First Strand cDNA Synthesis Kit (Thermo Scientific, Hudson, NH, USA) and RealStar Green Mixture (GenStar, Beijing, China), respectively. The qPCR assays were performed with a CFX96 ConnectTM Real-Time System (Bio-Rad, Hercules, CA, USA), and G6PDH gene was used as an internal reference59. The genes were knocked out according to a previously described method (Figure S3)60. Primers used for RT-qPCR and gene deletion are listed in Table S2.
Additional Information
How to cite this article: Yin, Z. et al. Horizontal gene transfer drives adaptive colonization of apple trees by the fungal pathogen Valsa mali. Sci. Rep. 6, 33129; doi: 10.1038/srep33129 (2016).
References
Howlett, B. J. & Oliver, R. P. Evolution of virulence in eukaryotic microbes (eds Sibley, D. et al.) Ch. 25, 473–484 (John Wiley & Sons, Inc. 2012).
Ochman, H., Lawrence, J. G. & Groisman, E. A. Lateral gene transfer and the nature of bacterial innovation. Nature 405, 299–304 (2000).
Jain, R., Rivera, M. C., Moore, J. E. & Lake, J. A. Horizontal gene transfer accelerates genome innovation and evolution. Mol Biol Evol 20, 1598–1602 (2003).
Schonknecht, G., Weber, A. P. & Lercher, M. J. Horizontal gene acquisitions by eukaryotes as drivers of adaptive evolution. Bioessays 36, 9–20, doi: 10.1002/bies.201300095 (2014).
Richards, T. A., Leonard, G., Soanes, D. M. & Talbot, N. J. Gene transfer into the fungi. Fungal Biol Rev 25, 98–110, doi: 10.1016/j.fbr.2011.04.003 (2011).
Soanes, D. & Richards, T. A. Horizontal gene transfer in eukaryotic plant pathogens. Annu Rev Phytopathol 52, 583–614, doi: 10.1146/annurev-phyto-102313-050127 (2014).
Metcalf, J. A., Funkhouser-Jones, L. J., Brileya, K., Reysenbach, A. L. & Bordenstein, S. R. Antibacterial gene transfer across the tree of life. Elife 3, e04266 (2014).
Lee, D. H., Lee, S. W., Choi, K. H., Kim, D. A. & Uhm, J. Y. Survey on the occurrence of apple diseases in Korea from 1992 to 2000. Plant Pathol J 22, 375 (2006).
Li, Z. et al. Survey of Apple Valsa canker in Weibei area of Shaanxi province. Acta Agriculturae Boreali-Occidentalis Sinica 1, 029 (2013).
Ke, X., Huang, L., Han, Q., Gao, X. & Kang, Z. Histological and cytological investigations of the infection and colonization of apple bark by Valsa mali var. mali. Australas Plant Pathol 42, 85–93 (2013).
Hirt, R. P., Alsmark, C. & Embley, T. M. Lateral gene transfers and the origins of the eukaryote proteome: a view from microbial parasites. Curr Opin Microbiol 23, 155–162, doi: 10.1016/j.mib.2014.11.018 (2015).
Yin, Z. et al. Genome sequence of Valsa canker pathogens uncovers a potential adaptation of colonization of woody bark. New Phytol 208, 1202–1216 (2015).
Nguyen, M., Ekstrom, A., Li, X. & Yin, Y. HGT-Finder: A new tool for horizontal gene transfer finding and application to Aspergillus genomes. Toxins 7, 4035–4053 (2015).
Podell, S. & Gaasterland, T. DarkHorse: a method for genome-wide prediction of horizontal gene transfer. Genome Biol 8, 1 (2007).
Keeling, P. J. & Palmer, J. D. Horizontal gene transfer in eukaryotic evolution. Nat Rev Genet 9, 605–618, doi: 10.1038/nrg2386 (2008).
Fitzpatrick, D. A. Horizontal gene transfer in fungi. FEMS Microbiol Lett 329, 1–8 (2012).
Ku, C. et al. Endosymbiotic origin and differential loss of eukaryotic genes. Nature 524, 427–432 (2015).
Alsmark, C. et al. Patterns of prokaryotic lateral gene transfers affecting parasitic microbial eukaryotes. Genome Biol 14, R19, doi: 10.1186/gb-2013-14-2-r19 (2013).
Ke, X. et al. Transcriptome profiling to identify genes involved in pathogenicity of Valsa mali on apple tree. Fungal Genet Biol 68, 31–38 (2014).
Sugiyama, M. & Kumagai, T. Molecular and structural biology of bleomycin and its resistance determinants. J Biosci Bioeng 93, 105–116 (2002).
Sugiyama, M. et al. Characterisation by molecular cloning of two genes from Streptomyces verticillus encoding resistance to bleomycin. Gene 151, 11 (1994).
Yan, X., Huang, L., Tu, X., Gao, X. & Kang, Z. Saccharothrix yanglingensis sp. nov., an antagonistic endophytic actinomycete isolated from cucumber plant. Antonie van Leeuwenhoek 101, 141–146 (2012).
Li, Z. et al. Saccharothrix yanglingensis strain Hhs. 015 is a promising biocontrol agent on apple valsa canker. Plant Dis, 100, 510–514, PDIS-02-15-0190-RE (2015).
Chisholm, V., Lea, H., Rai, R. & Cooper, T. Regulation of allantoate transport in wild-type and mutant strains of Saccharomyces cerevisiae. J Bacteriol 169, 1684–1690 (1987).
Drillien, R. & Lacroute, F. Ureidosuccinic acid uptake in yeast and some aspects of its regulation. J Bacteriol 109, 203–208 (1972).
Cai, H., Hauser, M., Naider, F. & Becker, J. M. Differential regulation and substrate preferences in two peptide transporters of Saccharomyces cerevisiae. Eukaryotic cell 6, 1805–1813 (2007).
Rai, R., Genbauffe, F., Lea, H. & Cooper, T. G. Transcriptional regulation of the DAL5 gene in Saccharomyces cerevisiae. J Bacteriol 169, 3521–3524 (1987).
Turoscy, V. & Cooper, T. Ureidosuccinate is transported by the allantoate transport system in Saccharomyces cerevisiae. J Bacteriol 169, 2598–2600 (1987).
Zabriskie, T. M. & Jackson, M. D. Lysine biosynthesis and metabolism in fungi. Nat Prod Rep 17, 85–97 (2000).
Liu, Y., White, R. H. & Whitman, W. B. Methanococci use the diaminopimelate aminotransferase (DapL) pathway for lysine biosynthesis. J Bacteriol 192, 3304–3310 (2010).
Fuchs, T. M., Schneider, B., Krumbach, K., Eggeling, L. & Gross, R. Characterization of a Bordetella pertussis diaminopimelate (DAP) biosynthesis locus identifies dapC, a novel gene coding for an N-Succinyl-L,L-DAP aminotransferase. J Bacteriol 182, 3626–3631 (2000).
Jaramillo, V. D., Sukno, S. A. & Thon, M. R. Identification of horizontally transferred genes in the genus Colletotrichum reveals a steady tempo of bacterial to fungal gene transfer. BMC genomics 16, 2 (2015).
Werren, J. H. Selfish genetic elements, genetic conflict, and evolutionary innovation. Proc Natl Acad Sci USA 108, 10863–10870 (2011).
Fagard, M. et al. Nitrogen metabolism meets phytopathology. J Exp Bot 65, 5643–5656 (2014).
Fernandez, J., Marroquin-Guzman, M. & Wilson, R. A. Mechanisms of nutrient acquisition and utilization during fungal infections of leaves. Annu Rev Phytopathol 52, 155–174, doi: 10.1146/annurev-phyto-102313-050135 (2014).
Wong, K. H., Hynes, M. J. & Davis, M. A. Recent advances in nitrogen regulation: a comparison between Saccharomyces cerevisiae and filamentous fungi. Eukaryotic cell 7, 917–925 (2008).
Richards, T. A. et al. Horizontal gene transfer facilitated the evolution of plant parasitic mechanisms in the oomycetes. Proc Natl Acad Sci USA 108, 15258–15263, doi: 10.1073/pnas.1105100108 (2011).
Ferri, S., Kim, S., Tsugawa, W. & Sode, K. Review of fructosyl amino acid oxidase engineering research: a glimpse into the future of hemoglobin A1c biosensing. J Diabetes Sci Technol 3, 585–592 (2009).
Dickson, N., Richard, T., Kozlowski, R. & Sobel, P. L. Composting to reduce the waste stream: a guide to small scale food and yard waste composting. (Northeast regional agricultural engineering service, 1991).
Yin, Z., Ke, X., Kang, Z. & Huang, L. Apple resistance responses against Valsa mali revealed by transcriptomics analyses. Physiol Mol Plant Pathol 93, 85–92 (2016).
Cipollone, R., Ascenzi, P., Tomao, P., Imperi, F. & Visca, P. Enzymatic detoxification of cyanide: clues from Pseudomonas aeruginosa rhodanese. J Mol Microbiol Biotechnol 15, 199–211 (2008).
Zagrobelny, M. et al. Cyanogenic glucosides and plant-insect interactions. Phytochemistry 65, 293–306 (2004).
Rajniak, J., Barco, B., Clay, N. K. & Sattely, E. S. A new cyanogenic metabolite in Arabidopsis required for inducible pathogen defence. Nature (2015).
Wybouw, N. et al. A gene horizontally transferred from bacteria protects arthropods from host plant cyanide poisoning. Elife 3, e02365 (2014).
Zhang, L. et al. Fungal endopolygalacturonases are recognized as microbe-associated molecular patterns by the arabidopsis receptor-like protein Responsiveness to Botrytis Polygalacturonases1. Plant Physiol 164, 352–364, doi: 10.1104/pp.113.230698 (2014).
Ma, Z. et al. A Phytophthora sojae glycoside hydrolase 12 protein is a major virulence factor during soybean infection and is recognized as a PAMP. The Plant cell 27, 2057–2072, doi: 10.1105/tpc.15.00390 (2015).
Rivas-San Vicente, M. & Plasencia, J. Salicylic acid beyond defence: its role in plant growth and development. J Exp Bot 62, 3321–3338 (2011).
Tanaka, S., Han, X. & Kahmann, R. Microbial effectors target multiple steps in the salicylic acid production and signaling pathway. Front Plant Sci 6, 349 (2015).
Rabe, F., Ajami-Rashidi, Z., Doehlemann, G., Kahmann, R. & Djamei, A. Degradation of the plant defence hormone salicylic acid by the biotrophic fungus Ustilago maydis. Mol Microbiol 89, 179–188 (2013).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30, 772–780 (2013).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. Iq-tree: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol, 32, 268–274 (2014).
Minh, B. Q., Nguyen, M. A. T. & von Haeseler, A. Ultrafast approximation for phylogenetic bootstrap. Mol Biol Evol, 30, 1188–1195 (2013).
Letunic, I. & Bork, P. Interactive tree of life v2: online annotation and display of phylogenetic trees made easy. Nucleic Acids Res, 39, W475–W478 (2011).
Finn, R. D. et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res, 44, D279–D285 (2015).
Marchler-Bauer, A. et al. CDD: NCBI’s conserved domain database. Nucleic Acids Res, 43, D222–D226 (2015).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res 44, D457–D462 (2016).
Petersen, T. N., Brunak, S., von Heijne, G. & Nielsen, H. SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8, 785–786 (2011).
Yin, Z. et al. Validation of reference genes for gene expression analysis in Valsa mali var. mali using real-time quantitative PCR. World J Microbiol Biotechnol 29, 1563–1571 (2013).
Li, Z. et al. Candidate effector proteins of the necrotrophic apple canker pathogen Valsa mali can suppress BAX-induced PCD. Front Plant Sci 6, 579 (2015).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Nos 31471732, 31301606), the Program for Agriculture (nyhyzx201203034-03) and the 111 Project (B07049).
Author information
Authors and Affiliations
Contributions
Z.Y., B.Z. and H.F. performed the bioinformatics analyses. B.Z. performed the experimental work. Z.Y. and L.H. wrote the manuscript. All authors have read and approved the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Yin, Z., Zhu, B., Feng, H. et al. Horizontal gene transfer drives adaptive colonization of apple trees by the fungal pathogen Valsa mali. Sci Rep 6, 33129 (2016). https://doi.org/10.1038/srep33129
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep33129
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.