Abstract
Rotation of bacterial flagellar motor is driven by the interaction between the stator and rotor and the driving energy is supplied by ion influx through the stator channel. The stator is composed of the MotA and MotB proteins, which form a hetero-hexameric complex with a stoichiometry of four MotA and two MotB molecules. MotA and MotB are four- and single-transmembrane proteins, respectively. To generate torque, the MotA/MotB stator unit changes its conformation in response to the ion influx and interacts with the rotor protein FliG. Here, we overproduced and purified MotA of the hyperthermophilic bacterium Aquifex aeolicus. A chemical crosslinking experiment revealed that MotA formed a multimeric complex, most likely a tetramer. The three-dimensional structure of the purified MotA, reconstructed by electron microscopy single particle imaging, consisted of a slightly elongated globular domain and a pair of arch-like domains with spiky projections, likely to correspond to the transmembrane and cytoplasmic domains, respectively. We show that MotA molecules can form a stable tetrameric complex without MotB and for the first time, demonstrate the cytoplasmic structure of the stator.
Similar content being viewed by others
Introduction
Various motor proteins are essential for different biological activities such as cell locomotion, cell morphogenesis, metabolism and material transport. Motor proteins convert various types of energy, such as ATP hydrolysis or electrochemical potential, into mechanical force for directional motion. Motor proteins are divided into two types: linear motors, such as myosin-actin and kinesin/dynein microtubule motors; and rotary motors, such as ATP synthase and bacterial flagellar motor. The bacterial flagellar motor is composed of about 20 different proteins and drives the rotation of the helical flagellar filament as a propeller for the locomotion of the bacterial cell. Torque of the flagellar motor is generated by the interaction between two components: the rotor and the stator. The rotor is composed of several rings with a central shaft. The stator is the energy-converting unit that works as an ion channel and interacts with the rotor. It is believed that the ion influx through the channel induces a conformational change in the stator, allowing its interaction with the rotor to generate torque1,2.
The stator is composed of two transmembrane proteins, MotA and MotB (Fig. S1). MotA has four transmembrane helices and a large cytoplasmic domain between the second and third transmembrane helices3. This cytoplasmic domain is thought to interact with a rotor protein, FliG4. MotB has a single transmembrane helix and a large periplasmic C-terminal domain, composed of a plug region for the channel, a peptidoglycan-binding region and a flexible linker that connects these two parts5,6,7. MotA and MotB form a MotA4MotB2 hetero-hexameric complex, containing an ion channel formed by the third and fourth transmembrane regions of MotA and a transmembrane region of MotB8,9,10. Previous studies on characterizing the stator were performed using purified MotA/MotB or its homologous proteins. PomA and PomB are homologs of MotA and MotB, respectively, in the sodium-driven flagellar motor of Vibrio alginolyticus. Purified PomA and PomB formed a PomA4PomB2 hexameric complex, whereas PomA alone formed a PomA2 homodimer when solubilized in detergent octyl β-D-glucopyranoside (OG)11,12. The MotA/MotB purified from the proton-driven motor of Escherichia coli also formed a MotA4MotB2 complex when solubilized in detergent n-dodecylphosphocholine (DPC)10. The purified PomA/PomB complex is functional as an ion channel when reconstituted into membrane vesicles12. The topology and arrangement of the transmembrane regions of the MotA/MotB complex were analyzed by disulfide crosslinking experiments8,9 and molecular dynamics simulation13. The crystal structures of the periplasmic peptidoglycan-binding domains of MotB and PomB show folds similar to those of other peptidoglycan-binding proteins14,15,16,17,18. The three-dimensional (3D) structure of the purified PomA/PomB complex, solubilized in detergent Cymal-5 and determined by electron microscopy (EM) single particle image analysis, revealed a globular shape with two periplasmic arms19.
The crystal structure of full length FliG from hyperthermophilic Aquifex aeolicus has been described previously20. FliG is thought to form a ring of approximately 30-fold rotational symmetry and to directly interact with MotA to generate torque4,20,21,22,23. Recently, we reported that the MotA/MotB complex of A. aeolicus can function in the motor of E. coli after two modifications: the replacement of the periplasmic region of MotB with that of E. coli; and the introduction of a point mutation at the C-terminal region of MotA (A225D)24. In the present study, we characterized the purified native MotA protein of A. aeolicus. We screened conditions to optimize the purification method and characterized the multimeric complex formation and structure by biochemical methods and EM image analysis.
Results
Screening of stator proteins and detergents
First, we cloned the genes encoding the stator proteins of the flagellar motor from various bacterial strains to identify highly expressing and stable stator proteins. The motA and motB genes or their orthologs were cloned from a marine bacterium, V. alginolyticus; three piezophilic bacteria, Photobacterium profundum, Shewanella benthica and Shewanella violacea; and two thermophilic bacteria, Thermotoga maritima and A. aeolicus, into two plasmids, pColdI and pBAD33. When motA/motB were cloned into pColdI, a six histidine (His6) tag was fused to the N-terminus of the produced MotA protein. E. coli cells were transformed with the pColdI-based plasmid and the production of MotA protein was tested by sodium dodecylsulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and immunoblotting using an anti-His antibody (Figs S2A and S2B). We also tested the function of the stator proteins expressed from the pBAD33-based plasmids in E. coli. We previously reported that the stator composed of A. aeolicus MotA and chimeric MotB functioned in E. coli flagellar motor24 and here we found that some of the stator from other species were also functional in the different hosts (Fig. S3 and Table S2). From the SDS-PAGE experiment, we found that MotAAa (MotA from A. aeolicus) showed the most stable and clear band (Fig. S2C) and therefore, decided to use this protein in this study. The whole-cell sample harboring the plasmid containing motAAa/motB1Aa was disrupted by sonication and separated into soluble cytoplasmic and insoluble membrane fractions by ultracentrifugation. Then, the membrane fraction was solubilized by detergent Cymal-5 and the MotAAa and MotBAa proteins were affinity-purified using a hexahistidine tag (Fig. S2C). Although both motAAa and motB1Aa were cloned into a plasmid as operons, only MotAAa was purified (Fig. S2D). Therefore, we used a plasmid expressing only motAAa (pColdI-motAAa) for the following experiments. A. aeolicus is known to have another motBAa gene, motB2Aa, immediately downstream of motB1Aa. We also cloned the motB2Aa gene alone and overproduced it in E. coli cells, but MotB2Aa was not soluble by Cymal-5 (Fig. S2E).
To screen the most effective detergents for solubilization of the MotAAa protein, the membrane fraction was treated with various detergents at a concentration of 1% (w/v) and the soluble and insoluble fractions were separated by ultracentrifugation as the supernatant and pellet fractions, respectively (Fig. 1). Among the nine detergents tested in this study, DPC, Cymal-5, DDM, DM and SMC were effective for the solubilization of MotAAa. Consequently, we decided to use Cymal-5, DDM and DM for the purification of the MotAAa protein in this study.
Purification of MotA
The MotAAa (hereafter, simply described as MotA) protein was solubilized in DDM as described above and purified using histidine-affinity beads and gel filtration chromatography. By gel filtration chromatography, three peaks were obtained for the MotA protein. The apparent sizes of the complexes in these three fractions were estimated from their elution peak positions: ca. 230 kDa, ca. 420 kDa and more than ca. 600 kDa (beyond the void volume) (Fig. 2A, top). When the fraction containing the 230-kDa complex was separated by gel filtration again, a single peak was detected with an estimated size of ca. 210 kDa (Fig. 2A, bottom). Coomassie brilliant blue (CBB) staining and immunoblotting analysis using anti-His antibody showed that this stable peak of ca. 210 kDa contained highly purified MotA protein (Fig. 2B). This size of 210 kDa is significantly larger than 28 kDa, the molecular mass of MotA with N-terminal His tag, suggesting that the purified MotA formed a stable multimeric complex.
Purification of MotA.
(A) Elution profile of purified MotA protein with 0.1% (w/v) DDM by Superdex 200 Increase 10/300 GL gel filtration chromatography. The upper panel shows the chromatogram obtained using samples collected just after purification by His-affinity beads and the lower panel shows a chromatogram obtained for samples from the fraction collected during the first run, whose elution volume was approximately 12 mL. (B) Results for gel electrophoresis obtained for the MotA protein purified in (A).
MotA alone can form at least a tetramer
To estimate the stoichiometry of MotA in the purified complex, we performed a chemical crosslinking experiment (Fig. 3). Purified protein was crosslinked by either BS3, a homo-bifunctional crosslinker that forms a bridge between amino groups, or EDC, a hetero-bifunctional crosslinker that forms a bridge between carboxyl and amino groups. Crosslinked products were analyzed by SDS-PAGE and immunoblotting. Without the chemical crosslink treatments, there was a clear single band with a molecular mass of ca. 25 kDa, which is in good agreement with that of monomeric His-MotA (28 kDa) (Figs 2B and 3A). Upon increasing the BS3 concentration, a band with a molecular mass of ca. 50 kDa appeared (Fig. 3A), probably corresponding to the MotA dimer. A similar result was observed with EDC, but with additional bands at ca. 75 kDa and ca. 100 kDa, corresponding to the MotA trimer and tetramer (Fig. 3B). Thus, purified MotA was found to form a multimeric complex composed of four or more MotA molecules.
Surface accessibility of amino acid residues in MotA
To examine the surface accessibility of amino acid residues of the MotA protein, cysteine substitutions were introduced into MotA (Fig. 4). We examined the motor function of the cysteine substitution mutants in E. coli using a hybrid mutant motor consisting of A. aeolicus MotA and chimeric MotB composed of A. aeolicus MotB and E. coli MotB24. The cysteine substitution mutants were functional except for R88C and E96C, which are mutations of conserved charged residues important for the interaction with the rotor protein FliG (Fig. 4B)4,25. The mutant MotA proteins were purified and treated with biotin maleimide and the bound biotin was detected using streptavidin-conjugated horseradish peroxidase (HRP). MotA-S85C, R88C and D121C were strongly labeled by biotin maleimide, MotA-E96C was moderately labeled, but MotA-F29C and G180C were not labeled at all. These results indicate that residues S85, R88, E96 and D121 are exposed at the surface of the MotA structure, whereas residues F29 or G180 may be buried inside the MotA multimeric complex.
Cys substitutions of MotA.
(A) Schematic representation of MotA displaying the mutated residues. (B) Motility assay of E. coli cells producing MotA (with or without mutation) and chimeric MotB of A. aeolicus in soft-agar plates. Overnight cultures were spotted on TB-0.25% agar plate containing 0.02% arabinose and 0.5 mM IPTG and incubated at 30 °C for 26 h. (C) Surface accessibility of mutated residues in MotA. Cys residues in mutant MotA protein were labeled with biotin-maleimide and detected with streptavidin-conjugated HRP.
Single particle image analysis of purified MotA
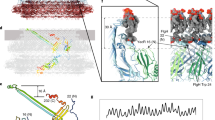
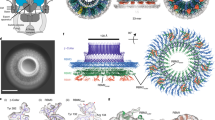
The 3D structure of the MotA multimeric complex purified from the 210 kDa fraction was analyzed by single particle EM image analysis. The sample was negatively stained with uranyl acetate, electron micrographs were recorded and a large number of single particle images were collected and analyzed by 3D image reconstruction, as described in the Methods section (Fig. S4). The stoichiometry of the MotA/MotB complex in E. coli has been estimated to be four molecules of MotA and two molecules of MotB10 and the 2-fold rotational symmetry of the PomA/PomB complex has been observed by electron tomography19. Consequently, 2-fold rotational symmetry was applied for single particle image analysis. The 3D image of the MotA complex was reconstructed at a 25 Å resolution (Fig. 5). The structure of the MotA complex is composed of two parts: a slightly elongated globular part (upper part); and a pair of arch-like domains with spiky projections (lower part). MotA is composed of a four-helix transmembrane domain and a relatively large cytoplasmic domain comprising 144 residues (residues 54–149 and 205–254). The height of the upper globular part is about 50 Å, which is equivalent to the thickness of the lipid bilayer, suggesting that this globular part corresponds to the transmembrane region of MotA. An atomic model of the transmembrane domain, which was generated by molecular dynamics simulation13, showed that the MotA tetramer complex fitted well into this region, taking into account that this compartment contains detergent molecules surrounding the transmembrane helices, forming a micelle structure (Fig. 5). The remaining arch-like domains should thus contain the cytoplasmic region of MotA. The total volume of the paired arch-like domains in the 3D map is approximately 84,000 Å3, which corresponds to a molecular mass of 68 kDa, assuming a protein density of 1.35 g/cm3. This is in good agreement with the molecular mass of 63 kDa calculated for the 576 residues, four times larger than the cytoplasmic 144 residues of MotA. Consequently, the upper and lower parts are likely to correspond to the transmembrane and cytoplasmic domains, respectively, as indicated in Fig. 5.
The 3D density map of the MotA multimeric complex by single particle EM image analysis.
(A) A side view. (B) Another side view, with a 90° rotation compared to the one shown in A. The yellow area labeled TM indicates the membrane regions. (C) A view from the cytoplasmic side. (D) A view from the periplasmic side. An atomic model of the transmembrane region of MotA tetramer complex13 was fitted into the transmembrane domain in A,B and D. Four MotA molecules are shown in Cα ribbon representation, each colored in cyan, red, brown and magenta.
Discussion
The stator is one of the most important parts for the proper functioning of the bacterial flagellar motor and is believed to work as an energy-converting unit that transduces electrochemical potential gradient across the cytoplasmic membrane into mechanical force. The interaction surfaces of the stator and the rotor have been well studied by mutational analyses4,26,27. However, the mechanism governing energy conversion remains unknown because of the lack of structural information on the stator.
Here, we cloned the stator genes from various bacteria and reproduced their expression in the E. coli system. Their patterns of expression, observed in the bands in the denaturing gel, were variable, which might be indicative of variations in their molecular sizes and structural stabilities (Fig. S2). Some of them were functional stators within the flagellar motor of E. coli or V. alginolyticus. A sequence similarity of more than 60% and 90% from native MotA (PomA) and MotB (PomB), respectively, might be required to provide the functional compensation (Table S2).
We overproduced and purified the stator proteins of a thermophilic bacterium, A. aeolicus, from the E. coli expression system. These stator proteins were chosen because of their observed stabilities. Purified MotA formed multimeric complexes and is likely to be composed of four MotA molecules. To date, MotA has been thought to form a tetramer through its interaction with the MotB dimer, located at the center of the transmembrane region of the MotA4MotB2 complex. However, the present study shows that MotA can form a tetramer even in the absence of MotB. Such robust complex formation might be important for the assembly and function of the stator in the motor. A previous study on the V. alginolyticus PomA protein showed that PomA formed a dimer instead of a tetramer11. This variation might be due to differences in detergents used to solubilize the protein from the membrane and discrepancies in protein stability. In the chemical crosslinking experiment, a strong band corresponding to the dimer was clearly observed, whereas those corresponding to the trimer and tetramer were weak (Fig. 3). It has been thought that four MotA molecules form a pair of dimers within one stator unit. It is possible that the interactions between MotA molecules within the dimer structure are stronger than those between dimers forming the tetrameric unit.
In the experiment aimed at probing surface accessibility, the cytoplasmic amino acid residues (S85C, R88C, E96C and D121C) were well labeled, whereas the periplasmic residues (F29C and G180C) were not (Fig. 4). In a previous study on the V. alginolyticus PomA protein, the periplasmic residues (F29C and G176C), which correspond to F29C and G180C of A. aeolicus, respectively, were treated with biotin maleimide in living cells28. G176C was labeled, unlike F29C. It was speculated that the F29 residue might be embedded in the pore region of the PomA complex, preventing binding with biotin maleimide. Furthermore, because we used detergent-solubilized MotA in this study, the detergent might have inhibited the reaction between biotin maleimide and the periplasmic residues in the vicinity of the membrane-spanning region. The globular shape of the MotA-complex transmembrane region was visualized by 3D EM image reconstruction (Fig. 5), showing that this region presumably contains a detergent micelle surrounding the transmembrane helices of the four MotA molecules.
Here, we report the first 3D structure of the MotA stator complex formed without MotB (Fig. 5). The shape of this complex is quite different from the corresponding PomA/PomB complex of V. alginolyticus, reported previously19. Although the resolution is still limited to 25 Å, our present density map clearly shows, for the first time, the shape of the cytoplasmic domain of the stator complex, which is distinctly separated from the transmembrane domain. Because this cytoplasmic domain is solely composed of the cytoplasmic regions of MotA, we measured its volume, estimated the corresponding molecular mass and found it to be in good agreement with that of the cytoplasmic regions of MotA. An atomic model of the transmembrane domain of the MotA tetramer complex, generated by molecular dynamics simulation13, also fitted well into the density map when taking into account the detergent micelle structure surrounding the transmembrane helices (Fig. 5). We are therefore convinced that our 3D density map reflects the actual structure of the MotA tetramer complex.
Until now, various models have been proposed for the molecular mechanism of the stator’s function. Among these is a power-stroke model29,30,31. In this model, the cytoplasmic region of the stator is thought to change its conformation under the action of an ion influx through its channel. Because the MotA complex we purified in this study does not contain MotB, the stator channel was not properly formed and consequently, the structure of the cytoplasmic domain might have been locked in a single conformation. However, the structure of the cytoplasmic domain should dynamically change under the coupled action of the ion influx through the stator channel and the stator-rotor interaction within the functional flagellar motor of living cells.
Methods
Preparation of membrane fractions containing MotAAa
The full-length motA gene from A. aeolicus (motAAa) was cloned into a pColdI vector (Takara Bio Inc., Kusatsu, Japan) as previously described24. MotAAa protein was expressed with an N-terminal histidine tag in E. coli BL21-CodonPlus(DE3)-RIPL (Agilent Technologies, Santa Clara, CA, USA) cells. The cells were grown in 1.5 L of SB medium [1.2% (w/v) Bacto tryptone, 2.4% (w/v) Bacto yeast extract, 0.5% (w/v) glycerol, 1.25% (w/v) K2HPO4, 0.38% (w/v) KH2PO4] containing 100 μg/mL ampicillin, at 37 °C, to an OD660 of 0.6–0.8 and 0.5 mM isopropyl-β-d-thiogalactopyranoside (IPTG) was subsequently added to the culture after cooling on ice for 30 min before the culture was prolonged for about 20 h at 16 °C. Cells were collected by centrifugation (7,000 × g) and then resuspended in 50 mL of TN buffer [50 mM Tris-HCl (pH 8.0], 200 mM NaCl) containing half a protease inhibitor cocktail tablet (Roche diagnostics) and about 10 mg of lysozyme (Wako, Osaka, Japan). Cells were then disrupted by sonication and ultracentrifuged at 100,000 × g for 30 min and the pellet (membrane fraction) resuspended in TN buffer.
Solubilization of MotAAa in detergent
The membrane fraction containing the His-MotAAa protein was solubilized in 1% (w/v) of CHAPS (Dojindo, Kamimashiki, Japan), Cymal-5 (Anatrace, Maumee, OH, USA), n-dodecyl β-d-maltoside (DDM) (Dojindo), n-decyl β-d-maltoside (DM) (Dojindo), n-dodecylphosphocholine (DPC) (Anatrace), n-octyl β-d-glucoside (OG) (Dojindo), sucrose monocaprate (SMC) (Dojindo), Triton-X100 (Wako) or Tween-20 (Wako) for about 30 min on ice. The detergent solubilized membrane sample was ultra-centrifuged at 100 000 × g for 30 min and separated into pellet (insoluble fraction) and supernatant (soluble fraction). The proteins were resolved by SDS-PAGE and subsequently detected by CBB staining and immunoblot analysis, using the anti-His antibody (MBL, Nagoya, Japan).
Purification of MotAAa
The membrane fraction containing the His-MotAAa protein was solubilized by 1% (w/v) Cymal-5 or DM for about 30 min on ice. The detergent-solubilized membrane sample was ultracentrifuged at 100,000 × g for 30 min and the supernatant was mixed with 3 mL of Ni-NTA agarose and then incubated on ice for 1 h. Protein-bound agarose was washed with TN buffer containing 0.1% (w/v) DDM or DM and 50 mM imidazole and subsequently, proteins were eluted with TN buffer containing 0.1% (w/v) DDM or DM and 500 mM imidazole. The proteins were concentrated using an Amicon Ultra 100 K device (Merck Millipore, Darmstadt, Germany) and His-MotAAa was purified using gel-filtration chromatography [Superdex 200 Increase 10/300 GL (GE Healthcare, Little Chalfont, UK)] with either TN or PN buffer [50 mM NaPO4 (pH 7.0), 150 mM NaCl] containing 0.1% (w/v) DDM or DM.
Chemical crosslinking
For the reaction with BS3 (Thermo Fisher Scientific, Waltham, MA, USA), about 0.5 mg/mL of purified His-MotAAa protein in PN buffer containing 0.1% (w/v) DM was incubated at 20 °C with 0, 0.2, 0.5, 1, 2 or 5 mM BS3, for 5 min. For the reaction with EDC (Thermo Fisher Scientific), about 0.5 mg/mL of purified His-MotAAa protein in PN buffer containing 0.1% (w/v) DM was incubated at 20 °C with 20 mM EDC for 0, 5, 15, 30, 60 or 120 min. The reactions were quenched by adding 200 mM Tris-HCl (pH 8.0), followed by a 15-min incubation step. The proteins contained in the reaction mixture were separated by SDS-PAGE and detected by CBB staining. The proteins separated by SDS-PAGE were also transferred to PVDF membranes and immunoblotting was performed using the anti-MotAAa antibody.
Introduction of mutations
Point mutations in motAAa gene on the plasmid were introduced by performing site-directed mutagenesis (Agilent Technologies). The mutations were confirmed by DNA sequencing.
Motility assay in soft-agar plates
Motility assays in soft agar plates were performed as previously described24. Two microliter of an overnight culture of RP6894 (E. coli ΔmotAB) cells harboring the plasmids pBAD24-motAAa-A225D (with or without mutations) and pSBETa-motB2AE, which expressed chimeric MotA/B complex function in E. coli24, were spotted on TB soft-agar plates [1% (w/v) Bacto tryptone, 0.5% (w/v) NaCl, 0.25% (w/v) Bacto agar] with 0.02% (w/v) arabinose and 0.5 mM IPTG and were incubated at 30 °C for 26 h.
Cysteine labeling by biotin maleimide
The membrane fraction of E. coli cells expressing the His-MotAAa protein (with or without Cys substitution mutation) and suspended in TN buffer was solubilized in 1% (w/v) DM, treated with 0.9 mM biotin-maleimide (Sigma-Aldrich, St. Louis, MO, USA) and incubated at 24 °C for 1 h. The biotin maleimide-treated His-MotAAa was then purified using Ni-NTA agarose and His-MotAAa proteins and biotin-labeled His-MotAAa proteins were checked using SDS-PAGE and CBB staining, followed by blotting analysis using streptavidin-conjugated horseradish peroxidase (HRP) (GE Healthcare).
Electron microscopy and image analysis
A 3-μL solution of MotA was applied on a glow-discharged continuous carbon grid. The excess solution was removed using a filter paper and the sample was subsequently stained on the carbon grid with 2% uranyl acetate. Electron microscopy images were recorded with a JEM-3200FSC transmission electron microscope (JEOL, Akishima, Japan) operated at 200 kV and equipped with a F415MP slow-scan CCD camera (TVIPS, Gauting, Germany) at a nominal magnification of 40,000x. About 2,000 particle images were selected manually from 50 electron micrographs and classified into 20 classes using Relion-1.4 (MRC, LMB, Cambridge, UK) and reference-free classification. The best class image in 20 class averages was used as a reference image for automated particle picking and in total, 54,532 particle images were selected. To reduce the computation time required for analysis, these images were separated into three groups and reference-free classification was carried out for each group separately. Finally, 5,419 particle images were selected from 150 class averages and these were used for 3D image reconstruction. The initial structure for the 3D image reconstruction was determined from six class averages by e2initialmodel.py in the EMAN2 package32. The final 3D structure was reconstructed using Relion-1.4.
Additional Information
Accession code: The density map of MotA has been deposited in the Electron Microscopy Data Bank, http://www.emdatabank.org/ (accession code: EMD-3417).
How to cite this article: Takekawa, N. et al. The tetrameric MotA complex as the core of the flagellar motor stator from hyperthermophilic bacterium. Sci. Rep. 6, 31526; doi: 10.1038/srep31526 (2016).
References
Kojima, S. Dynamism and regulation of the stator, the energy conversion complex of the bacterial flagellar motor. Curr. Opin. Microbiol. 28, 66–71 (2015).
Minamino, T. & Imada, K. The bacterial flagellar motor and its structural diversity. Trends Microbiol. 23, 267–274 (2015).
Zhou, J. D., Fazzio, R. T. & Blair, D. F. Membrane topology of the MotA protein of Escherichia coli. J. Mol. Biol. 251, 237–242 (1995).
Zhou, J. D., Lloyd, S. A. & Blair D. F. Electrostatic interactions between rotor and stator in the bacterial flagellar motor. Proc. Natl. Acad. Sci. USA. 95, 6436–6441 (1998).
De, Mot, R. & Vanderleyden, J. The C-terminal sequence conservation between OmpA-related outer membrane proteins and MotB suggests a common function in both gram-positive and gram-negative bacteria, possibly in the interaction of these domains with peptidoglycan. Mol. Microbiol. 12, 333–334 (1994).
Chun, S. Y. & Parkinson, J. S. Bacterial motility: membrane topology of the Escherichia coli MotB protein. Science 239, 276–278 (1988).
Hosking, E. R., Vogt, C., Bakker, E. P. & Manson M. D. The Escherichia coli MotAB proton channel unplugged. J. Mol. Biol. 364, 921–937 (2006).
Braun, T. F., Al-Mawsawi, L. Q., Kojima, S. & Blair, D. F. Arrangement of core membrane segments in the MotA/MotB proton-channel complex of Escherichia coli. Biochemistry 43, 35–45 (2004).
Braun, T. F. & Blair, D. F. Targeted disulfide cross-linking of the MotB protein of Escherichia coli: evidence for two H+ channels in the stator complex. Biochemistry 40, 13051–13059 (2001).
Kojima, S. & Blair, D. F. Solubilization and purification of the MotA/MotB complex of Escherichia coli. Biochemistry 43, 26–34 (2004).
Sato, K. & Homma, M. Multimeric structure of PomA, the Na+ -driven polar flagellar motor component of Vibrio alginolyticus. J. Biol. Chem. 275, 20223–20228 (2000).
Sato, K. & Homma, M. Functional reconstitution of the Na+ -driven polar flagellar motor component of Vibrio alginolyticus. J. Biol. Chem. 275, 5718–5722 (2000).
Nishihara, Y. & Kitao, A. Gate-controlled proton diffusion and protonation-induced ratchet motion in the stator of the bacterial flagellar motor. Proc. Natl. Acad. Sci. USA. 112, 7737–7742 (2015).
Kojima, S. et al. Stator assembly and activation mechanism of the flagellar motor by the periplasmic region of MotB. Mol. Microbiol. 73, 710–718 (2009).
Roujeinikova, A. Crystal structure of the cell wall anchor domain of MotB, a stator component of the bacterial flagellar motor: implications for peptidoglycan recognition. Proc. Natl. Acad. Sci. USA. 105, 10348–10353 (2008).
Zhu, S. et al. Conformational change in the periplamic region of the flagellar stator coupled with the assembly around the rotor. Proc. Natl. Acad. Sci. USA. 111, 13523–13528 (2015).
Hizukuri, Y., Morton, J. F., Yakushi, T., Kojima, S. & Homma, M. The peptidoglycan-binding (PGB) domain of the Escherichia coli pal protein can also function as the PGB domain in E. coli flagellar motor protein MotB. J. Biochem. 146, 219–229 (2009).
Kojima, S. et al. Insights into the stator assembly of the Vibrio flagellar motor from the crystal structure of MotY. Proc. Natl. Acad. Sci. USA 105, 7696–7701 (2008).
Yonekura, K., Maki-Yonekura, S. & Homma, M. The structure of the flagellar motor protein complex PomAB: Implications for the torque-generating conformation. J. Bacteriol. 193, 3863–3870 (2011).
Lee, L. K., Ginsburg, M. A., Crovace, C., Donohoe, M. & Stock, D. Structure of the torque ring of the flagellar motor and the molecular basis for rotational switching. Nature 466, 996–1000 (2010).
Suzuki, H., Yonekura, K. & Namba, K. Structure of the rotor of the bacterial flagellar motor revealed by electron cryomicroscopy and single-particle image analysis. J. Mol. Biol. 337, 105–113 (2004).
Thomas, D. R., Morgan, D. G. & DeRosier, D. J. Rotational symmetry of the C ring and a mechanism for the flagellar rotary motor. Proc. Natl. Acad. Sci. USA. 96, 10134–10139 (1999).
Thomas, D. R., Francis, N. R., Xu, C. & DeRosier, D. J. The three-dimensional structure of the flagellar rotor from a clockwise-locked mutant of Salmonella enterica serovar Typhimurium. J. Bacteriol. 188, 7039–7048 (2006).
Takekawa, N. et al. Sodium-driven energy conversion for flagellar rotation of the earliest divergent hyperthermophilic bacterium. Sci. Rep. 5, 12711 (2015).
Zhou, J. D. & Blair, D. F. Residues of the cytoplasmic domain of MotA essential for torque generation in the bacterial flagellar motor. J. Mol. Biol. 273, 428–439 (1997).
Morimoto, Y. V., Nakamura, S., Hiraoka, K. D., Namba, K. & Minamino, T. Distinct roles of highly conserved charged residues at the MotA-FliG interface in bacterial flagellar motor rotation. J. Bacteriol. 195, 474–481 (2013).
Takekawa, N., Kojima, S. & Homma, M. Contribution of many charged residues at the stator-rotor interface of the Na+ -driven flagellar motor to torque generation in Vibrio alginolyticus. J. Bacteriol. 196, 1377–1385 (2014).
Asai, Y., Shoji, T., Kawagishi, I. & Homma, M. Cystein-scanning mutagenesis of the periplasmic loop regions of PomA, a putative channel component of the sodium-driven flagellar motor in Vibrio alginolyticus. J. Bacteriol. 182, 1001–1007 (2000).
Berry, R. M. & Berg, H. C. Torque generated by the flagellar motor of Escherichia coli while driven backward. Biophys. J. 31, 580–587 (1999).
Mandadapu, K. K., Nirody, J., A., Berry, R. M. & Oster, G. Mechanics of torque generation in the bacterial flagellar motor. Proc. Natl. Acad. Sci. USA. 112, E4381–4389 (2015).
Kojima, S. & Blair, D. F. Conformational change in the stator of the bacterial flagellar motor. Biochemistry 40, 13041–13050 (2001).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Acknowledgements
We thank Dr. Chiaki Kato for providing chromosomal DNA of P. profundum, S. benthica and S. violacea. We thank Akio Kitao for providing the model of the transmembrane region of the MotA tetramer. This work was supported in part by Grants-in-Aid for scientific research from the Japan Society for the Promotion of Science (24117004 and 23247024 to M.H., 26840035 to N.Te., 26650021, 24117001, 24117004 to T.K. and 25000013 to K.N.), by Platform for Drug Discovery, Informatics and Structural Life Science from Japan Agency for Medical Research and Development (to A.H. and T.S.) and from the Japan Society for the Promotion of Science (13J02161 to N.Ta.). N.Ta. was partly supported by the Integrative Graduate Education and Research Program in Green Natural Sciences of Nagoya University.
Author information
Authors and Affiliations
Contributions
N. Takekawa, T.K., K.N. and M.H. designed the experiments; N. Takekawa, N. Terahara, M.G., K.M., A.H., Y.O., T.S. and T.K. performed the experiments; N. Takekawa, N. Terahara, T.K., S.K., T.S, K.N. and M.H. analyzed the data; and N. Takekawa, N. Terahara, T.K., K.N. and M.H. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Takekawa, N., Terahara, N., Kato, T. et al. The tetrameric MotA complex as the core of the flagellar motor stator from hyperthermophilic bacterium. Sci Rep 6, 31526 (2016). https://doi.org/10.1038/srep31526
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep31526
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.