Abstract
The bicarbonate ion (HCO3−) is involved in two major physiological processes in corals, biomineralization and photosynthesis, yet no molecular data on bicarbonate transporters are available. Here, we characterized plasma membrane-type HCO3− transporters in the scleractinian coral Stylophora pistillata. Eight solute carrier (SLC) genes were found in the genome: five homologs of mammalian-type SLC4 family members and three of mammalian-type SLC26 family members. Using relative expression analysis and immunostaining, we analyzed the cellular distribution of these transporters and conducted phylogenetic analyses to determine the extent of conservation among cnidarian model organisms. Our data suggest that the SLC4γ isoform is specific to scleractinian corals and responsible for supplying HCO3− to the site of calcification. Taken together, SLC4γ appears to be one of the key genes for skeleton building in corals, which bears profound implications for our understanding of coral biomineralization and the evolution of scleractinian corals within cnidarians.
Similar content being viewed by others
Introduction
Symbiotic cnidarians from the order Scleractinia form the foundation of coral reefs that constitute one of the most important biogenic structures worldwide. Coral reefs provide habitat and trophic support for myriad of marine species, the richness of which rivals the biological diversity of tropical rainforests1. Despite their environmental significance, key elements of coral physiology, such as the symbiotic interactions between the animal host and its intracellular photosynthetic dinoflagellates of the genus Symbiodinium, or the biomineralization process underlying the formation of the coral skeleton, are poorly understood. This is largely due to knowledge gaps in fundamental aspects of cnidarian cell biology2,3.
Photosynthesis by the symbionts and the biomineralization process both involve the use of dissolved inorganic carbon (DIC). In seawater, DIC exists in the form of chemically inter-convertible molecules that exist in a pH dependent equilibrium: the non-ionic form, CO2, with a concentration on the order of 10 μM at normal seawater pH of 8.1 and two ionic forms, HCO3− and CO32−, with concentrations of up to 200 times higher (i.e. ~2.4 mM). Photosynthesis and calcification, however, occur in compartments that are not in direct contact with seawater and thus need to be actively supplied with DIC. For instance, the intracellular symbionts are located in the endodermal tissue layer, separated from seawater by the ectoderm tissue layer. Further, symbionts are separated from the host cytoplasm by the perisymbiotic membrane4. In order to secure continuous provision of DIC despite these constraints, the coral has developed CO2-concentrating mechanisms to absorb and transfer DIC from the seawater to its symbionts for photosynthesis5. DIC uptake by the host involves an H+-ATPase that acidifies the ectodermal boundary layer where bicarbonate (HCO3−) is converted to CO2 by a membrane-bound isoform of carbonic anhydrase (CA). The uncharged CO2 molecule then diffuses into the epidermal cells. Once in the animal cytoplasm, another CA isoform is involved in the equilibration between CO2 and HCO3− according to the intracellular pH6, which prevents CO2 back-diffusion (for review see7). The mechanism of DIC transport through the remaining membranes to the symbionts is currently debated, but it is accepted that bicarbonate has to exit the ectodermal cells to subsequently enter the endodermal cells by a bicarbonate anion transporter (BAT) (for reviews, see5,8). Physiological experiments with radioactive tracers together with pharmacological experiments using classic bicarbonate transport inhibitors, such as 4,4’-Diisothiocyano-2,2’-stilbenedisulfonic acid (DIDS)9, support the contribution of BATs in the supply of DIC required for photosynthesis by the symbiont (see Fig. 1,5,8). However, no molecular data on BATs in cnidarians are currently available.
Structural histology and model of DIC transport through the different coral tissue layers (modified from Bertucci et al60). Dotted arrows represent CO2 diffusion, the projected involvement of BATs is indicated by red circles. Zoox.: zooxanthellae; M.: mitochondrion. CA: Carbonic Anhydrase; PMCA: plasma membrane Calcium ATPase. Unknown mechanisms of DIC transport are indicated with a question mark
The calcification process itself occurs at a site that is separated from seawater by four tissue layers: the coral oral ectoderm and endoderm, as well as the coral aboral endoderm and ectoderm. The major portion of the DIC used for CaCO3 formation comes from metabolic CO25,10, which is produced by the calcifying cells forming the aboral ectoderm (also called the calicoblastic ectoderm). Part of this metabolic CO2 may diffuse across the plasma membrane, whereas another part is hydrated into HCO3− due to the alkaline intracellular pH5,6. The proportion of CO2 diffusion versus CO2 hydration is currently unknown. However, inhibition of anion transport with DIDS also inhibits calcification5,11 indicating that BATs are likely involved in the transport of HCO3− across the calcifying cells to the site of calcification, but further data on the molecular characteristics of the transporter(s) involved is lacking (Fig. 1).
In Mammals, two distinct families of membrane BATs are differentiated: solute carrier 4 (SLC4) (for review,12,13) and solute carrier 26 (SLC26)14 transporters. The SLC4 family represents the majority of HCO3− transporters, whereas the SLC26 family consists of members that can transport diverse ions besides HCO3−. Mammalian BATs consist of 14 genes, nine SLC4 members and five from the SLC26 family15. SLC4 family members are separated into three functional groups: i) Na+-independent Cl−/HCO3− exchangers, mediating electroneutral exchange of Cl− for HCO3−; ii) Na+-HCO3− co-transporters mediating the co-transport of Na+ and HCO3− and iii) Na+-driven Cl−/HCO3− exchangers (NDCBE) mediating the electroneutral exchange of Cl− for Na+ and HCO3−. Within these groups, proteins are separated according to their phylogenetic position, tissue distribution, anion selectivity, regulatory properties and mechanism of action. For example, the first group contains SLC4A1 (also named Band 3 or AE1), SLC4A2 (or AE2) and SLC4A3 (or AE3); the second group contains SLC4A4 (or NBC1), SLC4A5 (or NBC4), SLC4A7 (or NBC3), SLC4A9 (or AE4) and SLC4A10 (or NCBE); NDCBE is only represented by SLC4A8. SLC4A11 (or BTR1) was originally reported as a sodium/borate co-transporter16, but more recently found to facilitate water and Na+ fluxes17,18. The human SLC26 family consists of 11 members, where SLC26A10 is likely a pseudogene19. Similar to SLC4 family members, SLC26 family members can be grouped into three groups: i) the SO42 transporters, ii) the Cl−/HCO3− exchangers (also called the SLC26 BATs) and iii) the selective Cl− channels (it should be noted that this last group has minimal HCO3− permeability19). The group of Cl−/HCO3− exchangers consists of either electroneutral (SLC26A3, SLC26A4 and SLC26A6) or electrogenic (SLC26A7 and SLC26A9) transporters15.
Besides a physiological understanding of the process of calcification, insights into the extent of conservation of gene families within Cnidaria might hold key insights for our understanding of the evolution of this phylum. For instance, an evolutionary analysis of the innate immunity gene repertoire among Scleractinia (i.e. Acropora millepora), Actinaria (i.e. Nematostella) and Hydrozoa (i.e. Hydra) revealed an extended Toll-like receptor (TLR) gene set in corals in line with the symbiotic lifestyle of corals20.
To gain insight into the composition and evolution of coral BATs, we performed genome data mining and monitored expression and localization of eight identified putative solute carrier (SLC) proteins in the coral Stylophora pistillata, the species from which many of the aforementioned physiological data were obtained. Our present study aimed at defining a comprehensive analysis of the repertoire of these transporters and to discuss their potential role in symbiosis and biomineralization. Further, we were interested in the distribution of these transporters across key cnidarian taxa to contribute to our understanding of the evolution of scleractinian corals within the phylum Cnidaria.
Results
Candidate HCO3− Transporters in Stylophora pistillata
Coral genomes have a remarkably high level of conservation with vertebrates in comparison to their invertebrate counterparts, such as flies and nematodes21. We identified eight candidate HCO3− anion transporter (BAT) genes (Fig. 2 and File S1) in the genome of S. pistillata (genome sequence provided by C.R.V. and M.A, personal communication, in collaboration with D. Z., S. T. and D.A.,22. Of these genes, five (referred to as SpiSLC4α, β, γ, δ and ε) and three (referred to as SpiSLC26α, β and γ) were similar to members of the mammalian SLC4A and SLC26A family, respectively. With the exception of SpiSLC4α, these genes were expressed and found in S. pistillata EST libraries23. In addition, we have identified homologs of these genes in the coral A. digitifera and the sea anemone N. vectensis (Table S2), two anthozoans for which a genome sequence is available20,24. Comparison between homologs and paralogs of the three species showed that, surprisingly, no homolog of the scleractinian SLC4γ was found in the N. vectensis genome (Table S3). Searches in the transcriptome25 and genome (which CEGMA pourcentage completeness is 96.72%) of another anemone, Aiptasia pallida (Baumgarten, S., Simakov, O., Esherick, L. Y., Liew, Y. J., Lehnert, E. M., Michell, C. T., Li, Y., Hambleton, E. A., Guse, A., Oates, M. E., Gough, J., Weis, V. M., Aranda, M., Pringle, J. R. & Voolstra, C. R. ; personal communication) , confirmed the absence of this SLC4γ homolog in actinarians. Phylogenetic reconstruction using human and cnidarian homologs (Fig. 3) indicated that the cnidarian SLC4 members fall into the categories previously described13,15. These categories were electroneutral (human SLC4A7, A8 and A10) and electrogenic (human SLC4A4, A5 and A9) Na+-coupled Cl−/HCO3− co-transporters, Na+-independent Cl−/HCO3− exchangers (human SLC4A1, A2 and A3) and Na+/borate co-transporters (human SLC4A11). Furthermore, evolutionary relationships indicate that the SLC26 members fall into the two phylogenetic groups described by Dorwart et al.19 consisting of Cl−/HCO3− exchangers and selective Cl− channels.
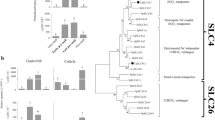
Phylogenetic relationships of human and anthozoan Bicarbonate Anion Transporter protein sequences inferred from Maximum Likelihood (ML) and Bayesian analyses. Bootstrap network of BAT sequences based on ML distances are estimated with a LG+G model (α = 0.749) using PHYML. Bayesian posterior probabilities are indicated in black whereas ML bootstrap values are in red.
Sequence analysis
Structure prediction showed that the S. pistillata SLC4 family members (in the following referred to as SpiSLC4s) exhibit the canonical three domain pattern shared in all SLC4 transporters13,26: a long N-terminal hydrophilic domain of 424, 380, 511, 496 and 362 amino acids for SpiSLC4α, β, γ, δ and ε respectively and a short C-terminal hydrophilic domain (Fig. 2). Both termini are known to be intracellular27. The central part consisted of 11-13 predicted transmembrane segments, although the exact number of transmembrane segments remains unknown due to possible re-entrant loops in the C-terminal half of some membrane domains13. With regard to post-translational modifications, Asn577 and Asn528 were predicted to be N-glycosylated in SpiSLC4α and SpiSLC4ε respectively. Potential N-glycosylation sites in the other proteins were located in the cytoplasmic part of the proteins and, consequently, did not support glycosylation. Furthermore, we predicted 31, 44, 56, 47 and 33 phosphorylation sites for SpiSLC4 α, β, γ, δ and ε, respectively. Of note, the prominent BAT inhibitors, the stilbene inhibitors DIDS and H2DIDS28, covalently label Lys539 and/or Lys851 in human SLC4A1; the first Lys residue was present only in SpiSLC4δ (amino acid 630), whereas the second was conserved in all the isoforms (amino acid 876, 827, 961 and 976 for SpiSLC4α, β, γ and δ respectively).
Structural analysis of SLC26 proteins showed that S. pistillata SLC26 members (in the following referred to as SpiSLC26s) exhibited shorter N-terminal domains than the SLC4 family (112, 121 and 45 amino acids for SpiSLC26α, β and γ respectively). The C-terminal domains of all the mammalian SLC26 proteins included a sulfate transporter domain and an anti-sigma factor antagonist (STAS) domain29. Similarly, the three S. pistillata proteins also shared the sulfate transporter domain (amino acids 234–511 for SpiSLC26α, amino acids 228–505 for SpiSLC26β and 131–433 for SpiSLC26γ) and the STAS domain (amino acids 567–738 for SpiSLC26α, amino acids 561–708 for SpiSLC26β and 476–569 for SpiSLC26γ), as shown by CDD blast analysis30. Potential glycosylation sites were located at the amino acid 294 for SpiSLC26α, amino acid 389 for SpiSLC26β and amino acid 291 for SpiSLC26γ. Phosphorylation site predictions identified 40, 42 and 20 phosphorylation sites for SpiSLC26α, β and γ respectively.
Gene structure
The genomic structure of the different BAT genes was established (Fig. 4 and Table 1) using the SIM4 algorithm31. Genomic sequences are given in File S4. It should be noted that SpiSLC4β and SpiSLC4γ are on the same scaffold and separated by 18017 bp and that SpiSLC26α and SpiSLC26β genes are also on the same scaffold and separated by 2491 bp.
Tissue distribution and localization of the different isoforms
Relative expression PCR was performed on colony-wide RNA (total) and oral discs RNA obtained by micro-dissection (32 and Fig. 5A). Experiments were performed on three independent colonies grown in the aquarium facilities of the Centre Scientifique de Monaco. Expression in the two fractions were normalized to acidic ribosomal phosphoprotein P0 expression (36B4), which is not affected by experimental conditions or tissue specificity33. SpiSLC26α and γ showed higher expression in oral tissues whereas SpiSLC26β was ubiquitously expressed in the different tissues of S. pistillata (Fig. 5B). With regard to the expression of the SLC4 family members, we did not detect expression of SpiSLC4α in coral tissues, as observed for transcriptomic data (see above). SpiSLC4β was expressed almost twice as high in oral tissue, while SpiSLC4δ showed ubiquitous expression. Intriguingly, SLC4γ was either absent or very weakly expressed in the oral fraction, suggesting a specific expression in the aboral tissues. The expression of SpiSLC4ε showed large inter-individual variation, which prevented further interpretation. In the following, we confirmed gene expression results with protein localization using specific antibodies raised against the ubiquitously expressed SpiSLC26β and the highly specifically expressed SpiSLC4γ. Paraffin-embedded cross-sections of S. pistillata encompassing the different tissues (see Fig. 1 and34,35 for histology details) were used to visualize the proteins using specific antibodies (Fig. 6). Sections of S. pistillata tissues labeled with the anti-SpiSLC26β antibody (Fig. 6 B) and magnification thereof (Fig. 6 C) confirmed that its location was ubiquitous, with signals in the ectoderm and endoderm of both oral and aboral tissues. In contrast, immunolocalization of SpiSLC4γ localized this protein to the calicoblastic ectoderm (i.e. the calcifying cell layer), clearly associated with the plasma membrane (Fig. 6 E and F). Negative controls, using preimmune serum showed no labeling (Fig. 6 A and D).
Relative expression of the different BAT proteins in coral tissues based on qPCR. (A) cDNA are prepared from total tissues (whole coral fragment), or from oral disc. (B) Gene expression normalized to 36B4 expression in total tissues (Total), or in oral disc (Oral). Grey bars represent different colonies. Error bars represent the SD of three technical replicates of each colony.
Immunolocalization of SpiSLC26β and SpiSLC4γ. Embedded cross-section of Stylophora pistillata tissues labeled by (A) preimmune serum for SpiSLC26β, (B) and (C) anti-SpiSLC26β, (D) preimmune serum for SpiSLC4γ and (E) and (F) anti- SpiSLC4γ antibody. Rows (A), (B), (D) and (E) are views of the four tissues composing the coral. (C) are magnifications of oral tissues and (F) are magnifications of aboral tissues. Nuclei are labeled in blue in first column (DAPI), streptavidin AlexaFluor 568 fluorescence appears in orange in second column, merged is in the third column. The background red color in cross-section (A) and (D) with preimmune serum corresponds to autofluorescence of coral tissues. AEnd = Aboral Endoderm; AT = Aboral Tissue; CEct = Calicoblastic Ectoderm; Co = Coelenteron; m = Mesoglea; OEct = Oral Ectoderm; OEnd = Oral Endoderm; OT = Oral Tissue; Sw = Seawater; Sk = Skeleton
Discussion
In this study, we provided a molecular characterization of BATs in the coral Stylophora pistillata and showed the presence of bicarbonate anion transporters families SCL4 and SCL26 in this coral. Further, we conducted a phylogenetic analysis of these protein families in other cnidarian taxa.
Analysis of partial sequences of the starlet anemone Nematostella vectensis, led to the suggestion that Na+-coupled HCO3− transporters SLC4 appeared first in Cnidaria13. The characterization of the full-length SpiSLC4δ and its Acropora homolog shows undoubtedly that these transporters are indeed present in Cnidaria. While the earlier analysis reported four members of SLC4 in the starlet anemone, we could identify five members in the corals S. pistillata and Acropora digitifera (Fig. 2 and 3). We hypothesize that this is due to a scleractinian specific gene duplication of a Cl−/HCO3− exchanger as shown in the phylogenic tree (Fig. 3). Interestingly, in A. digitifera, AdiSLC4β and AdiSLC4γ were present on the same scaffold separated by 64977 bp in a forward-reverse orientation, whereas in S. pistillata, SpiSLC4β and SpiSLC4γ were separated by 18017 bp in a forward-forward orientation. Gene duplication is a main mechanism through which new genetic material is generated during molecular evolution and can lead to evolutionary innovation36. Indeed, a SpiSLC4β homolog is present in sea anemones (N. vectensis and A. pallida) and corals (A. millepora and S. pistillata), whereas SpiSLC4γ was only present in hard corals. Thus, SpiSLC4β and SpiSLC4γ seem to be paralogs and should be referred as SpiSLC4β1 and SpiSLC4β2. However, since our findings suggest that SpiSLC4γ has acquired a specific role in skeleton building, we decided to keep distinct names for both genes. The assumed neofunctionalization of SpiSLC4γ is further supported by the highly specific expression and restricted distribution of this gene within coral tissues (see below).
Pharmacological experiments with BAT inhibitors are unable to discriminate whether SLC26 or SLC4 BATs are involved in calcification and photosynthesis. Stilbene dilfonates inhibit bicarbonate transport because of their dual hydrophobic/anionic character9. Some of these compounds (like DIDS) have covalently-reactive isothiocyanate groups that covalently label specific lysine residues in SLC4 family proteins. Stilbene disulfonates, however, also inhibit anion transporters non-covalently, meaning that that will also target SLC26 proteins lacking the conserved DIDS-reactive lysine residues. Assessment of BAT tissue expression patterns was thus necessary to define the BATs involved in calcification and symbiosis. Relative expression analysis (Fig. 5) of SpiSLC4α shows that this protein is not expressed in adult tissues, suggesting that it could be specific to the embryonic stages. SpiSLC4δ, the putative sodium-coupled transporter and SLC26β are ubiquitously expressed in all tissues. Therefore they are unlikely to play a specific role in photosynthesis or calcification, but may have a role in homeostatic control of processes, including cell pH, volume and bicarbonate metabolism. The higher relative expression of SpiSLC26α, SpiSLC26γ and SpiSLC4β in oral tissues (Fig. 5) suggests that these transporters rather play a role in symbiosis than in calcification. Indeed, as outlined in the introduction, bicarbonate is transported by ectodermal cells, then enters endodermal cells and finally is delivered to the dinoflagellate symbionts. All these steps of transcellular transport thus involve BATs and we suggest that SpiSLC4β is one of the proteins responsible for this function. Moreover the COOH-terminal tail of SLC4 proteins contain one or more acidic motifs that may serve as a binding site for the cytoplasmic carbonic anhydrase II (CAII)37. In SpiSLC4β, this domain is located between residues 861 and 865 (LDNEE). The cytoplasmic binding of CAII and the simultaneous interaction of SLC4 anion exchangers with carbonic anhydrases have been proposed to constitute a bicarbonate transport metabolon37,38. Carbonic anhydrases play a major role in carbon supply for photosynthesis in corals5 and further, a cytoplasmic CA is located in symbiotic endodermal cells39. This suggests that the BAT SpiSLC4β protein might be coupled to a cytoplasmic carbonic anhydrase, such as STPCA2 and could serve to accelerate transmembrane bicarbonate transport40.
Finally, in our data SpiSLC4γ was completely absent from oral coral tissues, it has not been found in sea anemones and it localizes to the ectodermal calcifying cells of corals. These data strongly suggest that SpiSLC4γ plays a key role in calcification and might represent one of the evolutionary key innovation genes for calcification in Scleractinia. The calcification process produces protons that need to be removed from the site of calcification in order to favor the precipitation of calcium carbonate41,42. It has been proposed that removal of protons occurs through a plasma membrane Ca2+-ATPase that is located in the membranes of the calcifying cells and which functions as a proton exchanger (Fig. 1)43. In addition, the observed increase in pH to values higher than seawater pH44 could be performed by supplying bicarbonate at the site of calcification. Such a role might be performed by the here-characterized SpiSLC4γ. Indeed, in mammals BATs are able to neutralize stomach acid entering the intestine by secreting high concentrations of bicarbonate45. Sodium dependent- and independent-SLC4, together with SLC26 have been proposed to be involved in this mechanism46. Our results suggest that BATs are involved in supplying bicarbonate to the site of calcification and possibly play a role in extracellular pH regulation.
Over recent decades, coral reefs have been impacted by the effects of global environmental change47 and are now threatened by ocean acidification48. Several experimental studies have shown that a low pH may have detrimental effects on calcification rates and skeletal growth of various coral species49,50,51,52. Transcripts corresponding to SLC4 members were up-regulated in response to a CO2-driven pH decrease experiment with Pocillopora damicornis53, suggesting that BATs may play a role in coral resilience when facing environmental acidification conditions. The emergence of the gene SLC4γ via gene duplication appears to be a key adaptation in the evolution of calcification in scleractinian corals. In this context, the role of HCO3− in supplying DIC for coral calcification, as well as further elucidation of the function and evolution of BATs and SLC4γ in particular, is fundamental for determining the response of coral reefs to ocean acidification.
Methods
Coral culture
Experiments were conducted in the laboratory using the zooxanthellate scleractinian coral Stylophora pistillata. Colonies were cultivated as indicated previously11.
Data mining
Sequences homologous to human SLC4 and SLC26 amino acid sequences from NCBI (http://www.ncbi.nlm.nih.gov/protein) were identified amongst Stylophora pistillata ESTs obtained with 454 pyrosequencing23 or Illumina22, using the TblastN algorithm. Furthermore, we used a draft assembly of the S. pistillata genome (unpublished data). A CEGMA analysis54 reports the conserved proteome to be 94.5% complete, which is similar to the A. digitifera genome. Acropora digitifera and Nematostella vectensis sequences were retrieved from the following web servers: http://marinegenomics.oist.jp/genomes/ and http://www.cnidariangenomes.org/download/nve.gene_models.vie130208/, respectively.
Sequence analysis
Putative transmembrane domains were predicted using the Phobius server55. Phosphorylation and N-glycosylation prediction analyses were performed with NetPhos and NetNGlyc respectively on the Center for Biological Sequence analysis prediction server (http://www.cbs.dtu.dk/services/).
Phylogenetic constructions
Phylogenetic trees were constructed with both Maximum Likelihood and Bayesian methods in order to assess result congruencies. ClustalW alignments of all amino acids sequences were performed using MultAlin56 with the Blosum62 default parameters. Based upon amino acid alignment, maximum likelihood estimates of the topology and branch length were obtained using PhyML v3.057 with the LG+G model of substitution as recommended by alignment analysis with ProtTest v3.458. Further, phylogenetic relationships were investigated using Bayesian methods as implemented in MrBayes v3.1.259 starting from a random tree, using the LG model of amino acid substitution generating trees for 6,000,000 generations with sampling every 1000 generations and with four chains in order to obtain the final (consensus) tree and to determine the posterior probabilities at the different nodes.
Oral disc dissection
Fragments of Stylophora pistillata colony were used for RNA extractions. They were set to rest in a glass petri dish filled with sea water until polyps were extended. Tricaine mesylate (MS-222, Sigma) dissolved in sea water to 0.4% was added into the petri dish to a final concentration of 0.04% and incubated under dimmed light for 15 min. Subsequently, viewed under a binocular microscope oral discs (the apparent portion of the polyp, Fig. 5A) were cut from the colony, using micro-dissection scissors with 5 mm blades (Vannas). Batches of 10–15 oral discs were collected and transferred into Trizol®. Dissections were stopped after a maximum of 45 min of MS-222 incubation to elude any potential secondary effect of the drug.
Real-time PCR
Total RNA extraction and cDNA synthesis were performed as described previously39. Briefly, cDNAs were synthesized using the Superscript®III kit (Invitrogen). qPCR runs were performed, as in33, on an ABi 7300 using “EXPRESS SYBR® GreenER® qPCR Supermix with Premixed ROX” for PCR amplification. Primers used are listed in Supplementary Table S5. Relative expressions were calculated using Biogazelle qbase +2.6™.
Custom made antibodies
Antibodies against SpiSLC26β and SpiSLC4γ were produced in rabbit using synthetic peptides (Eurogentec). Anti-SpiSLC26β was generated against the peptide MESSPGERSIHRQSPE - (amino acids 12-27) and against the peptide DKGNSNRGNPGSKPK (amino acids 624-638). Anti-SpiSLC4γ was raised against the peptide SESNYEGDHSHDDSR - (amino acids 25-39) and against the peptide VTEGFKPTQHDKRGW (amino acids 744-758). For each antibody, ten rabbits were initially screened for non-cross reactivity with S. pistillata proteins and two were selected for the Speedy program. Each selected antibody was affinity purified with peptide columns by Eurogentec before use.
Immunolocalization
Apexes of colonies were prepared for immunolocalization as described previously34. Briefly, apexes of S. pistillata were fixed in 3% paraformaldehyde in S22 buffer (450 mM NaCl, 10 mM KCl, 58 mM MgCl2, 10 mM CaCl2, 100 mM Hepes, pH 7.8) at 4 °C overnight and then decalcified, using 0.5 M ethylenediaminetetraacetic acid (EDTA) in Ca-free S22 at 4 °C. They were then dehydrated in an ethanol series and embedded in Paraplast. Cross-sections (6 μm thick) were cut and mounted on silane-coated glass slides. After cutting, deparaffinized sections of tissues were incubated for 1 h in saturating medium (1% BSA, 0.2% teleostean gelatin, 0.05% Tween 20 in Phosphate-buffered saline (PBS) pH 7.4) at 20 °C. Samples were then incubated with the anti-SpiSLC26β or SpiSLC4γ (10 μg/ml) as primary antibody. After rinsing in saturating medium, samples were incubated with biotinylated anti-rabbit antibodies as secondary antibodies. After rinsing with PBS, pH 7.4, samples were finally stained for 15 min with streptavidin AlexaFluor 568 (Molecular probes, 1:50 dilution) and 4’,6-diamidino-2-phenylindole, DAPI (Sigma, 2 μg mL−1). Samples were embedded in Pro-Long antifade medium (Molecular Probes) and observed with confocal laser-scanning microscope (Leica, SP5). Immunostaining experiment controls were performed with pre-immune serum for SpiSLC26β or SpiSLC4γ and then treated with biotinylated anti-rabbit antibodies and streptavidin AlexaFluor 568 as described above.
Additional Information
How to cite this article: Zoccola, D. et al. Bicarbonate transporters in corals point towards a key step in the evolution of cnidarian calcification. Sci. Rep. 5, 9983; doi: 10.1038/srep09983 (2015).
References
Reaka-Kudla, M. in Biodiversity II. Understanding and Protecting Our Natural Resources (eds ML Reaka-Kudla, DE Wilson, & EO Wilson ) 83–108 (Joseph Henry, National Academy Press, 1997).
Weis, V. & Allemand, D. Whar determines Coral Health? Science 324, 1153–1155, 10.1126/science.1172540 (2009).
Weis, V. M., Davy, S. K., Hoegh-Guldberg, O., Rodriguez-Lanetty, M. & Pringle, J. R. Cell biology in model systems as the key to understanding corals. Trends Ecol Evol 23, 369–376, 10.1016/j.tree.2008.03.004 (2008).
Roth, E., Jeon, K. & Stacey, G. in Mol genet of plant-microbe interactions (ed R. & Verma Palacios, D. ) 220–225 (The American Phytopatological Society, 1988).
Furla, P., Galgani, I., Durand, I. & Allemand, D. Sources and mechanisms of inorganic carbon transport for coral calcification and photosynthesis. J Exp Biol 203, 3445–3457 (2000).
Venn, A. A. et al. Imaging intracellular pH in a reef coral and symbiotic anemone. Proc Natl Acad Sci U S A 106, 16574–16579, 10.1073/pnas.0902894106 (2009).
Furla, P. et al. The Symbiotic Anthozoan: A Physiological Chimera between Alga and Animal. Integr Comp Biol 45, 595–604, 10.1093/icb/45.4.595 (2005).
Furla, P., Allemand, D. & Orsenigo, M. N. Involvement of H+-ATPase and carbonic anhydrase in inorganic carbon uptake for endosymbiont photosynthesis. Am J Physiol (Regul Integr Comp) 278, R870–R881 (2000).
Cabantchik, Z. I. & Greger, R. Chemical probes for anion transporters of mammalian cell membranes. Am J Physiol 262, C803–827 (1992).
Erez, J. Vital effect on stable-isotope composition seen in foraminifera and coral skeletons. Nature 273, 199–202, 10.1038/273199a0 (1978).
Tambutté, É., Allemand, D., Mueller, E. & Jaubert, J. A compartmental approach to the mechanism of calcification in hermatypic corals. J Exp Biol 199, 1029–1041 (1996).
Pushkin, A. & Kurtz, I. SLC4 base (HCO3−, CO32−) transporters: classification, function, structure, genetic diseases and knockout models. Am J Physiol Renal Physiol 290, F580–599, 10.1152/ajprenal.00252.2005 (2006).
Parker, M. D. & Boron, W. F. The divergence, actions, roles and relatives of sodium-coupled bicarbonate transporters. Physiol Rev 93, 803–959, 10.1152/physrev.00023.2012 (2013).
Alper, S. L. & Sharma, A. K. The SLC26 gene family of anion transporters and channels. Mol Aspects Med 34, 494–515, 10.1016/j.mam.2012.07.009 (2013).
Cordat, E. & Casey, J. R. Bicarbonate transport in cell physiology and disease. Biochem J 417, 423–439, 10.1042/BJ20081634 (2009).
Park, M., Li, Q., Shcheynikov, N., Zeng, W. & Muallem, S. NaBC1 is a ubiquitous electrogenic Na+ -coupled borate transporter essential for cellular boron homeostasis and cell growth and proliferation. Mol Cell 16, 331–341, 10.1016/j.molcel.2004.09.030 (2004).
Jalimarada, S. S., Ogando, D. G., Vithana, E. N. & Bonanno, J. A. Ion transport function of SLC4A11 in corneal endothelium. Invest Ophthalmol Vis Sci 54, 4330–4340, 10.1167/iovs.13-11929 (2013).
Vilas, G. L. et al. Transmembrane water-flux through SLC4A11: a route defective in genetic corneal diseases. Hum Mol Genet 22, 4579–4590, 10.1093/hmg/ddt307 (2013).
Dorwart, M. R., Shcheynikov, N., Yang, D. & Muallem, S. The solute carrier 26 family of proteins in epithelial ion transport. Physiology (Bethesda) 23, 104–114, 10.1152/physiol.00037.2007 (2008).
Shinzato, C. et al. Using the Acropora digitifera genome to understand coral responses to environmental change. Nature, 10.1038/nature10249 (2011).
Kortschak, R. D., Samuel, G., Saint, R. & Miller, D. J. EST Analysis of the Cnidarian Acropora millepora Reveals Extensive Gene Loss and Rapid Sequence Divergence in the Model Invertebrates. Curr Biol 13, 2190–2195, 10.1016/j.cub.2003.11.030 (2003).
Jin Liew, Y. et al. Identification of microRNAs in the coral Stylophora pistillata. PLoS One 9, e91101, 10.1371/journal.pone.0091101 (2014).
Karako-Lampert, S. et al. Transcriptome analysis of the scleractinian coral Stylophora pistillata. PLoS One 9, e88615, 10.1371/journal.pone.0088615 (2014).
Putnam, N. H. et al. Sea anemone genome reveals ancestral eumetazoan gene repertoire and genomic organization. Science 317, 86–94, 10.1126/science.1139158 (2007).
Lehnert, E. M. et al. Extensive differences in gene expression between symbiotic and aposymbiotic cnidarians. G3 (Bethesda) 4, 277–295, 10.1534/g3.113.009084 (2014).
Alper, S. L. Molecular physiology of SLC4 anion exchangers. Exp Physiol. 91, 153–161, 10.1113/expphysiol.2005.031765 (2006).
Zhu, Q., Lee, D. W. & Casey, J. R. Novel topology in C-terminal region of the human plasma membrane anion exchanger, AE1. J Biol Chem 278, 3112–3120, 10.1074/jbc.M207797200 (2003).
Okubo, K., Kang, D., Hamasaki, N. & Jennings, M. L. Red blood cell band 3. Lysine 539 and lysine 851 react with the same H2DIDS (4,4’-diisothiocyanodihydrostilbene-2,2’-disulfonic acid) molecule. J Biol Chem 269, 1918–1926 (1994).
Aravind, L. & Koonin, E. V. The STAS domain - a link between anion transporters and antisigma-factor antagonists. Curr Biol 10, R53–55, 10.1016/S0960-9822(00)00335-3 (2000).
Marchler-Bauer, A. et al. CDD: a Conserved Domain Database for the functional annotation of proteins. Nucleic Acids Res 39, D225–229, 10.1093/nar/gkq1189 (2011).
Florea, L., Hartzell, G., Zhang, Z., Rubin, G. M. & Miller, W. A computer program for aligning a cDNA sequence with a genomic DNA sequence. Genome Res 8, 967–974, 10.1101/gr.8.9.967 (1998).
Ganot, P. et al. Structural molecular components of Septate Junctions in cnidarians point to the origin of epithelial junctions in Eukaryotes. Mol. Biol. Evol. 32, 44–62 (2015).
Moya, A. et al. Cloning and use of a coral 36B4 gene to study the differential expression of coral genes between light and dark conditions. Mar Biotechnol (NY) 10, 653–663, 10.1007/s10126-008-9101-1 (2008).
Puverel, S. et al. Soluble organic matrix of two Scleractinian corals: partial and comparative analysis. Comp Biochem Physiol B Biochem Mol Biol 141, 480–487, 10.1016/j.cbpc.2005.05.013 (2005).
Tambutte, E. et al. Observations of the tissue-skeleton interface in the scleractinian coral Stylophora pistillata. Coral Reefs 26, 517–529, 10.1007/s00338-007-0263-5 (2007).
Conant, G. C. & Wolfe, K. H. Turning a hobby into a job: how duplicated genes find new functions. Nat Rev Genet 9, 938–950, 10.1038/nrg2482 (2008).
Sterling, D., Reithmeier, R. A. & Casey, J. R. A transport metabolon. Functional interaction of carbonic anhydrase II and chloride/bicarbonate exchangers. J Biol Chem 276, 47886–47894, 10.1074/jbc.M105959200 (2001).
Sterling, D., Alvarez, B. V. & Casey, J. R. The extracellular component of a transport metabolon. Extracellular loop 4 of the human AE1 Cl-/HCO3- exchanger binds carbonic anhydrase IV. J Biol Chem 277, 25239–25246, 10.1074/jbc.M202562200 (2002).
Bertucci, A., Tambutte, S., Supuran, C. T., Allemand, D. & Zoccola, D. A New Coral Carbonic Anhydrase in Stylophora pistillata. Mar Biotechnol (NY) 13, 992–1002, 10.1007/s10126-011-9363-x (2011).
Alper, S. L. Molecular physiology and genetics of Na+-independent SLC4 anion exchangers. J Exp Biol 212, 1672–1683, 10.1242/jeb.029454 (2009).
Allemand, D. et al. Biomineralisation in reef-building corals: from molecular mechanisms to environmental control. CR Palevol 3, 453–467, 10.1016/j.crpv.2004.07.011 (2004).
Tambutte, S. et al. Coral biomineralization: From the gene to the environment. J Exp Mar Biol Ecol 408, 58–78, 10.1016/j.jembe.2011.07.026 (2011).
Zoccola, D. et al. Molecular cloning and localization of a PMCA P-type calcium ATPase from the coral Stylophora pistillata. Biochim Biophys Acta 1663, 117–126, 10.1016/j.bbamem.2004.02.010 (2004).
Venn, A., Tambutte, E., Holcomb, M., Allemand, D. & Tambutte, S. Live tissue imaging shows reef corals elevate pH under their calcifying tissue relative to seawater. PLoS One 6, e20013, 10.1371/journal.pone.0020013 (2011).
Brown, C. D., Dunk, C. R. & Turnberg, L. A. Cl-HCO3 exchange and anion conductance in rat duodenal apical membrane vesicles. Am J Physiol 257, G661–667 (1989).
Shandro, H. J. & Casey, J. R. in Adv Mol Cell Biol Vol. 38 (eds E. Edward Bittar & Pusch Michael ) 279–328 (Elsevier, 2006).
Hughes, T. P. et al. Climate Change, Human Impacts and the Resilience of Coral Reefs. Science 301, 929–933, 10.1126/science.1085046 (2003).
Doney, S. C., Fabry, V. J., Feely, R. A. & Kleypas, J. A. Ocean acidification: the other CO2 problem. Ann Rev Mar Sci 1, 169–192, 10.1146/annurev.marine.010908.163834 (2009).
Houlbrèque, F. et al. Effects of increased pCO2 on zinc uptake and calcification in the tropical coral Stylophora pistillata. Coral Reefs 31, 101–109, 10.1007/s00338-011-0819-2 (2012).
Kaniewska, P. et al. Major cellular and physiological impacts of ocean acidification on a reef building coral. PLoS One 7, e34659, 10.1371/journal.pone.0034659 (2012).
Marubini, F., Ferrier-Pagès, C., Furla, P. & Allemand, D. Coral calcification responds to seawater acidification: a working hypothesis towards a physiological mechanism. Coral Reefs 27, 491–499, 10.1007/s00338-008-0375-6 (2008).
Ries, J. B., Cohen, A. L. & McCorkle, D. C. Marine calcifiers exhibit mixed responses to CO2-induced ocean acidification. Geol Soc Am, 1131–1134, 10.1130/G30210A.1 (2009).
Vidal-Dupiol, J. et al. Genes related to ion-transport and energy production are upregulated in response to CO2-driven pH decrease in corals: new insights from transcriptome analysis. PLoS One 8, e58652, 10.1371/journal.pone.0058652 (2013).
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–1067, 10.1093/bioinformatics/btm071 (2007).
Kall, L., Krogh, A. & Sonnhammer, E. L. Advantages of combined transmembrane topology and signal peptide prediction-the Phobius web server. Nucleic Acids Res 35, W429–432, 10.1093/nar/gkm256 (2007).
Corpet, F. Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res 16, 10881–10890, 10.1093/nar/16.22.10881 (1988).
Guindon, S. & Gascuel, O. A simple, fast and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 52, 696–704, 10.1080/10635150390235520 (2003).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27, 1164–1165, 10.1093/bioinformatics/btr088 (2011).
Huelsenbeck, J. P. & Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17, 754–755, 10.1093/bioinformatics/17.8.754 (2001).
Bertucci, A. et al. Carbonic anhydrases in anthozoan corals-A review. Bioorg Med Chem 21, 1437–1450, 10.1016/j.bmc.2012.10.024 (2013).
Acknowledgements
Thanks are due to Dominique Desgré for coral maintenance. This study was conducted as part of the Centre Scientifique de Monaco research program, supported by the Government of the Principality of Monaco. This project was partially funded by KAUST baseline funds to CRV and MA.
Author information
Authors and Affiliations
Contributions
DZ, PG, AB, ST designed and conceived the experiments. DA, ST and ET contributed reagents and materials. DZ, PG, AB, NCS and NT generated data. DZ, PG, AB, CRV, MA, DA, JRC and ST analyzed data and wrote the manuscript. All authors read and approved the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Zoccola, D., Ganot, P., Bertucci, A. et al. Bicarbonate transporters in corals point towards a key step in the evolution of cnidarian calcification. Sci Rep 5, 9983 (2015). https://doi.org/10.1038/srep09983
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep09983
This article is cited by
-
Shedding light on starvation in darkness in the plastid-bearing sea slug Elysia viridis (Montagu, 1804)
Marine Biology (2023)
-
Effects of light intensity and spectral composition on the growth and physiological adaptation of Acroporid corals
Coral Reefs (2023)
-
Differences in carbonate chemistry up-regulation of long-lived reef-building corals
Scientific Reports (2023)
-
Investigating calcification-related candidates in a non-symbiotic scleractinian coral, Tubastraea spp.
Scientific Reports (2022)
-
Coral calcification mechanisms in a warming ocean and the interactive effects of temperature and light
Communications Earth & Environment (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.