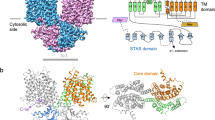
Abstract
The purine salvage pathway plays a major role in the nucleotide production, relying on the supply of nucleobases and nucleosides from extracellular sources. Although specific transporters have been suggested to be involved in facilitating their transport across the plasma membrane in mammals, those which are specifically responsible for utilization of extracellular nucleobases remain unknown. Here we present the molecular and functional characterization of SLC43A3, an orphan transporter belonging to an amino acid transporter family, as a purine-selective nucleobase transporter. SLC43A3 was highly expressed in the liver, where it was localized to the sinusoidal membrane of hepatocytes and the lung. In addition, SLC43A3 expressed in MDCKII cells mediated the uptake of purine nucleobases such as adenine, guanine and hypoxanthine without requiring typical driving ions such as Na+ and H+, but it did not mediate the uptake of nucleosides. When SLC43A3 was expressed in APRT/HPRT1-deficient A9 cells, adenine uptake was found to be low. However, it was markedly enhanced by the introduction of SLC43A3 with APRT. In HeLa cells, knock-down of SLC43A3 markedly decreased adenine uptake. These data suggest that SLC43A3 is a facilitative and purine-selective nucleobase transporter that mediates the cellular uptake of extracellular purine nucleobases in cooperation with salvage enzymes.
Similar content being viewed by others
Introduction
Nucleotides play vital roles in all living organisms both by forming the nucleic acids DNA and RNA, which store and implement genetic information and as individual monomers involved in biological signaling and energy cycling. The cellular pool of such nucleotides is strictly regulated by catabolism and biosynthesis; the latter employs both de novo and salvage pathways. Although the de novo pathway assembles purine and pyrimidine nucleotides from several fundamental molecules such as amino acids and glucose through multistep reactions, including those that produce nucleobases and nucleosides before their conversion to nucleotides, the salvage pathway performs the same task by simply reutilizing the nucleobases and nucleosides that are produced by the degradation of nucleotides within cells1. Nucleobases and nucleosides can also be salvaged extracellularly from dietary sources and from some tissues that produce excess nucleobases and nucleosides via the de novo pathway, thus supplementing the rather limited supply of degraded nucleotide products. In particular, in many types of cells exhibiting poor de novo synthesis activity, the salvage pathway has been suggested to play a major role in nucleotide production.
Extracellularly supplied nucleobases and nucleosides must be transported across the plasma membrane to be utilized by the salvage pathway. Because this process is generally difficult for this class of hydrophilic compounds to undergo by simple diffusion, the involvement of specific transporters has been suggested. For nucleosides, concentrative nucleoside transporters (CNTs/SLC28As) and equilibrative nucleoside transporters (ENTs/SLC29As) have been identified as involved in this process and their transport functions have been well characterized2,3,4,5. The CNT family includes CNT1/SLC28A1 and CNT2/SLC28A2, which operate unidirectionally for influx of nucleosides by a sodium-dependent secondary active mechanism at the apical membrane of the epithelial cells in several organs, typically the small intestine and kidney; the ENT family includes ENT1/SLC29A1 and ENT2/SLC29A2, which bidirectionally facilitate influx and efflux depending on the substrate concentration gradient at the basolateral membrane in epithelial cells of the same organs. The CNT family has one more member, CNT3/SLC28A3, which is expressed mainly in pancreas, trachea, bone marrow and mammary gland. The ENT family has two more members, ENT3/SLC29A3 and ENT4/SLC29A4, both of which are expressed in a wide variety of tissues and operate in a pH-dependent manner. The former is a nucleoside transporter that operates at the lysosomal membrane and the latter is redefined as plasma membrane monoamine transporter (PMAT), which operates multispecifically for the transport of monoamines and some other types of cationic compounds, as well as adenosine.
However, the molecular mechanisms underlying nucleobase transport are unresolved in mammals, although nucleobase transport systems have been suggested to occur in various cells and tissues. These hypothetical transport systems can be classified into secondary active transporters, which are coupled with sodium and facilitative transporters, just as nucleoside transporters are classified. ENT1 and ENT2 can mediate the transport of purine nucleobases, such as adenine, hypoxanthine and guanine, in a facilitative manner6,7; however, purine transport by ENTs, which can be inhibited by nucleosides, their principal substrates, cannot account for the purine nucleobase-selective transport system. Such a system has been observed in red blood cells, where it is involved in the facilitative uptake of adenine, which is inhibited competitively by hypoxanthine but not by nucleosides. Similar purine nucleobase-selective transport systems have been suggested to be present in several types of cells, such as the cell lines L9298, CHO8 and COR-L239, as well as in rabbit cornea epithelial cells10 and human cardiac microvascular endothelial cells11. These transport systems, such as those in red blood cells, are sodium independent, highly nucleobase selective in substrate recognition and insensitive to specific inhibitors of ENTs. Furthermore, the facilitative uptake of hypoxanthine and adenine has been observed even in cell lines that lack nucleoside transport systems12. These lines of evidence strongly suggest the presence of one or more facilitative nucleobase transporters in addition to ENTs. Although we have recently identified sodium-dependent nucleobase transporter 1 (SNBT1/Slc23a3) in rat as the first nucleobase transporter in mammals, this transporter differs from such facilitative nucleobase transporters in its sodium-dependent function and ability to transport uracil and guanine and their analogs13. Moreover, SNBT1 is pseudogenized in humans and some other higher primates. Thus, at least one facilitative nucleobase-specific transporter remains unidentified in mammals.
We here report that SLC43A3, an orphan transporter that is abundantly expressed in the liver and lung, can mediate the transport of purine nucleobases specifically. Based on the facilitative nature in its function, we have named it equilibrative nucleobase transporter 1 (ENBT1). A detailed functional analysis has shown that salvage enzymes cooperate in the cellular uptake of purine nucleobases mediated by ENBT1.
Results
Identification of SLC43A3 as a nucleobase transporter
To identify a candidate gene for purine nucleobase transport function, we conducted a comprehensive screen of human cDNA encoding putative transporters based on the uptake activity of [3H]adenine in HEK293 cells transiently expressing each cDNA. Among these cDNA, we observed that the clone encoding SLC43A3 induced an increase in the cellular uptake of [3H]adenine (Fig. 1). This purine nucleobase transport activity was also observed for [3H]hypoxanthine.
SLC43A3 is an orphan transporter belonging to an amino acid transporter family comprising three members, along with LAT3/SLC43A114 and LAT4/SLC43A215, both of which mediate the uptake of neutral amino acids such as leucine. Human SLC43A3 is 491 amino acid residues in length and predicted to have 12 transmembrane domains. The amino acid sequence of SLC43A3 is ~27% identical with those of LAT3 and LAT4 but not significantly similar to those of any other transporters, including nucleoside transporters. The orthologs of SLC43A3 are conserved among several mammalian species.
Functional characterization of SLC43A3
To characterize the transport function of SLC43A3 in detail, we generated MDCKII cells stably expressing human SLC43A3 (henceforth, SLC43A3 cells) and examined the uptake of various nucleobases and nucleosides in these cells. The uptake of purine nucleobases such as [3H]guanine and [3H]hypoxanthine as well as [3H]adenine were markedly greater in SLC43A3 cells than in mock cells, consistent with the purine nucleobase uptake of the HEK293 cells that transiently expressed SLC43A3, further suggesting that this transporter is a purine nucleobase transporter (Fig. 2A). Although [3H]xanthine is also a purine nucleobase, SLC43A3 cells demonstrated only minimal uptake of this nucleobase. Cellular uptake of [3H]uracil, a pyrimidine nucleobase, was also observed to be minimal in SLC43A3 cells. Nucleosides, such as [3H]adenosine, [3H]thymidine and [3H]uridine, did not exhibit increased uptake by SLC43A3 cells. Ascorbate was also included as a test compound because mammalian ascorbate transporters are orthologs of bacterial nucleobase transporters16, but this compound did not exhibit increased uptake by SLC43A3 cells. Among those compounds that appeared to be transported by SLC43A3, we selected adenine to be used as a probe substrate for further assessment of the transport function of SLC43A3 as it is one of the best characterized nucleobases exhibiting carrier-mediated transport in biological systems.
Functional characterization of ENBT1 stably expressed in MDCKII cells.
(A) Substrate specificity of ENBT1. ENBT1-mediated uptake was evaluated by dividing the uptake in the cells expressing ENBT1 by that in mock cells. The uptakes of 3H-labeled adenine, guanine, hypoxanthine, xanthine, uracil, adenosine, thymidine and uridine (10 nM for guanine and 5 nM for the others) and 14C-labeled ascorbate (50 μM) were evaluated at 37 °C for 1 min. The broken line is for unity. (B) Effect of extracellular ions on the ENBT1-specific uptake of [3H] adenine. NaCl in the control medium was replaced as indicated. The uptake of [3H]adenine (5 nM) was evaluated at 22 °C for 40 s in the cells expressing ENBT1 and mock cells and the ENBT1-specific uptake was calculated by subtracting the uptake in the latter from that in the former. (C) Effect of extracellular pH on the ENBT1-specific uptake of [3H] adenine. The uptake of [3H]adenine (5 nM) was evaluated at 22 °C for 40 s. (D) Effect of various compounds on the ENBT1-specific uptake of [3H]adenine. The uptake of [3H]adenine (5 nM) was evaluated at 22 °C for 40 s in the presence of an inhibitor (10 μM for decynium-22 and 200 μM for the others) or in its absence. *p < 0.05, n = 4.
The specific uptake of [3H]adenine by SLC43A3 cells was not altered by replacement of NaCl in the medium with any of the sodium or potassium salt of chloride or gluconate or mannitol, suggesting that this transporter does not require extracellular Na+ or Cl− for operation (Fig. 2B). In addition, uptake was not altered by changes in pH throughout a range of 5.0–8.0, suggesting that H+ is not required for its operation either (Fig. 2C). These results support the role of SLC43A3 as a facilitative nucleobase transporter permitting substrate nucleobases to rapidly equilibrate across the plasma membrane; accordingly, we named it ENBT1. This transporter also appears to transport purine nucleobases preferentially to uracil as a pyrimidine nucleobase, as suggested in Fig. 2A.
Effect of various compounds on ENBT1-mediated adenine transport
To further explore the substrate specificity of ENBT1, we examined the effect of various compounds (10 μM for decynium-22 and 200 μM for the others) on the ENBT1-mediated uptake of [3H]adenine in SLC43A3 cells (Fig. 2D). Purine nucleobases such as adenine, guanine and hypoxanthine, which were found to be substrates of SLC43A3 and hence, potential competitive inhibitors, inhibited [3H]adenine uptake of SLC43A3 cells. Moreover, purine and its several analogs, such as 6-thioguanine and mercaptopurine, showed inhibitory activities, suggesting that they are also substrates. In contrast, xanthine, a purine nucleobase and uracil, a pyrimidine nucleobase, did not exhibit any substantial inhibitory activity. Because these two molecules were transported by SLC43A3 minimally (Fig. 2A), they are likely to be recognized as very poor substrates with affinities extremely low to act as competitive inhibitors. The other pyrimidine nucleobases (thymine and cytosine) and 5-fluorouracil, an analog of uracil, did not exhibit any inhibitory activity either. Adenosine, a purine nucleoside and thymidine and uridine, which are pyrimidine nucleosides, did not exhibit any inhibitory activity, consistent with the finding that they are not transported by SLC43A3 (Fig. 2A). Several other purine and pyrimidine nucleosides did not exhibit any inhibitory activity either. Thus, ENBT1 did not recognize any purine or pyrimidine nucleosides as substrates (or competitive inhibitors). Nitrobenzylthioinosine (NBMPR), a potent inhibitor of ENT1, exhibited inhibitory activity, although its inhibition was not extensive. Dipyridamole, a potent inhibitor of both ENT1 and ENT2, did not exhibit inhibitory activity. Decynium-22, which has recently been reported as a specific inhibitor of a purine nucleobase transport system in PK15NTD cells17, was found to be a potent ENBT1 inhibitor that can inhibit [3H]adenine uptake almost completely at 10 μM. We also examined the effect of various amino acids (5 mM), as ENBT1 is a member of the amino acid transporter family14,15; however, none of them affected the adenine uptake activity of SLC43A3 cells (see Supplementary Table S1 online), supporting a previous study indicating that SLC43A3 expression did not induce any amino acid transport activity in Xenopus oocytes18.
Kinetic analysis
The time courses of [3H]adenine uptake were examined in both SLC43A3 cells and mock cells (Fig. 3A). The [3H]adenine uptake increased rapidly, exhibiting a linear increase for up to 1 min in SLC43A3 cells, whereas it was far slower in mock cells. Accordingly, we decided to evaluate the initial rate of nucleobase uptake at 40 s. Kinetic analysis indicated that the specific uptakes of [3H]adenine, [3H]guanine and [3H]hypoxanthine were saturable with the Km values of 0.94 μM, 1.70 μM and 1.32 μM, respectively (Fig. 3B). However, as shown in Fig. 3C, the adenine IC50 value for [3H]hypoxanthine uptake (13 μM) and the guanine and hypoxanthine IC50 values for [3H]adenine uptake (70 μM and 350 μM, respectively) were greater than the Km values for the respective nucleobases.
Kinetic analyses of the uptakes of purine nucleobase mediated by ENBT1 stably expressed in MDCKII cells.
(A) Time courses of the uptake of [3H] adenine (5 nM) was evaluated at 22 °C in the cells expressing ENBT1 (open circle) and mock cells (closed circle). (B) Concentration-dependent uptakes of adenine (•), guanine (○) and hypoxanthine (▼) were evaluated at 22 °C for 40 s in the cells expressing ENBT1 and mock cells. The ENBT1-specific uptake was calculated by subtracting the uptake in the latter from that in the former and was used for kinetic analysis. The values of Km (μM) are 0.94 ± 0.14, 1.70 ± 0.54 and 1.32 ± 0.26 for adenine, guanine and hypoxanthine, respectively, as the computer-fitted parameters with S.E. (n = 7–8) and those of Vmax (pmol/min/mg protein) are 107.8 ± 9.6, 98.2 ± 20.7 and 73.1 ± 9.2, respectively. Solid line, dotted line and dashed line are computer-fitted profiles for adenine, guanine and hypoxanthine, respectively. (C) Concentration-dependent inhibition of the ENBT1-mediated uptake of [3H]hypoxanthine by adenine (•) and that of [3H]adenine by guanine (○) and hypoxanthine (▼). The uptakes of [3H]hypoxanthine and [3H]adenine (5 nM for each) were evaluated at 22 °C for 40 s in the presence of a nucleobase as an inhibitor at specified concentrations or in its absence. The values of IC50 (μM) are 13.0 ± 3.7, 70.0 ± 46.3 and 350.0 ± 31.2 for adenine, guanine and hypoxanthine, respectively, as the computer-fitted parameters with S.E. (n = 8). Solid line, dotted line and dashed line are computer-fitted profiles for adenine guanine and hypoxanthine, respectively. All the data are presented as the means ± S.E. (n = 4).
When a substrate acts as a competitive inhibitor, its IC50 should be theoretically comparable to its inhibition constant (Ki) and its transport constant (Km) as long as IC50 is determined for the inhibition of the transport of a test substrate at a concentration much lower than its Km, as in the present study. Based on this, the discrepancies between Km and IC50 are unexpected findings.
However, this inconsistency can be explained by considering that each of the [3H]-labeled nucleobases is converted to its nucleotide by its specific salvage enzyme (adenine phosphoribosyltransferase (APRT) for adenine and hypoxanthine phosphoribosyltransferase 1 (HPRT1) for guanine and hypoxanthine) in SLC43A3 cells and thus accumulated in that form. This may maintain low concentrations of nucleobase forms within the cells, thus sustaining its inwardly directed concentration gradient across the plasma membrane, which is the driving force for facilitative transport mediated by ENBT1. Therefore, the kinetic parameters of the apparent uptake of [3H]-labeled nucleobases may be aggregates of the cooperative process that involves ENBT1 and the specific salvage enzymes. Because the Km of [3H]adenine uptake (0.94 μM) was comparable to those of APRT for adenine in the literature (0.6–0.9 μM), apparent [3H]adenine uptake may be rate-limited by APRT-mediated metabolism. Similarly, the Km of [3H]hypoxanthine uptake (1.32 μM) was comparable to that of HPRT1 for hypoxanthine (3 μM), suggesting that apparent [3H]hypoxanthine uptake could be rate-limited by HPRT1-mediated metabolism. On the other hand, within experiments evaluating the adenine IC50 value for [3H]hypoxanthine uptake, adenine acts on ENBT1 but not HPRT1. Therefore, the empirical IC50 value (13 μM) may represent the affinity of ENBT1 for adenine. This IC50 value was equal to the Km value for adenine uptake in human erythrocytes (also 13 μM19), which is likely to represent the transporter affinity. In addition, the IC50 value is much lower than the Km values of ENT1 (3.2 mM) and ENT2 (1.8 mM) for adenine in the literature6,7 and closer to adenine concentrations in plasma (~1 μM), suggesting that ENBT1 operates more effectively at physiologically relevant adenine concentrations than ENTs. Similarly, the guanine and hypoxanthine IC50 values (70 and 350 μM, respectively) for [3H]adenine uptake are likely to represent the affinity of guanine and hypoxanthine for ENBT1 because they act on ENBT1 but not APRT.
Cooperative operation of ENBT1 with salvage enzymes
To explore the effect of such salvage enzymes on nucleobase uptake mediated by ENBT1, we conducted a series of uptake experiments with mouse fibroblast-derived A9 cells, which are deficient in APRT and HPRT1. The uptake of [3H]adenine in A9 cells was low and not enhanced by the introduction of human ENBT1. However, the introduction of both human APRT and ENBT1 resulted in a significant increase in [3H]adenine uptake, suggesting their cooperation in adenine uptake (Fig. 4A). Although [3H]adenine uptake was also increased by the introduction of APRT alone, the increase could be due to the involvement a native nucleobase transporter present in A9 cells. The introduction of human HPRT1, instead of APRT, into A9 cells did not induce any increase in [3H]adenine uptake, regardless of the presence or absence of cointroduced ENBT1 (Fig. 4B), suggesting the necessity of APRT for adenine uptake in cooperation with ENBT1. It should be noted additionally that [3H]adenine uptake increased in proportion to time at least up to 2 min in A9 cells expressing APRT alone and those expressing APRT with ENBT1, indicating that the uptake period of 1 min was in that range and appropriate for the evaluation of initial uptake process (see Supplementary Fig. S2 online). Although [3H]adenine uptake in A9 cells expressing ENBT1 alone was too small to confirm such an initial uptake phase, [3H]adenine uptake was greater in A9 cells expressing APRT with ENBT1 than in those expressing ENBT1 alone at the earliest time of 20 s and linearly increased with time for a more prolonged period in the former than in the latter. Therefore, it is most likely that APRT worked for accelerating and enhancing [3H]adenine uptake by metabolic channelling in cooperation with ENBT1. Similarly to the results for [3H]adenine uptake, [3H]guanine uptake was enhanced by the introduction of HPRT1 alone and furthermore by the additional introduction of ENBT1, whereas it was not enhanced in any A9 cells with APRT or ENBT1 alone (Fig. 4C,D). Thus, HPRT1 appears to be necessary for guanine uptake in cooperation with ENBT1.
Cooperative operation of ENBT1 with salvage enzymes for purine nucleobases.
The uptake of [3H]adenine (5 nM) was evaluated at 37 °C for 1 min in APRT/HPRT1-deficient A9 cells transfected with a plasmid for ENBT1 and one for APRT (A) or HPRT1 (B) with 1: 1 ratio (1 μg of total plasmid). The uptake of [3H]guanine (5 nM) was evaluated similarly in A9 cells transfected with a plasmid for ENBT1 and one for APRT (C) or HPRT1 (D). *Significantly different from control at p < 0.05. †Significantly different from APRT or HPRT1 alone at p < 0.05. All the data are presented as the means ± S.E. (n = 3 or 4).
Similarly, silencing ENBT1 or APRT by RNA interference in HeLa cells, which exhibit native adenine uptake, substantially decreased [3H]adenine uptake (Fig. 5A), in accordance with reduced expression levels of each protein (Fig. 5B).
Effect of silencing of ENBT1 and APRT on [3H]adenine uptake in HeLa cells.
(A) The uptake of [3H]adenine (5 nM) was evaluated at 37 °C for 1 min in HeLa cells transfected with siRNA for ENBT1 or APRT. *p < 0.05, n = 4. (B) The endogenous protein levels for ENBT1, APRT and β-actin in those cells were analyzed by western blotting.
Collectively, these results suggest that ENBT1 is involved in the utilization of extracellular purine nucleobases through functional interactions with salvage enzymes.
Tissue distribution and cellular localization of ENBT1
To examine the expression pattern of ENBT1 in human tissues, we performed quantitative real-time RT-PCR. ENBT1 mRNA was expressed in all the tissues examined, although at the highest levels in the liver and lung, followed by the pancreas (Fig. 6A).
Tissue distribution and cellular localization of ENBT1.
(A) The mRNA level of ENBT1 was determined by real-time PCR and standardized by the mRNA level of GAPDH in each human tissue (n = 4). (B) The immunofluorescent image shows the subcellular localization of ENBT1 (green, white arrow) and P-glycoprotein (red, pink arrow) in the human liver.
Immunofluorescence staining of human liver showed that ENBT1 was localized to the basolateral (sinusoidal) membrane of human hepatocytes and was clearly distinguished from P-glycoprotein, which localized to the apical (canalicular) membrane (Fig. 6B). Basolateral localization of ENBT1 was also observed by confocal laser scanning microscopy in polarized MDCKII cells stably expressing ENBT1 fused with green fluorescent protein (see Supplementary Fig. S1 online).
Discussion
SLC43A3 was originally identified as embryonic epithelia gene 1 (EEG1), a gene upregulated in a cell culture model of kidney development20. Because its mRNA is also expressed in several embryonic tissues, such as lung and liver and its peak expression occurs during embryogenesis, it has been implicated in the processes of cellular proliferation and tissue development. In the present study, we have demonstrated that ENBT1/SLC43A3 is a facilitative and purine-selective nucleobase transporter that mediates the cellular uptake of extracellular purine nucleobases, a key step for their efficient utilization in the salvage pathway. This is the first report that has identified a purine-specific nucleobase transporter in mammals.
The presence of at least three types of transport systems for the uptake of hypoxanthine in human has been postulated based on differences in sensitivity to inhibitors21. Two of these hypothetical transports are inhibited by nucleosides competitively and also by dipyridamole; in conjunction with recent ENT studies, ENT1 and ENT2 have been identified as responsible for one of the mechanisms inhibited by NBMPR and the other NBMPR-insensitive mechanism6,7. The third system is not inhibited by either NBMPR or dipyridamole, but it is competitively inhibited by adenine but not nucleosides. These characteristics are in agreement with those of ENBT1, suggesting that ENBT1 is the transporter responsible for this nucleobase uptake system. Although adenine transport by ENBT1 was inhibited by NBMPR by approximately 40% at a 200 μM concentration (Fig. 2D), this fact implies that ENBT1 is far less sensitive to this inhibitor than ENT1, which is inhibited with a Ki of 2 nM.
The hypoxanthine uptake of human cardiac microvascular endothelial cells has been previously shown to be insensitive to NBMPR and dipyridamole but extensively inhibited by adenine with an IC50 value of 19 μM11, which is comparable with that of adenine IC50 for ENBT1 in the present study. Furthermore, a recent transcriptome analysis has revealed endothelial-specific SLC43A3 expression in broad types of microvasculatures22. As such, ENBT1 is likely to be functionally expressed in microvascular endothelial cells, where it may play an important role in the formation and homeostasis of blood vessels by regulating the cellular uptake of purine nucleobases. Furthermore, significant increases in SLC43A3 mRNA expression levels have been reported in the monocytes from blood of healthy volunteers infused with lipopolysaccharide23 and in thyroid carcinomas from patients exposed to radioactive iodine24, suggesting pathophysiological roles of ENBT1 in inflammation and carcinoma.
Among organs, the liver is the major one having de novo purine nucleobase synthesis activity. Therefore, high ENBT1 expression in the liver likely produces a net efflux of synthesized purine nucleobases, which could then be supplied to the other organs via the systemic circulation. For example, the lung receives all the systemic venous blood gathered to the heart and hence, may receive all the nucleobases from the liver before they are distributed to the other organs through the systemic arteries. Moreover, the lung is rich in salvage enzymes that can produce nucleotides from nucleobases in cooperation with ENBT1, whereas it contains relatively less amount of de novo enzymes25,26. Therefore, high ENBT1 expression in the lung potentially enables the uptake of nucleobases from the liver and the cooperative operation of ENBT1 and salvage enzymes could be the major mechanism for the production of nucleotides in the lung, which includes pulmonary endothelial cells, alveolar epithelial cells and residential lymphoid cells. This would be of particular importance in the repair and growth of lung tissue injured by oxidative stress, which may be induced by respiratory exposure to oxygen. Lung injury is often caused by an adverse event associated with the use of several drugs, such as methotrexate and leflunomide, which act on cells actively synthesizing nucleotides27, including hepatocytes, by inhibiting de novo nucleotide synthesis; a reduction of nucleobases supplied by the liver may be indirectly involved in this damage. Thus, it is likely that the lungs rely heavily on the supply of nucleobases from the liver to maintain their function and that ENBT1 plays a critical role in their trafficking.
On the other hand, Bodoy et al.18 recently demonstrated that mice harboring a nonsense mutation in Slc43a3 are phenotypically normal, without any sign of embryonic lethality. Other transporters, including nucleoside transporters, may have compensated for the functional defect of ENBT1. Further studies are needed to elucidate the role of ENBT1 in the utilization of purine nucleobases and homeostasis of nucleotides in vivo.
In conclusion, we have successfully identified the function of SLC43A3 as a novel, facilitative, purine-selective nucleobase transporter and demonstrated its cooperative operation with salvage enzymes in the cellular utilization of purine nucleobases. The functional characteristics of ENBT1 were observed to be almost completely in agreement with those of the previously described carrier-mediated transport system involved in the uptake of adenine and hypoxanthine in many types of cells and tissues. Therefore, it is most likely that SLC43A3 is its molecular entity. This transporter is apparently responsible for the cellular uptake of extracellularly supplied purine nucleobases, facilitating their salvage and subsequent use in the production of nucleotides. The present study provides new insights into the complexity of intracellular nucleotide level homeostasis and could lead to opportunities to develop new classes of nucleobase analogs for the treatment of cancer and viral infection and also optimal therapeutic strategies for such diseases.
Methods
Materials
[3H]adenine (25.0 Ci/mmol), [3H]hypoxanthine (27.0 Ci/mmol), [3H]thymidine (20.0 Ci/mmol), [3H]uridine (14.7 Ci/mmol), [3H]uracil (42.8 Ci/mmol), [3H]xanthine (12.8 Ci/mmol) and [3H]adenosine (39.2 Ci/mmol) were obtained from Moravek Biochemicals (Brea, CA), [14C]guanine (55 mCi/mmol) was from American Radiolabeled Chemicals (St. Louis, MO) and [14C]ascorbate (8.5 mCi/mmol) was from PerkinElmer Life Sciences (Boston, MA). Adenine, papaverine and dipyridamole were obtained from Sigma-Aldrich (St. Louis, MO) and hypoxanthine, guanine and nitrobenzylthioinosine were from Wako Pure Chemicals (Osaka, Japan). All other reagents were of analytical grade and commercially obtained.
Cell culture
HEK293, HeLa, A9 and MDCKII cells were maintained at 37 °C and 5% CO2 in Dulbecco’s modified Eagle’s medium supplemented with 10% fetal bovine serum, 100 units/ml penicillin and 100 μg/ml streptomycin.
Preparation of HEK293 cells transiently expressing ENBT1
HEK293 cells (1.5 × 105 cells/well initially) were grown on 24-well plates coated with poly-L-lysine for 12 h, transfected with 1 μg/well of the plasmid carrying the cDNA of ENBT1 by using 1.5 μl/well of Lipofectamine 2000 (Invitrogen, Carlsbad, CA) as a transfection reagent, according to the manufacturer’s instructions and cultured for 36 h for transient expression.
Transient expression of ENBT1, APRT and HPRT1 in A9 cells
For the transient expression of ENBT1 with APRT or HPRT1, A9 cells (2.0 × 105 cells/well initially) were seeded on 24-well plates, transfected with the plasmid carrying the cDNA of ENBT1 and the one carrying the cDNA of APRT or HPRT1 by using Lipofectamine LTX (Invitrogen) as a transfection reagent, according to the manufacturer’s instructions and cultured for 48 h. The cells were transfected with 1 μg/well of total plasmids with a ratio of 1:1 for the one for ENBT1 and the one for APRT, or HPRT1. For the expression of one of them alone, a half of the total amount of plasmids was replaced with the empty pCI-neo vector.
Preparation of MDCKII cells stably expressing ENBT1
MDCKII cells were transfected with the plasmid carrying the cDNA of ENBT1 by using Lipofectamine 2000 and cultured in DMEM supplemented with 10% FBS and 800 μg/ml Geneticin for 2 to 3 weeks. Antibiotic-resistant clones were selected and tested for the transport of [3H]adenine as a probe substrate.
Silencing of ENBT1 and APRT in HeLa cells
HeLa cells (2.5 × 104 cells/well initially) were grown on 24-well plates coated with poly-L-lysine for 6 h, transfected with 4 pmol/well of the siRNA (StealthSelect RNAiTM, Invitrogen) specific to the mRNA of ENBT1 or APRT by using 0.5 μl/well of Lipofectamine RNAi MAX (Invitrogen), according to the manufacturer’s instructions and cultured for 72 h for silencing of ENBT1 or APRT. For control, Stealth RNAiTM Negative Control Low GC Duplex (Invitrogen) was used. The sequences of siRNA were as follows: ENBT1 sense, 5′- AAU GUC AGA AGA AUG GCA AGC AUG A -3′; ENBT1 antisense, 5′- UCA UGC UUG CCA UUC UUC UGA CAU U -3′; APRT sense, 5′-AGA UGU CCC UGA AUA CCA CGC CUG G-3′; APRT antisense, 5′- CCA GGC GUG GUA UUC AGG GAC AUC U -3′.
Transport Study
MDCKII cells stably expressing ENBT1 (1.5 × 105 cells/well initially) were grown on 24-well plates for 72 h. The cells in each well were preincubated in 0.25 ml of substrate-free uptake buffer (140 mM NaCl, 5 mM KCl, 0.4 mM KH2PO4, 0.8 mM MgSO4, 1.0 mM CaCl2, 25 mM glucose and 10 mM HEPES, pH 7.4) for 5 min, unless otherwise indicated. Uptake assays were started by replacing the substrate-free uptake buffer for preincubation with one containing the radioisotope-labeled substrate (0.25 ml). All of the procedures were conducted at 22 °C or 37 °C. Assays were stopped by addition of ice-cold substrate-free uptake buffer (2 ml) and the cells were washed two times with 2 ml of the same buffer. The cells were solubilized in 0.5 ml of 0.2 M NaOH solution containing 0.5% SDS at room temperature for 1 h and the associated radioactivity was measured by liquid scintillation counting. Cellular protein content was determined by the method of Lowry et al.28 using bovine serum albumin as the standard. Uptake assays were also conducted in mock cells, which were transfected with empty pCI-neo vector, to estimate nonspecific uptake.
Uptake assays were similarly conducted in HEK293 cells transiently expressing ENBT1, A9 cells transiently expressing ENBT1 and/or APRT, or HPRT1 and HeLa cells treated for silencing ENBT1 or APRT.
Quantification of ENBT1 mRNA by real-time PCR
Real-time quantitative PCR was carried out using a THUNDERBIRD SYBR qPCR Mix (Toyobo) on a 7300 Fast Real-Time PCR System (Applied Biosystems, Forest City, CA). The expression levels were normalized with that of GAPDH. The sequences of primers were as follows: ENBT1 forward primer, 5′- ATT TTC CAA GTG CTC AAA CGC-3′; ENBT1 reverse primer, 5′-CTG CCA AGG CTA AGT GCA AGG -3′; GAPDH forward primer, 5′- CGG AGT CAA CGG ATT TGG TCG TAT -3′; GAPDH reverse primer, 5′- AGC CTT CTC CAT GGT GGT GAA GAC -3′.
Immunofluorescence staining
A specimen of human normal adult liver frozen tissue sections (BioChain Institute, Inc., Newark, CA) was washed three times with PBS and then incubated for 5 min at room temperature with 3% hydrogen peroxide in methanol to inactivate endogenous peroxidase. After washing three times with PBS, the tissues were incubated for 30 min at room temperature with 5% normal goat serum in PBS to block nonspecific signals. The tissues were washed three times with PBS and then incubated for overnight at 4 °C with the primary antibodies of anti-human SLC43A3 rabbit-polyclonal antibody and anti-human P-glycoprotein mouse-monoclonal antibody (GeneTex Inc.) in Can Get Signal ImmunoStain Solution B (Toyobo). The tissues were washed three times with 0.1% BSA in PBS and those primary antibodies were probed with anti-rabbit IgG F(ab’)2 Alexa Fluor 488 and anti-mouse IgG F(ab’)2 Alexa Fluor 555 (Cell signaling Technology, Beverly, MA), respectively, by incubating for 1 h at room temperature and visualized by using a confocal laser-scanning microscope (LMS510; Zeiss, Jena, Germany).
Data Analysis
The saturable transport of each substrate, adenine, guanine and hypoxanthine, by ENBT1 was analyzed by assuming Michaelis-Menten type carrier-mediated transport represented by the following equation: v = Vmax × s/(Km × s). The maximum transport rate (Vmax) and the Michaelis constant (Km) were estimated by fitting this equation to the experimental profile of the uptake rate (v) versus the substrate concentration (s), using a non-linear least-squares regression analysis program, WinNonlin (Pharsight Corp., Mountain View, CA) and the reciprocal of variance as the weight.
When s is much smaller than Km (s Km), the v in the presence of a competitive inhibitor can be described as follows: v = v0/(1 + (i/IC50)n). The half-inhibition concentration (IC50) was estimated together with the Hill coefficient (n) and v in the absence of inhibitors (v0) by fitting this equation to the experimental profile of v versus the inhibitor concentration (i).
Experimental data are presented as the means ± S.E. and statistical analysis was performed by using Student’s t test or, when multiple comparisons were needed, analysis of variance followed by Dunnett’s test, with p < 0.05 considered significant.
Western blot analysis, cDNA cloning and other detailed methods are provided in Supplementary Information.
Additional Information
How to cite this article: Furukawa, J. et al. Functional identification of SLC43A3 as an equilibrative nucleobase transporter involved in purine salvage in mammals. Sci. Rep. 5, 15057; doi: 10.1038/srep15057 (2015).
Change history
04 March 2016
A correction has been published and is appended to both the HTML and PDF versions of this paper. The error has not been fixed in the paper.
References
Murray, A. W. The biological significance of purine salvage. Annu. Rev. Biochem. 40, 811–826 (1971).
Gray, J. H., Owen, R. P. & Giacomini, K. M. The concentrative nucleoside transporter family, SLC28. Pflugers Arch. 447, 728–734 (2004).
Pastor-Anglada, M., Cano-Soldado, P., Errasti-Murugarren, E. & Casado, F. J. SLC28 genes and concentrative nucleoside transporter (CNT) proteins. Xenobiotica 38, 972–994 (2008).
Baldwin, S. A. et al. The equilibrative nucleoside transporter family, SLC29. Pflugers Arch. 447, 735–743 (2004).
Young, J. D., Yao, S. Y., Sun, L., Cass, C. E. & Baldwin, S. A. Human equilibrative nucleoside transporter (ENT) family of nucleoside and nucleobase transporter proteins. Xenobiotica 38, 995–1021 (2008).
Yao, S. Y. M. et al. Functional and molecular characterization of nucleobase transport by recombinant human and rat equilibrative nucleoside transporters 1 and 2. Chimeric constructs reveal a role for the ENT2 helix 5-6 region in nucleobase translocation. J. Biol. Chem. 277, 24938–24948 (2002).
Yao, S. Y. M., Ng, A. M. L., Cass, C. E., Baldwin, S. A. & Young, J. D. Nucleobase transport by human equilibrative nucleoside transporter 1 (hENT1). J. Biol. Chem. 286, 32552–32562 (2011).
Puziss, M. B., Wohlhueter, R. M. & Plagemann, P. G. Adenine transport and binding in cultured mammalian cells deficient in adenine phosphoribosyltransferase. Mol. Cell. Biol. 3, 82–90 (1983).
Marshman, E., Taylor, G. A., Thomas, H. D., Newell, D. R. & Curtin, N. J. Hypoxanthine transport in human tumour cell lines: relationship to the inhibition of hypoxanthine rescue by dipyridamole. Biochem. Pharmacol. 61, 477–484 (2001).
Majumdar, S., Tirucherai, G. S., Pal, D. & Mitra, A. K. Functional differences in nucleoside and nucleobase transporters expressed on the rabbit corneal epithelial cell line (SIRC) and isolated rabbit cornea. AAPS PharmSci 5, E15 (2003).
Bone, D. B. J. & Hammond, J. R. Nucleoside and nucleobase transporters of primary human cardiac microvascular endothelial cells: characterization of a novel nucleobase transporter. Am. J. Physiol. Heart Circ. Physiol. 293, H3325– H3332 (2007).
Ullman, B., Patrick, J. & McCartan, K. Expression of the high-affinity purine nucleobase transporter in mutant mouse S49 cells does not require a functional wild-type nucleoside-nucleobase transporter. Mol. Cell. Biol. 7, 97–103 (1987).
Yamamoto, S. et al. Identification and functional characterization of the first nucleobase transporter in mammals: implication in the species difference in the intestinal absorption mechanism of nucleobases and their analogs between higher primates and other mammals. J. Biol. Chem. 285, 6522–6531 (2010).
Babu, E. et al. Identification of a novel system L amino acid transporter structurally distinct from heterodimeric amino acid transporters. J. Biol. Chem. 278, 43838–43845 (2003).
Bodoy, S. et al. Identification of LAT4, a novel amino acid transporter with system L activity. J. Biol. Chem. 280, 12002–12011 (2005).
Gournas, C., Papageorgiou, I. & Diallinas, G. The nucleobase-ascorbate transporter (NAT) family: genomics, evolution, structure-function relationships and physiological role. Mol. Biosyst. 4, 404–416 (2008).
Hoque, K. M., Chen, L., Leung, G. P. H. & Tse, C. M. A purine-selective nucleobase/nucleoside transporter in PK15NTD cells. Am. J. Physiol. Regul. Integr. Comp. Physiol. 294, R1988–R1995 (2008).
Bodoy, S., Fotiadis, D., Stoeger, C., Kanai, Y. & Palacín, M. The small SLC43 family: facilitator system l amino acid transporters and the orphan EEG1. Mol. Aspects Med. 34, 638–645
Domin, B. A., Mahony, W. B. & Zimmerman, T. P. Purine nucleobase transport in human erythrocytes. Reinvestigation with a novel “inhibitor-stop” assay. J. Biol. Chem. 263, 9276–9284 (1988).
Stuart, R. O. et al. EEG1, a putative transporter expressed during epithelial organogenesis: comparison with embryonic transporter expression during nephrogenesis. Am. J. Physiol. Renal Physiol. 281, F1148– F1156 (2001).
Griffith, D. A. & Jarvis, S. M. Nucleoside and nucleobase transport systems of mammalian cells. Biochim. Biophys. Acta 1286, 153–181 (1996).
Wallgard, E. et al. Identification of a core set of 58 gene transcripts with broad and specific expression in the microvasculature. Arterioscler. Thromb. Vasc. Biol. 28, 1469–1476 (2008).
Sivapalaratnam, S. et al. Identification of candidate genes linking systemic inflammation to atherosclerosis; results of a human in vivo LPS infusion study. BMC Med. Genomics 4, 64 (2011).
Abend, M. et al. Iodine-131 dose dependent gene expression in thyroid cancers and corresponding normal tissues following the Chernobyl accident. PLoS One 7, e39103 (2012).
Natsumeda, Y., Prajda, N., Donohue, J. P., Glover, J. L. & Weber, G. Enzymic capacities of purine de Novo and salvage pathways for nucleotide synthesis in normal and neoplastic tissues. Cancer Res. 44, 2475–2479 (1984).
Mentzer, R. M., Rubio, R. & Berne, R. M. Release of adenosine by hypoxic canine lung tissue and its possible role in pulmonary circulation. Am. J. Physiol. 229, 1625–1631 (1975).
Roubille, C. & Haraoui, B. Interstitial lung diseases induced or exacerbated by DMARDS and biologic agents in rheumatoid arthritis: a systematic literature review. Semin. Arthritis Rheum. 43, 613–626 (2014).
Lowry, O. H., Rosebrough, N. J., Farr, A. L. & Randall, R. J. Protein measurement with the Folin phenol reagent. J. Biol. Chem. 193, 265–275 (1951).
Acknowledgements
This work was supported in part by JSPS KAKENHI grant No. C2459020 (to K.I.) and the research project grants from the Nakatomi Foundation (to K.I.) and the Aichi Cancer Research Foundation (to K.I.).
Author information
Authors and Affiliations
Contributions
J.F., K.I., J.M. and H.Y. designed research; J.F., J.M., T.Y. and K.O. performed research; Y.K., T.T. and H.M. contributed new reagents/analytic tools; J.F., K.I., J.M. and H.Y. analyzed data; J.F., K.I. and H.Y. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Furukawa, J., Inoue, K., Maeda, J. et al. Functional identification of SLC43A3 as an equilibrative nucleobase transporter involved in purine salvage in mammals. Sci Rep 5, 15057 (2015). https://doi.org/10.1038/srep15057
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep15057
This article is cited by
-
Neuroinflammatory astrocyte subtypes in the mouse brain
Nature Neuroscience (2021)
-
Model Analysis of the Apparent Saturation Kinetics of Purine Nucleobase Uptake in Cells co-Expressing Transporter and Metabolic Enzyme
Pharmaceutical Research (2021)
-
Quantitative comparative analysis of human erythrocyte surface proteins between individuals from two genetically distinct populations
Communications Biology (2019)
-
In-silico gene essentiality analysis of polyamine biosynthesis reveals APRT as a potential target in cancer
Scientific Reports (2017)
-
The influence of menstrual cycle and endometriosis on endometrial methylome
Clinical Epigenetics (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.