Abstract
We demonstrate that Magnetic Particle Imaging (MPI) enables monitoring of cellular grafts with high contrast, sensitivity and quantitativeness. MPI directly detects the intense magnetization of iron-oxide tracers using low-frequency magnetic fields. MPI is safe, noninvasive and offers superb sensitivity, with great promise for clinical translation and quantitative single-cell tracking. Here we report the first MPI cell tracking study, showing 200-cell detection in vitro and in vivo monitoring of human neural graft clearance over 87 days in rat brain.
Similar content being viewed by others
Introduction
The evaluation of cell transplantation efficacy and integration currently relies on destructive analytical methods like histology or functional improvements that can take months to manifest1. Inaccuracies in cell localization during initial delivery are common, with one clinical study reporting a 50% misadministration rate by experienced operators under ultrasound guidance2. Late detection of such outcomes can seriously impact the development of cell-based therapies. Clinical translation and adoption of cell therapies could be accelerated by safe, quantitative in vivo imaging methods3.
All standard preclinical imaging modalities have been used to track cells in vivo with varying degrees of success3. Bioluminescence and fluorescence imaging are the most common preclinical cell-tracking techniques; however, shallow optical penetration depth prevents linear quantification and clinical translation. Nuclear imaging techniques like Positron Emission Tomography (PET) have excellent tracer sensitivity and depth penetration, but face limitations in tracer half-life and radiation dose.
Clinically, most cell tracking studies have used super-paramagnetic iron oxide (SPIO) labeled cells, because SPIO-based methods have minimal effects on cell viability, proliferation and differentiation4,5,6,7, along with excellent depth penetration and in vivo persistence measured in months. However, the primary challenge for MRI-based SPIO cell tracking is that SPIOs induce MRI signal dropouts that are difficult to distinguish from tissues with naturally low MRI signal (e.g., bones, tendon, lungs, or any tissues near air). Moreover, MRI methods with positive contrast suffer from robustness and sensitivity challenges8,9. Until now, no technique has been able to combine high specificity with sensitive and quantitative imaging of the distribution and in vivo fate of SPIO-labeled cells3,4.
Magnetic Particle Imaging10,11 is a technique introduced in 2005 by Philips Research that directly images the intense magnetization of SPIOs rather than indirectly detecting SPIOs via MRI signal dropouts. Briefly, MPI uses a magnetic gradient field to saturate all SPIO magnetization outside a central field-free region, which we show as a “field-free line”, or FFL, in Fig. 1a. To produce an image, the FFL is rapidly rastered over an imaging volume via the application of a time varying magnetic field (called a “drive field”) produced by an electromagnet. As the FFL traverses a location containing SPIO nanoparticles, the ensemble SPIO magnetization responds by changing in magnitude and orientation12. This time-varying particle magnetization induces a voltage in the receiver coil, which can be assigned to the instantaneous FFL location to produce a MPI image11,13. Importantly, while the magnetization behavior of superparamagnets is nonlinear, the voltage signals generated in MPI are linearly proportional to the number of SPIOs at the instantaneous FFL location, enabling quantification of SPIO number13. The MPI induction signal is detectable with even a miniscule mass of tracer (5 ng Fe/voxel in our projection MPI scanner) because the SPIO magnetization saturates to 600 mT. By comparison, the nuclear paramagnetism of water in a 7 T MRI scanner is only 27 nT. That is, MPI images a magnetization that is 22 million times more intense than we image routinely in high-field MRI. Moreover, biological tissue does not generate or attenuate the low-frequency magnetic fields used in MPI (10 kHz to 1 MHz)14, giving the technique ideal contrast independent of source depth. The combination of these characteristics enables MPI to be uniquely suitable for high-contrast, radiation-free cell tracking in vivo15.
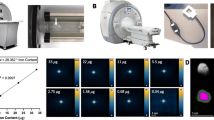
MPI is a tracer imaging modality that can image tagged cells using time varying magnetic fields.
(a) Diagram of a projection MPI scanner. Three superposed magnets (field-shifting magnets, gradient magnets and transmit coil) produce and translate a magnetic field-free line (FFL) across an imaging volume. As the FFL is rastered across a distribution of superparamagnetic iron oxide (SPIO) particles using a spatial imaging trajectory, the particle ensemble magnetization changes in magnitude and orientation in response. The changing particle magnetization is detected via a detector coil and reconstructed to form an image. (b) Plot of MPI signal from 9 SPIO-labeled cell populations ranging from 1 × 104 to 1 × 106 cells, shown with MPI system noise floor. SPIO signal in MPI is linear with cell number (R2 = 0.994, linear fit and 95% confidence bounds for fitted slope shown). (c) Detection threshold for MPI cell tracking in FFL scanner. Approximately 1,000 SPIO-labeled hESC-derived cells in a 100 μL in vitro suspension were imaged using MPI and compared to an empty scan. The signal-noise ratio of the image is estimated at above 5, giving a detection limit of approximately 200 cells. This represents the current detection sensitivity limit of our FFL MPI scanner, but the theoretical detection limit may be as low as a single cell. FOV: 6 cm × 8 cm. 20 second MPI acquisition. (d) MPI 1D line profile of Resovist-labeled cells and Resovist point sources. The FWHM resolution of the MPI signal is approximately 1.5-fold better for Resovist SPIO particles alone, at 5 mm, than for SPIO particles transfected into cells, at 7.7 mm. Scale bar: 1 cm. (e) MPI estimates for average cellular iron content (27.0 ± 3.3 pg/cell) correspond with ICP analysis (26.8 ± 0.3 pg/cell), making MPI useful for nondestructive iron quantification. n = 3 for ICP and n = 4 for MPI; results are mean ± SD.
To test MPI for tracking cells in vivo, we built two small animal MPI scanners, the construction of which are previously described and analyzed16,17. These include a projection-format FFL scanner with 2.35 × 2.35 T/m gradient, which achieves roughly 4 mm × 8 mm resolution along and orthogonal to the axis of the imaging bore respectively using undiluted Resovist tracer18. The FFL scanner’s projection-mode image acquisition enables fast scanning times on the order of seconds for a small animal-sized imaging volume. We also report the use of a field-free point (FFP) MPI scanner with a 7 × 3.5 × 3.5 T/m gradient, which enables roughly 2 mm × 4 mm × 2 mm resolution using Resovist and higher resolution using MPI-tailored SPIOs17,19. An example MPI image of labeled cells scanned using the FFP scanner is shown in Fig. 2. Both scanners use our group’s recent advances in MPI reconstruction that are necessary and sufficient for linear quantification13. We also note that the experimental detection sensitivity of the FFL scanner is, at roughly 5 ng Fe/voxel, 15-fold better than the FFP scanner, as expected due to a larger voxel volume. Based on the high sensitivity and faster scan times of the FFL scanner, we conducted all MPI imaging in this study, with the exception of the image shown in Fig. 2, using the FFL scanner.
FFP imaging of an acrylic phantom filled with six populations of SPIO-labeled cells.
The MPI signal from cell populations (ranging from 4 × 105 to 1.6 × 106 in number) corresponds linearly to iron oxide tracer concentration, enabling quantitative imaging. Imaging parameters: 3.5 min acquisition on a 7 T/m 3D MPI scanner, 5 cm × 4.5 cm × 3 cm FOV. Scale bar: 1 cm.
Results
Linear and Absolute Quantification
We first asked whether our MPI scanners could quantify SPIO-labeled cells in vitro. We modified an established MRI labeling protocol to label hESC-derived cells with Resovist SPIO tracer using a protamine sulfate transfection agent5. Both Resovist and protamine sulfate are approved for clinical use in the European Union and have no significant effect on cell proliferation, viability and differentiation5,6,7. Using the projection-mode MPI imager, we imaged nine labeled hESC-derived cell populations ranging from 1 × 104 to 1 × 106 cells. The normalized MPI signal was linear to the number of cells imaged (R2 > 0.994, Fig. 1b). In equivalent control cell populations without iron oxide labeling, no MPI signal was detected.
We further asked if MPI allows absolute quantification of iron oxide tracer. To determine this, we created a calibration curve using the MPI signal from measured volumes at 0, 0.3 and 1 μL of Resovist SPIO tracer. We then used the calibration curve to estimate average cellular SPIO content in MPI images of 1 × 106 labeled cells. These MPI cellular iron estimates, at 27.0 ± 3.3 pg/cell, were within 99% agreement (P = 0.923) with standard but destructive Inductively Coupled Plasma (ICP) measurements at 26.8 ± 0.3 pg Fe/cell (Fig. 1e).
Sensitivity
We next investigated the sensitivity threshold of the prototype FFL MPI scanner for imaging labeled cells. We imaged 1 × 103 in vitro tagged cells in a pellet using a 20 second projection MPI scan and reconstructed the final MPI image using only the second and third harmonics of the drive field frequency. The reconstructed image showed a signal-noise ratio (SNR) greater than 5, corresponding to a 200-cell (5.4 ng Fe) detection limit (Fig. 1c). This detection limit corresponds to 130 nM sensitivity for a measured voxel volume of 750 μL.
Resolution
To assess the effect of SPIO cell labeling on MPI resolution, we acquired and compared a MPI line scan of two point sources: (a) 8 × 106 labeled hESC-derived cells and (b) 20 μL of 10× diluted Resovist tracer (Fig. 1d). The point sources were placed 3.5 cm apart on a 3D printed sample holder. The point sources were small (1.3 mm width) compared to the native resolution of the FFL system. The axial dimension of the point sources did not affect resolution analysis because the axis of each vial was placed parallel to the FFL during imaging. After a native MPI image was acquired, a line profile was generated across the center of the sample image to assess image blurring (Fig. 1d, bottom panel). A comparison of the MPI signal widths of the two samples showed that the full-width-at-half-maximum (FWHM) resolution of diluted Resovist particles in solution was 5 mm, approximately 1.5-fold better than that of the labeled cells at 7.7 mm. The measured native resolution for Resovist at 5 mm matches well with theory11 and approaches for improving the resolution are explored in Discussion.
Longitudinal Cell Tracking
Last, we tested MPI for tracking neural progenitor cells (NPCs) in vivo, which have previously shown promise as treatments for Parkinson’s Disease, Huntington’s disease, epilepsy and ischemia20. It is known that NPCs survive, migrate and show reconstructive properties when implanted in rodent models of forebrain ischemia21,22. The high specificity of MPI for SPIO imaging would facilitate quantification of cell delivery and retention, leading to better understanding of dynamic events such as graft movement in such brain disease models.
To evaluate MPI for long-term tracking of neural implants, we stereotactically implanted 5 × 105 SPIO-tagged hESC-derived NPCs in 5 μL PBS solution in the forebrain of two immunosuppressed rats, representing a standard therapeutic dose for neural cell implants21,22,23. To assess MPI monitoring of cell graft mobility and clearance in vivo, we also implanted an equivalent number of labeled NPCs near the lateral ventricle of a third animal to facilitate translocation and potential clearance of the graft. 0.15 μL Resovist tracer diluted in 4.85 μL PBS was administered to the forebrain of a fourth animal as a control. Due to institutional policy, we began longitudinal imaging of the animals post-recovery at day 10 for animals 1–3 and day 4 for animal 4. We then monitored the graft signals using MPI imaging for a period of 87 days post-administration. Serial MPI images of cell grafts show high signal contrast with no detectable signal from surrounding anatomy (Fig. 3a). Figure 3b shows total MPI iron signal from the cell graft in animals 1-2 had non-significant decay over time. In contrast, the cell graft in animal 3 exhibited the presence of iron caudal/posterior to the implant site and significant iron clearance compared to animals 1-2, suggesting movement and clearance of NPCs throughout the ventricle. The control animal receiving SPIO tracer alone showed no long-term detectable MPI signal.
MPI quantitatively tracks NPC neural implants in rats over 87 days.
(a) Longitudinal MPI imaging of 5 × 105 SPIO-labeled human NPCs implanted in the forebrain cortex (Animals 1-2), near lateral ventricle (Animal 3) and equivalent SPIO-only tracer in the forebrain cortex as control (Animal 4). MPI imaging quantifies graft clearance and movement over time, with rapid total clearance of SPIO-only injection (MPI: 30 sec acquisitions, 9.3 cm × 6 cm FOV, color intensity in ng/mm2, MRI reference: 10 min acquisitions, 7.5 cm × 4 cm, 384 × 384 matrix). All images are equally scaled; scale bar: 1 cm. (b) Total iron MPI estimates for in vivo cell grafts are plotted as a function of time with exponential fit. In vivo iron in Animal 1 and 2 do not show significant decrease over time, while iron signal was significantly decreased in Animal 3 starting Day 10. Animal 4 showed no long-term persistent MPI signal.
Histological and MRI Validation
We then compared MPI cell tracking data to postmortem Prussian blue (PB) staining and MRI (Fig. 4). Animals 1 and 2 (Fig. 4a) showed the presence of SPIOs only at the injection site. In Animal 3, PB staining and MRI also revealed the presence of SPIOs adjacent to the lateral ventricle (Fig. 4c), which confirmed the MPI observation that tagged cells had migrated posteriorly throughout the ventricle. Additional postmortem PB staining did not show any residual iron in the brain of the animal that received tracer alone, indicating rapid tracer clearance seen in MPI images. Immunohistochemical staining against neural progenitor markers nestin and human-specific neural cell adhesion molecule (NCAM) and human cytoplasmic marker SC121 confirmed the presence of human SPIO-labeled cells at the administration site (Fig. 4b). Expression of the macrophage/microglial marker CD68 showed some immune cell infiltration within the grafts. CD68 staining also indicated the presence of iron-containing immune cells along the lateral ventricle in Animal 3, indicating immunogenicity as a mechanism of graft clearance.
Histological and MRI validation of iron location and quantification.
(a) Histological analysis of NPC grafts. Postmortem Prussian blue (PB) staining confirms presence of iron-labeled cells at administration site but not for SPIO-only control (Animal 4). (b) Representative immunohistochemical slices (top right panels) are shown for NPC marker nestin (top left, Animal 1), neural cell adhesion molecule (top right, Animal 3) and human-specific cytoplasmic marker SC121 (bottom left, Animal 2), which indicate iron label within NPC grafts. CD68 staining (bottom right, Animal 3) also indicates immune cells at administration site, suggesting immune-based graft clearance. Scale bars: 100 μm. (c) Postmortem axial MRI indicates iron in lateral ventricle in Animal 3 (arrow). PB and CD68 staining of lateral ventricle in Animal 3 shows SPIO uptake by immune cells. MRI: 20 min acquisition, 2.56 cm isotropic FOV, 2563 pixel matrix. Scale bars: 1 mm.
Discussion
In this study, we describe the use of longitudinal in vivo MPI to track and quantify implanted neural cell grafts over three months. We also demonstrate that the MPI signal is linear and can be used to quantify cell number in vivo and that our projection format MPI scanner has a detection sensitivity of as few as 200 cells in vitro. As a comparison, MRI, SPECT, PET and fluorescent imaging are reported to have robust detection limits greater than 104 cells, with only bioluminescent imaging able to detect 103 cells superficially implanted in small animals3. We note that in the described prototype MPI scanners, noise in the detector coil preamplifier, at around 1 nV/√Hz, is not optimized and can be further improved by 1-2 orders of magnitude. This preamplifier noise exceeds unwanted harmonic interference and is currently the dominant noise source in our MPI systems. Thus, with further instrumentation development, the true physical limit of in vivo detection for MPI may range between 1–10 cells at scanning times between seconds to minutes.
An area of promising research in MPI is improving the spatial resolution, now roughly 1 mm with a 7 T/m magnetic gradient. Here, we employed 2.35 T/m gradients for a FFL scanner, which yielded approximately 7 mm spatial resolution when imaging cells labeled with Resovist tracer. Spatial resolution in MPI is predicted to increase linearly with the gradient field strength and cubically with improved nanoparticles11. For example, previously published 22 nm particles enable double the image resolution of Resovist19, which behaves similar to a solid core 17 nm particle18. Many groups continue to develop MPI-optimized contrast agents for improved MPI resolution24,25,26,27,28.
We routinely observed an intriguing change in MPI spatial resolution between Resovist tracer alone and Resovist-labeled cells, an effect also noted in previous studies on the MPI signal of SPIO-labeled red-blood cells29,30. Possible mechanisms for this resolution change could include size-selective endocytosis of transfection agent-Fe complexes5,31, increased MPI relaxation effects due to increased intracellular viscosity32,33 and transfection agent- mediated nanoparticle aggregation that may cause changes to the bulk particle magnetization and relaxation properties34,35,36. A better understanding of the mechanism underlying this phenomenon would not only enable tailored nanoparticles for MPI cell tracking, but could also potentially be exploited to generate image contrast based on nanoparticle relaxation or clustering. For example, changes in particle relaxation due to changes in pH or viscosity may be used to differentiate between different physiological conditions in live and dead cell populations, detect cell apoptosis, or image in vivo uptake of SPIOs by immune cells.
It is clear that the direct detection of SPIO labels in MPI can enable studies that require longitudinal monitoring of cells in anatomy previously inaccessible to optical, MRI and nuclear techniques, such as the GI tract, pulmonary vasculature and near bone. MPI’s high image specificity for SPIOs enables visualizing systemic cell retention, migration and biodistribution post-administration3. These systemic monitoring applications could include non-invasive probing of the immune system, such as the monitoring of labeled macrophages migrating to sites of inflammation3,37,38 and monitoring the fate of labeled T-cells used in immunotherapies. Finally, the SPIO labels and magnetic fields used in MPI have been shown to be human-safe17, enabling a clear path to clinical MPI translation.
Methods
Magnetic labeling of differentiated human embryonic stem cells (hESCs) and determination of labeling efficiency
H1 hESCs (WiCell, Madison, WI) were cultured on Matrigel-coated cell culture plates (BD, Franklin Lakes, NJ) in X-Vivo medium (Lonza, Norwalk, CT) supplemented with 80 ng/mL FGF- 2 (PeproTech, Rocky Hill, NJ) and 0.5 ng/mL TGF-β1 (R&D Systems, Minneapolis, MN), as described previously39.
For embryoid body (EB) mediated differentiation, H1 colonies were enzymatically isolated from Matrigel-coated plates by 5-minute treatment with 1 mg/mL collagenase type IV (Worthington Biochemical Corporation, Lakewood, NJ) and partially dissociated by gentle pipetting. The resulting cell clusters were resuspended in hESC culture medium, X-Vivo without FGF-2 and TGF-β1 and transferred to ultra-low-attachment plates (Corning Incorporated, NY) for cell aggregation for 4 days. The generated EBs were then seeded on Matrigel-coated plates and allowed to differentiate for an additional 10–12 days in adherent conditions in X-Vivo medium.
For cell labeling, Resovist superparamagnetic iron oxide particles (Bayer Schering Pharma AG, Leverkusen, Germany; 0.5 mmol Fe/mL) at a concentration of 200 μg Fe/mL were incubated with 6 μg/mL protamine sulfate (American Pharmaceuticals Partner, Schaumburg, IL) in serum free X-Vivo medium (Lonza, Walkersville, MD) and allowed to form complexes on a shaker for 6 hr. The solution containing Fe-Pro complexes was added to differentiating hESCs in adherent culture condition. The cultures were incubated with Fe-Pro complexes overnight. Cultures after labeling were washed with 10 U/mL heparin (Sigma) in PBS three times to dissolve remaining surface-bound Fe-Pro complexes.
For assessment of labeling efficiency, Prussian blue cellular staining was performed by incubating cells, fixed with 4% paraformaldehyde, in a mixture of 4% potassium ferrocyanide and 3.7% HCl (Iron Stain Kit, Sigma) for 20 minutes to visualize intracellular iron particles. Cells were counted manually under a microscope to determine average cell labeling efficiency. Labeling efficiency was found to be 71% +/− 8%. A hemocytometer was used to determine all cell numbers.
MPI image acquisition
Two custom-built MPI systems were used in this study: 1) a projection MPI scanner with 2.35 T/m/μ0 field gradient18 and 2) a 3D MPI scanner with 7 T/m/μ0. Both systems used a drive field frequency of 20.05 kHz and excitation strength of 40 mTpp/μ0, with 30 second total scan time for projection MPI scanner and 3 minute scan time for 3D MPI scanner unless otherwise specified. MPI images were reconstructed using an x-space MPI reconstruction17,38,12, followed by light Wiener deconvolution using a measured point-spread function. All MPI images were windowed at 10% full scale. All data acquisition, control and data processing were performed with MATLAB (Mathworks). All MPI imaging was conducted using the projection FFL scanner with the exception of the cell phantom imaging study shown in Fig. 2.
Determination of linearity of MPI signal with cell number
To investigate the linearity of the MPI signal with imaged cell number, we imaged 9 populations of Resovist-labeled hESC-derived cell pellets containing 10 × 103, 25 × 103, 50 × 103, 100 × 103, 200 × 103, 400 × 103, 600 × 103, 800 × 103 and 1,000 × 103 cells using a projection-format MPI scanner as described above. Following image reconstruction, the maximum signal intensity for each MPI image was determined and the Pearson correlation coefficient was determined between normalized MPI signal and imaged cell number. Subsequent linear regression on the data and 95% confidence intervals for the slope of the regression line were determined.
Intracellular Iron Measurement
Average intracellular iron was determined using induction-coupled plasma optical emission spectrometry (Optima 5300 DV, PerkinElmer, Waltham, MA) on a population of 1 × 106 SPIO-labeled hESC-derived cells. Prior to measurement, cells suspended in 0.2 mL PBS solution were digested using a 1.8 mL mixture of 70% nitric acid and 30% hydrogen peroxide and diluted to 60 mL using deionized water. ICP analysis was performed in triplicate to determine intracellular iron as mean ± SD in picograms of iron per cell. Intracellular iron was also measured using MPI. A calibration curve for volume MPI signal per picogram iron was first generated using 0, 0.3 and 1.0 μL Resovist tracer. Subsequently, we performed MPI imaging of 1 × 106 labeled cells (N = 4 each) and determined intracellular iron using the calibration curve as mean ± SD in picograms of iron per cell. To evaluate the congruence between MPI and ICP intracellular measurements, we performed an unpaired two-sample T-test for mean intracellular iron, with significance level set to 0.05.
In vivo stem cell implantation and MPI imaging
All animal procedures were conducted according to the National Research Council’s Guide for the Care and Use of Laboratory Animals and approved by UC Berkeley’s Animal Care and Use Committee. Immunocompetent 8-week old female Fischer 344 rats weighing 145 g were used for in vivo imaging. All animals were fed on an ad libitum diet of Teklad Rodent Diet 2018 (Harlan, Indianapolis, IL).
For in vivo MPI cell tracking experiments, hESCs were differentiated to neural progenitor cell by dual inhibition of SMAD signaling40 using SB431542 (10 μM) and LDN-193189 (1 μM) in adherent conditions for 12 days. The generated NPCs were labeled with Resovist SPIO particles as previously described here. The cell suspension contained approximately 100 × 103 viable cells/μL.
Animals were anesthetized with an intraperitoneal injection of ketamine (100 mg/kg rat weight) and xylazine (7.5 mg/kg), positioned into a stereotactic frame. For two animals, a total of 500 × 103 SPIO-labeled cells in 5 μL were injected at the following stereotaxic coordinates: 1.0 mm AP (relative to Bregma), 2.0 mm lateral, 2.5 mm ventral (from dura) over 10 min using a Hamilton syringe at a rate of 0.5 μL/min. For Animal 3, the coordinates were adjusted to 1.0 mm AP (relative to Bregma), 1.5 mm lateral and 3.5 mm ventral (from dura). A control animal was injected using 0.15 μL of Resovist tracer diluted in 4.85 μL PBS using the same stereotactic coordinates as Animals 1-2. The needle was left in place for 3 min after injection and slowly removed. Animals were allowed to recover from surgery for up to ten days before MPI imaging, with 7 daily injections of cyclosporine-A starting 1 day prior to implantation at a dose of 15 mg/kg body weight.
Animals were imaged using MPI at days 10, 24, 35, 45, 59 and 87 post-implantation. Animal 4 was imaged at days 4 and 14 instead of days 10 and 24 with all other time points remaining the same. During in vivo MPI imaging, animals were induced into anesthesia using 2.5–3.0% isoflurane and maintained at 1.5% isoflurane at 2 L/min and placed in a custom-built animal bed in the MPI scanner. In vivo MPI scans were conducted on the projection MPI scanner using a 9.3 cm by 6 cm FOV with 30 second scan time.
For determination of in vivo SPIO clearance rates measured by MPI, a linear regression was performed on the logarithm of the time-varying volumetric MPI signal for each animal.
Postmortem MRI
Ex vivo brains were imaged on a 7T Bruker PharmaScan MRI scanner using a ZTE pulse sequence. MR scans had total acquisition times of 20 minutes with 2563 voxels and 100 μm3 voxel size at 2.9 degree flip angles and 4 ms TR. Anatomical MRI images were acquired with postmortem age-matched animals using a TurboRare pulse sequence with 10 minute acquisition and 3842 voxels, 7.5 cm × 4 cm FOV and 2.5 sec TR.
Stereology and Immunohistological Analysis
The brains were fixed in 4% paraformaldehyde overnight, then cryopreserved in 20% sucrose, frozen in dry ice and cut in 40 μm coronal sections using a microtome. For immunohistochemistry, the following primary antibodies were used: mouse anti-nestin (1:500, R&D system), SC121 (1:400, Stem Cells Inc.), mouse anti-CD68 (1:100, Abcam). Sections were then incubated with fluorescent-labeled secondary antibodies Alexa 488 (green) or Alexa 568 (red)-labeled goat IgG; 1:1000, (Life Technologies) for 2 hr at room temperature and then mounted on glass slides. Brain slides were counter-stained with DAPI and imaged using a Zeiss Axio Observer A1 inverted microscope or a confocal microscope (LSM 710, Zeiss).
Additional Information
How to cite this article: Zheng, B. et al. Magnetic Particle Imaging tracks the long-term fate of in vivo neural cell implants with high image contrast. Sci. Rep. 5, 14055; doi: 10.1038/srep14055 (2015).
References
Bjorklund, L. M. et al. Embryonic stem cells develop into functional dopaminergic neurons after transplantation in a Parkinson rat model. Proc. Natl. Acad. Sci. USA. (2002), 10.1073/pnas.022438099.
De Vries, I. J. M. et al. Magnetic resonance tracking of dendritic cells in melanoma patients for monitoring of cellular therapy. Nat. Biotechnol. 23, 1407–1413 (2005).
Nguyen, P. K., Riegler, J. & Wu, J. C. Stem cell imaging: from bench to bedside. Cell Stem Cell 14, 431–44 (2014).
Bulte, J. W. M. In vivo MRI cell tracking: clinical studies. AJR. Am. J. Roentgenol. 193, 314–25 (2009).
Arbab, A. S. et al. Efficient magnetic cell labeling with protamine sulfate complexed to ferumoxides for cellular MRI. Blood 104, 1217–1223 (2004).
Au, K.-W. et al. Effects of iron oxide nanoparticles on cardiac differentiation of embryonic stem cells. Biochem. Biophys. Res. Commun. 379, 898–903 (2009).
Crabbe, A. et al. Effects of MRI contrast agents on the stem cell phenotype. Cell Transplant. 19, 919–36 (2010).
Cunningham, C. H. et al. Positive contrast magnetic resonance imaging of cells labeled with magnetic nanoparticles. Magn. Reson. Med. 53, 999–1005 (2005).
Ahrens, E. T. & Bulte, J. W. M. Tracking immune cells in vivo using magnetic resonance imaging. Nat. Rev. Immunol. 13, 755–63 (2013).
Gleich, B. & Weizenecker, J. Tomographic imaging using the nonlinear response of magnetic particles. Nature 435, 1214–7 (2005).
Goodwill, P. W. & Conolly, S. M. The x-space formulation of the magnetic particle imaging process: three-dimensional signal, gridding, pulse sequences and reconstruction. IEEE Trans. Med. Imaging 64, 267–273 (2010).
Goodwill, P. W. & Conolly, S. M. Multidimensional X-space magnetic particle imaging. IEEE Trans. Med. Imaging 30, 1581–1590 (2011).
Lu, K., Goodwill, P. W., Saritas, E. U., Zheng, B. & Conolly, S. M. Linearity and shift invariance for quantitative magnetic particle imaging. IEEE Trans. Med. Imaging 32, 1565–1575 (2013).
Baker-Jarvis, J. & Kim, S. The Interaction of Radio-Frequency Fields with Dielectric Materials at Macroscopic to Mesoscopic Scales. Journal of Research of the National Institute of Standards and Technology 117, 1 (2012).
Bulte, J. W. M. et al. Developing Cellular Mpi: Initial Experience. Magn. Nanoparticles - Part. Sci. Imaging Technol. Clin. Appl. - Proc. First Int. Work. Magn. Part. Imaging 201–204 (2010), 10.1142/9789814324687_0028.
Konkle, J. J., Goodwill, P. W., Carrasco-Zevallos, O. M. & Conolly, S. M. Projection reconstruction magnetic particle imaging. IEEE Trans. Med. Imaging 32, 338–47 (2013).
Saritas, E. U. et al. Magnetic particle imaging (MPI) for NMR and MRI researchers. J. Magn. Reson. 229, 116–26 (2013).
Goodwill, P. W., Konkle, J. J., Zheng, B., Saritas, E. U. & Conolly, S. M. Projection x-space magnetic particle imaging. IEEE Trans. Med. Imaging 31, 1076–85 (2012).
Ferguson, R. et al. Magnetic Particle Imaging with Tailored Iron Oxide Nanoparticle Tracers. IEEE Trans. Med. Imaging 0062, 1–1 (2014).
De Feo, D., Merlini, A., Laterza, C. & Martino, G. Neural stem cell transplantation in central nervous system disorders: from cell replacement to neuroprotection. Curr. Opin. Neurol. 25, 322–33 (2012).
Chu, K., Kim, M., Jeong, S., Kim, S. U. & Yoon, B. Human neural stem cells can migrate, differentiate and integrate after intravenous transplantation in adult rats with transient forebrain ischemia. Neurosci. Lett. 343, 129–33 (2003).
Kelly, S. et al. Transplanted human fetal neural stem cells survive, migrate and differentiate in ischemic rat cerebral cortex. Proc. Natl. Acad. Sci. USA. 101, 11839–11844 (2004).
Guo, J. et al. In vivo evaluation of cerebral transplantation of resovist-labeled bone marrow stromal cells in Parkinson’s disease rats using magnetic resonance imaging. Appl. Biochem. Biotechnol. 163, 636–48 (2011).
Ferguson, R. M., Minard, K. R., Khandhar, A. P. & Krishnan, K. M. Optimizing magnetite nanoparticles for mass sensitivity in magnetic particle imaging. Med. Phys. 38, 1619 (2011).
Roca, A. G. et al. Progress in the preparation of magnetic nanoparticles for applications in biomedicine. J. Phys. D. Appl. Phys. 42, 224002 (2009).
Lak, A., Wawrzik, T., Ludwig, F. & Schilling, M. Synthesis of Single-Core Iron Oxide Nanoparticles as a Tracer for Magnetic Particle Imaging. Springer Proc. Phys. 140, 93–97 (2012).
Eberbeck, D., Trahms, L. & Kratz, H. Evaluation of Different Magnetic Particle Systems with Respect to Its MPI. Springer Proc. Phys. 140, 53–57 (2012).
Eberbeck, D., Wiekhorst, F., Wagner, S. & Trahms, L. How the size distribution of magnetic nanoparticles determines their magnetic particle imaging performance. Appl. Phys. Lett. 98, 182502 (2011).
Markov, D. E. et al. Human erythrocytes as nanoparticle carriers for magnetic particle imaging. Phys. Med. Biol. 55, 6461–73 (2010).
Rahmer, J. et al. Nanoparticle encapsulation in red blood cells enables blood-pool magnetic particle imaging hours after injection. Phys. Med. Biol. 58, 3965–77 (2013).
Sun, R. et al. Physical and biological characterization of superparamagnetic iron oxide- and ultrasmall superparamagnetic iron oxide-labeled cells: a comparison. Invest. Radiol. 40, 504–513 (2005).
Croft, L. R., Goodwill, P. W. & Conolly, S. M. Relaxation in x-space magnetic particle imaging. IEEE Trans. Med. Imaging 31, 2335–42 (2012).
Rauwerdink, A. M. & Weaver, J. B. Viscous effects on nanoparticle magnetization harmonics. J. Magn. Magn. Mater. 322, 609–613 (2010).
Van Buul, G. M. et al. Ferumoxides-protamine sulfate is more effective than ferucarbotran for cell labeling: Implications for clinically applicable cell tracking using MRI. Contrast Media Mol. Imaging 4, 230–236 (2009).
Kim, H. S. et al. The effects of clinically used MRI contrast agents on the biological properties of human mesenchymal stem cells. NMR Biomed. 23, 514–522 (2010).
Panizzo, R. A. et al. In vivo magnetic resonance imaging of endogenous neuroblasts labelled with a ferumoxide-polycation complex. Neuroimage 44, 1239–46 (2009).
Kircher, M. F., Gambhir, S. S. & Grimm, J. Noninvasive cell-tracking methods. Nat. Rev. Clin. Oncol. 8, 677–88 (2011).
Rogers, W. J., Meyer, C. H. & Kramer, C. M. Technology insight: in vivo cell tracking by use of MRI. Nat. Clin. Pract. Cardiovasc. Med. 3, 554–62 (2006).
Meng, Y. et al. Characterization of integrin engagement during defined human embryonic stem cell culture. FASEB J. 24, 1056–65 (2010).
Chambers, S. M. et al. Highly efficient neural conversion of human ES and iPS cells by dual inhibition of SMAD signaling. Nat. Biotechnol. 27, 275–80 (2009).
Acknowledgements
The authors would like to thank Prof. Jeff Bulte for valuable discussions and insight, Kuan Lu and Dr. Justin Konkle for MPI scanner assistance, Di Xiao and Neerav Dixit for animal imaging assistance, Paul Keselman and Dr. Mike Wendland for MRI and CT assistance and Analisa Nazari for assistance in histological analysis. We gratefully acknowledge funding from Siebel Scholars Foundation, NSF GRFP under grant number DGE 1106400, NIH R01 EB013689, CIRM RT2-01893, Keck Foundation 009323, NIH 1R24 MH106053 and NIH 1R01 EB019458 and ACTG 037829.
Author information
Authors and Affiliations
Contributions
B.Z., T.V., P.G., E.S., D.S. and S.C. contributed to critical discussions toward experimental designs. B.Z., P.G. and E.S. contributed to scanner operation and imaging studies. T.V. contributed to cell labeling. T.V. and A.C. contributed to cell implantation and animal handling. T.V. and A.V. contributed to histology studies. B.Z., T.V. and P.G. contributed to writing the paper. All authors reviewed the manuscript.
Ethics declarations
Competing interests
Drs. Conolly and Goodwill are stockholders in Magnetic Insight, Inc. Dr. Conolly is also a board member of the company; Dr. Goodwill is an employee of the company.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Zheng, B., Vazin, T., Goodwill, P. et al. Magnetic Particle Imaging tracks the long-term fate of in vivo neural cell implants with high image contrast. Sci Rep 5, 14055 (2015). https://doi.org/10.1038/srep14055
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep14055
This article is cited by
-
Counting cells in motion by quantitative real-time magnetic particle imaging
Scientific Reports (2024)
-
Omniparticle Contrast Agent for Multimodal Imaging: Synthesis and Characterization in an Animal Model
Molecular Imaging and Biology (2023)
-
In silico assessment of electrophysiological neuronal recordings mediated by magnetoelectric nanoparticles
Scientific Reports (2022)
-
Molecular imaging of cellular immunotherapies in experimental and therapeutic settings
Cancer Immunology, Immunotherapy (2022)
-
Tracking Neural Stem Cells in vivo: Achievements and Limitations
Stem Cell Reviews and Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.