Abstract
Crimean Congo hemorrhagic fever virus (CCHFV) is a highly virulent tick-borne pathogen that causes hemorrhagic fever in humans. The geographic range of human CCHF cases largely reflects the presence of ticks. However, highly similar CCHFV lineages occur in geographically distant regions. Tick-infested migratory birds have been suggested, but not confirmed, to contribute to the dispersal. Bats have recently been shown to carry nairoviruses distinct from CCHFV. In order to assess the presence of CCHFV in a wide range of bat species over a wide geographic range, we analyzed 1,135 sera from 16 different bat species collected in Congo, Gabon, Ghana, Germany, and Panama. Using a CCHFV glycoprotein-based indirect immunofluorescence test (IIFT), we identified reactive antibodies in 10.0% (114/1,135) of tested bats, pertaining to 12/16 tested species. Depending on the species, 3.6%–42.9% of cave-dwelling bats and 0.6%–7.1% of foliage-living bats were seropositive (two-tailed t-test, p = 0.0447 cave versus foliage). 11/30 IIFT-reactive sera from 10 different African bat species had neutralizing activity in a virus-like particle assay. Neutralization of full CCHFV was confirmed in 5 of 7 sera. Widespread infection of cave-dwelling bats may indicate a role for bats in the life cycle and geographic dispersal of CCHFV.
Similar content being viewed by others
Introduction
Crimean Congo hemorrhagic fever virus (CCHFV; Genus Nairovirus, Family Bunyaviridae) is a tick-borne pathogen that has caused more than 10,000 documented human infections worldwide, with a case fatality proportion of approximately 10–30%1,2. Clinical symptoms range from acute febrile illness to severe hemorrhagic fever1. To date, the majority of CCHFV cases have been documented in Asia (in particular Russia, Iran, Georgia, Turkey) and Southeast Europe (Greece)1,2. In Africa, approximately 100 human CCHFV cases have been reported, primarily in Mauritania and South Africa2,3,4. Serological surveys on animals, however, suggest enzootic circulation of CCHFV in different parts of Africa5,6. The Americas have not reported any autochthonous CCHFV cases1,2.
Within the genus Nairovirus CCHFV represents one of at least seven serogroups. Nairoviruses, classified in serogroups Hughes, Dera Ghazi Khan and Sakhalin, are associated with birds. Viruses in the serogroups Thiafora and Qalyub were identified in shrews and rodents, respectively. Serogroups CCHFV as well as Nairobi sheep disease virus (NSDV) are associated with ungulates2,7. Only CCHFV, NSDV and the NSDV-related Dugbe virus1 are considered human pathogens, while all other nairoviruses are restricted to certain vertebrate host taxa7.
CCHFV is known to be transmitted by ticks of the genera Hyalomma and Rhipicephalus2,6,8. The geographic distribution of CCHFV coincides well with the presence of ticks9. However, closely related CCHFV lineages occur in geographically distant regions that do not follow the habitat connectivity of tick hosts. Several studies suggested that tick-infested migratory birds may be responsible for the spatial distribution of CCHFV9,10,11.
Bats (Order: Chiroptera) represent the only migratory flying mammals. Bats are commonly infested with soft and hard ticks12,13, and were recently shown to carry nairoviruses7,14,15. Phylogenetic analyses as well as serological studies based on cross-reactivity in complement fixation tests suggest that all identified bat-associated nairoviruses belong to two novel serogroups that are distantly related to CCHFV7. One representative, termed Leopards Hill virus (LPHV), was successfully isolated from samples of the African bat species Hipposideros gigas (H. gigas). Phenotypic characterization revealed that LPHV causes hemorrhagic disease in mice15. Whether LPHV may be pathogenic for other mammals including humans is unknown.
The large diversity of viruses in bats16,17,18,19,20 together with previous findings of bat nairoviruses7,14,15 encourage a wider and more systematic assessment of the endemicity of nairoviruses in bats. Previous studies were predominantly based on nucleic acid detection and virus isolation, which can only detect viraemic animals. While we do not know the expected frequency of nairovirus viraemia in free-ranging bats, field studies on other RNA viruses with a viraemic infection pattern, including hepadnaviruses19, hepaciviruses17, filoviruses21,22,23, and bunyaviruses24,25, indicate very low virus detection rates. Unless very large sample collections are studied, virus detection by PCR lacks the power to reflect the host species range of a given virus, in particular because of a lack of negative predictive value17,18. Serological techniques are better suited for host range studies because seroprevalence is less sensitive to temporal and spatial variation of infection activity26. Serology is also more suitable for the limited sample size that can be achieved in bat-related studies, as most bat species are too small to be bled without destruction.
In the present study we screened 1,135 bat serum samples comprising 16 bat species (migratory and non-migratory) from five different countries for the presence of CCHF-like viruses. The sample was focused on African bat species as nairoviruses have been predominantly detected in the Old World bats14,15,16. We applied a staged set of serological assays including a recombinant glycoprotein (GP)-based indirect immunofluorescence test (IIFT), a novel pseudotype neutralization assay27 as well as full virus neutralization tests conducted under biosafety level 4 conditions. Additional molecular screening involved two modified generic nairovirus RT-PCRs based on previously established protocols28,29,30.
Results
Bats carry CCHFV GP-reactive antibodies
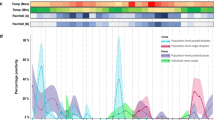
From 2003 through 2011 bats were sampled in Congo, Gabon, Ghana, Germany and Panama. The sample included 1,135 blood or serum specimens from 16 bat species pertaining to six different bat families with variable habitat preference and diet. To assess if bats harbor CCHF-like viruses, we screened bat serum samples for CCHFV GP-reactive antibodies by IIFT. In total, 114 of 1,135 (10.0%) sera from 12 of 16 bat species sampled between 2005 and 2009 in 4/5 countries reacted with recombinant CCHFV GP antigen (range 1.0–57.6%, Table 1, Fig. 1). An example of a reactive bat serum sample is shown in Fig. 2A. IIFT-positive detections were predominantly identified in cave-dwelling, migratory bats including the frugivorous species Rousettus aegyptiacus (24.4%; 48/197) as well as the insectivorous species Coleura afra (42.9%; 6/14), Hipposideros cf. caffer (6.3%; 3/48), Miniopterus inflatus (17.6%; 9/51) and Hipposideros gigas (24.8%; 32/129) from Congo and Gabon (Fig. 1). CCHFV seropositivity was significantly elevated for bat species that roost in caves versus those that roost in trees (Fig. 1; Supplementary Table 1; two-tailed t-test, p = 0.0447). In particular, bat species from the Batouala cave in Gabon, sampled in 2009, had highest seropositivity (range 18.4–57.6%; Table 1, Supplementary Figure). Age (p = 0.7434), dietary (p = 0.4622), gender (p = 0.4613), migration (p = 0.4788) and seasonality (p = 0.1605) were not associated with differences in seroprevalence (Supplementary Table 1). No antibodies were found in (n = 43) sera from New World bats, corresponding to the notion that CCHFV is an Old World virus2.
All bat serum samples were analyzed for CCHFV glycoprotein (GP)-reactive antibodies by a modified commercial IIFT (EUROIMMUN). Sera were tested at a dilution of 1:40. Secondary detection was performed with goat-anti bat immunoglobulin G (IgG, 1:1,000) followed by DyLight488–labeled donkey-anti goat Ig (1:100). Gray bars represent frugivorous bat species whereas black bars show insectivorous bat species. A two-tailed t-test was used to compare seropositivity of cave-dwelling versus foliage living bat species (p = 0.0447). Abbreviations: PAN, Panama, GHA, Ghana, GAB, Gabon, COG, Congo, GER, Germany. A. jam. = Artibeus jamaicensis, A. lit. = Artibeus lituratus, E. h. = Eidolon helvum, Ep. f. = Epomops franqueti, Hy. m. = Hypsignathus monstrosus, Mic. p. = Micropteropus pusillus, My. t. = Myonycteris torquata, R. a. = Rousettus aegyptiacus, C. a. = Coleura afra, H. c. = Hipposideros cf. caffer, H. g. = Hipposideros gigas, Mi. i. = Miniopterus inflatus, R. spe. = Rhinolophus species, M. das. = Myotis dasycneme, M. dau. = Myotis daubentonii, N. noc. = Nyctalus noctula.
(A) Endpoint antibody titration of a CCHFV-reactive bat serum sample by IIFT. A serum sample of Epomops franqueti (sample code GB2497) was applied in four different dilutions (range 1:40–1:1,280). Scale bar represents 20 μm. (B) Determination of anti-CCHFV neutralizing activity by VLP-based NT. CCHF VLPs were incubated for 1 h at 37 °C with antisera at a final dilution of 1:100. Cells were infected with serum-treated VLPs for 1 h at 37 °C. At 24 h post infection, luciferase activity was measured in cell lysates using the DLA kit. All CCHF VLP-derived Renilla values were normalized to an applied transfection control (Firefly luciferase values). An untreated CCHF VLP sample (without serum) was used as a negative control (−) and was set to 100% to calculate the relative percentage of luminescence activity. A reduction ≥80% of the normalized Renilla luciferase activity (dashed line) was considered as a significant VLP neutralizing activity (positive samples marked by black square). A convalescent mouse-anti CCHFV reference serum (+) was used as a positive control. Mean values and standard deviations were calculated from three independent experiments. Abbreviations: Negative control (−), Positive control (−), numbers 1–30 represent CCHFV IIFT positive bat sera: 1–4 (Coleura afra), 5–8 (Epomops franqueti), 9 (Eidolon helvum), 10–12 (Hipposideros cf. caffer), 13–16 (Hipposideros gigas), 17 (Hypsignathus monstrosus), 18–21 (Miniopterus inflatus), 22–26 (Micropteropus pusillus), 24–26 (Myonycteris torquata), 27–30 (Rousettus aegyptiacus). Numbers 31–40 show CCHFV IIFT negative sera: 31 (Coleura afra), 32 (Epomops franqueti), 33 (Eidolon helvum), 34 (Hipposideros cf. caffer), 35 (Hipposideros gigas), 36 (Hypsignathus monstrosus), 37 (Miniopterus inflatus), 38 (Micropteropus pusillus), 39 (Myonycteris torquata), 40 (Rousettus aegyptiacus). Detailed information is provided in Supplementary Table 2.
Detection of CCHFV neutralizing antibodies in African bat species
As antibody cross-reactivity between CCHFV and viruses from the related NSDV serogroup cannot be completely ruled out by IIFT31, specific virus neutralization tests (NT) were done. NT can prove previous infection with CCHFV because there is no cross-neutralization between serogroups31. Because NTs require relatively large volumes of serum that cannot be obtained from most bat species, only for 30 of the 114 IIFT-positive sera covering 10/12 bat species, NT assays could be performed. In addition, 10 CCHFV IIFT-negative samples with sufficient volume, representing all 10 assessed bat species, were tested. Sera were tested at a 1:100 dilution in a 96-well format using a recently established CCHF virus-like particle (VLP)-based NT27. Out of 30 IIFT-positive sera, 11 showed significant neutralizing activity defined as 80% reduction of luciferase luminescence signal (Fig. 2B, Supplementary Table 2). None of the 10 IIFT-negative control sera had neutralizing activity (Fig. 2B). In parallel, all 40 (30 IIFT-positive, 10 IIFT-negative) bat sera were tested in a Rift Valley fever (RVF) VLP-based NT32, all with negative results, ruling out nonspecific neutralization activity in bat sera (Fig. 3, Supplementary Table 2).
To rule out unspecific neutralizing activity of bat serum samples (the same CCHFV IIFT-positive and negative sera as in Fig. 2) RVF VLPs were used with the same serum dilution of 1:100. Incubations and read-outs were performed as described in Fig. 2. A reduction ≥80% of the specific luciferase activity (dashed line) was considered as VLP neutralisation. A convalescent mouse anti-RVFV reference serum was used as positive control (+). Mean values and standard deviations were calculated from three independent experiments. Abbreviations: Negative control (−), Positive control (+), numbers 1–30 represent CCHFV IIFT positive bat sera. Numbers 31–40 show CCHFV IIFT negative sera. For detailed information refer to Figure legend of Fig. 2 and Supplementary Table 2.
For 7 of 11 CCHF VLP-positive sera, enough material was available to conduct additional neutralization tests by a full virus CCHFV neutralization test under biosafety level 4 conditions. Endpoint titration by IIFT in these samples revealed high reciprocal titers between 160 and 1,280 (Table 2, Fig. 2A). Sera were thus titrated in 2-fold serial dilutions in a range of 1:40–1:1,280. The test confirmed full virus neutralizing activity in 5 of 7 sera, with reciprocal titers ranging between 40 (lowest testing dilution due to lack of serum) and 160 (Table 2).
Lack of evidence for CCHFV-related nucleotide sequences in bat serum samples
To identify CCHFV-related nucleotide sequences in serum samples, we combined three previously established generic RT-PCR protocols that were shown to detect all known nairoviruses28,29,30. Modifications were made to increase the sensitivity for the detection of CCHFV-related nucleotide sequences by applying low annealing temperatures and degenerated oligonucleotides. Viral RNA of CCHFV strain IbAr was used to test oligonucleotides from Lambert et al.29 in combination with primers from Wölfel and colleagues30. The endpoint RT-PCR detection limit of an RT-PCR formulation that used the best combination of primers was between 8 and 80 copies per reaction. An additional RT-PCR assay was developed based on a hemi-nested formulation using primers described in Honig et al.28 followed by a 2nd round RT-PCR step with novel primers. The sensitivity limit of this assay also ranged between 8 and 80 copies per reaction. Serum samples from Artibeus jamaicensis (n = 28), A. lituratus (n = 15, both from Panama) as well as Myotis dasycneme (n = 26, Germany) were not available for RT-PCR testing due to sample volume limitations (<25 μl). In the remaining 1,067 of 1,135 serologically-tested sera, no CCHFV RNA was found by both RT-PCR formulations. In parallel, these same bat serum sample-derived RNA extracts had successfully been used for the detection of novel paramyxo18-, hepe20-, hepadna19- and flaviviruses17, confirming that the material was appropriate for RNA virus testing.
Discussion
In this assessment of the potential host species range of CCHFV in bats we obtained strong serological evidence for bats constituting a putative host for CCHFV or a closely related virus belonging to the CCHFV serotype. Our failure to directly detect viral RNA may be caused by an overall low infection prevalence that precludes detection in spite of using sensitive RT-PCR assays on a rather large collection of serum samples from bats. We applied two generic pan-nairovirus RT-PCR assays that made use of low stringency amplification conditions to enable the detection of unknown CCHF-related viruses. These conditions led to a maximal 10-fold higher limit of detection (LOD: 8–80 copies/reaction) compared to previously established diagnostic CCHFV real time PCRs with LODs of 5–16 copies per reaction33,34,35,36. However, as CCHFV-infected viraemic humans, for example, have viral loads up to 109 viral RNA copies per ml37, we assume that viraemic animals would have been identified if present. Nonetheless, we cannot rule out that bats may not experience a pronounced viraemia as observed for birds38,39.
In addition, sampling time point and/or type of sample may have prevented amplification. In humans it was shown that viremia can be very short (6 days)37. Although there is, to our knowledge, no data on nairovirus viraemia in bats, we think that the chances to detect CCHFV RNA in bat samples may be very limited. Previous studies conducted by us and others have already shown that the detection frequency of viral RNA is generally very low (<3%)17,19.
As shown by Ishii et al. the bat-related nairovirus LPHV was primarily detected in lung samples15. Therefore, destructive sampling involving the collection of organ tissue of designated bat species may be more promising for nairovirus detection as virus might persist for prolonged periods in parenchymatous organs such as the lung or liver14,15. In the context of this retrospective study we did not test different organs from bats because most species are protected and were sampled without destruction or samples were exhausted from previous experiments40,41.
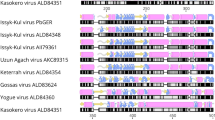
In spite of this limitation we consider the serological results from our study to be indicative of the presence of members of the CCHFV serogroup in bats. The current demarcation criteria for nairovirus taxa have to rely on the definition of serogroups as long as formal species definitions are not in place. Serogroups based on cross-reactivity in antibody affinity assays reflect phylogeny and can be used to classify novel nairoviruses as they are identified42,43. For example, complement fixation assays (CFA) and IIFT were applied to discriminate shrew nairoviruses from CCHFV and classify them into a separate serogroup termed Thiafora44. Phylogenetically, all members of the Thiafora serogroup form a monophyletic clade in sister relation to a clade containing all members of the CCHFV and NSDV serogroups7 (overview shown in Fig. 4). The NSDV serogroup contains NSDV (also named Ganjam virus), and Dugbe virus whose serological classification in one serogroup was again based on cross-reactivity in CFA and IIFT31. NT enables further serological discrimination between taxa in serogroups, exemplified by the presence of cross-reactivity but absence of cross-neutralization between Dugbe and NSDV31. These taxa might be referred to as two different serotypes within the NSDV serogroup (Fig. 4). The whole NSDV serogroup is discriminated from the CCHFV serogroup, its monophyletic sister taxon, based on absence of cross-reactivity in CFA and decreased IIFT endpoint titers31,43,45. Further sub-differentiation into serotypes based on NT has not been observed within the CCHFV serogroup42,46. Consequently, the combined reactivity in IIFT and NT suggests that bat species, tested in the present study, carry viruses which are less distinct from CCHFV than Dugbe virus is from NSDV.
The discrimination of nairovirus serogroups and serotypes is based on cross-reactivity patterns in different serological assays31,42,43. Absence of cross-reactivity in complement fixation assay (CFA) and discriminative endpoint titers in indirect immunofluorescence test (IIFT) are used to subdivide serogroups. A differentiation into serotypes can only be achieved by cross-neutralization tests. The detection of high IIFT endpoint titers and neutralizing antibodies in bat serum samples in the present study suggest that bat-associated CCHF-like viruses belong to the CCHFV serogroup.
Also the height of IIFT titers observed in our study, up to 1:1,280, corresponds to CCHFV-specific titers in experimentally infected wild mammals that reached endpoint titers of 128–1,024 by IIFT47. We conclude that African bat species have been infected with CCHFV or a closely related nairovirus in the same serogroup and probably the same species.
The identification of high neutralizing antibody titers (up to 1:160 based on full virus neutralization) indicates that bats experience infections followed by seroconversion, which in case of humans is correlated with clearance of the virus and survival of infection48. CCHFV might thus exemplify another highly pathogenic agent that is effectively controlled in bats.
As several cave-dwelling bat species migrate over various distances, bats might contribute to the geographic dispersal of CCHFV in similar ways as hypothesized for birds9,10,11. Future studies should aim to clarify whether the CCHFV strains carried by bats or bat ticks are identical or distinct from strains associated with birds or bird ticks.
Interestingly, we found a predominance of CCHFV antibody-positive bats among cave-dwelling species in particular in the Batouala cave in Gabon. The specific habitat conditions in caves (high population density, host availability, humidity, moderate temperature variations) could potentially enable a virus amplification cycle between bats and ticks12,13,49. Caves harbor many different arthropods including soft and hard ticks. Cave-dwelling animals like ruminants, bird and bats are highly exposed to blood-sucking parasites. Whereas in case of humans and ruminants it is known that CCHFV is predominantly transmitted by hard ticks2,6,8 the role of soft ticks in the CCHFV transmission cycle is less clear. However, soft ticks have been found to carry bunyaviruses50 and evidence from CCHFV-endemic countries suggests that certain soft tick species (Ornithodoros lahorensis, Argas reflexus) may as well serve as vectors for CCHFV51,52. Soft ticks are long-lived and blood meals of soft ticks often take only a short period of time (minutes to hours)53. Consequently, bats may not be infested by soft ticks during sampling. The low rate of viremia in bats points to a persistence of the virus in ticks rather than bats, encouraging targeted investigations into the pathogen ecology of CCHFV based on cave-associated ticks that can be studied in proximity to bat roosts.
Methods
Sampling sites, time points and ethics
In total, N = 1,135 bat serum specimens were collected in five countries (Congo, Gabon, Ghana, Germany, Panama) from 2003 through 2011. A detailed summary of sampling time points is shown in Supplementary Table 3. All animals were handled according to national and European legislation for the protection of animals (EU council directive 86/609/EEC). For all individual sampling sites, study protocols including trapping, sampling and testing of animals were approved by the responsible animal ethics committees as detailed below. Maximum efforts were made to leave animals unharmed or to minimize suffering of animals. All bats were caught with mist nets and blood was taken by vein or heart punctures by trained personnel in accordance with the approved guidelines of the respective authorities. Any surgical procedure was performed under sodium pentobarbital/ketamine anesthesia. Sampling, capturing and sample transport were done as described before17,18,19,20 under the following wildlife permits and ethical clearances: Panama (Ethics committee of the Smithsonian Tropical Research Institute (STRI); research permit: STRI2563 (PI VC)- IACUC 100316-1001-18; ethics permit: IACUC 100316-1001-18; export permits: SEX/A-30-11, SEX/A-55-11, SEX/A-81-10, SEX-A-26-10); Ghana (Ministry of Food and Agriculture, Wildlife Division, Forestry Commission, Accra; research permit A04957; ethics permit: CHRPE49/09; export permit: state contract between Ghana and Germany; Congo/Gabon (Ministry of Water and Forest, Libreville; ethics and export permit: 00021/MEFEPA/SG/DGEF/DFC); Germany (Landesamt für Naturschutz und Umwelt (LANUV); ethics permit: LANU 314/5327.74.1.6).
Indirect immunofluorescence test
CCHFV antibody detection was performed by a modified, commercial, indirect IIFT (EUROIMMUN AG, Lübeck, Germany) according to previous protocols18,54. EU90 cells (African green monkey kidney) were transfected with plasmids encoding the CCHFV GP (full length GPC precursor of strain IbAr10200). Sera were applied in a 1:40 dilution. Detection involved goat-anti bat IgG (Bethyl, Montgomery, AL, USA) diluted 1:1,000 and Dylight488 labeled donkey-anti goat IgG (1:100). Analysis was performed with a Motic Immunofluorescence microscope (Zeiss, Jena, Germany).
Neutralization test using CCHFV virus-like particles
Biosafe NT were done using CCHF VLP consisting of CCHFV structural proteins and a Renilla luciferase minigenome unable to produce infectious viral progeny27. In parallel, a pGL3-Firefly construct was applied as transfection control. VLPs were incubated for 1 h at 37 °C with antisera at a final dilution of 1:100. Cells were infected with serum-treated VLPs for 1 h at 37 °C. At 24 h post infection, luciferase activities were measured in cell lysates using the Dual-Luciferase Reporter Assay System (Promega, Mannheim, Germany). The same VLP methodology was described earlier for Rift Valley Fever virus (RVFV) and was used in parallel as a control antigen in this study32.
Neutralization test by CCHFV strain IbAr
Sera reactive against CCHF VLP at dilution 1:100 were confirmed by quantitative CCHFV NT in a biosafety level 4 facility. Serial two-fold dilutions (from 1:40–1:1,280) of serum were incubated with ~1,000 focus-forming units (FFU) of CCHFV (IbAr strain, originally isolated from a Nigerian camel tick5) for 1 h at 37 °C. Vero E6 cells were infected in triplicate with serum-treated CCHFV for 1 h at 37 °C. After 20 h cells were fixed and stained with rabbit polyclonal antibody raised against the CCHFV nucleoprotein and detected by Alexa Fluor 488 donkey anti rabbit conjugate (Thermo Scientific, Braunschweig, Germany).
Viral nucleic acid detection
For nucleic acid detection viral RNA was extracted with the QIAamp viral RNA mini kit (QIAGEN, Hilden, Germany). A combination of three previously established generic nairovirus PCR protocols were applied with modifications28,29,30. Oligonucleotides from Lambert et al.29 were used in multiple combinations with primers described by Wölfel and colleagues30. Tenfold dilution series of viral RNA of CCHFV strain IbAr were used to determine the sensitivity of the PCR assays. Optimal amplification of CCHFV RNA was identified using the oligonucleotide combinations Nairo-F (ATG ATT GC5 AAY AG5 AAY TTY AA) and Nairo-R (ACA GCA RTG 5AT 5GG 5CC CCA YTT, 1st round) and cc1a-c-F (GTG CCA CTG ATG ATG CAC AAA AGG ATT CCA TCT) and Nairo-R (2nd round) in a hemi-nested PCR protocol. The use of the nucleoside inosine is indicated by the number 5. In a second PCR assay, which was based on the Honig et al.28 protocol, the published 1st round of PCR amplification by oligonucleotides Nairo-F and Nairo-R was followed by a newly introduced 2nd round PCR using oligonucleotides Nairo-Fnest (CCA AGA AGY GT5 AGR AGY AAR GT) and Nairo-Rnest (TTG GGC CCC ACT T5G TRT TRT C5C C). The PCR protocol was as follows: 1st round PCR including reverse transcription 50 °C, 20 min, 94 °C 3 min, 10× (94 °C 15 s; annealing for 20 s at decreasing temperature starting at 60 °C to 50 °C; 72 °C 35 s), 40× (94 °C 15 s, 50 °C 20 s, 72 °C, 35 s). 2nd round PCR same condition as the 1st round without reverse transcription.
Statistics
A two-tailed t-test with equal variance (www.openepi.com) was used to correlate CCHFV seroprevalence in different bat species with designated features (age: adult/subadult versus juvenile; dietary: frugivorous versus insectivorous; gender: male versus female; migration: migratory versus resident; roosting: foliage versus cave-living; seasonality: wet versus dry). For all calculations only bat species with a positive CCHFV-IIFT result were included.
Additional Information
How to cite this article: Müller, M. A. et al. Evidence for widespread infection of African bats with Crimean-Congo hemorrhagic fever-like viruses. Sci. Rep. 6, 26637; doi: 10.1038/srep26637 (2016).
References
Ergonul, O. Crimean-Congo haemorrhagic fever. Lancet Infect Dis 6, 203–214, doi: 10.1016/S1473-3099(06)70435-2 (2006).
Bente, D. A. et al. Crimean-Congo hemorrhagic fever: history, epidemiology, pathogenesis, clinical syndrome and genetic diversity. Antiviral Res 100, 159–189, doi: 10.1016/j.antiviral.2013.07.006 (2013).
Swanepoel, R. et al. Crimean-Congo hemorrhagic fever in South Africa. Am J Trop Med Hyg 32, 1407–1415 (1983).
Shepherd, A. J. et al. A nosocomial outbreak of Crimean-Congo haemorrhagic fever at Tygerberg Hospital. Part V. Virological and serological observations. S Afr Med J 68, 733–736 (1985).
Causey, O. R., Kemp, G. E., Madbouly, M. H. & David-West, T. S. Congo virus from domestic livestock, African hedgehog, and arthropods in Nigeria. Am J Trop Med Hyg 19, 846–850 (1970).
Shepherd, A. J., Swanepoel, R., Shepherd, S. P., McGillivray, G. M. & Searle, L. A. Antibody to Crimean-Congo hemorrhagic fever virus in wild mammals from southern Africa. Am J Trop Med Hyg 36, 133–142 (1987).
Walker, P. J. et al. Genomic Characterization of Yogue, Kasokero, Issyk-Kul, Keterah, Gossas, and Thiafora Viruses: Nairoviruses Naturally Infecting Bats, Shrews, and Ticks. Am J Trop Med Hyg, doi: 10.4269/ajtmh.15-0344 (2015).
Shepherd, A. J., Swanepoel, R., Cornel, A. J. & Mathee, O. Experimental studies on the replication and transmission of Crimean-Congo hemorrhagic fever virus in some African tick species. Am J Trop Med Hyg 40, 326–331 (1989).
Mild, M., Simon, M., Albert, J. & Mirazimi, A. Towards an understanding of the migration of Crimean-Congo hemorrhagic fever virus. J Gen Virol 91, 199–207, doi: 10.1099/vir.0.014878-0 (2010).
Leblebicioglu, H. et al. Role of migratory birds in spreading Crimean-Congo hemorrhagic fever, Turkey. Emerg Infect Dis 20, 1331–1334, doi: 10.3201/eid2008.131547 (2014).
Lindeborg, M. et al. Migratory birds, ticks, and Crimean-Congo hemorrhagic fever virus. Emerg Infect Dis 18, 2095–2097, doi: 10.3201/eid1812.120718 (2012).
Klompen, J. S., Black, W. C. t., Keirans, J. E. & Oliver, J. H., Jr. Evolution of ticks. Annual review of entomology 41, 141–161, doi: 10.1146/annurev.en.41.010196.001041 (1996).
Arthur, D. R. The Ixodes ticks of Chiroptera (Ixodoidea, Ixodidae). J Parasitol 42, 180–196 (1956).
Dacheux, L. et al. A preliminary study of viral metagenomics of French bat species in contact with humans: identification of new mammalian viruses. PLoS One 9, e87194, doi: 10.1371/journal.pone.0087194 (2014).
Ishii, A. et al. A nairovirus isolated from African bats causes haemorrhagic gastroenteritis and severe hepatic disease in mice. Nat Commun 5, 5651, doi: 10.1038/ncomms6651 (2014).
Calisher, C. H., Childs, J. E., Field, H. E., Holmes, K. V. & Schountz, T. Bats: important reservoir hosts of emerging viruses. Clin Microbiol Rev 19, 531–545, doi: 10.1128/CMR.00017-06 (2006).
Drexler, J. F. et al. Evidence for novel hepaciviruses in rodents. PLoS Pathog 9, e1003438, doi: 10.1371/journal.ppat.1003438 (2013).
Drexler, J. F. et al. Bats host major mammalian paramyxoviruses. Nat Commun 3, 796, doi: 10.1038/ncomms1796 (2012).
Drexler, J. F. et al. Bats carry pathogenic hepadnaviruses antigenically related to hepatitis B virus and capable of infecting human hepatocytes. Proc Natl Acad Sci USA 110, 16151–16156, doi: 10.1073/pnas.1308049110 (2013).
Drexler, J. F. et al. Bats worldwide carry hepatitis E virus-related viruses that form a putative novel genus within the family Hepeviridae. J Virol 86, 9134–9147, doi: 10.1128/JVI.00800-12 (2012).
Towner, J. S. et al. Marburg virus infection detected in a common African bat. PLoS One 2, e764, doi: 10.1371/journal.pone.0000764 (2007).
Negredo, A. et al. Discovery of an ebolavirus-like filovirus in europe. PLoS Pathog 7, e1002304, doi: 10.1371/journal.ppat.1002304 (2011).
Leroy, E. M. et al. Fruit bats as reservoirs of Ebola virus. Nature 438, 575–576, doi: 10.1038/438575a (2005).
Weiss, S. et al. Hantavirus in bat, Sierra Leone. Emerg Infect Dis 18, 159–161, doi: 10.3201/eid1801.111026 (2012).
Guo, W. P. et al. Phylogeny and origins of hantaviruses harbored by bats, insectivores, and rodents. PLoS Pathog 9, e1003159, doi: 10.1371/journal.ppat.1003159 (2013).
Xu, G. J. et al. Viral immunology. Comprehensive serological profiling of human populations using a synthetic human virome. Science 348, aaa0698, doi: 10.1126/science.aaa0698 (2015).
Devignot, S., Bergeron, E., Nichol, S., Mirazimi, A. & Weber, F. A virus-like particle system identifies the endonuclease domain of Crimean-Congo hemorrhagic fever virus. J Virol 89, 5957–5967, doi: 10.1128/JVI.03691-14 (2015).
Honig, J. E., Osborne, J. C. & Nichol, S. T. The high genetic variation of viruses of the genus Nairovirus reflects the diversity of their predominant tick hosts. Virology 318, 10–16, doi: 10.1016/j.virol.2003.09.021 (2004).
Lambert, A. J. & Lanciotti, R. S. Consensus amplification and novel multiplex sequencing method for S segment species identification of 47 viruses of the Orthobunyavirus, Phlebovirus, and Nairovirus genera of the family Bunyaviridae. J Clin Microbiol 47, 2398–2404, doi: 10.1128/JCM.00182-09 (2009).
Wolfel, R. et al. Low-density macroarray for rapid detection and identification of Crimean-Congo hemorrhagic fever virus. J Clin Microbiol 47, 1025–1030, doi: 10.1128/JCM.01920-08 (2009).
Davies, F. G., Casals, J., Jesset, D. M. & Ochieng, P. The serological relationships of Nairobi sheep disease virus. J Comp Pathol 88, 519–523 (1978).
Habjan, M. et al. Efficient production of Rift Valley fever virus-like particles: The antiviral protein MxA can inhibit primary transcription of bunyaviruses. Virology 385, 400–408, doi: 10.1016/j.virol.2008.12.011 (2009).
Atkinson, B. et al. Development of a real-time RT-PCR assay for the detection of Crimean-Congo hemorrhagic fever virus. Vector Borne Zoonotic Dis 12, 786–793, doi: 10.1089/vbz.2011.0770 (2012).
Jaaskelainen, A. J. et al. Development and evaluation of a real-time RT-qPCR for detection of Crimean-Congo hemorrhagic fever virus representing different genotypes. Vector Borne Zoonotic Dis 14, 870–872, doi: 10.1089/vbz.2014.1577 (2014).
Drosten, C. et al. Rapid detection and quantification of RNA of Ebola and Marburg viruses, Lassa virus, Crimean-Congo hemorrhagic fever virus, Rift Valley fever virus, dengue virus, and yellow fever virus by real-time reverse transcription-PCR. J Clin Microbiol 40, 2323–2330 (2002).
Wolfel, R. et al. Virus detection and monitoring of viral load in Crimean-Congo hemorrhagic fever virus patients. Emerg Infect Dis 13, 1097–1100, doi: 10.3201/eid1307.070068 (2007).
Cevik, M. A. et al. Viral load as a predictor of outcome in Crimean-Congo hemorrhagic fever. Clin Infect Dis 45, e96–100, doi: 10.1086/521244 (2007).
Shepherd, A. J., Swanepoel, R., Leman, P. A. & Shepherd, S. P. Field and laboratory investigation of Crimean-Congo haemorrhagic fever virus (Nairovirus, family Bunyaviridae) infection in birds. Transactions of the Royal Society of Tropical Medicine and Hygiene 81, 1004–1007 (1987).
Zeller, H. G., Cornet, J. P. & Camicas, J. L. Experimental transmission of Crimean-Congo hemorrhagic fever virus by west African wild ground-feeding birds to Hyalomma marginatum rufipes ticks. Am J Trop Med Hyg 50, 676–681 (1994).
Maganga, G. D. et al. Bat distribution size or shape as determinant of viral richness in african bats. PLoS One 9, e100172, doi: 10.1371/journal.pone.0100172 (2014).
Maganga, G. D. et al. Identification of an unclassified paramyxovirus in Coleura afra: a potential case of host specificity. PLoS One 9, e115588, doi: 10.1371/journal.pone.0115588 (2014).
Bishop, D. H. et al. Bunyaviridae. Intervirology 14, 125–143 (1980).
Casals, J. & Tignor, G. H. The Nairovirus genus: serological relationships. Intervirology 14, 144–147 (1980).
Zeller, H. G. et al. Electron microscopic and antigenic studies of uncharacterized viruses. II. Evidence suggesting the placement of viruses in the family Bunyaviridae. Arch Virol 108, 211–227 (1989).
Davies, F. G., Jessett, D. M. & Otieno, S. The antibody response of sheep following infection with Nairobi sheep disease virus. J Comp Pathol 86, 497–502 (1976).
Dowall, S. D. et al. Hazara virus infection is lethal for adult type I interferon receptor-knockout mice and may act as a surrogate for infection with the human-pathogenic Crimean-Congo hemorrhagic fever virus. J Gen Virol 93, 560–564, doi: 10.1099/vir.0.038455-0 (2012).
Shepherd, A. J., Leman, P. A. & Swanepoel, R. Viremia and antibody response of small African and laboratory animals to Crimean-Congo hemorrhagic fever virus infection. Am J Trop Med Hyg 40, 541–547 (1989).
Burt, F. J., Leman, P. A., Abbott, J. C. & Swanepoel, R. Serodiagnosis of Crimean-Congo haemorrhagic fever. Epidemiol Infect 113, 551–562 (1994).
Sevcik, M., Kristofik, J., Uhrin, M. & Benda, P. New records of ticks (Acari: Ixodidae) parasiting on bats in Slovakia. Vespertilio 13–14, 139–147 (2010).
Oba, M. et al. A novel Bunyavirus from the soft tick, Argas vespertilionis, in Japan. The Journal of veterinary medical science/the Japanese Society of Veterinary Science, doi: 10.1292/jvms.15-0536 (2015).
Telmadarraiy, Z. et al. A survey of Crimean-Congo haemorrhagic fever in livestock and ticks in Ardabil Province, Iran during 2004–2005. Scandinavian journal of infectious diseases 42, 137–141, doi: 10.3109/00365540903362501 (2010).
Tahmasebi, F. et al. Molecular epidemiology of Crimean-Congo hemorrhagic fever virus genome isolated from ticks of Hamadan province of Iran. Journal of vector borne diseases 47, 211–216 (2010).
Vial, L. Biological and ecological characteristics of soft ticks (Ixodida: Argasidae) and their impact for predicting tick and associated disease distribution. Parasite 16, 191–202 (2009).
Muller, M. A. et al. Coronavirus antibodies in African bat species. Emerg Infect Dis 13, 1367–1370, doi: 10.3201/eid1309.070342 (2007).
Acknowledgements
We thank Stephan Kallies, Monika Eschbach-Bludau, Tobias Bleicker, Sebastian Brünink and Anette Klein for excellent technical assistance, Gudrun Wibbelt and Kristin Mühldorfer (IZW, Berlin) for providing bat serum samples, Antje Seebens (Noctalis) for field work in Ghana, and Eric Bergeron and Stuart Nichol for donating the original minireplicon plasmids. We thank Tasnim Suliman (Stellenbosch University) for critically reading the manuscript and Dennis Bente (UTMB, Galveston) for helpful comments. The CCHFV N antibody and CCHFV strain IbAr10200 were kindly provided by Ali Mirazimi, SMI, Sweden. The CCHFV positive serum was provided by Gülay Korukluoglu (Refik Saydam National Public Health Agency, Ankara, Turkey), and the negative human serum by Andreas Kaufmann (Philipps Universität Marburg, Germany). This work was supported by European Commission projects Antigone (contract no. 278976 to CD) and CCHFever Network (contract no. 260427 to FW). Samples and essential reagents were contributed by earlier projects funded by the Deutsche Forschungsgemeinschaft, including grants DR772/3-1, DR772/12-1, DR722/10-1 to CD, We 2616/7-1 to FW, MU 3564/1-1 to MAM, KA 1241/18-1 to E. Kalko and MT). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
M.A.M., S.D., F.W., J.F.D., E.M.L. and C.D. designed and conceived the work. M.A.M., S.D., E.L., V.M.C., G.D.M., F.G.-R, T.B., P.V., P.E., V.M.C., M.T., S.O., J.F.D., F.W. and E.M.L carried out the experiments and provided essential material. M.A.M. and C.D. wrote the main manuscript text. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Müller, M., Devignot, S., Lattwein, E. et al. Evidence for widespread infection of African bats with Crimean-Congo hemorrhagic fever-like viruses. Sci Rep 6, 26637 (2016). https://doi.org/10.1038/srep26637
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep26637
This article is cited by
-
Ornithodoros (Pavlovskyella) ticks associated with a Rickettsia sp. in Pakistan
Parasites & Vectors (2022)
-
The virome of German bats: comparing virus discovery approaches
Scientific Reports (2021)
-
Translating Predictions of Zoonotic Viruses for Policymakers
EcoHealth (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.