Abstract
Intestinal immune homeostasis requires dynamic crosstalk between innate and adaptive immune cells. Dendritic cells (DCs) exist as multiple phenotypically and functionally distinct sub-populations within tissues, where they initiate immune responses and promote homeostasis. In the gut, there exists a minor DC subset defined as CD103+CD11b− that also expresses the chemokine receptor XCR1. In other tissues, XCR1+ DCs cross-present antigen and contribute to immunity against viruses and cancer, however the roles of XCR1+ DCs and XCR1 in the intestine are unknown. We showed that mice lacking XCR1+ DCs are specifically deficient in intraepithelial and lamina propria (LP) T cell populations, with remaining T cells exhibiting an atypical phenotype and being prone to death and are also more susceptible to chemically-induced colitis. Mice deficient in either XCR1 or its ligand, XCL1, similarly possess diminished intestinal T cell populations and an accumulation of XCR1+ DCs in the gut. Combined with transcriptome and surface marker expression analysis, these observations lead us to hypothesise that T cell-derived XCL1 facilitates intestinal XCR1+ DC activation and migration and that XCR1+ DCs in turn provide support for T cell survival and function. Thus XCR1+ DCs and the XCR1/XCL1 chemokine axis have previously-unappreciated roles in intestinal immune homeostasis.
Similar content being viewed by others
Introduction
Intestinal immune homeostasis is regulated by a variety of innate and adaptive immune cells1, including dendritic cells (DCs)2,3. The DC population is heterogeneous, consisting of several subsets with specific functions, distinguished by characteristic patterns of surface marker expression4,5. In mice, CD103+CD11b− and CD103−CD11b+ DC subsets are present in the spleen and lymph nodes (LNs), while in the lamina propria (LP) of the intestine, an additional CD103+CD11b+ subset exists4,5. The roles of the various intestinal DC subsets are variously well-studied: CD103+CD11b+ DCs are the most abundant and are involved in generating Th17 and regulatory T (Treg) cells6,7,8,9 and anti-fungal immunity10; CD103−CD11b+ DCs are related to macrophages2 and play an immunoregulatory role via the secretion of IL-10 and TGF-β111,12; while in contrast, little is known of the role of CD103+CD11b− DCs in this site. CD103+CD11b− DCs also express the T cell co-receptor CD8α and the chemokine receptor XCR1 (lymphotactin receptor/G-protein-coupled receptor 5) and represent approximately 5% of murine intestinal DCs13,14,15,16. This subset (hereafter “XCR1+ DCs”) also exists in other tissues including spleen, LNs and skin, where it specializes in the uptake of dead cells and cross-presentation of antigen to CD8+ T cells17, which is important for protection against viruses, bacteria and parasites and for anti-tumour immunity14,16,18,19,20,21. XCR1+ DCs are also present in other mammalian species22,23; in humans, XCR1 is expressed on a DC subset that is present in various tissues, including skin and blood and possesses high cross-presenting activity24. How the XCR1+ DC subset functions in either the murine or human intestine is currently unknown.
The ligand for XCR1 is XCL1 in mice and XCL1 and XCL2 in humans25. In both species, XCL1 is produced mainly by natural killer (NK) and activated CD8+ T cells15,16,17,18,22,23. Mice lacking XCR1 or XCL1 show diminished CD8+ T cell responses against the antigens cross-presented by CD103+CD11b− DCs15; XCL1 is also involved in regulating medullary accumulation of thymic XCR1+ DCs and thymic generation of naturally-occurring regulatory T (Treg) cells26. Thus, the XCR1-XCL1 axis has the potential to modulate both the localization and function of T cells and DCs, though the extent to which this is relevant outside of the thymic environment is not yet clear.
In order to clarify the functions of XCR1+ DCs, we generated and analysed mutant mice in which these cells are constitutively ablated. These mice possessed significantly fewer intestinal T cells than their wildtype counterparts and the remaining T cells exhibited an atypical phenotype. Consistent with the regulatory roles of intestinal T cells, XCR1+ DC-deficient mice showed exaggerated manifestations during chemically-induced colitis. Alongside, in mice lacking either XCR1 or XCL1, a similar decrease in T cell populations was seen, accompanied by an accumulation of CD103+CD11b− DCs in the intestine. Thus, we have identified novel regulatory roles of XCR1+ DC and the XCR1-XCL1 axis in maintaining intestinal immune homeostasis and have proposed a hypothetical model on which to base future studies to define the underlying mechanisms.
Results
Intestinal T cell populations are specifically decreased in mice lacking XCR1+ DCs
We exploited the specific expression of XCR1 on this DC subset to generate a mouse model in which CD103+CD11b− XCR1+ DCs are constitutively ablated as a result of diphtheria toxin A subunit (DTA) expression (XCR1-DTA mice)(Supplementary Fig. 1A,B). We first confirmed the ablation of CD103+CD11b− DCs in the LP, mesenteric lymph nodes (MLNs) and spleens of XCR1-DTA mice and the retention of the other DC subsets across these tissues (Supplementary Fig. 1C). In the spleen, CD103+CD11b− DCs were more severely ablated than CD8α+CD11b− DCs, which is consistent with the finding that CD8α+CD11b− DC populations contain more XCR1− cells than CD103+CD11b− DC populations13,14 (Supplementary Fig. 1D).
We next asked whether T cell populations were affected by the absence of XCR1+ DCs. In thymus, spleen and MLNs, the size and composition of the T cell populations were comparable between control and XCR1-DTA mice (Supplementary Fig. 2A,B). Normally the intestine contains several distinct lymphocyte populations: within the LP, conventional TCRαβ+ CD4+ and CD8+ T cells predominate, while the intraepithelial lymphocyte (IEL) population additionally includes TCRγδ+ T cells and an atypical TCRαβ+ subset that expresses CD8αα instead of CD8αβ27. In the LP of XCR1-DTA mice there were approximately two-thirds fewer TCRαβ+ T cells than in control mice, with significant decreases across both CD4+ and CD8+ subpopulations (Fig. 1A,B); all IEL subsets were also significantly reduced compared to controls (Fig. 1C,D). Moreover, taking into consideration the lower T cell numbers, there was a further relative decrease in the more mature CD4+CD8αα+ IEL subset, accompanied by a relative accumulation of the less mature CD4+CD8− IELs in XCR1-DTA mice (Fig. 1C). The relative scarcity of TCRβ- expressing cells was similarly evident histologically (Fig. 1E). Thus, XCR1+ DCs are critical for keeping both intestinal LP T cells and IEL populations.
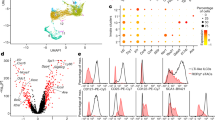
Intestinal T cell populations are decreased in mice lacking XCR1+ DCs.
(A–D) Percentages (A,C) and total numbers (B,D) of T lineage cells of LP (A,B) and IELs (C,D) from control (Xcr1+/cre) and XCR1-DTA mice. Dead cells and doublets were eliminated by FSC/SSC gating and LIVE/DEAD staining, with live cells subjected to further gating as indicated. (E) Immunofluorescent images of intestinal sections from control and XCR1-DTA mice labelled with anti-TCRβ and anti-Ep-CAM Abs. Scale bars, 500 μm. Means ± s.e.m. of five mice are indicated (B,D). (F) AnnexinV and 7-AAD staining of LP and intraepithelial TCRαβ+ cells, in control and XCR1-DTA mice. Percentage of double-positive cells is indicated. Results are representative of five (A-D) or two (E,F) independent experiments. (*P < 0.05; **P < 0.01; ***P < 0.001, Student’s t test).
Intestinal T cells are more prone to death in XCR1-DTA mice
The low numbers of intestinal T cells in XCR1-DTA mice might have been due to decreased proliferation or increased cell death. The proliferation marker Ki67 was expressed at a comparable frequency in LP T cell and IEL sub-populations from XCR1-DTA and control mice (Supplementary Fig. 3A,B). However, there were markedly more dead and/or dying intestinal TCRαβ+ T cells in mice lacking XCR1+ DCs, in particular in the IEL compartment, as assessed by frequency of Annexin V and 7-amino-actinomycin D (7-AAD) staining (Fig. 1F). The findings indicate that the T cells present in XCR1-DTA mice were not defective in proliferation, but were more prone to undergo cell death, likely contributing to the T cell deficiency observed in these mice. Thus, the absence of XCR1+ DCs compromises the survival of intestinal T cell populations.
LP T cells exhibit an abnormal phenotype in XCR1-DTA mice
We next compared the phenotype of the lymphoid tissue T cells from control and XCR1-DTA mice. In control mice, LP TCRαβ+ T cells expressed lower levels of the cell adhesion molecule CD62L and higher levels of the integrin CD103, compared with splenic and MLN TCRαβ+ T cells (Fig. 2A, shaded histograms). However, LP TCRαβ+ T cells from XCR1-DTA mice showed defects in this intestinal T cell phenotype (Fig. 2A). Expression of the gut-homing receptors α4β7 integrin and CCR928,29 was comparable between control and XCR1-DTA mice (Fig. 2A).
Intestinal T cells in mice lacking XCR1+ DCs exhibit an aberrant phenotype.
(A) Intensity of expression of CD62L, CD103, CCR9 and α4β7 integrin on CD4+ and CD8+ T cells in spleen, MLNs and LP from control (Xcr1+/cre, shaded histograms) and XCR1-DTA (open histograms with red lines) mice. Results are representative of four independent experiments. (B,C) Nanostring gene expression analysis of purified LP CD4+ T cells from control and XCR1-DTA mice. Scatter plot (B) shows means of normalized log intensities of individual probes with lines indicating the 2-fold difference threshold. Expression profiles (C) of indicated genes are shown as means ± s.e.m. of three independent experiments. Cells from three or four mice were pooled for each experiment.
We then asked whether LP T cells from XCR1-DTA and control mice further differed at the transcriptional level. LP CD4+ T cells from mice lacking XCR1+ DCs contained significantly more transcripts for Il17a, Il22 and Il4 and significantly fewer transcripts for Ifng and Il10 than those from control mice (Fig. 2B,C), suggesting polarization towards a Th2/17 profile, rather than the Th1 profile evident in control animals. Furthermore, it is notable that Runx3 expression in CD4+ T cells from XCR1-DTA mice was decreased compared with that of CD4+ T cells from control mice. A transcription factor, RUNX3, is required for transition of CD4+CD8− to CD4+CD8αα+ T cells30,31. Therefore, decrease of Runx3 expression might explain the observed block in the generation of CD4+CD8αα+ from CD4+CD8− T cells in XCR1-DTA mice (Fig. 1C). Thus, gene expression profile analysis revealed that LP T cell surface phenotype and gene expression profile is profoundly altered in mice lacking XCR1+ DCs.
Given the known role of XCL1 in the thymic generation of natural Treg cells26 we next investigated whether the intestinal Treg population was affected in XCR1-DTA mice. The proportion of Treg cells, defined as CD25+Foxp3+, among CD4+ T cells and the expression of the natural Treg marker, Nrp132, were comparable between control and XCR1-DTA mice (Supplementary Fig. 4A–C). In line with the overall CD4+ T cell decrease in XCR1-DTA mice, the number of LP Tregs was approximately half that of control mice (Supplementary Fig. 4C), but the remaining cells existed at the expected frequency and exhibited a normal phenotype. Therefore, in contrast to other LP T cell populations, the generation and maintenance of Tregs is not specifically affected in mice lacking XCR1+ DCs.
XCR1-DTA mice are more susceptible to DSS-induced colitis
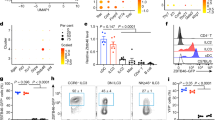
While mice lacking all CD11c+ DCs exhibit aberrant myeloid cell proliferation and autoimmunity33, the absence of CD103+CD11b− DCs alone does not induce any overt signs of dysregulated immunity under steady-state conditions34. However, during dextran sodium sulphate (DSS) -induced colitis, while mice lacking CD103+CD11b− DCs through deficiency in the transcription factor BATF3 (Basic leucine zipper transcription factor ATF-like 3) (BATF3) are as susceptible as their wild-type counterparts34, induced ablation of CD103+CD11b− DCs in Clec9A-diphtheria toxin receptor (DTR) mice resulted in significant exacerbation of colitis35. As XCR1-DTA mice also did not show clear signs of dysregulated intestinal immunity/intestinal inflammation in the steady state, we assessed their susceptibility to DSS-induced colitis. After administration of DSS, XCR1-DTA mice lost significantly more weight, achieved significantly higher disease activity scores and underwent significantly more colon shortening compared to control mice (Fig. 3). Thus, while the marked differences in the intestinal immune compartment in XCR1-DTA mice do not result in inflammatory changes under homeostatic conditions, these animals are rendered exquisitely sensitive to induced intestinal inflammation.
XCR1-DTA mice are more susceptible to DSS-induced colitis.
(A,B) Body weight changes (A) and disease activity index (B) after DSS exposure of control (open circles) and XCR1-DTA (filled circles) mice. Mice were given 1.5% DSS in drinking water for 7 days and clinically monitored. Means ± s.e.m. of control (n=8) and XCR1-DTA (n=6) mice are indicated. (C) Colon length after 10 days of DSS treatment. Means are shown as bars. Results are representative of three (A-C) independent experiments. (*P < 0.05; **P < 0.01, Student’s t test).
Mice lacking XCR1 or XCL1 exhibit similar defects in intestinal T cell populations as XCR1-DTA mice
We then asked whether it was possible that the XCR1-XCL1 axis itself was contributing to the observed defects in mice lacking XCR1+ DCs. We first generated XCL1-deficient mice (Supplementary Fig. 5A,B) and found that the size and composition of the T cell population in spleen and MLNs was comparable with control mice (Supplementary Fig. 6A,B). However, similar to XCR1-DTA mice, XCL1-deficient mice possessed significantly fewer TCRαβ+ T cells in their LP and intraepithelial compartment (Fig. 4A–D). All the LP and intraepithelial TCRαβ+ T cell subsets, but not TCRγδ+ T cells, were significantly decreased in XCL1-deficient mice compared to controls. Relative proportions of CD4+CD8− and CD4+CD8αα+ T cells were also increased and decreased, respectively within the IEL population (Fig. 4C), as in XCR1-DTA mice (Fig. 1C).
The XCR1-XCL1 axis is involved in maintenance of intestinal T cell populations.
(A–D) Percentages (A,C) and total numbers (B,D) of T lineage cells of LP and IELs from control (XCL1+/−) and XCL1-deficient (XCL1−/−) mice are shown. (E–H) Percentages (E,G) and total numbers (F,H) of T lineage cells of LP and IELs from control (XCR1+/venus) and XCR1-deficient (XCL1venus/venus) mice are shown. Dead cells and doublets were eliminated by FSC/SSC gating and LIVE/DEAD staining, with live cells subjected to further gating as indicated. Means ± s.e.m. of five mice are shown (B,D,F,H). Results are representative of four independent experiments. (*P < 0.05; **P < 0.01; ***P < 0.001, Student’s t test).
We then analysed XCR1-deficient mice16. These animals also possessed significantly fewer LP and intraepithelial TCRαβ+ T cells than controls (Fig. 4E–H), while splenic and MLN T cell populations were comparable between control and mutant mice (Supplementary Fig. 6C,D). Furthermore, all LP and intraepithelial TCRαβ+ T cell subsets, but not TCRγδ+ T cells, were decreased in XCR1-deficient mice.
In summary, these data show that mice lacking either XCL1 or XCR1 exhibit similar specific defects in intestinal T cell populations as XCR1-DC-deficient mice. Therefore it is possible that the XCR1-XCL1 axis itself is directly involved in intestinal T cell homeostasis, via a mechanism that might either involve interaction with XCR1+ DCs or be independent of this pathway.
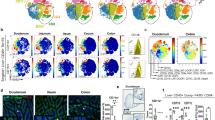
XCR1 deficiency alters the gene expression profile of intestinal DCs
To further dissect the direct effects of the XCR1-XCL1 axis on DCs in the gut, we compared the gene expression profiles of LP DC subsets from control and XCR1-deficient mice. The mutant CD103+CD11b− DCs contained significantly fewer transcripts for 62 of the 547 immune-related genes measured than did control CD103+CD11b− DCs. The most highly differentially-expressed genes within this subset were Xcr1, Ccr7, Cd40, Il12b, Ccl22 and Il6 (Fig. 5A). Ccr7 encodes the chemokine receptor, CCR7, which is required for DCs to migrate from the peripheral tissues to the draining LNs36; thus low expression of Ccr7 might impede emigration from the LP and thus result in abnormal patterns of DC distribution. To assess the biological significance of this finding, we measured the relative abundance of CD103+CD11b− DCs in the LP and MLNs of mice lacking either XCL1 or XCR1 and in control animals. As would be expected if DCs expressed insufficient CCR7 for effective migration to the LN, in both XCL1- and XCR1- deficient mice, the CD103+CD11b− DC population was significantly smaller in the MLN and significantly larger in the LP, compared to control mice (Fig. 5B–E). Meanwhile, the composition of the splenic DC compartment was unaffected by the absence of XCL1 or XCR1 (Supplementary Fig. 7A–D). Thus it appears that the XCR1-XCL1 axis is required for optimal gene expression to support normal migration of intestinal XCR1+ DCs to the MLN.
XCR1+ DC distribution is affected in the absence of XCR1 or XCL1.
(A) Nanostring gene expression analysis of sorted LP DC subsets pooled from eight Xcr1+/venus or eight Xcr1venus/venus mice. (B,C) Percentages (B) and total numbers (C) of MLN and LP DCs from control (XCL1+/−) and XCL1-deficient (XCL1−/−) mice are shown. (D,E) Percentages (D) and total numbers (E) of MLN and LP DCs of control (XCR1+/venus) and XCR1-deficient (XCL1venus/venus) mice are shown. Dead cells and doublets were eliminated by FSC/SSC gating and LIVE/DEAD staining, with live cells subjected to further gating as indicated. Results are representative of four (C,E) independent experiments. Means ± s.e.m. of five mice are indicated (C,E). (*P < 0.05; **P < 0.01; ***P < 0.001, Student’s t test).
As XCL1 is known to be produced by NK and CD8 T cells in other tissues15,16,17,18,22,23 and given the observed importance of XCL1 for optimal intestinal DC function, we then asked which cells within the intestinal compartment express Xcl1 and might therefore act as a source of the chemokine in vivo. Compared with splenic T cells, LP and intraepithelial T cell subsets from wildtype mice showed high expression of Xcl1 (Fig. 6A). Interestingly, we saw that Xcl1 expression in LP T cells of XCR1-DTA mice was decreased to approximately 20% of that in control mice (Fig. 2B), which might indicate that the high expression of Xcl1 in intestinal T cells is supported by the presence of XCR1+ DC. Immunofluorescence imaging of intestinal sections confirmed that XCR1+ DCs and T cells are closely associated in vivo (Fig. 6B). Furthermore, in similar to XCL1- or XCR1-deficient mice, RAG2-deficient mice, which lack any mature lymphocytes, showed the increase and decrease of CD103+CD11b− DCs in the LP and MLN, respectively (Fig. 6C). Taken together, these data suggest the possibility that intestinal T cells might provide XCR1+ DCs with XCL1 while closely associated in the gut.
Intestinal CD4 T cells are closely associated with XCR1+ DCs in vivo.
(A) Xcl1 expression in sorted T cells from spleen and LP, IELs and splenic B (CD3ε−B220+) and NK (CD3ε−CD49b+) cells pooled from eight wildtype mice, determined by quantitative real-time PCR. (B) Immunofluorescence imaging of intestinal sections from Xcr1+/venus mice labelled with anti-CD4 Ab and venus. Scale bars, 60 μm. (C) Percentages of MLN and LP DC subsets in Rag2+/− and Rag2−/− mice. Dead cells and doublets were eliminated by FSC/SSC gating and LIVE/DEAD staining, with live cells subjected to further gating as indicated. Results are representative of two independent experiments.
Discussion
Here we have generated and analysed XCR1-DTA mice which lack the CD103+CD11b−XCR1+ DC subset with the aim of understanding the role of this DC subset. While these mice did not exhibit any overt signs of pathology as a result of the absence of XCR1+ DCs, all LP T cell and IEL populations were specifically and significantly decreased, while the abundance of T lineage cells in thymus, spleen and MLNs was not affected. Thus the presence of XCR1+ DCs is necessary for normal numbers of T cells in the intestinal LP and IEL compartments.
In contrast to the absence of pathology in the steady-state, we found that the defects induced by the absence of XCR1+ DCs rendered XCR1-DTA mice significantly more susceptible to induced colitis compared with control mice. While the pathology of DSS colitis does not require the presence of T cells37, the condition is exaggerated in various mutant mice in which IELs, or in particular CD4+CD8αα+ T cells, are depleted27,38,39,40,41,42. Exacerbation of colitis in XCR1-DTA mice, with their profoundly diminished intestinal T cell populations, is consistent with these reports. Thus, while the absence of XCR1+ DCs does not cause inflammation per se, the associated quantitative and/or qualitative changes in the intestinal T cell compartment may reduce the ability of XCR1-DTA mice to control the response to DSS-induced gut inflammation.
Similar to our findings, mice lacking CD103+CD11b− DCs do not exhibit signs of spontaneous inflammation in the steady state34,35, but their susceptibility to DSS-induced colitis seems to vary depending on additional factors: models using deficiency in the transcription factor BATF3 to induce CD103+CD11b− DC deficiency are as susceptible as their wild-type counterparts34, while employing diphtheria-toxin-mediated ablation of CD103+CD11b− DCs in Clec9A-DTR mice renders them significantly more sensitive to induced colitis35. Alongside the difference in DC ablation method, susceptibility to DSS-induced colitis can be affected by differences in the commensal gut microbiota, mouse genetic background and source and dose of the DSS used. For example, Edelson et al. used 129S6/SvEv mice with 5% DSS from TDB Consultancy34, whereas we used C57BL/6 mice with 1.5% DSS from MP Biomedicals. Furthermore, it cannot be formally excluded that CD103+CD11b− DCs are compensatory generated during the course of DSS-induced colitis in BATF3-deficient mice43. Meanwhile, our results are consistent with enhanced manifestations of DSS-induced colitis in Clec9A-DTR mice, in which CD103+CD11b− DCs are inducibly ablated35. Muzaki et al. used BALB/c Clec9A-DTR mice treated with 2% DSS from MP Biomedicals and showed that CD103+CD11b− DCs were critical for anti-inflammatory responses, dependent on T cell-derived IFN-γ; although intestinal T cell populations were not investigated in their study, we also observed decreased Ifng expression in the remaining LP CD4+ T cells from XCR1-DTA mice prior to the induction of colitis (Fig. 2C).
In addition to the depletion of T cell populations and an inability to restrain induced inflammation, we detected further abnormalities in the intestinal T cell compartment of mice lacking XCR1+ DCs. Intestinal T cells normally express lower levels of CD62L and higher levels of CD103 than T cells in other tissues, however LP T cells from XCR1-DTA mice showed defects in this typical intestinal phenotype. Decreased expression of CD62L is indicative of the constitutive activation of intestinal T cells, most likely by antigens from the intestinal lumen, such as those of the commensal bacteria. Indeed, germ-free or antibiotic-treated mice also show decreased populations of intestinal T cells including IELs44,45. Therefore, the failure to downregulate CD62L of LP T cells from XCR1-DTA mice indicates a deficiency somewhere along the pathway leading to activation of these T cells, or in their ability to respond to activating stimuli. It is unknown whether XCR1+ DCs directly incorporate and present antigens in the gut, though there is evidence that CX3CR1+ macrophages can capture and transfer antigens to CD103+ DCs46. It will be intriguing to investigate further how the absence of XCR1+ DCs is leading to the emergence of a diminished T cell population with atypical activation characteristics.
The upregulated expression of CD103 by intestinal T cells is driven by the local cytokine milieu and particularly by TGF-β47. The absence of either CD103 or functional TGF-β signalling in T cells results in impaired development of intestinal T cell populations48,49, as seen in XCR1-DTA mice. TGF-β is normally produced by intestinal DCs, including XCR1+ DCs11, therefore we speculate that the lack of effective CD103 upregulation by intestinal T cells in XCR1-DTA mice might be caused by an absence of XCR1+ DC-derived TGF-β, which could also contribute to the decrease in T cell populations.
Intestinal T cells in XCR1-DTA mice had poor survival capacity and a tendency to express genes related to Th2 or Th17 differentiation, rather than Th1. It is unclear how the absence of XCR1+ DC is linked to this defect, however BATF3-dependent DCs are known to be involved in Th1 cell differentiation20, so it is possible that XCR1+ DCs might also drive T cell polarization. Meanwhile, Treg cells were numerically reduced in line with the overall decrease in T cell populations, but we did not detect any abnormalities in their phenotype in XCR1-DTA mice. Similarly, BATF3-deficient mice maintained a normal population of Treg cells in their LP and MLNs34.
Absence of XCR1+ DCs also led to further relative decrease in CD4+CD8αα+ IELs, accompanied by a relative accumulation of the CD4+CD8− T cells in XCR1-DTA mice. The generation requires certain MHC class II-expressing antigen presenting cells and Runx3 expression in CD4+CD8− T cells30,31. Because Runx3 expression in CD4+CD8− T cells was decreased in XCR1-DTA mice, we assume that XCR1+ DCs are responsible for the generation of CD4+CD8αα+ T cells by keeping Runx3 expression levels. Runx3 expression is driven by TGF-β and retinoic acids, which are generated by aldehyde dehydrogenase from vitamin A30,31. XCR1+ DCs express both Tgf-b1 and aldh1a211. Although further studies are necessary to clarify whether the decrease of CD4+CD8αα+ T cells in XCR1-DTA mice is caused by developmental block and/or impaired expansion, it is possible that XCR1+ DCs are required for the generation of CD4+CD8αα+ T cells by providing TGF-β and/or retinoic acids.
While the removal of XCR1+ DCs clearly induces profound changes in the intestinal immune compartment, the question remains: to what extent is the XCR1-XCL1 axis involved, or are the observed effects more likely to be due to other, XCR1-XCL1-independent, effects? While this study was not designed to answer this question specifically, we found that mice lacking XCR1 or XCL1 recapitulated important features of the T cell insufficiencies and imbalances present in XCR1-DTA mice. Moreover, intestinal T cells from wild type mice abundantly expressed Xcl1 and were observed in close association with XCR1+ DCs in tissue sections, while LP T cells from XCR1-DTA mice expressed significantly lower levels of Xcl1 than those from control mice (Fig. 2B). Thus it is possible that intestinal T cells provide XCL1 to local XCR1+ DCs. We also noted that in XCR1- and XCL1-deficient mice, CD103+CD11b− DCs accumulated in the LP and were reduced in number in the MLN: in XCR1-deficient mice at least, this could be due to the lower than normal level of Ccr7 expression in CD103+CD11b− DCs, as CCR7 is required for intestinal DCs to migrate to the MLN50.
Taken together, our findings lead us to propose a hypothetical model of XCR1-DC-T cell crosstalk in the intestine as a basis for further study and discussion (Fig. 7): activated intestinal T cells produce XCL1, which specifically attracts nearby XCR1+ DC; the ensuing DC-T cell crosstalk supports T cell survival, promotes the typical CD103high/CD62Llow intestinal phenotype and leads to maintenance of intraepithelial and LP T cells; continuing XCL1 expression by the T cells in turn enables DC maturation and in particular CCR7 expression necessary for migration from the LP to the MLN.
Hypothetical model of the possible crosstalk between XCR1+ DCs and intestinal T cells.
Activated intestinal T cells produce XCL1, which attracts nearby XCR1+ DC; the ensuing DC-T cell crosstalk supports T cell survival, promotes upregulation of CD103 expression with downregulation of CD62L expression and leads to maintenance of intraepithelial and LP T cells; continuing XCL1 expression by the T cells in turn enables DC maturation, including upregulated CCR7 expression to enable migration from the LP to the MLN.
In conclusion, our findings highlight the potential significance of the XCL1-XCR1 axis and in particular the interaction of XCR1+ DCs and the intestinal T cell compartment, in maintaining gut homeostasis and ensuring appropriate control of inflammation. This both advances our understanding of the fundamental workings of the intestinal immune system and identifies the XCR1-XCL1 axis as a novel potential therapeutic target for the treatment of human intestinal immune disorders. Further work to identify the key molecular mediators and underlying mechanisms will be necessary to fully understand this intriguing system.
Methods
Mice
C57BL/6J mice were purchased from CLEA Japan. All mice were bred and maintained in the Animal Facility of RIKEN Research Institute for Allergy and Immunology (Yokohama, Japan), Animal Resource Center for Infectious Diseases, Research Institute for Microbial Diseases and Immunology Frontier Research Center, Osaka University (Suita, Japan) and Institute for Animal Experimentation, Wakayama Medical University (Wakayama, Japan) under specific pathogen-free conditions and were used at 7–14 weeks of age according to the institutional guidelines of RIKEN Institute, Osaka University and Wakayama Medical University. All animal experiments were approved by the animal research committees and were carried out in accordance with approved guidelines of the animal care committees of of RIKEN Yokohama Research Institute, Osaka University and Wakayama Medical University.
Generation of mutant mice
XCR1-DTA mice were generated as follows: first, the entire murine XCR1 coding sequence was replaced with a gene encoding the Cre recombinase to generate mice in which Cre recombinase is expressed in XCR1-expressing cells (Supplementary Fig. 1A). The Cre recombinase gene was derived from pPGK-Cre-bpA (kindly provided by Dr. Klaus Rajewsky, Max-Delbrück-Center for Molecular Medicine, Berlin, Germany) and carried a polyadenylation signal derived from the bovine growth hormone gene (bGHpA). A neomycin-resistance gene driven by the MC1 promoter and flanked by yeast FRT sequences was used as a selection marker. A herpes simplex virus thymidine kinase gene (HSV-TK) was inserted for negative selection. The C57BL/6-derived embryonic stem (ES) cell line, Bruce4 (kindly provided by Drs. Colin L. Stewart, Institute of Medical Biology, Singapore, through Masaki Hikida, Kyoto University, Japan), was transfected with the linearized targeting vector by electroporation and selected with G418 (Nacalai Tesque) and ganciclovir (Mitsubishi Tanabe Pharma). Doubly-resistant clones were screened for homologous recombination by PCR and verified by Southern blot analysis. Germline-transmitting chimeras were generated by injection of targeted ES clones into blastocysts from BALB/c mice which were bred with C57BL/6J mice. Xcr1+/cre mice were further backcrossed with C57BL/6J mice for 2–6 generations. Xcr1+/cre mice were crossed with R26:lacZbpAfloxDTA mice (kindly provided by Dr. Dieter Riethmacher, University of Southampton, UK, through Dr. Shigekazu Nagata, Kyoto University, Japan) carrying the DTA gene, which is designed to be expressed specifically in the cells or tissues expressing Cre recombinase under the control of the ubiquitously-active Rosa26 promoter51. This crossing generated the XCR1-DTA mice, in which XCR1+ cells are ablated by DTA expression. Xcr1+/cre littermates were used as controls.
Xcr1+/venus mice were generated by knocking a gene encoding a fluorescent protein, venus, into the Xcr1 locus16. Xcr1venus/venus mice were therefore rendered XCR1-deficient.
To generate XCL1-deficient mice, the targeting vector was designed to replace the murine XCL1 coding sequences with a neomycin resistance gene driven by the MC1 promoter, flanked by the yeast FRT sequence (Supplementary Fig. 5A). Bacteriophage P1 loxP sequences and the DTA gene were inserted for conditional ablation and negative selection, respectively. The targeted ES cells were generated as described above, but were selected with G418 alone. The neomycin resistance gene was removed by crossing with CAG-cre transgenic mice (kindly provided by Jun-ichi Miyazaki, Osaka University, Osaka, Japan)52 and the resultant Xcl1+/− mice were then backcrossed with C57BL/6J mice for 4 generations. The resulting Xcl1−/− mice were thus XCL1 deficient.
Cell preparations
Thymocyte and splenocyte suspensions were prepared by grinding the appropriate organs through mesh filters. MLNs were digested with 400 units/ml collagenase III (Worthington) and 100 μg/ml deoxyribonuclease (DNase) I (Sigma-Aldrich) at 37 °C for 20 min. Intestinal immune cells were prepared from the small intestine as follows: fat and Peyer’s patches were dissected away before the small intestine was opened longitudinally and stirred in RPMI containing 2% FCS, 2 mM EDTA for 20 min at 37 °C. IELs were isolated from the resulting supernatant using a 40–75% Percoll density gradient (GE Healthcare) and by collecting the cells that layered between the 40 and 75% fractions. Following supernatant collection the intestinal tissue was stirred for an additional 20 min at 37 °C in RPMI containing 2% FCS, before mincing and further stirring in 400 units/ml of collagenese D (Roche Diagnostics) and 10 μg/ml DNase I for 20 min at 37 °C. Floating cells were collected and the collagenase digestion was repeated twice more. The pooled cell suspensions were then centrifuged on a 40–70% Percoll density gradient and the cells that layered between the 40–75% fractions were collected as LP cells.
Flow cytometric analysis
Single cell suspensions were incubated with anti-CD16/32 Ab (TONBO Biosciences) to block non-specific binding of Abs. The cells were then labelled with fluorochrome-conjugated Abs and/or biotinylated Abs against murine CD4 (GK1.5), CD8α (53-6.7), CD8β (H35-17.2), CD11b (M1/70), CD11c (N418), CD25 (PC61), CD62L (MEL-14), CD103 (M290), CCR9 (CW-1.2), α4β7 (DATK32), TCRβ (H57-597), TCRγδ (GL3), I-A/I-E (M5/114.15.2) or Foxp3 (FJK-16s). Biotinylated Abs were visualized via fluorochrome-conjugated streptavidin. Abs and fluorochrome-conjugated streptavidin were purchased from BD Biosciences, eBioscience, BioLegend and TONBO biosciences. Biotinylated anti-Nrp1 Ab and normal goat IgG were purchased from R&D. Foxp3 was detected intracellularly using Foxp3 Staining Buffer Set (eBioscience). Dead cells were excluded using the LIVE/DEAD Fixable Dead Cell Stain Kit (Invitrogen). Cells were analysed on a FACSVerse or FACSAria II (BD Biosciences) and data analysed with FlowJo software (TreeStar).
Histology
For Fig. 1E, small intestines were embedded in FSC22 frozen section compound (Leica Microsystems). Five μm cryosections were labelled with rat anti-human Ep-CAM (G8.8; BioLegend) Ab and biotinylated armenian hamster anti-TCRβ (H57-597; BioLegend) Ab and then labelling was visualised with streptavidin-Alexa Fluor 594 (Invitrogen) and Alexa Fluor 647-conjugated anti-rat IgG (H+L) Ab (Invitrogen). For Fig. 6B, small intestines from Xcr1+/venus mice were first fixed with 4% paraformaldehyde and then embedded in FSC22 frozen section compound. Five μm cryosections were labelled with rat anti-mouse CD4 Ab (BioLegend) and labelling visualised with goat anti-Rat IgG Ab conjugated-Alexa Fluor 594 (Invitrogen). Sections were examined using the FV10i microscope (Olympus).
Cell death and proliferation analysis
For analysis of cell death, LP cells and IELs were washed with annexin-binding buffer (10 mM HEPES, 140 mM NaCl, 2.5 mM CaCl2, pH 7.4) and incubated with fluorochrome-conjugated annexin-V and 7-AAD Viablity Staining Solution (eBioscience) for 15 min at room temperature. For analysis of cell proliferation, cells were surface-labelled with the appropriate conjugated Abs, fixed, permeabilized using Foxp3 Staining Buffer Set (eBioscience) and subsequently incubated with anti-Ki67-FITC (B56; BD Biosciences) for 30 min. After washing, cells were analysed on a FACSAria II.
Gene expression analysis
Gene expression profiles of LP CD4+ T cells (Fig. 2B) and DC subsets (Fig. 5A) were analysed with the Mouse Immunology Kit containing 547 immune-related genes and the NanoString nCounter gene expression system (NanoString Technologies, Seattle, WA). Briefly, 10,000 cells of each T cell or DC subset were lysed in RLT buffer (Qiagen, Valencia, CA). The lysates were hybridized for 16 h with the Mouse Immunology Kit and loaded into the nCounter prep station followed by quantification using the nCounter Digital Analyzer (NanoString Technologies). The nCounter data were normalized in two steps: firstly using the positive spiked-in controls provided by the nCounter instrument (NanoString Technologies) and secondly, according to the expression of 14 control genes (Alas1, Eef1g, G6pdx, Gapdh, Gusb, Hprt, Oaz1, Polr1b, Polr2a, Ppia, Rpl19, Sdha, Tbp and Tubb5).
Colitis model
Mice were allowed free access to filtered drinking water containing 1.5% DSS (MW = 36,000–50,000; MP Biomedicals) for 1 week. Mice were weighed at indicated days. Severity of colitis was assessed using the disease activity index based on presence of diarrhea and bleeding53. At day 10 all mice were euthanized and colons were dissected for measurement of length.
Quantitative real-time PCR
Total RNAs were obtained using the Sepasol RNA-I Super G (Nacalai tesque), reverse transcribed and analysed by an ABI PRISM 7000 (Applied Biosystems). TaqMan probes (TaqMan Gene Expression Assay; Applied Biosystems) were used for 18S rRNA (internal control). The following SYBR Green primers were used: Xcl1 (Forward 5′- TTTGTCACCAAACGAGGACTAAA-3′, Reverse 5′-CCAGTCAGGGTTATCGCTGTG-3′). Gene expression was normalized to that of 18S rRNA and is represented as the ratio relative to the indicated reference samples. All primers were validated for linear amplification.
Statistical analysis
The data were analysed using two-tailed unpaired Student’s t test with Welch’s correction in cases of unequal variance. P values < 0.05 were considered statistically significant. All statistical analysis was performed using GraphPad Prism software (GraphPad).
Additional Information
How to cite this article: Ohta, T. et al. Crucial roles of XCR1-expressing dendritic cells and the XCR1-XCL1 chemokine axis in intestinal immune homeostasis. Sci. Rep. 6, 23505; doi: 10.1038/srep23505 (2016).
References
Mowat, A. M. & Agace, W. W. Regional specialization within the intestinal immune system. Nat. Rev. Immunol. 14, 667–685 (2014).
Grainger, J. R., Askenase, M. H., Guimont-Desrochers, F., da Fonseca, D. M. & Belkaid, Y. Contextual functions of antigen-presenting cells in the gastrointestinal tract. Immunol. Rev. 259, 75–87 (2014).
Mortha, A. et al. Microbiota-dependent crosstalk between macrophages and ILC3 promotes intestinal homeostasis. Science 343, 1249–1288 (2014).
Liu, K. & Nussenzweig, M. C. Origin and development of dendritic cells. Immunol. Rev. 234, 45–54 (2010).
Merad, M., Sathe, P., Helft, J., Miller, J. & Mortha, A. The dendritic cell lineage: ontogeny and function of dendritic cells and their subsets in the steady state and the inflamed setting. Annu. Rev. Immunol. 31, 563–604 (2013).
Sun, C. M. et al. Small intestine lamina propria dendritic cells promote de novo generation of Foxp3 T reg cells via retinoic acid. J. Exp. Med. 204, 1775–1785 (2007).
Coombes, J. L. et al. A functionally specialized population of mucosal CD103+ DCs induces Foxp3+ regulatory T cells via a TGF-beta and retinoic acid-dependent mechanism. J. Exp. Med. 204, 1757–1764 (2007).
Persson, E. K. et al. IRF4 transcription-factor-dependent CD103(+)CD11b(+) dendritic cells drive mucosal T helper 17 cell differentiation. Immunity 38, 958–969 (2013).
Schlitzer, A. et al. IRF4 transcription factor-dependent CD11b+ dendritic cells in human and mouse control mucosal IL-17 cytokine responses. Immunity 38, 970–983 (2013).
Welty, N. E. et al. Intestinal lamina propria dendritic cells maintain T cell homeostasis but do not affect commensalism. J. Exp. Med. 210, 2011–2024 (2013).
Denning, T. L. et al. Functional specializations of intestinal dendritic cell and macrophage subsets that control Th17 and regulatory T cell responses are dependent on the T cell/APC ratio, source of mouse strain and regional localization. J. Immunol. 187, 733–747 (2011).
Kayama, H. et al. Intestinal CX3C chemokine receptor 1(high) (CX3CR1(high)) myeloid cells prevent T-cell-dependent colitis. Proc. Natl. Acad. Sci. USA 109, 5010–5015 (2012).
Bachem, A. et al. Expression of XCR1 Characterizes the Batf3-Dependent Lineage of Dendritic Cells Capable of Antigen Cross-Presentation. Front. Immunol. 3, 214 (2012).
Becker, M. et al. Ontogenic, Phenotypic and Functional Characterization of XCR1(+) Dendritic Cells Leads to a Consistent Classification of Intestinal Dendritic Cells Based on the Expression of XCR1 and SIRPalpha. Front. Immunol. 5, 326 (2014).
Dorner, B. G. et al. Selective expression of the chemokine receptor XCR1 on cross-presenting dendritic cells determines cooperation with CD8+ T cells. Immunity 31, 823–833 (2009).
Yamazaki, C. et al. Critical roles of a dendritic cell subset expressing a chemokine receptor, XCR1. J. Immunol. 190, 6071–6082 (2013).
Shortman, K. & Heath, W. R. The CD8+ dendritic cell subset. Immunol. Rev. 234, 18–31 (2010).
Murphy, T. L., Tussiwand, R. & Murphy, K. M. Specificity through cooperation: BATF-IRF interactions control immune-regulatory networks. Nat. Rev. Immunol. 13, 499–509 (2013).
Tailor, P., Tamura, T., Morse, H. C. 3rd & Ozato, K. The BXH2 mutation in IRF8 differentially impairs dendritic cell subset development in the mouse. Blood 111, 1942–1945 (2008).
Hildner, K. et al. Batf3 deficiency reveals a critical role for CD8alpha+ dendritic cells in cytotoxic T cell immunity. Science 322, 1097–1100 (2008).
Briseno, C. G., Murphy, T. L. & Murphy, K. M. Complementary diversification of dendritic cells and innate lymphoid cells. Curr. Opin. Immunol. 29, 69–78 (2014).
Bachem, A. et al. Superior antigen cross-presentation and XCR1 expression define human CD11c+CD141+ cells as homologues of mouse CD8+ dendritic cells. J. Exp. Med. 207, 1273–1281 (2010).
Crozat, K. et al. The XC chemokine receptor 1 is a conserved selective marker of mammalian cells homologous to mouse CD8alpha+ dendritic cells. J. Exp. Med. 207, 1283–1292 (2010).
Haniffa, M. et al. Human tissues contain CD141hi cross-presenting dendritic cells with functional homology to mouse CD103+ nonlymphoid dendritic cells. Immunity 37, 60–73 (2012).
Zlotnik, A. & Yoshie, O. Chemokines: a new classification system and their role in immunity. Immunity 12, 121–127 (2000).
Lei, Y. et al. Aire-dependent production of XCL1 mediates medullary accumulation of thymic dendritic cells and contributes to regulatory T cell development. J. Exp. Med. 208, 383–394 (2011).
Cheroutre, H., Lambolez, F. & Mucida, D. The light and dark sides of intestinal intraepithelial lymphocytes. Nat. Rev. Immunol. 11, 445–456 (2011).
Agace, W. W. T-cell recruitment to the intestinal mucosa. Trends Immunol. 29, 514–522 (2008).
Wurbel, M. A. et al. Mice lacking the CCR9 CC-chemokine receptor show a mild impairment of early T- and B-cell development and a reduction in T-cell receptor gammadelta(+) gut intraepithelial lymphocytes. Blood 98, 2626–2632 (2001).
Mucida, D. et al. Transcriptional reprogramming of mature CD4(+) helper T cells generates distinct MHC class II-restricted cytotoxic T lymphocytes. Nat. Immunol. 14, 281–289 (2013).
Reis, B. S., Rogoz, A., Costa-Pinto, F. A., Taniuchi, I. & Mucida, D. Mutual expression of the transcription factors Runx3 and ThPOK regulates intestinal CD4(+) T cell immunity. Nat. Immunol. 14, 271–280 (2013).
Weiss, J. M. et al. Neuropilin 1 is expressed on thymus-derived natural regulatory T cells, but not mucosa-generated induced Foxp3+ T reg cells. J. Exp. Med. 209, 1723–1742, S1721 (2012).
Bar-On, L. & Jung, S. Defining dendritic cells by conditional and constitutive cell ablation. Immunol. Rev. 234, 76–89 (2010).
Edelson, B. T. et al. Peripheral CD103+ dendritic cells form a unified subset developmentally related to CD8alpha+ conventional dendritic cells. J. Exp. Med. 207, 823–836 (2010).
Muzaki A. R. et al. Intestinal CD103+CD11b− dendritic cells restrain colitis via IFN-γ-induced anti-inflammatory response in epithelial cells. Mucosal Immunol. [Epub ahead of print]
Forster, R. et al. CCR7 coordinates the primary immune response by establishing functional microenvironments in secondary lymphoid organs. Cell 99, 23–33 (1999).
Dieleman, L. A. et al. Dextran sulfate sodium-induced colitis occurs in severe combined immunodeficient mice. Gastroenterology 107, 1643–1652 (1994).
Chen, Y., Chou, K., Fuchs, E., Havran, W. L. & Boismenu, R. Protection of the intestinal mucosa by intraepithelial gamma delta T cells. Proc. Natl. Acad. Sci. USA 99, 14338–14343 (2002).
Das, G. et al. An important regulatory role for CD4+CD8 alpha alpha T cells in the intestinal epithelial layer in the prevention of inflammatory bowel disease. Proc. Natl. Acad. Sci. USA 100, 5324–5329 (2003).
Kuhl, A. A. et al. Aggravation of intestinal inflammation by depletion/deficiency of gammadelta T cells in different types of IBD animal models. J. Leukoc. Biol. 81, 168–175 (2007).
Li, Y. et al. Exogenous stimuli maintain intraepithelial lymphocytes via aryl hydrocarbon receptor activation. Cell 147, 629–640 (2011).
Tsuchiya, T. et al. Role of gamma delta T cells in the inflammatory response of experimental colitis mice. J. Immunol. 171, 5507–5513 (2003).
Tussiwand, R. et al. Compensatory dendritic cell development mediated by BATF-IRF interactions. Nature 490, 502–507 (2012).
Kunisawa, J., Takahashi, I. & Kiyono, H. Intraepithelial lymphocytes: their shared and divergent immunological behaviors in the small and large intestine. Immunol. Rev. 215, 136–153 (2007).
Jiang, W. et al. Recognition of gut microbiota by NOD2 is essential for the homeostasis of intestinal intraepithelial lymphocytes. J. Exp. Med. 210, 2465–2476 (2013).
Mazzini, E., Massimiliano, L., Penna, G. & Rescigno, M. Oral tolerance can be established via gap junction transfer of fed antigens from CX3CR1(+) macrophages to CD103(+) dendritic cells. Immunity 40, 248–261 (2014).
El-Asady, R. et al. TGF-{beta}-dependent CD103 expression by CD8(+) T cells promotes selective destruction of the host intestinal epithelium during graft-versus-host disease. J. Exp. Med. 201, 1647–1657 (2005).
Schon, M. P. et al. Mucosal T lymphocyte numbers are selectively reduced in integrin alpha E (CD103)-deficient mice. J. Immunol. 162, 6641–6649 (1999).
Konkel, J. E. et al. Control of the development of CD8alphaalpha+ intestinal intraepithelial lymphocytes by TGF-beta. Nat. Immunol. 12, 312–319 (2011).
Bogunovic, M. et al. Origin of the lamina propria dendritic cell network. Immunity 31, 513–525 (2009).
Brockschnieder, D., Pechmann, Y., Sonnenberg-Riethmacher, E. & Riethmacher, D. An improved mouse line for Cre-induced cell ablation due to diphtheria toxin A, expressed from the Rosa26 locus. Genesis 44, 322–327 (2006).
Sakai, K. & Miyazaki, J. A transgenic mouse line that retains Cre recombinase activity in mature oocytes irrespective of the cre transgene transmission. Biochem. Biophys. Res. Commun. 237, 318–324 (1997).
Maxwell, J. R., Brown, W. A., Smith, C. L., Byrne, F. R. & Viney, J. L. Methods of inducing inflammatory bowel disease in mice. Curr. Protoc. Pharmacol./editorial board, SJ Enna Chapter 5, Unit5 58 (2009).
Acknowledgements
We thank Y. Matsuhisa, S. Haraguchi and A. Matsumura for secretarial assistance and I. Ogahara and Y. Tanaka for technical assistance. We also thank Dr. K. Rajewsky for pPGK-Cre-bpA vector, Drs. C. Stewart and M. Hikida for an ES cell line, Bruce4, Drs. D. Riethmacher and S. Nagata for R26:lacZbpAfloxDTA mice, Dr. J. Miyazaki for CAG-cre transgenic mice and Dr. L. Robinson of Insight Editing London for assistance in manuscript preparation. This work was in part supported by the Kishimoto Foundation; a Grant-in-Aid for Scientific Research (B, C), Challenging Exploratory Research and Young Scientists from Japan Society for the Promotion of Science (JSPS), a Grant-in-Aid for Scientific Research on Innovative Areas and on Priority Areas from The Ministry of Education, Culture, Sports, Science and Technology (MEXT) and the Uehara Memorial Foundation. T. Ohta was supported by Grant-in-Aid for JSPS Fellows. M.S., C.Y. and I.S. were supported by a RIKEN Junior Research Associate Grant. F.G. was supported by the Singapore Immunology Network Core fund.
Author information
Authors and Affiliations
Contributions
T.Oh, M.S., H.H., S.O., I.S., Y.F. and T.Or performed the experiments and analysed the data; C.Y. and K.H. generated Xcr1+/cre mice; M.S. and K.H. generated XCL1-deficient mice; T.Oh., K. J. I., K.H., F.G. and T.K. interpreted the data and designed the study; T.Oh and T.K. conceived the project and wrote the manuscript; T.K. supervised the research.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ohta, T., Sugiyama, M., Hemmi, H. et al. Crucial roles of XCR1-expressing dendritic cells and the XCR1-XCL1 chemokine axis in intestinal immune homeostasis. Sci Rep 6, 23505 (2016). https://doi.org/10.1038/srep23505
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep23505
This article is cited by
-
Pro-regenerative biomaterials recruit immunoregulatory dendritic cells after traumatic injury
Nature Materials (2024)
-
Markers of immune dysregulation in response to the ageing gut: insights from aged murine gut microbiota transplants
BMC Gastroenterology (2022)
-
The kinase p38α functions in dendritic cells to regulate Th2-cell differentiation and allergic inflammation
Cellular & Molecular Immunology (2022)
-
High-dimensional and single-cell transcriptome analysis of the tumor microenvironment in angioimmunoblastic T cell lymphoma (AITL)
Leukemia (2022)
-
Conventional type 1 dendritic cells protect against age-related adipose tissue dysfunction and obesity
Cellular & Molecular Immunology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.