Abstract
Plasmodiophora brassicae causes clubroot, a major disease of Brassica oil and vegetable crops worldwide. P. brassicae is a Plasmodiophorid, obligate biotrophic protist in the eukaryotic kingdom of Rhizaria. Here we present the 25.5 Mb genome draft of P. brassicae, developmental stage-specific transcriptomes and a transcriptome of Spongospora subterranea, the Plasmodiophorid causing powdery scab on potato. Like other biotrophic pathogens both Plasmodiophorids are reduced in metabolic pathways. Phytohormones contribute to the gall phenotypes of infected roots. We report a protein (PbGH3) that can modify auxin and jasmonic acid. Plasmodiophorids contain chitin in cell walls of the resilient resting spores. If recognized, chitin can trigger defense responses in plants. Interestingly, chitin-related enzymes of Plasmodiophorids built specific families and the carbohydrate/chitin binding (CBM18) domain is enriched in the Plasmodiophorid secretome. Plasmodiophorids chitin synthases belong to two families, which were present before the split of the eukaryotic Stramenopiles/Alveolates/Rhizaria/Plantae and Metazoa/Fungi/Amoebozoa megagroups, suggesting chitin synthesis to be an ancient feature of eukaryotes. This exemplifies the importance of genomic data from unexplored eukaryotic groups, such as the Plasmodiophorids, to decipher evolutionary relationships and gene diversification of early eukaryotes.
Similar content being viewed by others
Introduction
Clubroot caused by Plasmodiophora brassicae, is a spreading soil-borne disease with high economical impact on Brassica oil- and vegetable crops and production of other valuable species within the family Brassicaceae worldwide1. P. brassicae is distinct from other plant pathogens, such as fungi or oomycetes, as it is an obligate biotroph protist in the Plasmodiophorids within the eukaryote supergroup Rhizaria. Rhizaria share a common ancestor with Stramenopiles and Alveolata forming the S-A-R supergroup, which together with Plantae build the SARP megagroup2,3. Plasmodiophorids are parasites of plants and oomycetes and a sister group to Phagomyxids, pathogens of sea grass, diatoms and brown algae4. Other agriculturally important Plasmodiophorid pathogens are Spongospora subterranea, the causal agent of potato powdery scab and root galls (Supplementary Fig. S1) and vector for the Potato Mop Top Virus and the virus transmitting Polymyxa betae and P. graminis infecting sugar beets and cereals.
Plasmodiophorids have a complex, not completely understood, life cycle5 consisting of different zoosporic stages, formation of plasmodia inside host cells and resting spore formation (Fig. 1). The haploid resting spore releases a zoospore, which infect the root hairs and form multinucleate plasmodia. Those develop several single-nucleate secondary zoospores, which are released into the soil. A fusion of those secondary zoospores is assumed to occur occasionally. Zoospores invade the root cortex, in which secondary multinucleate plasmodia develop. How and when karyogamy is taking place is not exactly known, but meiosis appears to occur in the plasmodia before resting spore formation.
The life cycle of Plasmodiophora brassicae.
A biflagellate primary zoospore (PZ) is released from each resting spore (RS). When the PZ reaches the surface of a root hair (RH) (a), it penetrates the cell wall and forms primary plasmodia in the root hairs. After a number of nuclear divisions the plasmodia cleaves into zoosporangia (ZS), developing and releasing secondary zoospores (SZ) (b). Occasional fusion of SZ has been suggested. PZ or SZ can penetrate root cortex cells, in which the pathogen develops into secondary plasmodia (SP) (c–f). Proliferation of SP is associated with cellular hypertrophy, causing a swollen root or club phenotype (Supplementary Fig. S1). After a number of nuclear divisions, the secondary plasmodia develop into multinuclear plasmodia and finally into resting spores (f–i), which are released into the surrounding soil when plants decay. RS = Resting spore, PZ = Primary zoospore, RH = root hair, ZS = Zoosporangia, SZ = Secondary zoospore, SP = Secondary plasmodia. PC = Plant cell, PCW = Plant cell wall
Plasmodia provoke abnormal cell enlargement (hypertrophy) and uncontrolled cell division (hyperplasia) leading to the development of galls (Supplementary Fig. S1) that obstruct nutrient and water transport. In infected root tissue, different developmental stages of the Plasmodiophorids occur simultaneously. For example, host cells containing plasmodia neighbor cells filled with resting spores. The only host-free stages are the short-lived bi-flagellated zoospores and resting spores. Therefore molecular and genomic data for Plasmodiophorids are rare as they only grow inside living host cells and remain uncultivable on their own.
Furthermore, the Rhizaria is the least studied group of eukaryotes6,7 with only the genomes of the chlorarachniophyte Bigelowiella natans8 and the foraminifer Reticulomyxa filosa9 available until now. To fill this gap, we sequenced, assembled and analyzed the genome and transcriptomes of P. brassicae, to create the first genome draft of a pathogenic Rhizaria. The single-spore isolate e310, for which no sexual recombination has been observed, was used to avoid complication due to potential polyploidy and chromosome polymorphism. We emphasized on generating transcriptomes of life-stage specific materials from P. brassicae which enabled analyses of developmental stage specific gene expression. The genome information of P. brassicae was further used to identify and analyze transcript data from the related potato pathogen S. subterranea.
Resting spores of P. brassicae are extremely resilient to harsh environmental conditions and contaminate arable land for decades. This feature makes it impossible to eradicate the organism via any known chemical or alternative soil treatment. The resting spores which are formed at the end of the life cycle inside the root contain chitin in their cell walls11. Besides being an important component of the Plasmodiophorid survival structure, chitin and other glycans are known as important signatures for plant immune activation or for the establishment of beneficial symbioses12. We analyzed the chitin-related proteins detected in the Plasmodiophorids in more detail and also the secretome for effector candidates that could interfere with microbe-triggered host responses.
Results and Discussion
Genome sequencing and Rhizaria genome comparisons
We sequenced the P. brassicae genome from genomic DNA of resting spores from the single spore isolate e3 (Supplementary Table S1). The 24 Mb genome was de novo assembled into 165 scaffolds with an N50 size of 473 kb and an average 500-fold coverage (Supplementary Fig. S2; Supplementary Table S2). The details of the sequencing analysis are provided in the supplementary material (Supplementary Note). The GC-content (58.5%) is higher than for B. natans (46%) and R. filosa (35%). The genome is small compared to the free-living Rhizaria B. natans (~100 Mb) and R. filosa (~320 Mb) but larger than earlier suggested (20.3 Mb)13. A total of 9,730 genes were predicted with support from seven transcriptome libraries (Supplementary Table S3) representing 66% of the genome with an average of 404 genes per Mb. P. brassicae genes contain an average of 4 introns per gene of 60bp length. With only 5,4% the detected content of repetitive and transposable elements, including new identified and previously described repeats (AF296444, AF296445, AF537105)10, is low (Supplementary Fig. S3; Supplementary Table S4). The low number of repeat is concurrent with a total genome size estimation of 25.5 Mb based on 17-mer analysis. The gene density of P. brassicae (Supplementary Figs. S3, S4), lower number of genes (Supplementary Fig. S5) and low content of repetitive elements, contribute to the smaller genome size compared to the free-living B. natans and R. filosa. Similar genome reductions are observed for other eukaryotic parasites compared to free-living species14.
CEGMA evaluation15 revealed that 92% of the eukaryotic core genes were of full length (94% partially), which is slightly higher compared to the other two Rhizarian genomes (Supplementary Table S5). The S. subterranea transcriptome was assembled into a total of 12,732 protein models of which 7,490 were full-length, covering 69% of CEGMA genes. The higher number compared to P. brassicae is likely due to partial assembled genes or might reflect current or recent transposon activity as suggested earlier16.
OrthoMCL17 analysis defined a Rhizaria core set of 1,863 protein families common to P. brassicae, R. filosa and B. natans of which 1,338 families were shared with S. subterranea (Fig. 2). The specific set of P. brassicae contained 2,161 protein families out of 5,605 proteins with no orthologues in R. filosa and B. natans. KOG-term analysis showed that high proportions of the proteins in the Plasmodiophorids are poorly characterized and the majority of the P. brassicae specific genes have no assigned function (Fig. 2; Supplementary Fig. S5). The P. brassicae-specific genes were diversely expressed in the different developmental stages, whereas the Rhizarian core genes were most active in the plasmodial stage, when P. brassicae was already established inside the plant root (Supplementary Fig. S6).
Protein families in Rhizaria and P. brassicae.
(a) Venn diagramm of OrthoMCL protein families predicted for P. brassicae, S. subterranea and the the non-pathogenic Rhizarian B. natans and R. filosa. A Rhizarian core set was defined by common familes of B. natans, R. filosa and P. brassicae (groups E + F). Groups A-D build P. brassicae specific proteins. The S. subterranea predicted models were not regarded for the definition of the core set and P. brassicae specific set as no complete genome information is available. Numbers in the table show the number of OrthoMCL families and in brackets the number of P. brassicae proteins in each family. (b) Functional annotation of the P. brassicae proteins according to KOG functional categories. Rhizaria core set and P. brassicae-specific genes are defined as in (a).
Compared to R. filosa and B. natans, fungi and plant pathogenic oomycetes, the two Plasmodiophorids were enriched for the carbohydrate esterase family 4 (CE4, polysaccharide deacetylases), galactose-binding lectins, ankyrin domains, the carbohydrate/chitin binding domain CBM18 and RAP RNA-binding domain proteins (Supplementary Fig. S7; Supplementary Table S6). The Gal-lectin domain is proposed to be of animal origin and acquired by the fish pathogenic oomycete Saprolegnia parasitica via horizontal gene transfer18. However, our analyses of Gal-lectin domains including SARP and Amoebozoa data suggesting a more ancient origin of these proteins (Supplementary Fig. S8). Unlike P. brassicae, the S. subterranea transcriptome was also enriched in transposon and virus-related genes (Supplementary Table S6), which has been observed earlier16.
It is not known if RNA-directed gene silencing occurs in Plasmodiophorids. P. brassicae and S. subterranea contain argonautes but only one gene each encoding for a protein with two RNaseIII domains and a dsRNA-binding motif (Supplementary Fig. S9). Those proteins resemble the Dicer-like protein 1 from the ciliate protozoan Tetrahymena thermophile and Drosha proteins of Drosophila melanogaster involved in miRNA processing. Any predicted RNA-dependent RNA polymerase and RNA polymerase IV were not detected in P. brassicae.
Metabolism shows biotrophic signatures
Similar to other eukaryotic biotrophic plant pathogens19,20,21 P. brassicae and S. subterranea are reduced in several metabolic pathways. Both Plasmodiphorids lack genes encoding for proteins involved in sulfur and nitrogen uptake as well as genes for histidine, tryptophan and threonine biosynthesis. Arginine and lysine synthesis were also compromised in P. brassicae (Supplementary Figs. S10, S11). Thiamine biosynthesis genes were also reduced in the Plasmodiophorids (Supplementary Fig. S12), suggesting a dependence of their host plants for those metabolites. No fatty acid synthase was identified in P. brassicae (Supplementary Fig. S13). The inability for fatty acid synthesis has been described for obligate biotrophic bacterial pathogens of insects and plants22 and kinetoplast parasites such as Trypanosoma species. The P. brassicae gene content for fatty acid biosynthesis was similar as for the kinetoplasts, which use an alternative parasitic-specific fatty acid synthesis, the microsomal elongase pathway23. Our data suggest that P. brassicae metabolize fatty acids from the host by elongating and modifying molecules of four or more carbons. Protein and transcript data showed that fatty acids were synthesized in the plasmodia and degradation occurred in the spores (Supplementary Figs. S14, S15). Seven putatively secreted lipases in the P. brassicae genome were among the highest expressed genes in the plasmodial stage providing the pathogen with fatty acid starter molecules. However, in the potato scab transcriptome, we detected two potential fatty acid synthase genes and no secreted lipases, suggesting that S. subterranea synthesizes fatty acids de novo.
We predicted carbohydrate conversion to mainly occur via glycolysis, glyoxylate, pyruvate metabolism, pentose phosphate pathway and the TCA cycle (Supplementary Figs. S14–16). Proteins and transcripts of the glyoxylate cycle were highly abundant in the germinating spore sample indicating a high conversion of sugars and lipids in this active life-stage. The P. brassicae genome contained trehalose biosynthesis genes expressed in all transcriptome sets (Supplementary Fig. S17), which corroborates to earlier findings of trehalose accumulation by P. brassicae in infected plants24. It has been speculated that trehalose production by plant pathogens interferes with the sugar sensing pathways of their host and thereby manipulate plant metabolism to their favor or regulate energy and nitrogen supply during colonization25. Besides affecting the host, trehalose in the resting spores could be important for their long-term survival as an energy resource or an osmoprotectant against the effects of desiccation. A trehalase-encoding gene of P. brassicae was primarily expressed in spores indicating a release of glucose as an energy resource in the germination process.
Plant hormone homeostasis
Plant tissue infected with P. brassicae shows a modified hormone balance26,27 that induces hypertrophy and cell divisions in host roots leading to gall formation. P. brassicae can potentially modify the plant hormone homeostasis of at least four plant hormones: cytokinins26 and salicylic acid (SA)28 and, as shown here, auxins and jasmonic acid (JA). The P. brassicae genome, but not the S. subterranea transcripts, contained an auxin-responsive Gretchen Hagen 3 (GH3) protein (PbGH3). GH3-proteins are known to maintain hormonal homeostasis by conjugating plant hormones to amino acids29. The PbGH3 predicted protein structure resembles plant GH3-proteins (Fig. 3a). The heterologous expressed PbGH3 protein conjugated auxins and JA with proteinogenic amino acids in vitro, although it is phylogenetically not related to plant GH3-like proteins (Fig. 3b, Supplementary Fig. S18). Both Plasmodiophorids contained two isopentenyl-transferases (IPTs) belonging to DMAPP:tRNA-IPTs classes I and II (Supplementary Fig. S19). Class I DMAPP:tRNA-IPTs include members from Physcomitrella patens and Arabidopsis thaliana that are involved in cis-zeatin synthesis30. Cytokinin homestasis can also be inferred by a P. brassicae cytokinin oxidase, although the expression in our transcript sets was low (Supplementary Fig. S19). P. brassicae produces trans-zeatin26 and the contribution of the Plasmodiophorid IPTs and the potential cytokinin oxidase to the host cytokinin homeostasis remains to be shown. The gall phenotype might be a result of pathogen-driven interference of the plant root phytohormone system combined with reprogramming of existing meristematic activity31. The recently described secreted methyl transferase (PbBSMT) can reduce the accumulation of SA in infected roots28. PbBSMT expression in spores and plasmodia was low, but high expression levels in clubroots of Brassica napus, B. oleracea and B. rapa host plants highlights its important role (Supplementary Fig. S20). A putative homolog was also found in S. subterranea indicating a conserved Plasmodiophorid strategy to interfere with plant host SA-defense responses.
Structure prediction and activity of the PbGH3 protein.
(a) Modeling of PbGH3 on plant GH3-proteins for which crystal structures are available. An overlay of the obtained protein model for PbGH3 (gray) with a monomer of the A. thaliana AtGH3.12 (PDB database entry 4EPM (http://www.rcsb.org/pdb/home/home.do) protein (magenta) is shown. The model was built in SWISS MODEL65 and the overlaid with the standard settings of Chimera66. (b) Conjugation activity of PbGH3 with IAA, SA and JA and amino acids (AA). Activity is indicated by black boxes (high activity) to white boxes (no activity).
The Plasmodiophorid secretome
Microbes secrete effector proteins to manipulate host processes in order to facilitate colonization. We predicted 553 secreted proteins in P. brassicae and 613 in S. subterranea. Both secretomes were smaller than that of B. natans but similar in numbers to other biotrophic fungi and oomycete plant pathogens (Supplementary Fig. S21). An OrthoMCL comparison of the two secretomes revealed that most secreted proteins were not shared between the two species (Supplementary Fig. S21) but were enriched in ankyrin and protein domains detected in effectors of other plant pathogens32, such as small cysteine-rich secreted proteins, chitin recognition proteins, proteases and protease inhibitors (Supplementary Table S7). The secretome of P. brassicae was further enriched in leucine–rich repeat domains and pathogenesis-related (PR) proteins, such as PR-5 thaumatin like-proteins. A cysteine-rich secretory protein with BLASTP hits to PR-1-like proteins with no homologues in the non-plant pathogenic Rhizarians, was also present in S. subterranea and identified in the Plasmodiophorid P. betae (Pbef2)33.
Oomycete effectors frequently code for host-targeting sequence motifs, such as RXLR, LXLFLAK and CHXC34. The secreted proteins of P. brassicea and S. subterranea do not contain such overrepresented amino acid motifs (Supplementary Table S8). Instead, we categorized the P. brassicae secretome into three groups based on their gene expression to identify potential effectors: 1) the PLeff genes - highest expressed in plasmodia, 2) the Heff genes – highest expressed in the transcriptomes obtained from the hosts and 3) the Leff genes – highest expressed late in the life cycle during spore formation (Supplementary Fig. S21; Supplementary Table S9). Each group contains proteins that could reprogram the plant metabolism and interfere with the plant defense on a protein level. Highly expressed PLeff genes contained ankyrin domains, which are frequent in all Rhizarian8,9, but are effector candidates for bacterial pathogens and symbionts35,36. PLeff genes also included a secreted lipase, leucine-rich repeats proteins, PBRA_T005609, that shows similarities to effector candidates of biotrophic smut fungi in the Ustilagomycetes37 and RING finger domain containing proteins. Secreted RING finger domain proteins interfere with the host ubiquitation system of the human intracellular pathogen T. cruzi38, a strategy also seen for plant pathogens32. PbBSMT is one of the highest expressed genes in the Heff group, likely reducing the SA content in infected host tissue. The Leff candidates include CE4-domain polysaccharide acetylases, proteases and protease inhibitors including Kazal-like and papain protease inhibitors and fascilin domain proteins.
Carbohydrate-Active enZymes (CAZymes)
Like other biotrophic plant pathogens19,20,21,39 the Plasmodiophorids have a reduced set of CAZymes (Supplementary Fig. S22). The total number of CAZymes was similar to those identified in the genomes of Hyaloperonospora arabidopsidis, Ustilago maydis or the arbuscular mycorhizal fungus Rhizophagus irregularis. However, the arsenal for both potential plant cell wall (PCW) modifying and chitin cell wall modifying enzymes, differed substantially from fungal and oomycete species (Fig. 4). In the proteins models of P. brassicae and S. subterranea only a few domains were identified that could degrade hemicellulose. Predicted CAZymes with domains involved in cellulose, lignin or pectin degradation or their modifications were also rare, compared to selected fungi, oomycetes, B. natans and R. filosa (Fig. 4a). Only one protein with a cellulose-binding motif (CBM1) was found. The Plasmodiophorids contain several members of the cellulase containing GH5-hydrolase family. However, the GH5 hydrolases of the Plasmodiophorids that we could classify based on phylogeny belonged to the ceramidase GH5_12 and GH5_27 subfamilies40 (Supplementary Fig. S23).
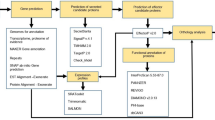
Distribution of (a) plant cell wall degrading enzymes and (b) chitin related enzymes in Rhizarian and selected fungal and oomycete genomes. The heatmap represents the abundance of CAZyme domains in each genome. Clustering of gene families is based on Pearson correlation of gene numbers in each family in each species. *Indicates CAZymes with a possible effect on polysaccharides contained in both chitinous and/or plant cell walls. Species abbreviations: HARA = Hyaloperonospora arabidopsidis, PINF = Phytophthora infestans, PSOJ = Phytophthora sojae, PULT = Pythium ultimatum. BNAT = Bigellowiella natans, PBRA = Plasmodiophora brassicae, SSUB = Spongospora subterranea, RFIL = Reticulomyxae filosa, ANIG = Aspergillus nigra, NCRA = Neurospora crassa, VDAL = Verticillium dahliae, LBIC = Laccaria bicolor, MLAR = Melampsora larici-populina, PGRA = Puccinia graminis f. sp. tritici, UMAY = Ustilago maydis, RIRR = Rhizophagus irregularis.
The most prominent chitin-related CAZy domains in the protein models belonged to the GH18 chitinases, CE4 and CBM18 (Fig. 4b). P. brassicae contained 16 GH18 domain-containing chitinases and their evolutionary history shows whole gene duplications and internal GH18 domain duplication events (Supplementary Fig. S24). GH18 domains can be separated into three phylogenetic clusters A, B and C41. The Plasmodiophorid GH18 chitinase domains grouped to cluster A and the bacterial cluster C previously not reported to contain eukaryotic members. In the cluster A, the Plasmodiophorid GH18 s formed a specific subgroup (Pd1). These Pd1-chitinase genes were mainly expressed during spore formation and germination suggesting a specific role for those proteins in resting spore cell wall modification and degradation. The phylogeny also revealed a new oomycete-specific cluster of GH18 domains.
Recognition of microbial cell walls including chitin and chitin degradation products can be recognized by plants and trigger defense responses12. In a microarray assay27 the plant receptor genes for this type of recognition were suppressed after P. brassicae infection (Supplementary Table S10). The CBM18 domain and the LysM harboring CBM50 domain are known to bind chitin. Plants contain LysM domains in receptors involved in microbe recognition. Fungi can escape host recognition via LysM effectors, proteins that consist of a signal peptide and multiple LysMs, but no catalytic domain12. In the datasets of the two Plasmodiophorids only P. brassicae contained one protein with a single CBM50 domain. However, the CBM18 domain, which is reduced in biotrophic fungal lineages39,42 and absent in B. natans and R. filosa and plant pathogenic oomycetes, is frequent in both Plasmodiophorids (Fig. 5b). Similarly CBM18-domain proteins are enriched in the aquatic fungus Batrachochytrium dendrobatadis and protect the fungus from breakdown by chitinases43. However, if the CBM18 proteins have similar functions in the Plasmodiophorids remains to be shown.
Interrelationships of eukaryotic chitin synthases (CHS) and model for CHS evolution in eukaryotes.
(a) Unrooted Bayesian phylogenetic reconstruction of chitin synthase CHS domains (Chitin_synth_2; PF03142). Node support as represent posterior probabilities. Black circles indicate nodes supported of parallel ML analysis (RAxML, LG model, 500 replicates) with a bootstrap >70. The detailed tree is shown in Supplementary Fig. S25 and proteins used are listed in Supplementary Dataset 1. (b) A proposed model for the evolution of CHS in the eukaryotic kingdom based on current available data.
CE4-domain proteins can convert chitin into its derivate chitosan and were frequent in the Plasmodiophorids but absent in the other available Rhizaria genomes. Some CE4-domain proteins also included the chitin binding CBM18 domain, suggesting that these proteins bind to the Plasmodiophorid chitin in order to promote modification into chitosan, a weaker inducer of immune responses than chitin in many plants12.
Plasmodiophorids contain two ancient chitin synthase (CHS) families
CHS are essential for the chitin synthesis. In P. brassicae 13 CHS-encoding genes and six in S. subterranea were identified. Whereas B. natans and R. filosa contained no predicted CHS, transcript data from Leptophrys, Sorites and Euglypha species include CHS domains, suggesting that chitin is also synthesized by Rhizarian outside the Plasmodiophorids. CHS are divided into two paralogous families, with family 1 represented by fungal CHS classes IV-VII and family 2 by fungal CHS classes I–III44. We extended previous phylogenetic analyses of CHS by including CHS domains from Rhizarian, Stramenopiles, Alveolata and from green algae (Chlorella variabilis, Picochlorum species) (Supplementary Fig. S25). Plasmodiophorid harbor both CHS families. No Plasmodiophorid CHS acquired additional protein domains, unlike Metazoa and fungi CHS where fusion with myosin domains have occurred45. The protein domain structure and phylogeny of the Plasmodiophorid family 1 CHS are related to fungal class VI CHS. A family previously only detected in fungal species (Fig. 5a; Supplementary Figs. S25, S26). In contrast to recent analyses45, animal CHS regrouped with fungal CHS classes IV–VII in family 1, together with SARP CHS of Plasmodiophorids and diatoms. The CHS family 2 comprised fungal classes I-III and CHS of oomycetes, plants, Aveolata and Rhizaria, including Plasmodiophorid in the SARP.
It was proposed that a branch of β-glycosyl-transferases developed into both CHS families after the plant kingdom split from the evolutionary line of Metazoa and fungi44. Our results revealed a different evolutionary scenario where the two lineages had already diverged in the last common ancestor of the SARP and Amorphea. This makes the absence of family 2 in Metazoa and family 1 in Plants and Alveolata all secondary losses (Fig. 5b). Family 2 was lost in the Metazoa concurrent with the divergence from Fungi in the Amorphea. In the SARP, Family 1 was lost in the Alveolata when they diverged from Rhizaria and Stramenopiles. Subsequently in the Stramenopiles, the loss of family 2 in diatoms and family 1 in oomycetes occurred when those groups separated. In the Rhizaria both families are present the Plasmodiophorids. At least family 2 is also contained in Sorites, Euglypha and Leptophrys, whereas CHS appear to been lost in R. filosa and B. natans. The ancient origin of the CHS families suggests that synthesis of chitin or chitin-like structures has been a driving force in the evolution and diversification of early eukaryotes.
Conclusion
We provide the de novo assembled and extensively annotated genome sequence of P. brassicae, the first pathogenic genome in the eukaryotic supergroup Rhizaria. Our data identified new evolutionary histories, as shown for the CHS proteins. This emphasizes the importance of genome data from neglected eukaryotic kingdoms such as the Rhizaria7 for a better understanding of the evolutionary events in Eukaryotes. We give an important first comprehensive insight into the life cycle events of this uncultivable intracellular pathogen, including basic metabolism, secreted proteins, CAZymes and phytohormones (Supplementary Fig. S27). Similar biotrophic features of Plasmodiophorids and phylogenetically unrelated eukaryotic biotrophic plant pathogens suggest a functional converging evolution. It remains challenging to elucidate the function of proteins in these uncultivatable protists. We provide Plasmodiophorid effector candidates which will be in focus of future studies to battle the agricultural important diseases caused by P. brassicae and other Plasmodiophorid pathogens.
Methods
DNA and RNA isolations
The P. brassicae single spore isolate e310 was used in this work. Several centrifugation steps followed by surface sterilization, antibiotic treatments, achieved resting spore isolation and purification. Prior DNA extraction resting spores were treated with DNaseI to minimize host DNA contamination. DNA was extraction from purified P. brassicae resting spores using a modified CTAB method46 with an additional phenol:chloroform step after lysis or using the IIlustra phytopure DNA extraction kit. Prior the lysis step, the spores were incubated in 5 mg/ml Trichoderma lysing enzyme in 0.64 M KCl and 0.2 M CaCl2 for 12–16 h at room temperature to weaken the resting spore cell walls and grinded in liquid nitrogen using a mortar and pestle. DNA was further purified using the gDNA Zymo purification kit or magnetic beads. Total RNA was obtained from surface sterilized clubroots of B. rapa, B. napus and B. oleracea var. capitata, 5 weeks post infection. Life-stage specific RNA was generated from germinating resting spores, maturing resting spores and plasmodia. S. subterranea RNA was extracted from potato root galls from potatoes grown for 2 months in natural infested soil. All RNA extractions were performed using the Spectrum™ Plant Total RNA Kit (Sigma-Aldrich).
Genome sequencing and assembly
The P. brassicae genome was assembled by combining Roche 454 FLX and lllumina Hiseq 2100 sequencing data. A standard and a mate pair library with 3 kb inserts were constructed and analyzed by Macrogen, Korea using their 454 platform. Illumina libraries used included a 5 kb mate-pair library sequenced at the Bejing Genome Institute, Hong Kong, China and a 200bp pair-end library sequenced at SciLife Laboratory, Stockholm, Sweden.
A first de novo genome draft of P. brassicae was assembled from 454 reads using Newbler v2.947 with default settings. Homopolymer errors were corrected by the Nesoni pipeline v0.109. High quality Illumina genomic reads were used to scaffold the 454 assembly with SSPACE v2.248 and to fill assembly gaps using GapFiller v1.1149. Scaffolds derived from contaminating host DNA were identified by BLASTN searches and their Illumina coverage and were removed from the assembly.
The P. brassicae genome assembly and the S. subterranea transcript data were checked for completeness by analysis of the single copy core eukaryotic orthologous genes using CEGMA 2.413. Genome size estimation of P. brassicae was based on k-mer analysis of raw Illumina genomic reads. 17-mer coverage distribution was inferred using jellyfish50, using only reads that mapped to the P. brassicae assembly. The peak depth was at 478x. The peak of 17-mer frequency (M) in reads is correlated with the real sequencing depth (N), read length (L) and kmer length (K), their relations can be expressed in the formula: M = N * (L – K + 1)/L. The total sequence length (1447 Gb) divided by the real sequencing depth estimated a total genome size of 25.47 Mb.
Transcriptome sequencing and filtering
RNA samples were sequenced with Illumina sequencing technology at SciLife Laboratory, Stockholm, Sweden and low-quality reads were removed. The transcriptome libraries of P. brassicae and S. subterranea were quality-filtered filtered using Condetri v.2.251. P. brassicae libraries were mapped to the genome assembly using Tophat v2.0.952 and analyzed using Cufflinks v 2.1.153, with default settings and an expected intron length of 5 to 5 kb. S. subterranea transcripts were de novo assembled using Trinity54. Potato host transcripts were removed from the subsequent analyses, if the assembled transcripts had a BLASTN match to either the Solanum tuberosum genome55 or A. thaliana (TAIR 9, www.arabidopsis.org) and no BLASTN match to the P. brassicae assembly at an e-value cut-off <10−9. We removed transcripts with BLASTN matches to the potential contaminant Potato virus A/Y (11 transcripts) and transcripts masked as retroelement (1288 transcripts) or transposons (251 transcripts), detected using RepeatMasker ( http://www.repeatmasker.org), combined with the repeat database, RepBase56. Only transcripts coding for peptides of minimal length of 100 amino acids were subsequently analyzed.
Gene content, annotation and expression
Repeats were annotated using RepeatModeler and RepeatMasker ( http://www.repeatmasker.org), combined with the repeat database, RepBase56 and the transposable element database of MAKER57. RepeatModeler was used to predict the novel repeat families and these families were combined with RepBase to produce the final library, from which RepeatMasker was used to call the consensus repeat sequences. For subsequent gene annotations, the genome was masked from these repeat regions, except for simple repeats. We annotated the repeat masked P. brassicae genome to predict protein-coding genes with combined annotation methods of ab initio gene annotation, homology-based gene prediction and transcriptome information using MAKER57. All ab initio predictors in the MAKER pipeline, except GeneMark-ES, were initially trained with the CEGMA set and iterated retrained. The strand-specific RNA-sequencing reads of P. brassicae mapped to the assembly were used as expressed sequence tag (ESTs) evidence. The P.brassicae ESTs, tBLASTX hits (e-value<10−10) to the UniProt/Swissprot database, to B. natans8 and R. filosa9 and to Rhizarian ESTs from the NCBI database, were combined to generate an protein-coding gene set using Maker. All predicted gene models were if necessary manually adjusted using Apollo58. Gene models with insufficient evidence from either transcript coverage or BLASTP searches were removed. For protein family classification analysis, we used Interproscan v5.44.059 to query multiple protein sequence and protein domain databases (Pfam v27.0, SMART v6.2, Gene3D v3.5.0, PANTHER v8.1, Phobius v1.01, SUPERFAMILY v1.75, PRINTS v42.0, ProSiteProfiles v20.89, ProSitePatterns). Gene Ontology terms were assigned with Blast2GO v2.560. Biological pathway information for the P. brassicae genome and S. subterranea transcriptome, was obtained using the KEGG Automatic Annotation Server (KAAS) ( www.genome.jp/tools/kaas/) with the non-organism specific option. Orthologous and close paralogous genes were identified using OrthoMCL v2.217. RNA sequencing reads mapped to the genome by TopHat52 were used for P. brassicae gene expression analysis by calculating the expected fragments per kilobase of transcript per million fragments (FPKM) using Cufflinks v 2.1.153. Gene expressions were visualized in heat maps as log10 transformed FPKM values and normalized by calculating Z-scores for each gene across all transcriptome libraries.
Secretome and selection of effector candidates
Proteins with a secretion signal predicted using SignalP-4.061 and no transmembrane domain predicted using TMHMM v2.062 were considered to be secreted. We identified 416 predicted secreted proteins below 450 amino acids in length. Number of candidates was reduced to 300 proteins as only genes that were expressed with a minimum value of at least 10 FPKM in at least one transcriptome library were considered. The transcriptome from the germinating spores was the only sample of P. brassicae without host contact. As effectors should interact with the plant we only selected protein models which genes had FPKM log2 fold change >5 in at least one other transcriptome library compared to the germinating spore library. This reduced the number of effector candidates to 92 (Supplementary Table 9).
Identification and analysis of carbohydrate-related proteins
Carbohydrate active enzymes (CAZymes) in the protein models of P. brassicae and the S. subterranea were predicted with dbCAN “DataBase for automated Carbohydrate-active enzyme Annotation” annotation pipeline63. The protein models for all other compared organism were analyzed equally.
Cloning, recombinant expression and conjugate synthetase test of the PbGH3 gene
The PbGH3 gene was cloned in the expression vector pGEX 4T3 to give a glutathione S-transferase (GST) fusion protein (GST::PbGH3). The GST::PbGH3 protein was recombinant expressed in Escherichia coli and the protein purified. Quality and quantity of the eluted fusion protein were verified using 12% SDS-PAGE gel electrophoresis. As controls E. coli cells transformed with empty vector were used. Conjugation activity tests were performed as previously described27,64.
Additional Information
Accession Codes: Data retrieved in this study are deposited at European Nucleotide Archive (ENA) under the projects PRJEB8376 (Accessions CDSF01000001-CDSF01000165) for the P. brassicae genomic data and PRJEB9159 (Accession HACM01000001-HACM01012732) for the S. subterranea transcriptome data.
How to cite this article: Schwelm, A. et al. The Plasmodiophora brassicae genome reveals insights in its life cycle and ancestry of chitin synthases. Sci. Rep. 5, 11153; doi: 10.1038/srep11153 (2015).
References
Dixon, G. R. The occurrence and economic impact of Plasmodiophora brassicae and clubroot disease. J. Plant Growth Regul. 28, 194–202 (2009).
Burki, F. et al. Evolution of Rhizaria: new insights from phylogenomic analysis of uncultivated protists. BMC Evol. Biol. 10, 377 (2010).
He, D. et al. An alternative root for the eukaryote tree of life. Curr. Biol. 24, 465–470 (2014).
Neuhauser, S., Kirchmair, M., Bulman, S. & Bass, D. Cross-kingdom host shifts of phytomyxid parasites. BMC Evol. Biol. 14, 33 (2014).
Kageyama., K. & Asano, T. Life cycle of Plasmodiophora brassicae. J. Plant Growth Regul. 28, 203–211 (2009).
Burki, F. & Keeling, P. J. Rhizaria. Curr. Biol. 24, R103–107 (2014).
Del Campo, J. et al. The others: our biased perspective of eukaryotic genomes. Trends Ecol. Evol. 29, 252–259 (2014).
Curtis, B. A. et al. Algal genomes reveal evolutionary mosaicism and the fate of nucleomorphs. Nature 492, 59–65 (2012).
Glöckner, G. et al. The genome of the foraminiferan Reticulomyxa filosa. Curr. Biol. 24,11–18 (2014).
Fähling, M., Graf, H. & Siemens, J. Characterization of a single-spore isolate population of Plasmodiophora brassicae resulting from a single club. J. Phytopathol. 152, 438–444 (2004).
Moxham, S. E. & Buczacki, S. T. Chemical composition of the resting spore wall of Plasmodiophora brassicae. Trans. Br. Mycol. Soc. 80, 297–304 (1983).
Sánchez-Vallet, A., Mesters, J. R. & Thomma, B. P. H. J. The battle for chitin recognition in plant-microbe interactions. FEMS Microbiol. Rev. 39,171–183 (2014).
Graf, H., Sokolowski, F., Klewer, A., Diederichsen, E., Luerssen, H. & Siemens, J. Electrophoretic karyotype of the obligate biotrophic parasite Plasmodiophora brassicae Wor. J. Phytopathol. 149, 313–318 (2001).
Keeling, P. J. & Slamovits, C. H. Causes and effects of nuclear genome reduction. Curr. Opin. Genet. Develop. 15, 601–608 (2005).
Parra, G., Gradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23,1061–1067 (2007).
Bulman, S. et al. Genomics of biotrophic, plant-infecting plasmodiophorids using in vitro dual cultures. Protist 162, 449–461 (2011).
Li, L., Stoeckert, C. J. Jr. & Roos, D. S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003).
Jiang, R. H. Y. et al. Distinctive expansion of potential virulence genes in the genome of the oomycete fish pathogen Saprolegnia parasitica. PLoS Genet. 9, e1003272 (2013).
Spanu, P. D. et al. Genome expansion and gene loss in powdery mildew fungi reveal tradeoffs in extreme parasitism. Science 330, 1543–1546 (2010).
Baxter, L. et al. Signatures of adaptation to obligate biotrophy in the Hyaloperonospora arabidopsidis genome. Science 330, 1549–1551 (2010).
Kemen, E. et al. Gene gain and loss during evolution of obligate parasitism in the white rust pathogen of Arabidopsis thaliana. PLoS Biol. 9, e1001094 (2011).
Oshima, K. et al. Reductive evolution suggested from the complete genome sequence of a plant-pathogenic phytoplasma. Nature Genet. 36, 27 – 29 (2004).
Lee, S. H., Stephens, J. L., Paul, K. S. & Englund, P. T. Fatty acid synthesis by elongases in trypanosomes. Cell 126, 691–699 (2006).
Brodmann, D. et al. Induction of trehalase in Arabidopsis plants infected with the trehalose-producing pathogen Plasmodiophora brassicae. Mol. Plant-Microbe Interact. 15, 693–700 (2002).
Wilson, R. A., Gibson, R. P., Quispe, C. F., Littlechild, J. A. & Talbot, N. J. An NADPH-dependent genetic switch regulates plant infection by the rice blast fungus. Proc. Natl. Acad. Sci. USA 107, 21902–21907 (2010).
Ludwig-Müller, J., Prinsen, E., Rolfe, S. A. & Scholes, J. D. Metabolism and plant hormone action during clubroot disease. J. Plant Growth Reg. 28, 229–244 (2009).
Siemens, J. et al. Transcriptome analysis of Arabidopsis clubroots indicates a key role for cytokinins in disease development. Mol. Plant-Microbe Interact. 19, 480–494 (2006).
Ludwig-Müller, J. et al. A novel methyltransferase from the intracellular pathogen Plasmodiophora brassicae participates in methylation of salicylic acid. Mol. Plant Pathol. 10.1111/mpp.12185 (2014).
Staswick, P. E. et al. Characterization of an Arabidopsis enzyme family that conjugates amino acids to indole-3-acetic acid. Plant Cell 17, 616–627 (2005).
Lindner, A. C. et al. Isopentenyltransferase-1 (IPT1) knockout in Physcomitrella together with phylogenetic analyses of IPTs provide insights into evolution of plant cytokinin biosynthesis. J. Exp. Bot. 65, 2533–2543 (2014).
Malinowski, R. et al. Gall formation in clubroot infected Arabidopsis results from an increase in existing meristematic activities of the host but is not essential for the completion of the pathogen life cycle. Plant J. 71, 226–238 (2012).
Ökmen, B. & Doehlemann, G. Inside plant: biotrophic strategies to modulate host immunity and metabolism. Curr. Opin. Plant Biol. 20, 19–25 (2014).
Desoignies, N. et al. Molecular interactions between sugar beet and Polymyxa betae during its life cycle. Ann. Appl. Biol. 164, 244–256 (2014).
Wawra, S. et al. Secretion, delivery and function of oomycete effector proteins. Curr. Opin. Microbiol. 15, 685–691 (2012).
Pan, X., Lührmann, A., Satoh, A., Laskowski-Arce, M. A. & Roy, C. R. Ankyrin repeat proteins comprise a diverse family of bacterial type IV effectors. Science 320,1651–1654 (2008).
Fan, L. et al. Functional equivalence and evolutionary convergence in complex communities of microbial sponge symbionts. Proc. Natl. Acad. Sci. USA 109, E1878–1887 (2012).
Hashimoto, M., Murata, E. & Aoki, T. Secretory protein with RING finger domain (SPRING) specific to Trypanosoma cruzi is directed, as a ubiquitin ligase related protein, to the nucleus of host cells. Cell Microbiol. 12, 19–30 (2010).
Laurie, J. D. et al. Genome comparison of barley and maize smut fungi reveals targeted loss of RNA silencing components and species-specific presence of transposable elements. Plant Cell 24, 1733–1745 (2012).
Zhao, Z. T., Liu, H. Q., Wang, C. F. & Xu, J. R. Correction: comparative analysis of fungal genomes reveals different plant cell wall degrading capacity in fungi. BMC Genom. 15, 6 (2014).
Aspeborg, H., Coutinho, P. M., Wang, Y., Brumer, H. & Henrissat, B. Evolution, substrate specificity and subfamily classification of glycoside hydrolase family 6 (GH5). BMC Evol. Biol. 12, 186 (2012).
Karlsson, M. & Stenlid, J. Evolution of family 18 glycoside hydrolases: diversity, domain structures and phylogenetic relationships. J. Mol. Microbiol. Biotechnol. 16, 208–223 (2009).
Soanes, D. M. et al. Comparative genome analysis of filamentous fungi reveals gene family expansions associated with fungal pathogenesis. PLoS One 3, e2300 (2008).
Liu, P. & Stajich, J. E. Characterization of the carbohydrate binding module 18 gene family in the amphibian pathogen Batrachochytrium dendrobatidis. Fungal Genet. Biol. 77, 31–39 (2015).
Ruiz-Herrera, J., González Prieto, M. J. & Ruiz, Medrano,. R. Evolution and phylogenetic relationships of chitin synthases from yeasts and fungi. FEMS Yeast Res. 1, 247–256 (2002).
Zakrzewski, A. C. et al. Early divergence, broad distribution and high diversity of animal chitin synthases. Genome Biol. Evol. 6, 316–325 (2014).
Russel, J. & Bulman, S. The liverwort Marchantia foliacea forms a specialized symbiosis with arbuscular mycorrhizal fungi in the genus Glomus. New Phytol. 165, 567–579 (2005).
Wijaya, E., Frith, M. C., Suzuki, Y., & Horton, P. Recount: expectation maximization based error correction tool for next generation sequencing data. Genome Inform. 23,189–201 (2009).
Boetzer, M., Henkel, C. V., Jansen, H. J., Butler, D. & Pirovano, W. Scaffolding pre-assembled contigs using SPACE. Bioinformatics 27, 578–579 (2011).
Boetzer, M. & Pirovano, W. Toward almost closed genomes with GapFiller. Genome Biol. 13, R56 (2012).
Marcais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770 (2011).
Smeds, L. & Künstner, A. ConDeTri– a content dependent read trimmer for Illumina data. PLoS One 6, e26314 (2011).
Trapnell, C., Pachter, L. & Salzberg, S. L. Tophat: discovering splice junctions with RNA-seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Grabherr, M. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
The Potato Genome Sequencing Consortium (TPGSC). Genome sequence and analysis of the tuber crop potato. Nature 475, 189–197 (2011).
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenet. Genome Res. 110, 462–467 (2005).
Cantarel, B. L. et al. MAKER : An easy- to-use annotation pipeline designed for emerging model organism genomes. Genome Res. 18, 188–196 (2008).
Lee, E., Harris, N., Gibson, M., Chetty, M. & Lewis, S. Apollo: a community resource for genome annotation editing. Bioinformatics 25, 1836–1837 (2009).
Hunter, S. et al. InterPro in 2011: new developments in the family and domain prediction database. Nucleic Acids Res. 40, D306–D312 (2012).
Conesa, A. et al. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676 (2005).
Petersen, T. N., Brunak, S., von Heijne, G. & Nielsen, H. SignalP4.0: discriminating signal peptides from transmembrane regions. Nat. Methods 8, 785–786 (2011).
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001).
Yin, Y. et al. dbCAN: a web resource for automated carbohydrate-active enzyme annotation, Nucleic Acids Res. 40, W445–451 (2012).
Ludwig-Müller, J., Jülke, S., Bierfreund, N. M., Decker, E. L. & Reski, R. Moss (Physcomitrella patens) GH3 proteins act in auxin homeostasis. New Phytol. 181, 323–338 (2009).
Arnold, K., Bordoli,L., Kopp, J. & Schwede, T. The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22, 195–201 (2006).
Pettersen, E. F. et al. UCSF chimera—a visualization system for exploratory research and analysis. J Comput. Chem. 25, 1605–1612 (2004).
Acknowledgements
The work was funded by the BioSoM program, the Swedish Research Councils (Formas & VR), the Swedish University of Agricultural Sciences (SLU), the Golden Seed Project (No. 213002-04-3-SB110), the Korean Ministries of Agriculture, Food and Rural Affairs (MAFRA) and of Oceans and Fisheries (MOF), the Rural Development Administration (RDA) and the Korea Forest Service (KFS). The authors would like to thank Salla Marttila for SEM photos, Roberto Sierra (University of Geneva, Switzerland) for additional CHS sequences data, Tim Kamber for comments and support from Science for Life Laboratory, the National Genomics Infrastructure (NGI), Sweden, the Knut and Alice Wallenberg Foundation and UPPMAX for providing assistance in DNA sequencing and computational infrastructure.
Author information
Authors and Affiliations
Contributions
A.S. and C.D. managed the project. A.S. prepared the DNA and RNA samples. A.S. and J.F. designed the analysis. J.F. performed the genome assembly. J.F. and T.L. performed the genome annotation. A.S. and A.K. prepared samples for peptide analyses and A.K. and A.Sh. performed peptide sequencing and analyses. S.J. and J.L.M. performed the PbGH3 cloning and enzymatic testing. G.B.R., M.K., J.F. and A.S. performed the phylogenetic analyses. J.F. performed data submission and database construction. A.S. wrote the paper with substantial contributions of J.L.M., J.F. Y.P.L. and C.D. All of the other authors contributed annotation, analyses or data throughout the project. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Schwelm, A., Fogelqvist, J., Knaust, A. et al. The Plasmodiophora brassicae genome reveals insights in its life cycle and ancestry of chitin synthases. Sci Rep 5, 11153 (2015). https://doi.org/10.1038/srep11153
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep11153
This article is cited by
-
The conserved GOLD domain in the Plasmodiophora brassicae effector Pb257 is required for triggering cell death and root swelling
Phytopathology Research (2024)
-
Advancements in Spongospora subterranea: Current Knowledge, Management Strategies, and Research Gaps
Potato Research (2024)
-
Clubroot (Plasmodiophora brassicae) Suppression Under Biocontrol Agents in Pak choi with Variations in Physiological, Biochemical, and Bacterial Diversity
Journal of Plant Growth Regulation (2023)
-
Genotypic characterization of Plasmodiophora brassicae in the paddy-field weed Cardamine occulta and symptomology reveal a distinct pathogen population in Japan
Journal of General Plant Pathology (2023)
-
Protocol: rhPCR and SNaPshot assays to distinguish Plasmodiophora brassicae pathotype clusters
Plant Methods (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.