Abstract
In multimeric membrane receptors the cooperative action of the subunits prevents exact knowledge about the operation and the interaction of the individual subunits. We propose a method that permits quantification of ligand binding to and activation effects of the individual binding sites in a multimeric membrane receptor. The power of this method is demonstrated by gaining detailed insight into the subunit action in olfactory cyclic nucleotide-gated CNGA2 ion channels.
Similar content being viewed by others
Introduction
A large number of membrane receptors are multimeric proteins composed of a well-defined number of equal or different subunits. When one or more ligands bind to a multimeric receptor, it exerts a conformational change, resulting either in modulating a conductance for ions or a changed readiness to interact with other proteins. Up to now for no multimeric receptor the molecular events upon activation have been fully elucidated. The main reason for this lack of exact knowledge is that receptor function is governed by subunits cooperating with each other in a complex manner1.
For methodical approaches, ligand-gated ion channels provide the unique feature that the final conformational change, the pore opening, can be directly excellently quantified by the patch-clamp technique2, often by using single-channel analysis and hidden Markovian state models3.
CNG channels are tetrameric membrane receptors that generate a non-selective conductance for cations4,5. The channels are activated by the cytosolic cyclic nucleotides cAMP or cGMP, binding to specific cyclic nucleotide binding domains (CNBDs) in each subunit6. Wild-type olfactory CNG channels are composed of three types of subunits7 out of which only the CNGA2 subunits can form functional homotetrameric channels on their own when expressed in heterologous systems8,9. Homotetrameric CNGA2 channels are very useful model systems for studying elementary biophysical processes10,11, though also for these channels the subunit action upon channel activation is still a mysteriously complex and cooperative process, resulting in respectively branched Markovian models10,12. Other experimental findings further substantiate the complexity of the activation process, including the facts that Hill coefficients are two13 or even higher14 and that the pore opening and closure is a one-step process though four subunits are involved. To shed further light into the activation process, it is therefore still highly desirable to develop new methods monitoring the operation of the individual subunits in a functional channel.
Here we present a powerful and straightforward approach to analyze both the binding affinity and the gating effect of the individual subunits in a functional multimeric ionotropic receptor channel. CNGA2 channels were functionally expressed in Xenopus oocytes and currents induced by the cGMP binding were measured in inside-out patches. We also used subunits with a disabled binding site by inferring the mutation R538E (mut)15 which shifted the apparent affinity in the concentration-activation relationship of homotetrameric mut-channels (4×mut) by a factor of 315 to higher concentrations with respect to homotetrameric wt-channels (4×wt; Fig. 1a) leaving the Hill coefficient unaffected (Supplementary Table 1a). We next constructed tetrameric concatamers with either four wt- or four mut-subunits and tested whether these wt-wt-wt-wt and mut-mut-mut-mut channels are activated closely similar to 4×wt and 4×mut channels, respectively. This was substantiated by the following results: (1) The respective concentration-activation relationships match each other (Fig. 1a). (2) Co-expressing wt-wt-wt-wt with mut-mut-mut-mut channels (cRNA ratio 1:1) produced a concentration-activation relationship containing two well determined components with characteristics specific for each channel entity, suggesting that the concatameric channels operate independently (Fig. 1b, Supplementary Table 1b). (3) At respectively saturating cGMP, the amplitude of the single-channel current in both wt-wt-wt-wt and mut-mut-mut-mut channels was indistinguishable from that in 4×wt-channels (Supplementary Fig. 1a). (4) The open probability at saturating cGMP in wt-wt-wt-wt (0.99 ± 0.002; 100 μM cGMP; N = 4) and mut-mut-mut-mut channels (0.99 ± 0.003; 5 mM cGMP; N = 5) was similarly close to unity as in 4×wt channels (0.99 ± 0.002; 100 μM cGMP; N = 4).
(a) Effect of an increasing number of mut-subunits. The channels are either formed by four monomers (4×wt, 4×mut) or by tetrameric concatamers (wt-wt-wt-wt, mut-wt-wt-wt, mut-mut-wt-wt, mut-mut-mut-wt, mut-mut-mut-mut). All fit parameters are provided by Supplementary Table 1a. (b) Concatamers assemble as tetrameric channels. Expression of either wt-wt-wt-wt or mut-mut-mut-mut channels alone or together with a cRNA ratio 1:1 (N = 14–21). All fit parameters are provided by Supplementary Table 1b. (c) The position of wt-subunits is irrelevant for the concatamer function. The concentration-activation relationships of four concatamers with two wt-subunits and four concatamers with one wt-subunit are plotted. The fit parameters are given in Supplementary Table 1c. The relationships for the concatamers with one wt-subunit were indistinguishable as were the relationships for the concatamers with two wt-subunits (multidimensional t-test with Holm correction, p-value = 1).
We then constructed concatamers with increasing numbers of mutated subunits in the background of wild-type subunits, i.e. mut-wt-wt-wt, mut-mut-wt-wt, and mut-mut-mut-wt. The concentration-response relationships for these channels were systematically shifted to higher cGMP concentrations in proportion to the number of incorporated mut-subunits (Fig. 1a). For concatamers with either one or two wt-subunits we also showed that, surprisingly, the channel function does not depend on the position of the respective subunits (Fig. 1c, Supplementary Table 1c). In the following analysis we included as representative concatamers mut-mut-wt-wt and mut-mut-mut-wt.
In the concentration-activation relationships of the respective channels the slope systematically decreased when decreasing the number of wt-subunits from four to one (Fig. 1a). Apart from mut-mut-mut-wt channels, all concentration-activation relationships could be described reasonably by a single Hill function (Equation 1) whereas mut-mut-mut-wt channels required the sum of two Hill functions (Equation 2), presumably mirroring the action of the only wt-subunit on the pore opening (high affinity component (EC50,H)) and that of the three mut-subunits (low affinity component (EC50,L), respectively.
To test whether or not the concentration-activation relationships considered so far represent exclusively effects of cGMP on the channel gating, and not on the unitary current, single-channel experiments were performed with mut-mut-mut-wt channels. The amplitude of the unitary currents (Supplementary Fig. 2) matched those of 4×wt, wt-wt-wt-wt, and mut-mut-mut-mut channels at saturating cGMP (Supplementary Fig. 1). Similar results were obtained with two wt-subunits in mut-mut-wt-wt (Supplementary Fig. 3). These results suggest that the channel pore operates as a whole and that an increasing number of occupied binding sites only increases the open probability. This conclusion is consistent with kinetic models containing only one concerted opening step of all subunits10,16.
Based on the above results, the concentration-activation relationships of the five properly operating concatameric channels (Fig. 1a) contain complex information about the intricate gating of the subunits. To adequately uncover this information we performed a global fit analysis by five Markovian submodels, one for each concatamer (4wt-submodel through 0wt-submodel; Fig. 2a). In each of these submodels four ligands can bind, quantified by the association constants KAx (x = 1…4), which are, according to the number of wt- and mut-subunits, subdivided in high-affinity constants, KAxH, and low-affinity constants, KAxL, respectively. At each degree of liganding the channel can open by a single step, quantified by an equilibrium constant for the closed-open isomerization Ex (x = 0…4). The five submodels are intimately coupled via their equilibrium constants. Due to microscopic reversibility17 the global fit contained only six free parameters (Supplementary Methods). Most favorably, an active effect of each subunit is available at two distant ligand concentrations, providing a considerable surplus of information.
(a) Markovian submodels describing the activation gating of five concatamers. The closed-open isomerizations, including their equilibrium constants, E0….E4, are indicated only once at the bottom. They are the same for each equally liganded state. KA1H, KA2H, KA3H, KA4H, KA1L, KA2L, KA3L, and KA4L are the equilibrium association constants for the four high and low affinity binding sites, respectively. E0 and E4 were set to 1.7 × 10−5 and 9.9 × 101 according to the single-channel experiments. (b) Global fit of the data points of the five concatamers shown in a. The values of the equilibrium constants are provided by Table 1. (c) Occupancy of the states (P) predicted by the 1wt-submodel as function of the [cGMP].
The coupled submodels fully describe the series of concentration-activation relationships (Fig. 2b; for parameters see Table 1). E0 and E4 were determined separately in single-channel experiments at zero and saturating cGMP in wt-wt-wt-wt channels. Because the fit did not permit to distinguish KA3H from KA4H, KA3L from KA4L and E3 from E4, we set these respective constants equal. With this restriction the equilibrium constants were determined with formidable accuracy. These results show to what extent KAxH and Ex depend on the number of ligands. In line with a previous approach10, binding of the first ligand generates already noticeable channel opening (E1 = 3.59 × 10−1).
Knowing the equilibrium constants for all binding and open-closed isomerizations provides the possibility to evaluate for all concatamers the occupancy of the states as function of the cGMP concentration. Consider e.g. the occupancies in the 1wt-submodel (Fig. 2c). The probabilities to occupy a binding site are extended along the abscissa but do also considerably overlap. Moreover, the strong disabling effect of the mutation R538E in the mut-subunits causes that, apart from the empty states (C0, O0), only states with an occupied wt-subunit are relevantly populated. Another result is that up to ~3 μM cGMP only the wt-subunit is occupied, generating measurable channel activity.
In summary, we propose a powerful method to learn more about how individual subunits in a multimeric ion channel work in the presence of the other subunits. The asset of the method comes from the fact that the different number of four to zero mut-subunits does not only extend the concentration-activation relationships along the concentration axis but also characteristically shapes them, thereby providing valuable information.
To some extent this analysis is related to the well-established double-mutant cycle analysis18 that analyzes Gibbs free energy differences for two single and the corresponding double mutant to specify free energies of interaction19. In case of ion channels the EC50 values are regularly used to estimate these free energies of interaction20. Compared to such analyses, our approach is more comprehensive by including single, double, triple, and quadruple mutated channels as well as the specific effects of these mutations on the curvature of the concentration-activation relationships.
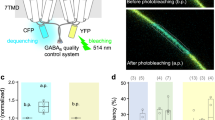
The proposed method bears potential to unravel also the properties of native heterotetrameric olfactory 2×CNGA2:CNGA4:CNGB1b channels, which would permit to address the question which of the subunits operates the channels at physiologically low cyclic-nucleotide concentrations. Besides CNG channels the method should be applicable also to any other channel. It seems to be also possible to adapt this method to analyze single-channel activity. And, beyond electrophysiological approaches, even metabotropic multimeric membrane receptors should be accessible for this approach if an appropriate read-out for the receptor activation is available, e.g. by inferring donor and acceptor fluorophores in the subunits and evaluating Förster resonance energy transfer (FRET)21.
Methods
Oocyte Preparation and cRNA Injection
Oocytes were obtained surgically under anesthesia (0.3% 3-aminobenzoic acid ethyl ester) from adult females of Xenopus laevis11. The procedures had approval from the authorized animal ethical committee of the Friedrich Schiller University Jena. The methods were carried out in accordance with the approved guidelines.
Molecular biology
The rCNGA2 tetrameric concatamers were obtained by joining the coding sequences of rat CNGA2 subunits via the short linker sequence GSA into a single ORF. To facilitate the concatenation procedure, we first introduced by PCR a BclI site in front of the CNGA2 (wt) start codon in vector pGEMHEnew22, and an in-frame BamHI site in front of the respective stop codon. These sites were also introduced into the pGEMHEnew vector coding for the R538E mutant (mut). In a second step, we constructed various dimers by inserting a BclI/EcoRI fragment of wt or mut into the BamHI/EcoRI sites of the modified pGEMHEnew vector containing either the full-length wt or mut sequences. Finally, two dimers were coupled using the same cloning strategy. The sizes of the dimeric and tetrameric DNA were confirmed by restriction enzyme mapping and gel electrophoresis. The linkers and flanking sequences were sequenced to verify that there were no secondary mutations and that the R538E mutation was correctly inserted. The cRNAs were transcribed from cDNAs in vitro with the mMESSAGE mMACHINE T7 Kit (Ambion, Austin, TX).
Electrophysiology
For obtaining steady-state concentration-activation relationships macroscopic currents were recorded from inside-out patches with a standard patch-clamp technique. The patch pipettes were pulled from quartz tubing (P-2000, Sutter Instrument, Novato, USA) with an outer and inner diameter of 1.0 and 0.7 mm (VITROCOM, New Jersey, USA). The pipette resistance was 0.5–1.7 MΩ. The bath and pipette solution contained (in mM): 150 KCl, 1 EGTA, 10 Hepes (pH 7.4 with KOH). Recordings were performed with an Axopatch 200A amplifier (Axon Instruments, Foster City, CA). Electrophysiology was controlled by the ISO2 hard- and software (MFK, Niedernhausen, Germany). The sampling rate was 5 kHz and the filter of the amplifier (4-pole Bessel) was set to 2 kHz. For single-channel measurements, the patch pipettes were pulled from quartz tubing with an outer and inner diameter of 1.0 and 0.5 mm (VITROCOM, New Jersey, USA). The pipette resistance was 5.0–12.0 MΩ. The sampling rate was 20 kHz and the filter of the amplifier (4-pole Bessel) was set to 5 kHz. The pipette solution contained (in mM): 150 KCl, 1 EGTA, 5 Hepes (pH 7.4 with KOH) and 200 μM niflumic acid to block endogenous chloride channels. The recording voltage was +100 mV.
Data analysis
If not otherwise noted, concentration-activation relationships were fitted with OriginPro 8G (Northampton, USA) by

I is the actual current amplitude, and Imax the maximum current amplitude at saturating cGMP specified for each patch. The saturating cGMP concentrations ranged from 100 μM to 5 mM. EC50 is the cGMP concentration generating the half maximum current and n the Hill coefficient.
Some concentration-activation relationships required the sum of a high (H) and a low affinity (L) component:

A is the fraction of the high affinity component.
Amplitude histograms of the single channel activity were fitted by respective sums of Gaussian functions. The global fit of the concentration-activation relationships is described in Supplementary Methods.
Statistics. Data are given as mean ± s.e.m.
Additional Information
How to cite this article: Wongsamitkul, N. et al. Quantifying the cooperative subunit action in a multimeric membrane receptor. Sci. Rep. 6, 20974; doi: 10.1038/srep20974 (2016).
Change history
28 June 2018
A correction has been published and is appended to both the HTML and PDF versions of this paper. The error has not been fixed in the paper.
28 June 2018
Scientific Reports 6; Article number: 20974; published online: 09 February 2016; updated: 28 June 2018 The Acknowledgements section in this Article is incomplete. “We would like to thank G. Ditze, G. Sammler, F. Horn, M. Händel, K. Schoknecht, S. Bernhardt, and A. Kolchmeier for technical assistance. This work was supported by grants of the Deutsche Forschungsgemeinschaft to K.
References
Whitty, A. Cooperativity and biological complexity. Nat Chem Biol 4, 435–439, 10.1038/nchembio0808-435 (2008).
Hamill, O. P., Marty, A., Neher, E., Sakmann, B. & Sigworth, F. J. Improved patch-clamp techniques for high-resolution current recording from cells and cell-free membrane patches. Pflugers Arch 391, 85–100 (1981).
Lape, R., Colquhoun, D. & Sivilotti, L. G. On the nature of partial agonism in the nicotinic receptor superfamily. Nature 454, 722–727 (2008).
Kaupp, U. B. & Seifert, R. Cyclic Nucleotide-Gated Ion Channels. Physiol Rev 82, 769–824 (2002).
Zagotta, W. N. & Siegelbaum, S. A. Structure and function of cyclic nucleotide-gated channels. Annu Rev Neurosci 19, 235–263 (1996).
Kaupp, U. B. & Seifert, G. Molecular Diversity of Pacemaker Ion Channels. Annu Rev Physiol 63, 235–257 (2001).
Sautter, A., Zong, X., Hofmann, F. & Biel, M. An isoform of the rod photoreceptor cyclic nucleotide-gated channel beta subunit expressed in olfactory neurons. Proc Nat Acad Sci USA 95, 4696–4701 (1998).
Bönigk, W. et al. The native rat olfactory cyclic nucleotide-gated channel is composed of three distincts subunits. J Neurosci 19, 5332–5347 (1999).
Matulef, K. & Zagotta, W. N. Cyclic nucleotide-gated ion channels. Annu Rev Cell Dev Biol 19, 23–44 (2003).
Biskup, C. et al. Relating ligand binding to activation gating in CNGA2 channels. Nature 446, 440–443, 10.1038/nature05596 (2007).
Nache, V. et al. Activation of olfactory-type cyclic nucleotide-gated channels is highly cooperative. The Journal of physiology 569, 91–102, 10.1113/jphysiol.2005.092304 (2005).
Nache, V., Eick, T., Schulz, E., Schmauder, R. & Benndorf, K. Hysteresis of ligand binding in CNGA2 ion channels. Nature communications 4, 2864, 10.1038/ncomms3866 (2013).
Dhallan, R. S., Yau, K. W., Schrader, K. A. & Reed, R. R. Primary structure and functional expression of a cyclic nucleotide-activated channel from olfactory neurons. Nature 347, 184- 187 (1990).
Bonigk, W. et al. The native rat olfactory cyclic nucleotide-gated channel is composed of three distinct subunits. J Neurosci 19, 5332–5347 (1999).
Tibbs, G. R., Liu, D. T., Leypold, B. G. & Siegelbaum, S. A. A state-independent interaction between ligand and a conserved arginine residue in cyclic nucleotide-gated channels reveals a functional polarity of the cyclic nucleotide binding site. J Biol Chem 273, 4497–4505 (1998).
Goulding, E. H., Tibbs, G. R. & Siegelbaum, S. A. Molecular mechanism of cyclic-nucleotide-gated channel activation. Nature 372, 369–374 (1994).
Song, L. & Magleby, K. L. Testing for microscopic reversibility in the gating of maxi K+ channels using two-dimensional dwell-time distributions. Biophys J 67, 91–104, 10.1016/S0006-3495(94)80458-8 (1994).
Carter, P. J., Winter, G., Wilkinson, A. J. & Fersht, A. R. The use of double mutants to detect structural changes in the active site of the tyrosyl-tRNA synthetase (Bacillus stearothermophilus). Cell 38, 835–840 (1984).
Kusch, J. et al. Role of the S4-S5 linker in CNG channel activation. Biophys J 99, 2488–2496, 10.1016/j.bpj.2010.07.041 (2010).
Hidalgo, P. & MacKinnon, R. Revealing the Architecture of a K+ channel pore through mutant cycles with a peptide inhibitor. Science 268, 307–310 (1995).
Hlavackova, V. et al. Sequential inter- and intrasubunit rearrangements during activation of dimeric metabotropic glutamate receptor 1. Sci Signal 5, ra59, 10.1126/scisignal.2002720 (2012).
Nache, V. et al. Differential regulation by cyclic nucleotides of the CNGA4 and CNGB1b subunits in olfactory cyclic nucleotide-gated channels. Sci Signal 5, ra48, 10.1126/scisignal.2003110 (2012).
Acknowledgements
We would like to thank G. Ditze, G. Sammler, F. Horn, M. Händel, K. Schoknecht, S. Bernhardt, and A. Kolchmeier for technical assistance. This work was supported by grants of the Deutsche Forschungsgemeinschaft to K.B. and V.N. and of the Friedrich Schiller University to V.N.
Author information
Authors and Affiliations
Contributions
N.W., V.N. and J.S. carried out the measurements and analyzed the experimental data, apart from the global fit. V.N. prepared the figures. T.Z. inferred the mutations and constructed the concatamers. T.E., S.H. and E.S. performed the global fits. R.S. performed part of analyses. K.B. planned the project, designed the experiments and the global fit, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wongsamitkul, N., Nache, V., Eick, T. et al. Quantifying the cooperative subunit action in a multimeric membrane receptor. Sci Rep 6, 20974 (2016). https://doi.org/10.1038/srep20974
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep20974
This article is cited by
-
Enlightening activation gating in P2X receptors
Purinergic Signalling (2022)
-
Single-molecule imaging with cell-derived nanovesicles reveals early binding dynamics at a cyclic nucleotide-gated ion channel
Nature Communications (2021)
-
Unravelling the intricate cooperativity of subunit gating in P2X2 ion channels
Scientific Reports (2020)
-
Rhinella marina oocytes: a suitable alternative expression system for functional characterization of aquaglyceroporins
Scientific Reports (2019)
-
Hydrophobic alkyl chains substituted to the 8-position of cyclic nucleotides enhance activation of CNG and HCN channels by an intricate enthalpy - entropy compensation
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.