Abstract
Inflammatory cells, an integral component of tumor evolution, are present in Glioblastomas multiforme (GBM). To address the cellular basis and dynamics of the inflammatory microenvironment in GBM, we established an orthotopic syngenic model by grafting GL261-DsRed cells in immunocompetent transgenic LysM-EGFP//CD11c-EYFP reporter mice. We combined dynamic spectral two-photon imaging with multiparametric cytometry and multicolor immunostaining to characterize spatio-temporal distribution, morphology and activity of microglia and blood-derived infiltrating myeloid cells in live mice. Early stages of tumor development were dominated by microglial EYFP+ cells invading the tumor, followed by massive recruitment of circulating LysM-EGFP+ cells. Fluorescent invading cells were conventional XCR1+ and monocyte-derived dendritic cells distributed in subpopulations of different maturation stages, located in different areas relative to the tumor core. The lethal stage of the disease was characterized by the progressive accumulation of EGFP+/EYFP+ monocyte-derived dendritic cells. This local phenotypic regulation of monocyte subtypes marked a transition in the immune response.
Similar content being viewed by others
Introduction
Glioblastomas (GBM), the most aggressive form of glioma, are highly vascularized infiltrating brain tumors. They are resistant to current therapies and invariably fatal1. Strategies aiming at targeting GBM cells have so far proved largely unsuccessful. Increasing evidence shows that tumors progress by interacting with host cell types that reside in or are recruited to the tumor microenvironment2,3. GBM are notoriously immune-suppressive and GBM cells likely manipulate immune cells to support tumor expansion (for review4).
Inflammatory myeloid and lymphoid cells make up a substantial part of the GBM tumor mass; by some estimates as high as one third of the cells5. Cells with macrophage activity, usually called tumor-associated macrophages, frequently accumulate in the tumor stroma4. They are proposed to play an important role in suppressing immune responses to cancer cells, but their origin and heterogeneity have not been fully documented. Within the central nervous system (CNS) inflammatory reactions differ substantially from those of other tissues in several ways6,7. First, the CNS has its own resident innate immune cells composed of microglial cells8,9, recently shown to have a potential for self-renewal without the contribution of peripheral myeloid cells10. Second, the CNS parenchyma lacks conventional resident dendritic cells (DCs). Perivascular macrophages, vascular pericytes and subpopulations of microglia subsume the functions of mature DCs11,12,13. Third, the permeability of microvessels for extravasation of large molecules and blood cells is reduced by the blood–brain barrier (BBB) in the CNS in comparison to the rest of the body. Hence, CNS innate immune cells do not recruit the machinery of the adaptive immune response as efficiently as do conventional DCs in peripheral tissues. It is therefore crucial to obtain a more detailed understanding of the cascade and kinetics of immune responses inside the tumor bearing CNS as targeting of tumor-associated host cell subpopulations may represent a therapy to be explored. The phenotypic classification of GBM-associated immune cells is challenging. Attributing functions in GBM progression specifically to microglial cells14 or monocyte-derived cells, is a matter of debate due to the lack of specific markers for each subpopulation and their plastic phenotypes. Significant confusion in the current literature of myeloid subpopulations seems to result from inappropriate grouping of cells or from lack of a common method for distinguishing the various subpopulations. However, recent additions to the repertoire of antibody markers of DCs versus macrophages versus monocytes provided significant clarity to this situation15. Moreover, no research techniques have for example permitted unambiguous distinction between microglia and infiltrated monocytes in CNS tissue. Indeed, irradiation chimerism16 or transient parabiosis between the circulatory systems of a green fluorescent protein, GFP, labeled animal and of a wild type littermate17,18 are two heavy methodologies suffering from intrinsic limitations19. Recent breakthroughs in intravital cellular imaging have opened the way to longitudinal and functional characterization of the immune cell populations in various pathological contexts20.
We devised a clinically relevant syngenic GL261 GBM model in C57BL/6 mice that showed pronounced evolution of immune profile during tumor development. In order to gain insight in the respective involvement of resident microglia and circulating leucocytes across the different stages of tumor development, this model was implemented using multicolor fluorescent reporter mice. We analyzed the immune response with three complementary approaches: intravital spectral two-photon microscopy, multi-parametric cytometry and immunohistochemistry. Their combination enabled us to link information on cell dynamics, distribution, density and morphology to information on phenotypes and frequencies of immune cell populations during GBM progression.
Results
The LysM-EGFP//CD11c-EYFP mouse model permits the study of the dynamics of innate immunity during GBM progression
We developed a syngenic orthotopic GBM model in immunocompetent C57BL/6 mice. This model recapitulates the typical features of the tumor and is suitable for intravital two-photon imaging and multiparametric flow cytometry analyses. We crossed LysM-EGFP21 and CD11c-EYFP22 mice to obtain LysM-EGFP//CD11c-EYFP double-transgenic animals to facilitate visualization of peripheral myeloid cells, microglia and DCs, respectively. Among the described murine models to study GBM initiation and development23, we selected a reproducible graft model in which the tumor develops in the brain at a depth accessible to in vivo two-photon imaging. This model employs the stereotaxic injection of a spheroid of fluorescent GL261 glioma cells stably expressing DsRed into the superficial layers of the parietal cortex (approximately 250 μm deep) of LysM-EGFP//CD11c-EYFP and occasionally of Thy1-CFP//LysM-EGFP//CD11c-EYFP mice to highlight subsets of cortical pyramidal neurons. The dura-mater is sealed with an agarose hemi-bead and the bone above the injection site is replaced by a glass window (4–5 mm in diameter). The cranial window allows a non-invasive follow-up of the events throughout the disease progression24.
We evaluated the effect of our grafting protocol on cortical architecture and inflammation by performing the whole surgical protocol except the injection of the GL261-DsRed glioma spheroid (sham mice). We showed by two-photon imaging of the Thy1-CFP//LysM-EGFP sham mice, that the adjacent neuronal cyto-architecture was not affected by the surgery (Suppl. Fig. 1). Imaging of the LysM-EGFP and CD11c-EYFP sham mice from day 15 (D15) to D33–34 post-surgery revealed that LysM-EGFP+ cells were virtually absent from the brain parenchyma whereas rare CD11c-EYFP+ cells could be seen. A few LysM-EGFP+ and CD11c-EYFP+ cells were present as a monolayer at the surface of the sealing agarose hemi-bead (Suppl. Fig. 1). Their density did not vary over time and the CD11c-EYFP+ cells in the parenchyma did not move towards the hemi-bead during the 20-day observation period.
We then implanted GL261-DsRed glioma spheroids in both wild type and LysM-EGFP//CD11c-EYFP transgenic mice. Tumor growth was measured by fluorescence macroscopy every 3 to 4 days from D15 until D33, the final stage of the disease at which all mice lost weight and some died. Tumor expansion (n = 40) fits with an exponential curve (R2 = 0.8378) (Fig. 1a) with slow growth until D15 followed by rapid expansion afterwards. No significant difference in tumor growth was seen between double transgenic (n = 13, Tau = 0.18; R2 = 0.8619) and wild-type (n = 9, Tau = 0.17 R2 = 0.783) mice (Fig. 1b), indicating that the introduction of fluorescent tags did not modify tumor development. Interestingly, although on average tumor surface increased exponentially over 4 weeks, the daily progression rate significantly decreased from 1.2 fold at D21 to 1.09 at D28 (n = 17, p < 0.001).
A syngenic glioblastoma mouse model allows intravital two-photon imaging of disease progression.
(a) DsRed+ Glioma261 growth curve measured by fluorescence macroscopy (n = 40 mice). (b) Tumor growth curves in WT (black, diamonds, n = 9 mice) and transgenic (red, squares, n = 13 mice) animals. (c) Intravital two-photon imaging of GBM cells and tumor vasculature at day 26 (D26) post-graft. Red: tumor cells; cyan: vasculature. Scale-bar: 100 μm. (d) Intravital two-photon imaging of GBM cells and brain vasculature on the same animal at D19 and D22 post-graft. Red: tumor cells; blue: vasculature; asterisk: Sephadex hemi-bead; arrows: stable vascular landmarks, dotted line: tumor margin. Scale-bar: 100 μm. (e) Intravital two-photon imaging of GBM and surrounding brain parenchyma at D22 post-graft. Note the escape of the tumor cells that invade the brain parenchyma at tumor margin. Red: tumor cells; blue: vasculature; cyan: neurons. Inset: zoom on escaping cells. Scale-bar: 100 μm. (f) Haematoxylin-Eosin staining of a GL261 tumor. Asterisk: necrotic area; arrows: peri-necrotic pseudopalissades. Scale-bar: 50 μm.
Recurrent two-photon imaging of GL261-DsRed grafted mice receiving an injection of Cascade blue to highlight blood vessels revealed that the tumor elicited vascular remodeling in its core (Fig. 1c,d), while individual tumor cells invading the adjacent healthy brain parenchyma can be observed (Fig. 1e). Histo-pathological examinations also revealed a necrotic core in the neoplasm (Fig. 1f).
Altogether these data showed that our syngenic orthotopic model in LysM-EGFP//CD11c-EYFP mice mimics GBM physiopathology (angiogenesis, invasiveness of parenchyma, necrotic core) and permits investigation of the changes in immunological parameters during GBM development by recurrent in vivo two- photon spectral imaging.
Imaging LysM-EGFP and CD11c-EYFP cells accumulation dynamics and distribution during GBM progression
To obtain information on temporal fidelity and statistical significance using a minimal number of animals, we repeatedly imaged the same LysM-EGFP//CD11c-EYFP mice every 3 to 4 days from D15 to D33 post-grafting (n = 12 to 6, depending on the time-point) (Fig. 2). Whatever the time post-graft, CD11c-EYFP+ cells were never observed in the circulation. At D19 of GBM progression, CD11c-EYFP+ cells were found to accumulate intermingled with tumor cells inside the tumor (224 ± 26 cells/mm2) (Fig. 2a,d). They were scarce in the parenchyma surrounding the tumor. We observed CD11c-EYFP+ cells wrapped around GBM cells (Fig. 2a). By contrast, LysM-EGFP+ cells, absent from healthy tissue, were observed in the circulation and tumor blood vessels suggesting that they were recruited from the periphery to the tumor site. In the pathological brain, LysM-EGFP+ cell density was low (70 ± 15 cells/mm2) until D22, with approximately two third of the cells observed at the periphery of the tumor in a non-clustered manner (Fig. 2a,e).
Intravital two-photon imaging of CD11c-EYFP and LysM-EGFP cells dynamics during GBM progression.
(a,b) Intravital two-photon imaging of tumor (red), CD11c-EYFP (yellow, arrowheads) and LysM-EGFP (green, arrows) cells at (a) early stage (D19) and (b) terminal stage (D33) of GBM progression. Scale-bar: 100 μm. (c) Amoeboid LysM-EGFP cells (green) accumulation in and around blood vessels (blue) at late stage (D29) of GBM progression. Scale-bar: 100 μm. (d,e) Evolution of CD11c-EYFP (d) and LysM-EGFP (e) cells densities inside the tumor during GBM progression (n = 12 to 6 mice depending on the timepoint). *p < 0.05.
At the latest stages (>D26) of GBM progression, CD11c-EYFP+ cells were still present, although their density decreased by half inside the tumor (D19, 224 ± 26 cells/mm2; D29, 100 ± 17 cells/mm2; Fig. 2b,d). On the other hand, the density of LysM-EGFP+ cells increased strongly during GBM progression, resulting in a 4-fold increase in density between D22 and D33 (D22, 70 ± 15 cells/mm2; D33, 297 ± 62 cells/mm2) (Fig. 2e). LysM-EGFP+ cells invaded the tumor core (Fig. 2b), most likely due to extravasation from localized intraluminal accumulations of cells (Fig. 2c). These changes in the relative densities of fluorescent cells were rather sharp and preceded the lethal phase of the disease. This dynamic recruitment was triggered by the tumor development, as it was not observed in sham-operated LysM-EGFP//CD11c-EYFP mice. In the tumor, the CD11c-EYFP+ and the LysM-EGFP+ cells seemed to have reciprocal dynamics, as the highest densities of CD11c-EYFP+ cells corresponded to the lowest densities of LysM-EGFP+ cells and vice-versa.
Phenotyping innate immune responses during GBM progression
To document the nature of the immune response and the degree of heterogeneity found in the LysM-EGFP+ and CD11c-EYFP+ populations during tumor progression, we performed multi-parametric flow cytometry analyses. We collected and dissociated tumors and neighboring cortex tissues of PBS perfused individual mice taken before (D21) and after (D28) tumor invasion by LysM-EGFP+ cells. Sham-operated mice were analyzed in parallel to tumor-implanted animals.
We validated a 19-parameter flow cytometry panel (Suppl. Table 1, Suppl. Figs 2 and 3) compatible with the simultaneous detection of the two EGFP and EYFP fluorescent reporter proteins. Analysis of living cells with the surface markers CD45 and CD11b allowed discrimination of three main populations, denoted as populations I (CD45high/CD11b−), II (CD45high/CD11b+) and III (CD45low/CD11b+, corresponding to microglial cells25) (Fig. 3a). In sham-operated mice (Fig. 3c), the explanted tissue was dominated by microglial cells (>90% of analyzed cells), whereas populations I and II accounted for <10% of the analyzed cells. At D21, the brain immune profile of tumor bearing animals, determined by flow cytometry was essentially the same as for control and sham-operated animals, both in term of cells numbers and fluorescent protein expression (Fig. 3d). The late stage of tumor development was characterized by a dramatic change of immune profile. Indeed, at D28, the representation of population III decreased in favor of populations I and II, concomitantly with the five-fold increase in the number of CD45+ infiltrating cells (I + II) when compared to tumor bearing mice at D21 (Fig. 3a–c).
Flow cytometric analysis of brain infiltrating leukocytes during tumor progression.
(a). CD45+/low leukocytes extracted from the brain before (D0), or 21 (T-D21) or 28 (T-D28) days after tumor grafting were analyzed for the expression of CD11b and CD45 and subdivided into CD45+CD11b− (denoted as I), CD45+CD11b+ (denoted as II) and CD45 lowCD11b+ (denoted as III) populations. The percentage of cells in each population is indicated. (b). Histogram depicting the number of brain infiltrating cells corresponding to pooled populations I and II (see a) in control (D0), sham-operated (S-D21 and S-D28) or and tumor implanted (T-D21 and T-D28) mice. Pooled data from individual mice are represented as mean ± SEM (D0: n = 5, S-D21: n = 7, S-D28: n = 5, T-D21: n = 12 and T-D28: n = 13). Statistical significance was evaluated by a Mann Whitney test: ***p < 0.001, ns: non significant. (c) Relative representation of populations I, II and III (see a) in brain in the different groups of mice. Data are presented as the mean ± SEM in each group. (d) Percentages of LysM-EGFP+ (green), CD11c-EYFP+ (orange) and LysM-EGFP+/CD11c-EYFP+ (blue) cells among populations I, II and III (see a) found in the brain of sham-operated (S-D21 and S-D28) or tumor implanted (T-D21 and T-D28) mice. The surface of the pie charts is proportional to the absolute cell number in the specified population. Cell numbers are indicated under each pie chart as mean ± SEM (S-D21: n = 4, S-D28: n = 4, T-D21: n = 6 and T-D28: n = 6).
Approximately 95% (90.9% S-D21, 97.6% S-D28) of the CD11c-EYFP+ cells in sham-operated mice belonged to population III. They were therefore identified as a subpopulation of resident microglia representing roughly 12% (9.6% S-D21, 14.9% S-D28) of the total microglial population (Fig. 3d). At D21, more than 90%, (91.2 ± 1.3%) of EYFP+ cells co-expressed Iba1, a marker for microglia in the healthy CNS (Suppl. Figs 4 and 5c). On the other hand, 80% of the LysM-EGFP+ cells (80.4% S-D21, 78.3% S-D28) belonged to population II where they accounted for roughly 50% of the cells. They were 10-fold less numerous in population I and accounted for less than 1% of population III (Fig. 3d).
The massive infiltration of peripheral immune cells at late tumor stages was accompanied by a diversification of phenotypes of the fluorescent cells as indicated by the detection of CD11c-EYFP+ cells in populations I (10.4%) and II (10%). We also observed the emergence of double-labeled cells essentially in population II (11.4%) (Suppl. Fig. 2) despite the absence of such cells in the pool of circulating cells.
Altogether these data confirmed that the resident microglia (population III) is the major population of immune cells involved in the early phase of the tumor growth and that the number of blood-derived infiltrating cells remains low. They also highlighted the delayed infiltration of cell populations I and II during the fourth week of tumor growth, in agreement with the invasion of the tumor core by LysM-EGFP+ cells (population II) that was observed by two-photon imaging from D26 onward (Fig. 2e). This correlation of terminal progression of the tumor with expansion of populations I and II raised issues about their cell composition and function.
To this end, successive gating on 13 membrane proteins in addition to CD45 and CD11b (Suppl. Figs 3 and 4, Suppl. Table 1), allowed us to untangle and quantify the corresponding populations of leucocytes (Fig. 4a). Comparing sham-operated and tumor-bearing mice at D28, we observed significant increases in T lymphocytes (three-fold increase), natural-killer cells and in all DC subpopulations (XCR1+ DCs, CD11b+ DCs and plasmacytoid DCs, monocyte-derived DC P2 and P3 as well as in their precursor P1). Microglial cells outnumbered DCs by 45-fold until D21 but were only 2.7-fold more numerous at D28.
Multiparametric analysis of brain infiltrating cells allowed identification of all infiltrating leukocytes.
(a) Absolute numbers of lymphocytes (T: T cell, B: B cell and NK: NK cell), granulocytes (Ne: Neutrophils and Eo: eosinophils), DCs (P: pDC, Xc:XCR1+DC and Cd: CD11b+DC) and monocytes/macrophages (Mi: microglia and P1, P2, P3, P4, P5 cells as defined in15 in the brain of sham-operated (S-D21 and S-D28 are grouped) and tumor-implanted (T-D21; T-D28) mice at D21 and D28. Data are presented as mean ± SEM (n = 12). Multiple comparisons were made using one-way ANOVA. Statistically significant differences are observed between T-D28 and S-D21/D28 or T-D21 with a ***p < 0.001 for most of the populations with the exception of B cells, neutrophils, eosinophils, P4, P5 and microglia. No statistically significant differences are observed between S-D21/28 and T-D21. (b) Frequency distribution and differentiation chart of the precursor P1 population, the mo-DC population (P2–P3) and the Macroph. population (P4-P5) at D21 and D28. mo-DC: monocyte-derived DCs; Macroph.: Macrophages. *p < 0.01 P2/3 vs. P4/5; $p < 0.01 D21 vs. D28, Mann-Whitney test.
It is noteworthy that one third of the total DCs present in the brain during tumor progression were monocyte-derived DCs (24% at D21, 37% at D28), distributed in subpopulations of different maturation stages, whose size and fate was likely influenced by the tumor microenvironment. The precursor P1 subpopulation differentiated into a predominant CCR2+ population composed of P2 and P3 monocyte-derived DCs and into a smaller CCR2− population composed of P4 and P5 macrophages. Differences between the densities of monocyte-derived DCs and macrophages were more pronounced during the latest stage of the disease (Fig. 4b).
Uncovering the immunophenotype of transgenically labeled cells
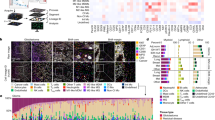
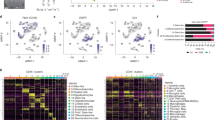
We used the expression profile of EYFP+ and EGFP+ cells determined by flow cytometry analysis among the different cell populations combined with immunostaining to describe their patterned distribution during tumor development (Fig. 5). At D28 multiparametric data indicated that in the brain of tumor-bearing mice EYFP expression was mainly encountered in a subset of the main microglial population (20%. of the whole microglial population, T-D28 in Fig. 3d), as well as in two-thirds of the XCR1+ DCs (60% of the whole XCR1+ DC population) and in nearly half of CD11b+ DCs (40% of the whole CD11b+ DC population) (Fig. 5a). Given the relative incidence of these cell populations, we inferred that the non-microglial EYFP+ cells we observed at D28 were mostly XCR1+ DCs since they outnumbered CD11b+ DCs by nearly eight-fold (Fig. 5b).
Immunophenotyping of LysM-EGFP and CD11c-EYFP cells.
(a) Representative evaluation of the percentage of LysM-EGFP and CD11c-EYFP expressing fluorescent cells in each of the populations specified in Fig. 4a, in the brain of tumor bearing mice at D28. Values are expressed as cumulated percentages. (b) Distribution of the different immune populations in the whole EYFP+, EYFP+/EGFP+ and EGFP+ populations at D28. (c) Multicolor immunostaining of EYFP+ (yellow), Iba1+ (cyan) and MHCII+ (magenta) cells in sagittal sections of tumor bearing brains at D21 (top) and D28 (bottom). (m) meninges; (cc) corpus callosum. Dotted-lines highlight tumor margins. Dotted-squares: EYFP+ cells in the (cc). Arrow: Flow of EYFP+ cells arising for deep brain structures such as ventricles. Scale-bar: 200 μm. (d) Dura-mater specific morphologies as observed in-vivo by two-photon imaging of EYFP+ (yellow, arrow) and EGFP+ (green, arrowhead) cells above the tumor at D15 post-graft. Note the second-harmonic signal (blue) characteristic of the fibers of the meningeal layer. (e) Main morphologies of EYFP+ (yellow) cells imaged in vivo in peri-tumoral and tumor areas. Red: tumor cells; blue: vasculature. (f ) Distribution of the three main morphologies (star-like elongated and amoeboid) of EYFP+ cells observed in vivo by two photon imaging at D21 and D28. Scale bar: 20 μm. (g) In vivo two-photon image of a CD11c-EYFP+/LysM-EGFP+ cell taken at D26 post-graft (asterisk: CD11c-EYFP+/LysM-EGFP+ cell, arrowhead: CD11c-EYFP+ cell, arrow: LysM-EGFP+ cell). Scale-bar: 100 μm.
Moreover, quantitative analysis of multicolor immunostained saggital brain slices at D28 showed that EYFP+/MHCII+ DCs (mainly XCR1+ DCs) represented 31% of the intratumoral EYFP+ cells. They were densely accumulated in the superficial tumor layer (Fig. 5c). On the other hand, DCs exhibiting a lower level of staining for MHCII were grouped along the corpus callosum that invaded the deepest part of the cortical tumor (Fig. 5c,d). In the tumor core, 75% of EYFP+ cells observed were Iba1+, which supported their microglial identity. Their star-shape and the multiple cytoplasmic extensions observed by two-photon imaging also identified them as microglia (Fig. 5e). The cells we observed inside the tumor were less ramified than in the tumor periphery and became amoeboid during tumor development, as expected from a progressive evolution of their activation state (Fig. 5e,f). Immunostaining indicated that 20% of these cells expressed MHCII at D28, thus revealing their antigen presentation ability. It was noteworthy that MHCII expression was more frequent in the non-fluorescent microglial population since 45% of the non fluorescent Iba1+ population was MHCII+ at D28. By contrast at D21 MHCII was present in less than 2% of EYFP+ microglia and in 5% of the non labeled Iba1+ population (Fig. 5c). Therefore, despite the lack of selectivity of the transgenic EYFP labeling, our data indicate that EYFP+ cell subpopulations can nevertheless be identified in the living animal based on their patterned distributions relative to the tumor core.
Similarly, Ly6G+ immunostaining data indicated that at D21 more than 80% of the EGFP+ cells accumulating in the tumor were monocyte-derived cells (Suppl. Fig. 5) and not EGFP+ neutrophils despite the high number of neutrophils extracted from the tissue (Fig. 4a).
Two-photon microscopy and FACS data further suggested that the subpopulations of EGFP+ monocyte-derived cells (P1 to P5) could be differentiated by the graded co-expression of EYFP with EGFP (Fig. 5b,g). In agreement, double labeling was absent from the immature P1 population but found in P2 and P3 monocyte-derived DCs that represented altogether 43% of the double-labeled EGFP+/EYFP+ cells at the latest stage. The low representation of the P4 and P5 populations at D28 over the P2 and P3 population (Figs 4b and 5b) prevented the clear identification of these cell phenotypes in the double-labeled population during two-photon microscopy observations. The accumulation of these double-labeled cells inside the tumor could be considered as a biomarker of a transition in the immune response (Fig. 5b). Moreover, absence of these double-labeled cells in the blood stream strongly suggested that the up-regulation of EYFP expression was triggered by the tumor environment.
Use of dynamic intravital imaging to gain insight into the function of LysM-EGFP+ and CD11c-EYFP+ cells
To gain insight into the functional diversity of the fluorescent cell populations described by FACS and immunostaining, we performed intravital time-series imaging at D21 and D28, respectively. Acquisitions were performed from 0 to 200 μm below the dura-mater inside the tumor every 5 min for 1 to 3 hours. We took advantage of time-color coded projections to highlight the mobility of fluorescent cells. At D21 we observed that amoeboid EGFP+ cells were highly motile (157 ± 16 μm/h, n = 5 mice and 25 cells) and that they could move randomly inside the tumor (Fig. 6a,c, Suppl. Fig. 6, Suppl. Movie 1). By contrast, CD11c-EYFP+ somas were immobile over a one-hour observation period (n = 5 mice and 24 cells, Fig. 6a,c, Suppl. Movie 1) and only their fine processes were dynamically and quickly scanning the environment (Fig. 6d) as reported for microglia. At D28, we evidenced a modest but significant increase in the average movement of CD11c-EYFP somas in the tumor (Fig. 6b,c, Suppl. Movie 2), correlated with the increase in the number of XCR1 DCs. We also observed a 30% decrease in the velocity of LysM-EGFP+ cells (110 ± 13 μm/h, n = 24 p < 0.05), concomitant with the diversification of cell phenotypes.
CD11c-EYFP and LysM-EGFP cells morphology and motility during GBM progression.
(a,b) Time-color coded images of LysM-EGFP+, CD11c-EYFP+ and tumor cells taken at (a) D21 and (b) D28 post-graft over a 55-min observation period. Highly motile cells and structures appear in color, immobile cells and structures appear in white. Scale-bar: 100 μm. (c) CD11c-EYFP+ and LysM-EGFP+ cells velocities at D21 (orange, n = 5 mice) and D28 (red, n = 4 mice) post-graft. Thy1+ cells (neurons) were used to assess the lower sensitivity threshold of the measurements (dotted-line). *p < 0.05. (d) Time-color coded image of a CD11c-EYFP+ star-shaped cell taken in the peri-tumoral area at D21 over a 55 min observation period. Note the highly motile cytoplasmic protrusions (color) and the immobile cell body (white). Scale-bar: 20 μm. (e,f) Intravital two-photon imaging of a (e) CD11c-EYFP+ and (f ) LysM-EGFP+ cell that have phagocytized a DsRed expressing tumor cell fragment (red, arrowhead). Scale-bar: 20 μm.
These average migration speeds resulted from phases of acceleration and arrest that might correspond to interactions with other immune cells types (Suppl. Fig. 6, Suppl. Movies 1, 3). Indeed, such episodes were reminiscent of cellular interaction events described in the case of adaptive immune cells26. We occasionally (n = 3) also observed lasting interactions between EGFP+ monocyte-derived DCs and EYFP+ microglia at D28. Contacts between EGFP+ or EYFP+ cells and DsRed GL261 cells were frequent and in some instances resulted in phagocytic events as attested by the presence of red inclusions in the cytoplasm of some of these cells at D28 (Fig. 6e,f). Such red inclusions were rare in LysM-EGFP+ cells, but frequent in the population of CD11c-EYFP+ cells (30% of the cells n = 3 mice at D21 and D28, 5 ROI/mouse), as expected from the reported phagocytic competence of microglia27.
Expression, distribution and morphology of LysM and CD11c in human GBM
To expand the potential knowledge gained in mice into an ability to translate it to humans, we searched for the expression of LysM, CD11c, CD11b and Iba1 markers in human GBM biopsies. Indeed, data showed that it is possible to correlate the subsets of mononuclear phagocyte cells found in the mouse with those present in the human28. Immunohistochemical analysis of the expression of these markers was performed on a cohort of GBM tumors (n = 123). Despite a lack of significant prognostic value of these markers within our cohort, it was of interest that we found that 50% of human GBM expressed LysM and that 58% expressed CD11c (Fig. 7a). As observed by imaging (Fig. 2), we noticed a differential distribution of the immune cells between the core (defined as the most contrasted areas following gadolinium injection) and the periphery, with higher densities of LysM+ and of CD11c+ cells inside the tumor compared to the tumor margin (Fig. 7b,c). Among LysM positive GBM samples, 61.5% expressed LysM only in the tumor core, 27% expressed LysM both in the tumor core and its periphery, while 11.5% expressed LysM only inside the vessels (Fig. 7d). LysM was preferentially expressed in biopsies presenting a significant level of CD11b immunostaining (p = 0.044), which supported the existence of a double positive LysM+/CD11b+ population in human GBM as shown in mice. Twenty seven per cent of patients were positive both for LysM and CD11c. The highest CD11c densities corresponded to weak LysM densities as expected from the mutual exclusion of the LysM+ and CD11c+ populations observed in mice. CD11c+ cells had either ramified or round morphologies (Fig. 7e). Similar ramified and round morphologies were also observed among the Iba1+ cells (Fig. 7f) in 95% of human biopsies (Fig. 7a). Taken together these results showed that LysM and CD11c are expressed in the human GBM microenvironment and most likely reflect the presence of similar cell populations as those found in the mouse.
Immunohistochemistry of inflammation markers in a cohort of 123 patients with GBM tumors.
(a) Number of negative and positive immuno-labeled samples for LysM, CD11c, Iba1 and CD11b inflammation markers. Overall survival (OS) and progression free survival (PFS) were calculated for negative and positive groups for each marker and were compared using the log-rank test (p-value). N.S.: not significant. (b) Representative LysM distribution in GBM tumor. LysM staining was denser in the core (T, top) than in the periphery of the tumor (P, bottom). (c) Representative CD11c distribution in GBM tumor. CD11c staining was denser in the core (T, top) than in the periphery of the tumor (P, bottom). (d) Representative picture of LysM labeling with positivity in the tumor core (top) and inside vessels (bottom). (e) CD11c+ cell morphologies: round (top); star-shaped (bottom) (f ) Representative pictures of Iba1+ cell morphologies: round (right) and star shaped (left). Scale-bars: 50 μm.
Discussion
In this study we detailed the dynamics of the innate immune response associated with GBM progression in immunocompetent transgenic reporter mice by combining the most advanced cortical imaging methodology29 with high content flow cytometry analysis15. Our results demonstrate the complementarity of multiparametric cytometry to untangle the diversity of cell phenotypes involved in the innate immune response and microscopy to delineate the dynamics and compartmentalization of this immune response between intratumoral and peritumoral areas. For example at early stages, MHCII+ cells identified by cytometry were mainly observed in the meninges but rarely inside the tumor mass. Conversely, the significant accumulation of EYFP+ microglia in the tumor as well as the subsequent reduction of this density at late disease stages were highlighted only by microscopic imaging. The microglial cells present in the healthy surrounding tissue collected for cytometry most likely attenuated and masked the changes of intratumoral densities.
Microscopic imaging can thus characterize the tumor microenvironment, particularly the representation and dynamic distribution of immune cells, as a regulator of malignancy. The specificity of our fluorescent markers in the early stages of the disease as well as characteristic morphological and kinematic features of the different populations of immune cells have allowed us to distinguish between resident microglia and neutrophils or monocyte-derived populations.
In our reporter mouse model EYFP+ cells were nearly all CD45low/CD11bhigh in the sham control and in early phases of tumor growth. They were never observed in the blood stream. They were also positive for Iba1, had ramified morphology and were thus considered as brain microglial resident cells8. EYFP+ microglia represented a small fraction of the total microglia, likely in a weakly activated state. Indeed MHCII+ was weakly expressed at D21 and at D28 was preferentially encountered in the non-labeled population. In immunostained images, only 20% were EYFP+ Iba1+ MHCII+, characterizing activated microglia. Moreover, of the three typical classes of morphologies reported for microglial activation states30, we indeed found mainly star shaped microglia. However, microglial activation was boosted during tumor development as indicated by the progressive prevalence of the amoeboid phenotype despite the significant decline of microglial cell densities. Phagocytic events by EYFP+ microglia were equally observed at all stages. Our data did not support previous observations suggesting that microglia lose their capacity to express MHCII within the GBM microenvironment31.
By contrast, at D28, cytometric analysis identified 40% of the EYFP+ cells as peripheral CD45high/CD11blow/CD11c+/MHCII+ cells rather than microglia. This is consistent with the massive recruitment of the XCR1+ DCs usually reported in tumors32. MHCII immunostaining indicated that XCR1+ DCs were exclusively located in the meninges and superficial tumor areas at D21 then accumulated at the tumor margins by D28. Such meningeal and peritumoral localization is consistent with the proposed role of XCR1+ DCs that hyperactivate CD8+ T cells during tumor development33, while T cells accumulate in the meningeal and choroid plexus vessels, potential gateways for immune cells into the brain34. Interestingly, we also observed a stream of Iba1−/MHCIIlow/EYFP+ cells in the corpus callosum that targets the deepest areas of the tumor in the cortex. This would be compatible with an infiltration by immature peripheral DCs originating from the ventricles of the depth of the tumor and their differentiation into XCR1+/MHCII+ phenotype along their migration through the tumor toward the meninges. This differential distribution pattern in the CNS is reminiscent of data reported for the injured spinal cord describing different functional phenotypes for meningeal (inflammatory M1) and ventricular (anti-inflammatory M2)macrophages35. Therefore, our results strongly suggest that the combination of transgenic EYFP labeling and 3D intratumoral localization can serve the identification of immune cells in the brain at all stages of tumor development.
Similarly, EGFP was confirmed as a marker of cells of myeloid origin by our cytometry experiments. At early time points, neutrophils outnumbered monocytes/macrophages in brain cell suspensions but their density inside the tumor was in fact always at least five-fold lower than that of monocytes/macrophages as indicated by local dynamic intravital imaging and immunostaining data. The fastest moving cells observed in the tumor were most likely these few neutrophils that are recruited in a wide variety of murine tumor models36. Nevertheless, monocytes were the main population of circulating cells that accumulated inside the tumor when microglial cells density decreased. They represented the source of peripheral macrophages (P4/P5) and monocyte-derived DCs (P2/P3) inside the tumor according to the characterization of similar cell populations in the skin15. Being extremely plastic in their function and phenotype according to the microenvironment37,38,39, the long-lived monocytic cells are thus likely the key regulators of anti-tumor immune responses. In agreement, imaging showed that fluorescent cells acquired distinct morphologies and migratory properties depending on the time and location of the immune responses. This suggests that the analysis of cell morphologies and migration signatures based on the direction, speed, persistence and percentage of moving cells, could have broad implications for advancing our understanding of the inflammatory response.
It is likely that the tumor microenvironment progressively impeded the maturation of monocyte-derived cells, a possible cause for the fatal evolution of the tumor disease40,41. If brain monocytes can no longer exert direct elimination of tumor cells and cross-presentation of antigens to T-cells, tumor progression can be facilitated42.
We show here that co-expression of EYFP and EGFP in monocyte-derived cells could be used as a biomarker of this immune transition in order to stage disease progression. The highest incidence of co-expression occurred in the most mature subpopulations, namely P3 antigen presenting and P5 phagocytic monocyte-derived DCs. However, approximately 20% of the predominant less mature P2 monocyte-derived DC subpopulation also co-expressed EYFP and EGFP at D28.
It was noteworthy that the inflammatory environment induced by a GBM tumor in the CNS, like dermis inflammation in the ear15, favored the differentiation of CCR2+ monocytes into a CCR2+ DC P2 or P3 phenotype. This is in contrast with the differentiation into a macrophagic CCR2− P4 or P5 phenotype encountered in the inflammatory microenvironment of a sterile liver injury43. A possible explanation could be that the antigen-free environment does not require an adaptative immune response, in contrast to tumors. Monocyte–derived DCs exhibit poorer T cell priming abilities than conventional DCs15, yet they accounted for 30% of the total DC population and therefore possibly impacted the adaptive antitumor immune response.
In the future, the targeted and specific ablation of XCR1+ DCs and monocyte-derived subpopulations in vivo should help to pinpoint their exact functions in brain immunity. This should aid in our understanding of the disease and actions of potential therapies on these processes. In support of the potential translational interest of our observations, we also showed that 50% of human GBM expressed LysM and that 58% expressed CD11c with spatial distributions in accord with observations made at different stages of tumor development in the mouse.
Methods
In vivo experiments
Animal care guidelines
All experimental procedures were performed in accordance with the French legislation and in compliance with the European Community Council Directive of November 24, 1986 (86/609/EEC) for the care and use of laboratory animals. The research on animals was authorized by the Direction Départementale des Services Vétérinaires des Bouches-du-Rhône (license D-13-055-21) and approved by the National Committee for Ethic in Animal Experimentation (Section N°14; project 87-04122012).
C57Bl6 mice (n = 97) were housed in cages with food and water ad libitum in a 12 h light/dark cycle at 22 ± 1 °C.
Cell culture and gene transfection
GL261 cells, a murine GBM cell line44 (National Cancer Institute, Charles River Labs), were transfected with a plasmid encoding for DsRed (pDsRed2-N1, Clontech, Mountain View, USA). Cells that stably express DsRed were selected using Geneticin (0.5 mg/ml, Gibco) and cultured as monolayers in RPMI1640 + GlutaMAX-1 (Gibco 61870) supplemented with 10% Fetal Calf Serum (Thermo Scientific) and Geneticin (0.5 mg/ml, Gibco). Cells were kept at 37 °C in a 5% CO2 atmosphere. Confluent cells were plated on Petri dishes coated with 1% soft agarose to induce spheroid formation as described in24.
Animal model
LysM-EGFP21, CD11c-EYFP22 and Thy1-CFP45 mice were used alone or crossed to obtain double transgenic mice.
The GBM animal model was realized as previously described24,46. Briefly, mice were anaesthetized by an intraperitoneal injection of a mixture of Xylazine/Ketamine (10 mg/kg and 100 mg/kg, respectively) and positioned on a stereotactic frame. A 3–4 mm diameter craniotomy was performed on the left parietal bone. The dura-mater was incised by a 31G needle and a 200–250 μm spheroid of GL261-DsRed murine GBM cells was injected into the cerebral cortex approximately 250 μm below the brain surface. A Sephadex hemi-bead with diameter fitting with the dura-mater opening was inserted in the injection wound and glued using histo-compatible acrylic glue (Cyanolit). A round glass coverslip (diameter: 5 mm) was then sealed on the adjacent bone and fixed to the skull by dental cement. Animals were left to recover for 15 days post-surgery before the first imaging session.
Experimental timetable
The experimental timetable is summarized in Suppl. Fig. 7. Mice were imaged every 3 to 4 days from day 15 until day 33 post-surgery. When animals showed a body weight loss of more than 15% of their original body weight, they were euthanized prior to this endpoint. For timelapse and cytometry experiments, animals were observed at day 21 and day 28.
Microscopy
Prior to each imaging session, mice were anaesthetized by an intraperitoneal injection of a mixture of Xylazine/Ketamine (10 mg/kg and 100 mg/kg, respectively), injected intravenously with 100 μl of a Cascade Blue conjugate dextran (70 kDa) solution (20 mg/ml in phosphate saline buffer (Sigma)) and positioned on a stereotactic frame allowing movements in the three-directions. Repositioning of the mice was realized using visual vascular landmarks to ensure that the same area is observed at each imaging session. The anesthesia is sufficient for a one hour observation. However, for timelapse experiments, the anesthesia was switched to a continuous inhalation of isoflurane (0.2 to 0.75% in humidified air) after the first hour of observation.
We used a Zeiss LSM 7MP two-photon microscope home-modified to allow animal positioning below the 20X water immersion objective (1.0 NA) and coupled to a femtosecond pulsed infrared tunable laser (Mai-Tai, Spectra Physics). After two-photon excitation, signals were epicollected and separated by dichroic mirrors and filters on 5 independent non-descanned detectors (collecting range: 390–485 nm, 500–520 nm, 520–549 nm, 555–605 nm and 605–678 nm)29. Images were acquired sequentially using an excitation wavelength tuned at 800 nm and then at 940 nm. Gains and offsets were identical for all the detectors, except for the red channels whose gain was reduced by 30% to compensate for the very strong expression of DsRed in tumor cells.
For CD11c-EYFP and LysM-EGFP cells recruitment experiments, images were typically acquired over a depth of 500 μm using 20 μm steps. Laser intensity was linearly increased with depth. Images were acquired as mosaics in order to cover the whole tumor surface with the healthy peritumoral tissue that surrounds it.
For time-lapse experiments, stacks of images were acquired from 20 to 200 μm below the dura-matter with a z-step of 3 μm. The acquisition was realized over 2 to 3 hours with time steps of 5 min. Ultra-fast timelapse experiments were also realized over a period of 10 min at a 0.1 Hz repetition rate.
Data analysis
For tumor growth measurements, the red channel (555–605 nm) from each mosaic z-stack was projected and the resulting area was measured as an index of the tumor size. Tumor size was plotted against time and an exponential curve was fitted. Fits were considered as valid if the determination coefficient (R2) was >0.546. The mean cell density (number of cells/mm2) was calculated at each time points from the CD11c-EYFP and LysM-EGFP cells. Counts were obtained in 3 to 5 representative regions of interest inside the tumor selected at different depths from 0 to 250 μm below the dura-matter.
Timelapse z-stacks were processed using the open-source software Fiji and dedicated macros. Hyperstacks were first registered in 3D and cells of interest were manually tracked in 3D using the plugin MTrackJ47. Time-color coded images were generated using the temporal-color code plugin of Fiji. The resulting stack was then projected to obtain an image where immobile cells appear in white whereas mobile cells appear in color.
Cytometry
Animals were deeply anaesthetized by an intraperitoneal injection of a mixture of Xylazine/Ketamine and were perfused by cardiac injection of 20 mL of PBS 1X. Brains were extracted and stored in ice-cold PBS. Olfactory bulbs and cerebellum were discarded and the cerebral hemisphere containing the tumor was excised. Cerebral tissues located below the corpus callosum including ventricles were removed with forceps. Then, tumor and surrounding healthy cerebral cortex including dura-mater were stored in a tube with DNAse I, Collagenase D (Roche Applied Science) and Collagenase V (Sigma). The dissociation was realized on a GentleMACS Octo Dissociator (Miltenyl Biotec), the suspension was then filtered while enzymatic reaction was blocked with EDTA. The cell suspension was centrifuged in an 80%/40% Percoll density gradient and rinsed before incubation with 2.4G2. Finally, the cell suspension was incubated 20 min with a mix of antibodies (see Suppl. Table 1) and labeled with Sytox Blue (Life technology). Acquisition was realized on a 5 lasers BD LSRFortessaTM FACS equipment and data were extracted using the BD FACSDiva software . All acquisition are done in a standardized way using application settings. The gating strategy used to extract immune cells is described in Suppl. Fig. 3.
Patient characteristics and tumor samples
One hundred and twenty three patients (80 males and 43 females) with GBM (GBM, WHO grade IV) were included in this retrospective study. Patient age at the time of surgery ranged from 20 to 81 years (mean age at surgery: 59.2 years ± 14.3 years). Complete or subtotal surgical excision was performed in 72 cases, incomplete surgical excision (partial or biopsy) in 49 cases, excision quality was unknown in 2 cases. Tumor specimens were obtained according to a protocol approved by the local institutional review board and ethics committee and conducted according to national regulations. All the patients provided written informed consent. GBM samples provided from the AP-HM tumor bank (authorization AC-2013-1786 and 2014-A00585-42) were pooled on a tissue-microarray (TMA) for high throughput screening. The histology of the paraffin-embedded samples was confirmed by a pathologist (DFB).
Immunohistochemistry
For mice, brains were extracted and fixed overnight in 4% paraformaldehyde. Then, they were cryoprotected by a 24 h bath in 30% sucrose. 25 μm thick slices were realized on a cryotome and permeabilized in a 0.5% Triton solution. A one-hour blockage of the non-specific sites was realized (bovine serum albumin 2%, goat serum 2%, donkey serum 2%, Triton 0.1%) and primary antibodies were incubated overnight at 4 °C in PBS (rabbit anti-Iba1, 1/200, Wako (019-19741); rat anti-MHCII, 1/50, Ebiosciences (145321); rat anti-Ly6G, 1/50, Ebiosciences (145931)). Secondary antibodies (goat anti-rabbit conjugated to Dylight405, 1/100, Thermo (35551); donkey anti-rat conjugated to Cy5, 1/100, Jackson Immunoresearch (712-175-150) were incubated 1 h 30 at room temperature. Slices were then mounted with Vectashield. Slices were washed in PBS in between the different steps. Observations were performed on a Zeiss LSM780 confocal microscope in the spectral mode, using 405, 488, 543 and 633 nm excitation wavelength and spectral deconvolution processing of the images.
For human biopsies, after steam-heat-induced antigen retrieval, 5-μm sections of formalin-fixed paraffin-embedded samples were tested for the presence of LysM (EPR2994(2), 1/1000, rabbit IgG, GeneTex, Inc, Irvine, USA), CD11c (EP1347Y, 1/400, rabbit IgG, Abcam, Paris, France), Iba1 (1/1000, rabbit IgG, Wako Chemicals GmbH, Germany) and CD11b (EP1345Y, 1/100, rabbit IgG, Abcam). A Benchmark Ventana autostainer (Ventana Medical Systems SA, Illkirch, France) was used for detection and slides were simultaneously immunostained to avoid inter-manipulation variability. Slides were then scanned (Nanozoomer 2.0-HT, Hamamatsu Photonics SARL France, Massy, France) and images processed in NDP.view2 software (Hamamatsu). Results of the stainings were determined from three areas of the tumor for each patient and then expressed as negative or positive.
Statistical analysis
All data are expressed as mean ± SEM. Mann-Whitney U-test or one-way ANOVA were used to test differences in between time points. p < 0.05 was used as a criterion for *significance, while p < 0.001 was used as a criterion for ***significance. All statistical analyses were performed with Microsoft Excel and Graphpad Prism.
To analyze data on the patient cohort, analyses were conducted using the statistical package SPSS software v.17 (SPSS Inc, Chicago, USA). To compare patients’ groups according to immunoexpressed inflammation markers, we used Pearson’s chi-square test and Fisher’s exact test as appropriate. Progression free survival (PFS) and overall survival (OS) of the patients were calculated from the date of surgery until disease progression, death or last follow-up (censured data). Survival was estimated using the Kaplan–Meier method and curves were compared using the log-rank test. All the tests were two-sided and p-values of less than 0.05 were considered significant for each statistical analysis.
Additional Information
How to cite this article: Ricard, C. et al. Phenotypic dynamics of microglial and monocyte-derived cells in glioblastoma-bearing mice. Sci. Rep. 6, 26381; doi: 10.1038/srep26381 (2016).
References
Woehrer, A., Bauchet, L. & Barnholtz-Sloan, J. S. Glioblastoma survival: has it improved? Evidence from population-based studies. Curr Opin Neurol 27, 666–674 (2014).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Friedl, P. & Alexander, S. Cancer invasion and the microenvironment: plasticity and reciprocity. Cell 147, 992–1009 (2011).
Glass, R. & Synowitz, M. CNS macrophages and peripheral myeloid cells in brain tumours. Acta Neuropathol 128, 347–362 (2014).
Charles, N. A., Holland, E. C., Gilbertson, R., Glass, R. & Kettenmann, H. The brain tumor microenvironment. Glia 60, 502–514 (2012).
Ousman, S. S. & Kubes, P. Immune surveillance in the central nervous system. Nat Neurosci 15, 1096–1101 (2012).
Dunn, G. P. & Okada, H. Principles of immunology and its nuances in the central nervous system. Neuro Oncol 17 Suppl 7, vii3–vii8 (2015).
Ginhoux, F. et al. Fate mapping analysis reveals that adult microglia derive from primitive macrophages. Science 330, 841–845 (2010).
Ginhoux, F., Lim, S., Hoeffel, G., Low, D. & Huber, T. Origin and differentiation of microglia. Front Cell Neurosci 7, 45 (2013).
Bruttger, J. et al. Genetic Cell Ablation Reveals Clusters of Local Self-Renewing Microglia in the Mammalian Central Nervous System. Immunity 43, 92–106 (2015).
Wlodarczyk, A., Lobner, M., Cedile, O. & Owens, T. Comparison of microglia and infiltrating CD11c+ cells as antigen presenting cells for T cell proliferation and cytokine response. J Neuroinflammation 11, 57 (2014).
D’Agostino, P. M., Gottfried-Blackmore, A., Anandasabapathy, N. & Bulloch, K. Brain dendritic cells: biology and pathology. Acta Neuropathol 124, 599–614 (2012).
Hickey, W. F. & Kimura, H. Perivascular microglial cells of the CNS are bone marrow-derived and present antigen in vivo. Science 239, 290–292 (1988).
Li, W. & Graeber, M. B. The molecular profile of microglia under the influence of glioma. Neuro Oncol 14, 958–978, doi: 10.1093/neuonc/nos116 (2012).
Tamoutounour, S. et al. Origins and functional specialization of macrophages and of conventional and monocyte-derived dendritic cells in mouse skin. Immunity 39, 925–938 (2013).
Mildner, A. et al. Microglia in the adult brain arise from Ly-6ChiCCR2+ monocytes only under defined host conditions. Nat Neurosci 10, 1544–1553 (2007).
Ajami, B., Bennett, J. L., Krieger, C., Tetzlaff, W. & Rossi, F. M. V. Local self-renewal can sustain CNS microglia maintenance and function throughout adult life. Nat Neurosci 10, 1538–1543 (2007).
Ajami, B., Bennett, J. L., Krieger, C., McNagny, K. M. & Rossi, F. M. V. Infiltrating monocytes trigger EAE progression, but do not contribute to the resident microglia pool. Nat Neurosci 14, 1142–1149 (2011).
Ransohoff, R. M. Microgliosis: the questions shape the answers. Nat Neurosci 10, 1507–1509 (2007).
Bousso, P. & Moreau, H. D. Functional immunoimaging: the revolution continues. Nat Rev Immunol 12, 858–864 (2012).
Faust, N., Varas, F., Kelly, L. M., Heck, S. & Graf, T. Insertion of enhanced green fluorescent protein into the lysozyme gene creates mice with green fluorescent granulocytes and macrophages. Blood 96, 719–726 (2000).
Lindquist, R. L. et al. Visualizing dendritic cell networks in vivo. Nat Immunol 5, 1243–1250 (2004).
Huszthy, P. C. et al. In vivo models of primary brain tumors: pitfalls and perspectives. Neuro Oncol 14, 979–993 (2012).
Ricard, C., Stanchi, F., Rougon, G. & Debarbieux, F. An orthotopic glioblastoma mouse model maintaining brain parenchymal physical constraints and suitable for intravital two-photon microscopy. J Vis Exp 86, doi: 10.3791/51108 (2014).
Clausen, B. H. et al. Interleukin-1beta and tumor necrosis factor-alpha are expressed by different subsets of microglia and macrophages after ischemic stroke in mice. J Neuroinflammation 5, 46 (2008).
Herz, J., Johnson, K. R. & McGavern, D. B. Therapeutic antiviral T cells noncytopathically clear persistently infected microglia after conversion into antigen-presenting cells. J Exp Med 212, 1153–1169 (2015).
Hussain, S. F. et al. The role of human glioma-infiltrating microglia/macrophages in mediating antitumor immune responses. Neuro Oncol 8, 261–279 (2006).
Crozat, K. et al. Comparative genomics as a tool to reveal functional equivalences between human and mouse dendritic cell subsets. Immunol Rev 234, 177–198 (2010).
Ricard, C. & Debarbieux, F. C. Six-color intravital two-photon imaging of brain tumors and their dynamic microenvironment. Front Cell Neurosci 8, 57 (2014).
Yang, I., Han, S. J., Kaur, G., Crane, C. & Parsa, A. T. The role of microglia in central nervous system immunity and glioma immunology. J Clin Neurosci 17, 6–10 (2010).
Tran, C. T. et al. Differential expression of MHC class II molecules by microglia and neoplastic astroglia: relevance for the escape of astrocytoma cells from immune surveillance. Neuropathol Appl Neurobiol 24, 293–301 (1998).
Gurka, S., Hartung, E., Becker, M. & Kroczek, R. A. Mouse Conventional Dendritic Cells Can be Universally Classified Based on the Mutually Exclusive Expression of XCR1 and SIRPalpha. Front Immunol 6, 35 (2015).
Broz, M. L. et al. Dissecting the tumor myeloid compartment reveals rare activating antigen-presenting cells critical for T cell immunity. Cancer Cell 26, 638–652 (2014).
Louveau, A. et al. Structural and functional features of central nervous system lymphatic vessels. Nature 523, 337–341 (2015).
Shechter, R. et al. Recruitment of beneficial M2 macrophages to injured spinal cord is orchestrated by remote brain choroid plexus. Immunity 38, 555–569 (2013).
Gregory, A. D. & Houghton, A. M. Tumor-associated neutrophils: new targets for cancer therapy. Cancer Res 71, 2411–2416 (2011).
Stout, R. D. & Suttles, J. Functional plasticity of macrophages: reversible adaptation to changing microenvironments. J Leukoc Biol 76, 509–513 (2004).
Schouppe, E., De Baetselier, P., Van Ginderachter, J. & Sarukhan, A. Instruction of myeloid cells by the tumor microenvironment: Open questions on the dynamics and plasticity of different tumor-associated myeloid cell populations. Oncoimmunology 1, 1135–1145 (2012).
Caronni, N., Savino, B. & Bonecchi, R. Myeloid cells in cancer-related inflammation. Immunobiology 220, 249–253 (2015).
Youn, J.-I., Nagaraj, S., Collazo, M. & Gabrilovich, D. I. Subsets of myeloid-derived suppressor cells in tumor-bearing mice. J Immunol 181, 5791–5802 (2008).
Schmid, M. C. & Varner, J. A. Myeloid cells in the tumor microenvironment: modulation of tumor angiogenesis and tumor inflammation. J Oncol 2010, 201026 (2010).
Kushchayev, S. V. et al. Monocyte-derived cells of the brain and malignant gliomas: the double face of Janus. World Neurosurg 82, 1171–1186 (2014).
Dal-Secco, D. et al. A dynamic spectrum of monocytes arising from the in situ reprogramming of CCR2+ monocytes at a site of sterile injury. J Exp Med 212, 447–456 (2015).
Gazdzinski, L. M. & Nieman, B. J. Cellular imaging and texture analysis distinguish differences in cellular dynamics in mouse brain tumors. Magnetic resonance in medicine 71, 1531–1541, doi: 10.1002/mrm.24790 (2014).
Feng, G. et al. Imaging neuronal subsets in transgenic mice expressing multiple spectral variants of GFP. Neuron 28, 41–51 (2000).
Ricard, C. et al. Dynamic quantitative intravital imaging of glioblastoma progression reveals a lack of correlation between tumor growth and blood vessel density. PLoS One 8, e72655 (2013).
Meijering, E., Dzyubachyk, O. & Smal, I. Methods for cell and particle tracking. Methods Enzymol 504, 183–200 (2012).
Acknowledgements
We thank Cecile Meunier for assistance with histology and mouse colony management, Carole Colin for help in treating statistical data, Alexandre Jaouen and Lisa Lamasse at CERIMED and the PicsL staff for imaging technical assistance. This work was supported by Institut National du Cancer (G.R., D.F.B., A.T. and F.D.) (PLBIO14-197), by Canceropole PACA (fellowship to C.R.), by Site de Recherche Integrée en Cancérologie (INCA-DGOS-INSERM 6038) (D.F.B., G.R. and F.D.), by GEFLUC (F.D.) (BEVAC802), by Fondation de Recherche Médicale (F.D.) (ING20140129149), by Agence Nationale Recherche ANR15-CE16-0009-01 (F.D. and M.M.) and core support from AMU, CNRS and INSERM.
Author information
Authors and Affiliations
Contributions
F.D. and C.R. designed the animal model, the imaging experiments and interpreted the results. M.M. and H.L. designed the cytometry experiments and interpreted the results. A.T. and D.F.B. designed and performed the experiments on the human glioma cohort and, interpreted the data C.R., P.G., A.T., H.L. and F.D. performed the experiments and data analysis. F.D., G.R. and M.M. wrote the paper with input from all of the authors and C.R. and M.M. made all the data figures.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ricard, C., Tchoghandjian, A., Luche, H. et al. Phenotypic dynamics of microglial and monocyte-derived cells in glioblastoma-bearing mice. Sci Rep 6, 26381 (2016). https://doi.org/10.1038/srep26381
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep26381
This article is cited by
-
In vivo imaging of inflammatory response in cancer research
Inflammation and Regeneration (2023)
-
Type I interferon response in astrocytes promotes brain metastasis by enhancing monocytic myeloid cell recruitment
Nature Communications (2023)
-
Immune tumoral microenvironment in gliomas: focus on CD3+ T cells, Vδ1+ T cells, and microglia/macrophages
Immunologic Research (2022)
-
Effects of VEGF blockade on the dynamics of the inflammatory landscape in glioblastoma-bearing mice
Journal of Neuroinflammation (2019)
-
Siglec-15 as an immune suppressor and potential target for normalization cancer immunotherapy
Nature Medicine (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.