Abstract
The MIEC-SVM approach, which combines molecular interaction energy components (MIEC) derived from free energy decomposition and support vector machine (SVM), has been found effective in capturing the energetic patterns of protein-peptide recognition. However, the performance of this approach in identifying small molecule inhibitors of drug targets has not been well assessed and validated by experiments. Thereafter, by combining different model construction protocols, the issues related to developing best MIEC-SVM models were firstly discussed upon three kinase targets (ABL, ALK, and BRAF). As for the investigated targets, the optimized MIEC-SVM models performed much better than the models based on the default SVM parameters and Autodock for the tested datasets. Then, the proposed strategy was utilized to screen the Specs database for discovering potential inhibitors of the ALK kinase. The experimental results showed that the optimized MIEC-SVM model, which identified 7 actives with IC50 < 10 μM from 50 purchased compounds (namely hit rate of 14%, and 4 in nM level) and performed much better than Autodock (3 actives with IC50 < 10 μM from 50 purchased compounds, namely hit rate of 6%, and 2 in nM level), suggesting that the proposed strategy is a powerful tool in structure-based virtual screening.
Similar content being viewed by others
Introduction
Virtual screening (VS) exhibits undefeatable advantage in today’s drug discovery campaign1,2,3, which shows short development time, low financial cost, whereas high production ratio4,5. Roughly, the VS approaches can be divided into two categories: ligand-based and structure-based strategies6. The ligand-based VS approaches employ ligand properties, such as molecular weight, number of hydrogen bond donors/acceptors, solvent accessible surface area, various molecular fingerprinting, etc., to construct prediction models according to known actives. Whereas the structure-based VS approaches additionally employ the target information for the predictions of actives, such as molecular docking, which can give the binding information of ligands upon their targets, i.e. the binding poses, the binding affinities, and even the residue-ligand interaction details. In principle, the compounds derived from the ligand-based VS approaches may usually be limited in the scope of similar core fragment molecules due to the reason that the ligand-based VS approaches can only depend on known actives. Whereas, the structure-based VS approaches may find complete new lead compounds by considering the target information. Thereafter, the structure-based VS approaches are more feasible in finding new lead compounds compared with the ligand-based VS approaches7,8,9,10.
Up to date, numerous strategies based on ligand-based and structure-based approaches have been proposed for VS. For instance, Sato et al. put forward a ligand-based VS strategy by combining three-dimensional molecular shape overlap method and support vector machine (SVM) to evaluate 15 drug targets and gained much better results compared with other two-dimensional structure-similarity based VS strategies11. Kong et al. developed a biologically relevant spectrum by considering the structures of the primary metabolites of organisms12, and found it effective in classifying launched drug from other phase candidates13. Our group has proposed a structure-based VS strategy by combining multiple protein structures, including crystallized structures and structures generated by molecular dynamics (MD) simulations, and machine leaning approaches6,14. Besides, we have also developed a unique structure-based VS approach by combining residue-ligand interaction matrix (also known as Molecular Interaction Energy Components, MIEC) and SVM to discriminate the binding peptides from the non-binders for protein modular domains15, and the prediction results have been validated by various experiments16,17. Since the residue-ligand interaction network can totally reflect the binding specificity of a ligand to the target, we can construct the classification models based on machine learning approaches to discriminate small molecular actives from non-actives. Fortunately, some pioneering work have engaged in this subject, for example, Ding et al. have evaluated the performance of MIEC-SVM in discriminating strong inhibitors of HIV-1 protease from a large database (ZINC database)18 and they have successfully predicted the binding of a series of HIV-1 protease mutants to drugs19. Nevertheless, the performance of MIEC-SVM needs to be assessed by the predictions to more drug targets and validated by real experiments. Moreover, this approach is parameter-dependent, and therefore the strategy to generate the best MIEC-SVM model needs to be addressed. Here, in conjunction with molecular docking, ensemble minimization, MM/GBSA free energy decomposition, and parameters tuning of SVM kernel function, we discussed how to construct a highly performed MIEC-SVM model in three kinase targets (Fig. 1). The best performed MIEC-SVM model for the ALK system was then used for VS, and the experimental results showed that the optimized MIEC-SVM model had markedly improved screening performance compared with the traditional molecular docking method.
(a) molecular docking, the most contributed residues were colored in orange; (b) residue decomposition, two strategies were used here: the top 1 docking pose was directly used for energy decomposition; and the top three docking poses were at first rescored by MM/GBSA approach, and then the best rescored docking pose was used for the decomposition analysis; (c) MIEC matrix construction, different combinations of energy components and top contributed residues were used for the matrix construction; (d) hyper-parameters optimization, c and γ were tuned using the grid searching approach and the corresponding MCC values were colored from blue (bad performance) to red (good performance); (e) model evaluation, the ROC curve, inhibitor probability, and Pearson correlation coefficient were employed for the model evaluation; (f) experimental testing, compound activity enrichment, enzyme inhibitory rate distribution, and the IC50 curves were used for the comparison of the methodologies.
Materials and Methods
Dataset Preparation and Processing
To summarize the best strategy for the MIEC-SVM construction, three tyrosine kinase targets were at first used for the evaluation, namely ABL (Abelson tyrosine kinase), ALK (Anaplastic lymphoma kinase), and BRAF (v-Raf murine sarcoma viral oncogene homolog B). The crystal structures of 2HYY (for ABL)20, 3LCS (for ALK)21, and 3IDP (for BRAF)22, were employed for the evaluation due to the good performance of Autodock in reproducing the binding modes of their co-crystallized ligands as shown in Table S1 in Supporting Information. All the inhibitors with IC50 (Ki) < 10 μM were obtained from the BindingDB database23. In total, 286, 342, and 402 inhibitors were collected for ABL, ALK, and BRAF, respectively. Although DUD dataset24 and other strategies25 have been proposed for the decoys construction, we did not try to use these strategies to avoid constructing models with limited range of the chemical properties. Thereby, 7000 compounds randomly chosen from the ChemBridge database by using the Find Diverse Molecules protocol in Discovery Studio 2.5 were used as non-inhibitors (or background molecules). The structural diversity was shown in Figure S1, where the structural similarity was calculated between the training dataset and the test dataset for the known inhibitors (Figure S1 A–C) and non-inhibitors (Figure S1G), and also between the known inhibitors and non-inhibitors for each target (Figure S1D–F). To the end, the ratio between inhibitors and non-inhibitors is approximate 1:24, 1:20, and 1:17 for ABL, ALK, and BRAF, respectively.
Molecular Docking
Autodock 4.226 with Lamarckian genetic algorithm (LGA)27 was employed for the docking mode selection due to its good performance of reproduction capability28,29,30, Before molecular docking, the protein targets were prepared with the Structure Preparation Tool module in Sybyl-X1.1, which added hydrogen atoms, repaired side-chains of the imperfect crystallized residues, and optimized the steric hindrance of side-chains. The protonation states of the proteins were determined by using PROPKA (version 3.1)31. Autodock4 atomic radii and Gasteiger partial charges32 were assigned to the macromolecules and the small molecules in molecular docking. The conformation selection space of a ligand was set to 18.75 × 18.75 × 18.75 Å3 (corresponding to 50 × 50 × 50 grids, with each grid 0.375 Å in length) around the binding pocket for each target. Each ligand was docked for 10 times to meet the demand of retrieving the top three docking poses with sufficient selection space.
Molecular Mechanics Optimization
Prior to molecular mechanics optimization, the ligand-protein systems were constructed with antechamber and tleap modules33 in Amber12 simulation package34. AM1-BCC charges35 were calculated for the small molecules by using sqm module in Amber12 due to its good performance and low computational cost36,37. The cutoff value was set to 8 Å to handle the short range electrostatic and van der Waals interactions, while the Particle mesh Ewald (PME) algorithm was employed to deal with the long-range electrostatic interactions38. Amber03 force filed39 and General Amber force field (GAFF)40 were used for the proteins and small molecules, respectively. Counter-ions of Na+and Cl− were added to neutralize the unbalanced charges of the systems. Octahedral-shaped TIP3P water box41 was added for each ligand-protein complex with 5 Å extended out of the solute to save the computational resources. Three phases of minimization were used to optimize each ligand-protein system. In the beginning, 50 kcal/mol · Å2 elastic constant was used to constrain the backbone atoms of protein for 1000 cycles (500 cycles of steepest descent and 500 cycles of conjugate gradient minimization); then, the elastic constant was decreased to 10 kcal/mol · Å2 for 1000 cycles; finally, the whole system was relaxed without any constrain for 3000 cycles. The optimized structure was submitted for free energy decomposition to derive the energy components.
Molecular Interaction Energy Components (MIEC) Matrix Calculation
The classifiers based on MIEC have been found effective in discriminating the known binding peptides from non-binders for protein modular domains in the previous studies16,18,42,43, Here, the MM/GBSA free energy decomposition approach was employed for the MIEC matrix construction based on the optimized structures. The residue-ligand interactions can be expanded as following:

where ΔGresidue-ligand denotes the total interaction energy between a residue and a ligand, which is composed of four terms: the van der Waal interaction (ΔGvdW), the electrostatic interaction (ΔGele), the polar part of solvation energy (ΔGGB), and the non-polar part of solvation energy (ΔGSA). The modified GB model developed by Onufriev et al. was employed for the polar solvation energy calculation44, and the ICOSA algorithm was employed to estimate the non-polar part of solvation energy45. Due to the good performance of a relatively higher interior dielectric constant in kinase systems46,47, all the free energy decompositions were performed under the interior dielectric constant of 4 (εin = 4). Here, the MIEC matrices were constructed by using two strategies based on the top 1 docking pose and the best of the top three docking poses due to the fact that the rescoring process may re-rank the originally scored binding modes of the small molecules48,49. For the later strategy, MM/GBSA was at first used for rescoring of the top three docking poses (εin = 4)47, and then, the best rescored binding pose (of the three docking poses) of each system was employed for MM/GBSA decomposition.
MIEC-SVM Model Construction
The energy components were selected and combined (such as the combinations of ΔGele and ΔGvdW; ΔGGB and ΔGSA; ΔGele and ΔGGB; ΔGvdW and ΔGSA; and ΔGele, ΔGvdW, ΔGGB, and ΔGSA, as shown in Tables S2–S4) to construct the MIEC matrices, where the energy components of the vital residues for the inhibitor binding, i.e. the top 20, 25, and 30 contributed residues (that is, 40, 50, and 60 vectors for the combinations of ΔGele and ΔGvdW, ΔGGB and ΔGSA, ΔGele and ΔGGB, ΔGvdW and ΔGSA, and 80, 100, and 120 vectors for the combination of ΔGele, ΔGvdW, ΔGGB, and ΔGSA; and the top residues were chosen by adding all the residue-ligand interaction pairs for all the known inhibitors, averaging the total energies for each residue-ligand pair according to the number of known inhibitors, and sorting the averaged residue-ligand pair for each target), were selected as the eigenvectors for the classification of inhibitors and non-inhibitors. Herein, the SVM algorithm50,51 implemented in libsvm package52 was employed for the model construction. Although SVM has been widely used in the scope of drug design due to the high accuracy of nonlinear fitting53,54,55, it may miss the best model if without the hyper-parameters optimization, such as when using the Radial Basis Function (RBF) as the kernel function, the parameters cost (c, which scales the penalty of classifying the samples to a false group) and gamma (γ, which regulates the inverse radius of influence of the training samples) should be adjusted carefully because they may significantly affect the classification performance56. Herein, RBF was also used as the kernel function, and grid searching was employed for the parameter tuning for the purpose of finding the best Matthews correlation coefficient (MCC) of the training dataset. The c and γ values were designed exponentially growing against 2, namely 2n, where n denotes c or γ and goes from −2 to 10 and −10 to 2, respectively. The grid space was set to 0.5 for both c and γ. Thereby, a total of 525 models were constructed for each protocol (Fig. 2, which will be discussed below). In searching different values of c and γ, different MCC values will be produced (as shown in Fig. 2), and the c and γ values corresponding to the highest MCC value were termed as best c and best γ, respectively, which were used for the external test dataset validation (Tables S2–S4). Here, half of the molecules (half inhibitors and non-inhibitors for each target) were randomly selected as the training set for model construction and the remaining molecules were used as the external test set for model verification. To assess the statistical significance of the classifiers, 5-fold cross-validation was employed for the validation of each model (or grid). Due to the unbalance of the inhibitors and non-inhibitors (~1:20), a higher weight (1.2) was set for the inhibitors to balance the classification42.
To evaluate the performance of the MIEC-SVM models, besides the MCC values, the sensitivity (SE), specificity (SP), prediction accuracies for inhibitors (Q+) and non-inhibitors (Q−) shown in Equations (2)~(6) were also considered for the comparison.





where TP and FP denote the number of true positives (inhibitors) and false positives, and TN and FN represent the number of true negatives (non-inhibitors) and false negatives. Moreover, the AUC value (area under curve) of the ROC (receiver operating characteristic) curve was also employed to quantitatively evaluate the prediction accuracy of each model based on the probability of a molecule to be an inhibitor given by the support vectors. The workflow of the data processing, model construction, and experimental testing was briefly summarized in Fig. 1.
Virtual Screening and Compounds Selection
To evaluate the performance of the optimized MIEC-SVM model in real experiments, the ALK system was used for the experimental testing. Here, the best performed MIEC-SVM model (model 15 in Table S3 with top 1 docking pose) for the ALK system was used to virtually screen the Specs database (containing ~220,000 compounds). Due to the high computational cost of the MIEC matrix calculation, herein, a hierarchical strategy was used for virtual screening: (1) all the compounds in Specs were docked into the binding site of ALK by Autodock 4.2 and scored by the Autodock scoring function; (2) the top 30,000 molecules ranked by the Autodock score were extracted for the MIEC matrix calculation and scored by the optimized MIEC-SVM model; (3) the top 300 compounds ranked by the MIEC-SVM model (inhibitor-probability derived from the SVM algorithm) and Autodock score were then respectively filtered by Lipinski’s “rules of five” (compounds with violation number ≥ 2 were eliminated) and the drug-likeness model developed by our previous study57,58; (4) in order to maximize the chemical diversity of the collected compounds for bioassays, the remaining compounds were structurally clustered, and the compounds with the Tanimoto similarity matrix computed from the MACCS structural keys higher than 0.80 were clustered into the same group59. The top molecule in each group was then sorted by the inhibitor-probability based on the MIEC-SVM model or the docking score based on Autodock. Finally, the top 50 compounds in each group (ranked by the MIEC-SVM probability and the Autodock docking score) available from Specs were purchased for experimental testing (purity ≥ 95%, confirmed by Specs, Table S5).
Reagents and Materials for Bioassays
All reagents and anhydrous solvents were obtained from commercial sources and used as received. The positive control inhibitors of ALK, crizotinib and ceritinib, were purchased from Bangshunda Technology and Selleck Chemicals, respectively. The compounds were dissolved in 100% dimethyl sulfoxide (DMSO) as a 10 mM stock solution. The final DMSO concentration in each reaction was less than 1%. Purified recombinant human ALK protein (Catalog number: PV3867) as well as reagents for TR-FRET assay, including Lantha Screen™ Tb-PY20 (Catalog number: PV3552), Fluorescein-Poly GT (Catalog number: PV3610), ATP (Catalog number: PV3227), Kinase Quench Buffer (Catalog number: P2832), Kinase Buffer (Catalog number: PV3189), and Antibody Dilution Buffer (Catalog number: PV3574), were all obtained from Life Technologies Inc.
In vitro Inhibitory Activity Assay of ALK
Lantha Screen™ kinase assay based on TR-FRET technology was used to measure the inhibitory activity of the screened compounds. All the assays were carried out in 384-well plate format. The 4 × test compounds were firstly prepared before the enzyme reaction starts. The 4 × recombinant human ALK protein and 2 × Substrate/ATP mixture were separately prepared in 50 mM HEPES (pH = 7.5), 0.01% BRIJ-35, 10 mM MgCl2, 4 mM MnCl2, 1 mM EGTA, and 2 mM DTT. The final 10 μL kinase reaction consists of 5 μL 1 × Substrate/ATP mixture (0.2 μM substrate and 5 μM ATP), 2.5 μL ALK (5 ng/ml ALK protein), and 2.5 μL 1 × test compounds with desired concentration. The assay plate was shook on a plate shaker for 30 seconds to mix the reactions thoroughly. After 1 hour kinase reaction incubation at room temperature (20 ~ 25 °C), 10 μL of pre-prepared 20 mM EDTA and 4 nM Tb-labeled antibody solution was then added to terminate the kinase reactions and to initiate antibody binding, and the assay plate was incubated for another 1 hour at room temperature. Then, the assay plate was placed into a fluorescence plate reader (BioTek Synergy™ 4) to measure both fluoresce in and terbium emission signals (excitation: 340 nm; emission: 520 and 495 nm, respectively) with 100 μs delay time and 200 μs integration time. To determine the IC50 values, the resulted inhibitory activity calculated from TR-FRET emission ratio (i.e. fluoresce in emission intensity/terbium emission intensity) was plotted against the concentration of inhibitor, and the data was fitted to a dose-response curve with a variable slop.
Results and Discussion
Vital Residues for Model Construction
To construct an effective classification model, distinguishable features should exist between the positive and negative samples, though it may be hard to be discriminated by simple observations. In the spirit of MIEC-SVM model, it considers only the binding specificity of the known inhibitors and non-inhibitors rather than the chemical structures of the known inhibitors. The model can discriminate the specific energetic spectra of the systems with even very little difference, such as a series of similar drug derivatives against a same protein60,61,62, a same drug against homologous proteins (drug selection)63,64 or protein mutants (drug resistance)65,66,67, etc. Therefore, the MIEC-SVM model is superior to most of the ligand/structure-based methods that usually bias the models to screen some known structures in the training set.
Here, the energetic contributions based on vital ligand-residue pairs were employed as the classification features to discriminate the inhibitors from non-inhibitors. The energy contributions of the top 11 contributed residues (within ~4 Å of the co-crystallized ligands for the three targets) to the inhibitors and non-inhibitors for the three targets were averaged and shown in Figure S2. Apparently, the energy contributions of the vital residues to the inhibitors (red bar) are always larger than those to the non-inhibitors (green bar) for all the three systems, implying that it is intrinsically distinguishable between the inhibitors and non-inhibitors by using the molecular interaction energy components contributed from the vital residues.
Importance of Tuning Hyper-parameters in MIEC-SVM Model Construction
As has been discussed above, two hyper-parameters (c and γ) may significantly affect the prediction performance of the models when using the RBF kernel function56. However, numerous studies ignored this key step by just using the default parameters (such as in libsvm c = 0 and γ = log2(1/n_features), with 2n in unit)18,19,42,43,54, and thereafter may loss the best model. Here by using the grid searching approach, the c and γ values were optimized by finding the grid with the highest MCC value to construct the best prediction models. Afterward, the best models were validated by the predictions to the prior prepared test sets. As shown in Fig. 2, remarkable difference was observed of the MCC values when using different combinations of the c and γ values for the training set. The MCC values of the training set vary from 0.2 (blue grids) to 0.7 (red grids) for the different groups of c and γ (grids were not colored when the predicted MCC less than 0.2). By using different combinations of energy components and docking poses (will be discussed in the following), 30 best performed MIEC-SVM models were constructed for each target as shown in Tables S2–S4. Interestingly, there is no best models located in the grid of c = 0 (or 2c = 1), implying that it may miss the best choice to construct the SVM classifiers based on the default parameters.
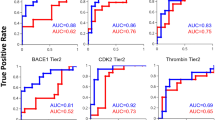
The correlation of the top 30 models for each target is plotted in Figure S3. Although the MCC values are large for the models based on the different combinations of the selected classification features (as illustrated in Tables S2–S4), high correlation coefficients were found across all the systems (r = 0.7 ~ 0.9), meaning that the models in each combination are stable and not over-fitted. Thereby, it is reliable to analyze the predicted results of the external test set based on the best model for each target (herein, the best model was chosen based on the highest training set MCC values for each target). The probability of a molecule to be an inhibitor was estimated by the SVM model as shown in Fig. 3A1–A3, where model 15 of ABL (based on the top 1 docking pose strategy), model 5 of ALK (based on the best of the top three docking poses strategy), and model 15 of BRAF (based on the best of the top three docking poses strategy) are plotted. It can be found that most of the non-inhibitors (blue cycles) are located at the bottom of the figures (Fig. 3A1–A3), suggesting that a large part of the molecules are recognized as in-actives, which is consistent with the fact that only few compounds natively show activities to the drug target. The corresponding ROC curve for each test set was also calculated based on the probability as shown in Fig. 3B1–B3, where the inflection points are shown in green dot lines (the inflection points were measured by 1% false positive rate of the test set as the fact that actives always exist in the chemical background, though the ratio is very low, usually < 1%).
The probability of a molecule to be an inhibitor is plotted in panel (A), where the inhibitors and the non-inhibitors are colored in red and blue, respectively. The ROC curves based on SVM probabilities (the inflection points are shown in green dot line) and docking scores (the inflection points are shown in red dot line) are illustrated in panel (B,C), respectively. The inflection points were measured by 1% false positive rate (Considering the test set contains 3500 non-inhibitors, the point with 35 non-inhibitors classified into the inhibitor group was used to determine the inflection point position).
To give a comparison, the ROC curve for the test set based on the top 1 docking scores of the inhibitors and non-inhibitors were also plotted for each target (Fig. 3C). As shown in Fig. 3B,C, the AUC values based on the best MIEC-SVM models (0.866 for ABL, 0.937 for ALK, and 0.892 for BRAF, Fig. 3B1–B3) are all significantly higher than the corresponding results based on the top 1 docking scores (0.848 for ABL, 0.898 for ALK, and 0.816 for BRAF, Fig. 3C1–C3), with the AUC values increased by ~2% for ABL, ~4% for ALK, and ~8% for BRAF. Besides, it shows that the inflection points of the ROC curves based on the SVM probabilities (green dot lines in Fig. 3B) are also much higher than the corresponding docking results (red dot lines in Fig. 3C) (0.4 ~ 0.5 versus 0.1 ~ 0.3 for the MIEC-SVM models and molecular docking, respectively), meaning that there is more opportunity to find more inhibitors with much lower false positive rate by using the hyper-parameters-tuned MIEC-SVM models. Taken all, considering that a little improvement of the model accuracy (i.e. 1%) will remarkably decrease the false positive rate of VS using a large database, it should be a good choice to use the hyper-parameters-tuned MIEC-SVM models for structure-based VS.
High Quality Model Construction by using More Energy Components
As the fact that the performance of a prediction model is usually affected by multiple factors such as the selection of different combination of feature vectors14,42,56,68, apart from tuning the kernel function parameters, we also optimized the MIEC-SVM models by considering different combinations of the feature components, such as different number of most contributed residues (top 20, 25, and 30 residues), different docking poses (top 1 docking pose and the best of the top three docking poses), and different energy components (ΔGele and ΔGvdW; ΔGGB and ΔGSA; ΔGele and ΔGGB; ΔGvdW and ΔGSA; and ΔGele, ΔGvdW, ΔGGB, and ΔGSA). Although Ding et al. found that the use of the top 30 most contributed residues may be the best choice to construct the MIEC-SVM models for the HIV-1 protease18, we show here that there is no remarkable difference when using the top 20, 25, or 30 most contributed residues to construct the MIEC-SVM models for the tyrosine kinase systems. As shown in Fig. 2, similar distribution patterns of the MCC values were found across all the groups using different numbers of top contributed residues (here, the three panels in each line within a same target, containing the top 20, 25 and 30 most contributed residues, was considered as a group), indicating that the MIEC-SVM models are not too sensitive to how many top contributed residues are used for model construction (at least for the case that more than 20 top contributed residues were used).
Different from the issue of selecting how many most contributed residues for model construction, the MIEC-SVM models are more sensitive to which energy components are used. As shown in Tables S2–S4, the MCC values are quite different for the training sets. The models based on the combinations using all the four energy components, namely ΔGele, ΔGvdW, ΔGGB, and ΔGSA, have better performance than those based on the other combinations of energy components. For instance, in the system of BRAF, the MCC values increase from 0.44 for the combination of ΔGGB and ΔGSA (model 4 in Table S4) to 0.63 for the combination of the four energy components (model 5 in Table S4). Moreover, as shown in Tables S2–S4, the combinations of ΔGele and ΔGvdW always yield better models than those of ΔGele and ΔGGB across all the constructed models. The reason why the combinations of ΔGele and ΔGGB always perform worse than those of ΔGele and ΔGvdW may be attributed to the high correlation between the electrostatic interactions (ΔGele) and the polar part of solvation energies (ΔGGB) upon the binding of small molecules. As shown in Figure S4A1–C1, very high negative correlations (r = −0.80 ~ −0.95) between the electrostatic interactions and the polar part of solvation energies exist for most contributed residues among all the three systems (the energies were calculated by summing up the energy components of the most contributed residues in Figure S2 for all the inhibitors and non-inhibitors), implying that much information derived from the features is redundant and it is not sufficient to use just the two features (ΔGele and ΔGGB) to construct models. On the contrary, there is no obvious correlations (r = −0.11 ~ 0.16) between the electrostatic (ΔGele) and van der Waals (ΔGvdW) interactions for the three systems as shown in Figure S4A2–C2, thereby supplying sufficient information for model construction. Nevertheless, the more energy components are used, the more information of features can be included, and the higher quality models can be constructed. One may argue that it may be a better way to normalize or reduce dimensions (such as using principle component analysis, PCA) for the model construction. Considering that the energies calculated here are all based on the same method (or calculating framework) with the same unit (kcal/mol), to keep the explicit physical meaning, we would like not to use any scaling or dimension reduction methods for the model construction.
The Performance of Using More Docking Poses for Models Construction
As the rescoring process (MM/GBSA) may re-rank the docking poses derived from the original docking results47,49, we rescored the top three docking poses by the MM/GBSA approach (with the top rescored poses used for model construction). It is well known that docking with multiple pocket conformations derived from different crystal structures is usually superior to one using a single pocket conformation6,14. However, it will be much time consuming to construct MIEC-SVM models in considering multiple crystal structures. Nevertheless, the optimization of the ligand-receptor complex in the MIEC-SVM model construction actually introduced induced-fit effect upon the ligand binding, which may be superior to the models constructed based on rigid receptor docking. Thereby, herein, two strategies were used for model construction: (1) the top 1 docking poses derived from Autodock were directly decomposed into residue-ligand pairs for model construction; and (2) the top three docking poses were rescored by the MM/GBSA approach at first, and then, the top 1 rescored poses were submitted to energy decomposition for model construction. As shown in the lower half of Fig. 2, which denotes the models derived from the best of the top three docking poses, similar patterns of the MCC distributions were found for the models based on the top 1 docking poses (the upper half of Fig. 2), meaning that the parameter optimization will not depend on which docking pose is used, but the MCC values are very different when using different docking poses. For instance, the best models derived from the top 1 docking pose strategy are mostly better than those derived from the best of the top three docking poses strategy in the system of ABL (Table S2), whereas, in the system of BRAF, the best models derived from the multiple docking poses strategy always perform better than those derived from the top 1 docking pose strategy (Table S4). Thereby, it may be system-specific whether to use multiple docking poses for MIEC-SVM model construction.
Comparison of the Optimized MIEC-SVM Model and the Traditional Molecular Docking Method based on Experimental Testing
The above issues have discussed how to generate the best MIEC-SVM models for virtual screening. To test whether the optimized MICE-SVM model is really better than the traditional molecular docking methods, the best-performed MIEC-SVM model for the ALK system (model 15 in Table S3) was employed to virtually screen the Specs database. Before experimental testing, we analyzed and compared the distributions of three important molecular properties (molecular weight, octanol/water partition coefficient, and aqueous solubility) for the top 300 molecules predicted by MIEC-SVM and those predicted by Autodock. As shown in Fig. 4, the molecular properties of the top 300 molecules predicted by MIEC-SVM are much closer to the averaged properties of the known ALK inhibitors (red dot lines in Fig. 4). Besides, we analyzed the violation number of Lipinski’s “rules of five” for the top molecules predicted by MIEC-SVM and Autodock. Apparently, as shown in Table 1, most of the top 300 molecules predicted by MIEC-SVM are drug-like compounds (86% with the violation number ≤ 1), while most of the top 300 molecules predicted Autodock are non-drug-like compounds (with only 30.3% exhibiting violation number ≤ 1), suggesting that the molecules predicted by MIEC-SVM are more drug-like than those predicted by Autodock.
The averaged values of the corresponding chemical properties of the known ALK inhibitors were shown in red dot lines. The distributions of the molecular weight (MW), the predicted octanol/water partition coefficient (logP), and the predicted aqueous solubility (logS) were shown in panels (A–C), respectively.
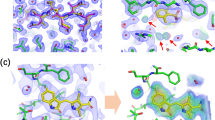
Thereafter, the top 50 molecules in each strategy (MIEC-SVM and Autodock methods) remained from the drug-likeness filtering and structural clustering were purchased and submitted to enzyme-based bioassay. Overall, 7 out of 50 (14%) tested compounds chosen by the MIEC-SVM model exhibited remarkable ALK inhibitory activity with IC50 < 10 μM (4 molecules showed nM level of activity as shown in Table 2), which is significantly higher than those chosen by Autodock (3 out of 50 molecules, namely hit rate of 6%, and 2 molecules in nM level of activity as shown in Table 2) and consistent with the conclusion that MIEC-SVM model usually goes with higher enrichment ratio (Fig. 3). Moreover, the averaged ALK inhibitory ratio of the tested molecules chosen by MIEC-SVM is also significantly higher than that chosen by Autodock (15% versus 9.7%). The binding modes, chemical structures, and IC50 curves of the 10 actives (IC50 < 10 μM) are shown in Fig. 5.
Besides, to evaluate the novelty of these identified inhibitors with respect to known ALK inhibitors, the pairwise Tanimoto similarity indices based on the FCFP_6 fingerprints for these inhibitors in Table 2 with the known ALK inhibitors obtained from the BindingDB database were calculated through the Find Similar Molecules by Fingerprints protocol in Discovery Studio 2.5. It can be found in Table 2 that most of the inhibitors identified by the MIEC-SVM model have low Tanimoto similarity to the known ALK inhibitors (most molecules < 0.2, with only one molecule > 0.3, Table 2), exactly exhibiting the fact that the MIEC-SVM model considers more about the binding specificity of the small molecules rather than the chemical structure of the known inhibitors.
Conclusion
By using multiple parameters tuning strategy, we systemically evaluated the performance of MIEC-SVM models in discriminating small molecule kinase inhibitors from non-inhibitors. We found that the optimization of the hyper-parameters embedded in the kernel function of SVM is always necessary since the default parameters cannot give the best result for any case of the studied systems. However, it is system-specific whether to use multiple docking poses for MIEC-SVM model construction. Besides, the MIEC-SVM models are not too sensitive to how many energy components are used, such as the models based on the MIEC matrices generated from of the top 20, 25, or 30 residues yield similar prediction capabilities. Whereas, the models are very sensitive to which energy components are employed for model construction, for example, the models based on the combinations of ΔGele, ΔGvdW, ΔGGB, and ΔGSA performed much better than those based on the other reduced combinations. By using the hyper-parameters-tuned MIEC-SVM model, we successfully found 7 significant inhibitors of ALK (IC50 < 10 μM) in 50 purchased compounds (with 4 in nM level), suggesting that the hyper-parameters-tuned MIEC-SVM model is a powerful tool for structure-based virtual screening.
Additional Information
How to cite this article: Sun, H. et al. Constructing and Validating High-Performance MIEC-SVM Models in Virtual Screening for Kinases: A Better Way for Actives Discovery. Sci. Rep. 6, 24817; doi: 10.1038/srep24817 (2016).
References
Hou, T. & Xu, X. Recent development and application of virtual screening in drug discovery: an overview. Curr. Pharm. Des. 10, 1011–1033 (2004).
Yan, C. et al. Discovery and characterization of small molecules that target the GTPase Ral. Nature 515, 443–447 (2014).
Keserü, G. M. & Makara, G. M. The influence of lead discovery strategies on the properties of drug candidates. Nat. Rev. Drug Discov. 8, 203–212 (2009).
Bajorath, J. Integration of virtual and high-throughput screening. Nat. Rev. Drug. Discov. 1, 882–894 (2002).
Klebe, G. Virtual ligand screening: strategies, perspectives and limitations. Drug Discov. Today 11, 580–594 (2006).
Tian, S. et al. Development and evaluation of an integrated virtual screening strategy by combining molecular docking and pharmacophore searching based on multiple protein structures. J. Chem. Inf. Model. 53, 2743–2756 (2013).
Chen, S. et al. Identifying Novel Selective Non-Nucleoside DNA Methyltransferase 1 Inhibitors through Docking-Based Virtual Screening. J. Med. Chem. 57, 9028–9041 (2014).
Xu, L. et al. Discovery of novel inhibitors targeting the macrophage migration inhibitory factor via structure-based virtual screening and bioassays. J. Med. Chem. 57, 3737–3745 (2014).
Ma, D.-L. et al. Virtual screening and optimization of Type II inhibitors of JAK2 from a natural product library. Chem. Commun. 50, 13885–13888 (2014).
Shen, M. et al. Discovery of Rho-kinase inhibitors by docking-based virtual screening. Mol. Biosyst. 9, 1511–1521 (2013).
Sato, T. et al. Application of support vector machine to three-dimensional shape-based virtual screening using comprehensive three-dimensional molecular shape overlay with known inhibitors. J. Chem. Inf. Model. 52, 1015–1026 (2012).
Kong, D.-X. et al. Do biologically relevant compounds have more chance to be drugs? J. Chem. Inf. Model. 49, 2376–2381 (2009).
Deng, Z.-L. et al. Exploring the biologically relevant chemical space for drug discovery. J. Chem. Inf. Model. 53, 2820–2828 (2013).
Tian, S. et al. Assessing an ensemble docking-based virtual screening strategy for kinase targets by considering protein flexibility. J. Chem. Inf. Model. 54, 2664–2679 (2014).
Hou, T. et al. Characterization of domain-peptide interaction interface: a case study on the amphiphysin-1 SH3 domain. J. Mol. Biol. 376, 1201–1214 (2008).
Hou, T. et al. Characterization of Domain-Peptide Interaction Interface A Generic Structure-based Model to Decipher the Binding Specificity of SH3 Domains. Mol. Cell Proteomics. 8, 639–649 (2009).
Xu, Z. et al. Proteome-wide detection of Abl1 SH3-binding peptides by integrating computational prediction and peptide microarray. Mol. Cell Proteomics. 11, O111. 010389 (2012).
Ding, B. et al. Characterization of small molecule binding. I. Accurate identification of strong inhibitors in virtual screening. J. Chem. Inf. Model. 53, 114–122 (2013).
Ding, B. et al. Characterizing Binding of Small Molecules. II. Evaluating the Potency of Small Molecules to Combat Resistance Based on Docking Structures. J. Chem. Inf. Model. 53, 1213–1222 (2013).
Cowan-Jacob, S. W. et al. Structural biology contributions to the discovery of drugs to treat chronic myelogenous leukaemia. Acta Crystallogr. D Biol. Crystallogr. 63, 80–93 (2006).
Lee, C. et al. Crystal structure of the ALK (anaplastic lymphoma kinase) catalytic domain. Biochem. J. 430, 425–437 (2010).
Smith, A. L. et al. Selective inhibitors of the mutant B-Raf pathway: discovery of a potent and orally bioavailable aminoisoquinoline. J. Med. Chem. 52, 6189–6192 (2009).
Liu, T. et al. BindingDB: a web-accessible database of experimentally determined protein–ligand binding affinities. Nucleic Acids Res. 35, D198–D201 (2007).
Mysinger, M. M. et al. Directory of useful decoys, enhanced (DUD-E): better ligands and decoys for better benchmarking. J. Med. Chem. 55, 6582–6594 (2012).
Lagarde, N. et al. Benchmarking Data Sets for the Evaluation of Virtual Ligand Screening Methods: Review and Perspectives. J. Chem. Inf. Model. 55, 1297–1307 (2015).
Morris, G. M. et al. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 30, 2785–2791 (2009).
Morris, G. M. et al. Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J. Comput. Chem. 19, 1639–1662 (1998).
Sun, H.-Y. et al. Structural and energetic analyses of SNPs in drug targets and implications for drug therapy. J. Chem. Inf. Model. 53, 3343–3351 (2013).
Sun, H.-Y. et al. Finding chemical drugs for genetic diseases. Drug Discov. Today 19, 1836–1840 (2014).
Hou, X. et al. How to improve docking accuracy of AutoDock4.2: A case study using different electrostatic potentials. J. Chem. Inf. Model. 53, 188–200 (2013).
Søndergaard, C. R. et al. Improved treatment of ligands and coupling effects in empirical calculation and rationalization of pKa values. J. Chem. Theory Comput. 7, 2284–2295 (2011).
Gasteiger, J. & Marsili, M. Iterative partial equalization of orbital electronegativity—a rapid access to atomic charges. Tetrahedron 36, 3219–3228 (1980).
Wang, J. et al. Automatic atom type and bond type perception in molecular mechanical calculations. J. Mol. Graph. Model. 25, 247–260 (2006).
Case, D. et al. AMBER 12. University of California, San Francisco (2012).
Jakalian, A. et al. Fast, efficient generation of high-quality atomic charges. AM1-BCC model: II. Parameterization and validation. J. Comput. Chem. 23, 1623–1641 (2002).
Xu, L. et al. Assessing the Performance of MM/PBSA and MM/GBSA Methods. 3. The Impact of Force Fields and Ligand Charge Models. J. Phys. Chem. B. 117, 8408–8421 (2013).
Tsai, K.-C. et al. A comparison of different electrostatic potentials on prediction accuracy in CoMFA and CoMSIA studies. Eur. J. Med. Chem. 45, 1544–1551 (2010).
Darden, T. et al. Particle mesh Ewald: An N log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Wang, J. et al. How well does a restrained electrostatic potential (RESP) model perform in calculating conformational energies of organic and biological molecules? J. Comput. Chem. 21, 1049–1074 (2000).
Wang, J. et al. Development and testing of a general amber force field. J. Comput. Chem. 25, 1157–1174 (2004).
Jorgensen, W. L. et al. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Hou, T. et al. Characterization of domain–peptide interaction interface: prediction of SH3 domain-mediated protein–protein interaction network in yeast by generic structure-based models. J. Proteome Res. 11, 2982–2995 (2012).
Li, N. et al. Characterization of PDZ domain-peptide interaction interface based on energetic patterns. Proteins: Struct., Funct., Bioinf. 79, 3208–3220 (2011).
Onufriev, A. et al. Exploring protein native states and large-scale conformational changes with a modified generalized born model. Proteins: Struct., Funct., Bioinf. 55, 383–394 (2004).
Gohlke, H. et al. Insights into protein-protein binding by binding free energy calculation and free energy decomposition for the Ras-Raf and Ras-RalGDS complexes. J. Mol. Biol. 330, 891–914 (2003).
Sun, H. et al. Assessing the Performance of MM/PBSA and MM/GBSA Methods. 4. Accuracies of MM/PBSA and MM/GBSA Methodologies Evaluated by Various Simulation Protocols using PDBbind Data Set. Phys. Chem. Chem. Phys. 16, 16719–16729 (2014).
Sun, H. et al. Assessing the performance of MM/PBSA and MM/GBSA methods. 5. Improved docking performance using high solute dielectric constant MM/GBSA and MM/PBSA rescoring. Phys. Chem. Chem. Phys. 16, 22035–22045 (2014).
Sgobba, M. et al. Application of a post-docking procedure based on MM-PBSA and MM-GBSA on single and multiple protein conformations. Eur. J. Med. Chem. 58, 431–440 (2012).
Zhang, X. et al. Towards fully automated high performance computing drug discovery: A massively parallel virtual screening pipeline for docking and MM/GBSA rescoring to improve enrichment. J. Chem. Inf. Model. 54, 324–337 (2013).
Vapnik, V. The nature of statistical learning theory. Springer Science & Business Media, (2013).
Cortes, C. & Vapnik, V. Support-vector networks. Machine learning 20, 273–297 (1995).
Chang, C.-C. & Lin, C.-J. LIBSVM: a library for support vector machines. ACM T. Intel. Syst. Tec. 2, 1–27 (2001).
Han, L. Y. et al. Support vector machines approach for predicting druggable proteins: recent progress in its exploration and investigation of its usefulness. Drug Discov. Today 12, 304–313 (2007).
Li, X. et al. In silico prediction of chemical acute oral toxicity using multi-classification methods. J. Chem. Inf. Model. 54, 1061–1069 (2014).
Hou, T. et al. Predicting drug resistance of the HIV-1 protease using molecular interaction energy components. Proteins: Struct., Funct., Bioinf. 74, 837–846 (2009).
Zang, Q. et al. Binary classification of a large collection of environmental chemicals from estrogen receptor assays by quantitative structure-activity relationship and machine learning methods. J. Chem. Inf. Model. 53, 3244–3261 (2013).
Walters, W. P. et al. Virtual screening-an overview. Drug Discov.Today. 3, 160–178 (1998).
Tian, S. et al. Drug-likeness analysis of traditional Chinese medicines: prediction of drug-likeness using machine learning approaches. Mol. Pharm. 9, 2875–2886 (2012).
Apweiler, R. et al. The universal protein resource (UniProt) in 2010. Nucleic Acids. Res. 38, 142–148 (2010).
Kong, X. et al. Importance of protein flexibility in ranking inhibitor affinities: modeling the binding mechanisms of piperidine carboxamides as Type I1/2 ALK inhibitors. Phys. Chem. Chem. Phys. 17, 6098–6113 (2015).
Shen, M. et al. Discovery of Novel ROCK1 Inhibitors via Integrated Virtual Screening Strategy and Bioassays. Sci. Rep. 5 (2015).
Sun, H. et al. Directly-binding rather than Induced-fit dominated binding affinity difference in (S) and (R)-crizotinib bound MTH1. J. Chem. Theory Comput. 12, 851–860 (2016).
Lin, Y.-L. & Roux, B. Computational Analysis of the Binding Specificity of Gleevec to Abl, c-Kit, Lck, and c-Src Tyrosine Kinases. J. Am. Chem. Soc. 135, 14741–14753 (2013).
Kong, X. et al. Molecular principle of the cyclin-dependent kinase selectivity of 4-(thiazol-5-yl)-2-(phenylamino) pyrimidine-5-carbonitrile derivatives revealed by molecular modeling studies. Phys. Chem. Chem. Phys. 18, 2034–2046 (2016).
Sun, H. et al. Insight into Crizotinib Resistance Mechanisms Caused by Three Mutations in ALK Tyrosine Kinase using Free Energy Calculation Approaches. J. Chem. Inf. Model. 53, 2376–2389 (2013).
Sun, H. et al. P-loop conformation governed crizotinib resistance in G2032R-mutated ROS1 tyrosine kinase: clues from free energy landscape. PLoS Comput. Biol. 10, e1003729 (2014).
Sun, H. Y. & Ji, F. Q. A molecular dynamics investigation on the crizotinib resistance mechanism of C1156Y mutation in ALK. Biochem. Biophys. Res. Commun. 423, 319–324 (2012).
Li, D. et al. ADMET evaluation in drug discovery. 13. Development of in silico prediction models for p-glycoprotein substrates. Mol. Pharm. 11, 716–726 (2014).
Acknowledgements
This study was supported by the National Science Foundation of China (21575128 and 81502982), and the National Science Foundation for Post-doctoral Scientists of China (2015M581953, 2015T80586, and 2015M581862).
Author information
Authors and Affiliations
Contributions
T.H. conceived and designed the experiments. H.S., P.P., S.T. and L.X. performed the experiments. H.S., P.P., S.T. L.X., X.K. and D.L. analyzed the data. H.S., Y.L. and T.H. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Sun, H., Pan, P., Tian, S. et al. Constructing and Validating High-Performance MIEC-SVM Models in Virtual Screening for Kinases: A Better Way for Actives Discovery. Sci Rep 6, 24817 (2016). https://doi.org/10.1038/srep24817
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep24817
This article is cited by
-
Comprehensive machine learning boosts structure-based virtual screening for PARP1 inhibitors
Journal of Cheminformatics (2024)
-
The impact of cross-docked poses on performance of machine learning classifier for protein–ligand binding pose prediction
Journal of Cheminformatics (2021)
-
Discovery of novel natural compound inhibitors targeting estrogen receptor α by an integrated virtual screening strategy
Journal of Molecular Modeling (2019)
-
Efficient iterative virtual screening with Apache Spark and conformal prediction
Journal of Cheminformatics (2018)
-
Performance of machine-learning scoring functions in structure-based virtual screening
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.