Abstract
G-protein coupled receptor 43 (GPR43) recognizes short chain fatty acids and is implicated in obesity, colitis, asthma and arthritis. Here, we present the first full characterization of the GPR43 promoter and 5′-UTR. 5′-RACE of the GPR43 transcript identified the transcription start site (TSS) and a 124 bp 5′-UTR followed by a 1335 bp intron upstream of the ATG start codon. The sequence spanning -4560 to +68 bp relative to the GPR43 TSS was found to contain strong promoter activity, increasing luciferase reporter expression by >100-fold in U937 monocytes. Stepwise deletions further narrowed the putative GPR43 promoter (−451 to +68). Site-directed mutagenesis identified XBP1 as a core cis element, the mutation of which abrogated transcriptional activity. Mutations of predicted CREB, CHOP, NFAT and STAT5 binding sites, partially reduced promoter activity. ChIP assays confirmed the binding of XBP1 to the endogenous GPR43 promoter. Consistently, GPR43 expression is reduced in monocytes upon siRNA-knockdown of XBP1, while A549 cells overexpressing XBP1 displayed elevated GPR43 levels. Based on its ability to activate XBP1, we predicted and confirmed that TNFα induces GPR43 expression in human monocytes. Altogether, our findings form the basis for strategic modulation of GPR43 expression, with a view to regulate GPR43-associated diseases.
Similar content being viewed by others
Introduction
Studies on knockout mice have identified Free Fatty Acid Receptor 2 (FFAR2 or GPR43) as a critical gene in the prevention of obesity, colitis, asthma and arthritis1,2,3,4,5,6,7,8. GPR43 is a G protein coupled-receptor that is activated by mid-micromolar concentrations of short-chain fatty acids (SCFAs) - namely acetate, propionate and butyrate. While the liver metabolism of ethanol can generate micromolar concentrations of acetate in the blood, by far the most abundant source of SCFAs in the human body is the colonic lumen, where hundreds of millimolars are continuously being produced during the anaerobic fermentation of dietary fibre by saccarolytic gut bacteria9,10. These gut SCFAs have been found to beneficially modulate blood glucose and lipid levels, the colonic environment and immune functions11,12,13. As an SCFA receptor, GPR43 has already been shown to mediate some of these beneficial effects, with knockout mice studies confirming a role in obesity and inflammation (Table 1).
Consistent with its role as an SCFA receptor, GPR43 expression is found in cells that are exposed to the highest concentrations of SCFAs. These include the cells of the distal ileum, colon and adipose tissue, with the highest expression found in immune cells such as monocytes and neutrophils14,15,16. In addition, GPR43 expression appears to be modulated during inflammation since immune challenge by lipopolysaccharide (LPS) or treatment with granulocyte-macrophage colony stimulating factor (GM-CSF) raises GPR43 transcript levels in human monocytes17. This tissue-specificity suggests that GPR43 expression is tightly regulated and may be important for its function. Indeed, the compelling outcomes of Gpr43 knockout (Table 1) imply that proper regulation of GPR43 expression is pertinent to the normal functioning of a range of physiological processes and consequently, targeting of GPR43 expression in diseases would provide new therapeutic potential. Thus, details on the factors involved in cell-type specific expression of the GPR43 gene under pathophysiological conditions remain an unexplored and intriguing area of investigation. Here, we characterized the human GPR43 gene, identifying the promoter and enhancer sequence elements, as well as the critical transcription factors and signalling pathways that regulate GPR43 expression.
Results
PMA-differentiated U937 monocytes are a suitable model for GPR43 expression
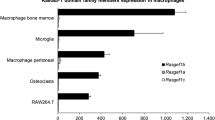
To understand the mechanisms underlying the specific expression of GPR43, we first confirmed the cell type with the highest level of GPR43 mRNA. Consistent with previous studies14,15,16, we found human peripheral blood monocytes and neutrophils to express the highest level of GPR43 mRNA (Fig. 1). PMA-mediated monocytic differentiation of the human promonocytic cell line, U93718,19, led to a 100-fold increase in the transcription of GPR43, yielding mRNA levels that was comparable to the peripheral blood monocytes and neutrophils. Hence, we reasoned that the U937 cell line would be a suitable model to study GPR43 transcriptional regulation in leukocytes.
GPR43 transcription start site is located at 1459 bp upstream of the ATG start codon
To define the GPR43 putative promoter site, we first performed a 5′-RACE of the GPR43 transcript, generating an approximately 450 bp product which was sequenced to reveal a 124 bp 5′-UTR upstream of the GPR43 ATG start codon (Fig. 2a). Mapping of this sequence with the human genome database (NCBI Ref Seq: NC_000019.10) revealed the presence of a 1335 bp intron flanked by the GPR43 5′-UTR and the start codon (Fig. 2b). Thus, the transcription start site (TSS) is located 1459 bp upstream of the ATG start codon. While it was difficult to judge the presence of other 5′-RACE products (Fig. 2a, Lane 1), we note that a previously reported northern blot analysis (Senga et al., 200317, Fig. 2h), detected two GPR43 transcripts, with the shorter (by about a few hundred bases) product being many times more abundant than the longer transcript. It is possible that the two transcript sizes described by Senga et al., 200317 are due to different 5′-UTR lengths, which can possibly result from alternate splicing20 or promoters21. The transcript detected by the 5′ RACE here is likely the shorter, more abundant, transcript. Our attempts to detect the presence of other possible 5′-UTR species via RACE were thus far unsuccessful (data not shown), presumably due to the low abundance. Overall, these results characterize the 5′ region of the human GPR43 gene and allow the putative core promoter to be identified for subsequent analysis of transcriptional elements.
The GPR43 transcription start site is mapped to 1459 bp upstream of the ATG start codon.
(a) 5′-RACE of GPR43 transcripts from U937 cells. Total RNA from U937 cells was extracted and full-length GPR43 mRNA was reverse transcribed using a gene specific primer complementary to its open reading frame. The resulting cDNA was ligated to a known oligonucleotide sequence at the 5′ end. Nested PCR using primers flanking the 5′ end was then performed to procure an approximately 450 bp product encompassing the 5′ untranslated region (5′-UTR) and part of the GPR43 coding sequence (lane 1). As positive controls, amplification of 246 bp of the GPR43 coding region (lane 2), as well as a 900 bp-long 5′ end of the β-actin gene (lane 3) were performed. (b) Schematic mapping the GPR43 5′-UTR region and protein coding region on the human chromosome 19. The PCR product from (a, lane 1) was sequenced to reveal a 124 bp long 5′-UTR exon upstream of the GPR43 ATG start codon.
The core and proximal promoter of GPR43 are located within −451 to −33 from the TSS
Upon identification of the TSS, we cloned a putative promoter region (spanning −4560 to +68 bp relative to the TSS) from primary human monocyte genomic DNA and analyzed its promoter activity using dual luciferase reporter system. This putative promoter contained aligned sequences with greater than 70% sequence identity to the corresponding mouse sequence (Fig. 3a), suggesting conservation with involvement in transcriptional regulation22. Indeed, compared to the vector alone, the promoter insert raised the luciferase reporter activity by >100-fold (Fig. 3b). Deletion from positions −4560 to −451 revealed no significant change (p-values > 0.05) in luciferase activity, implying that this region lacks regulatory elements, which is consistent with the general lack of sequence conservation observed (Fig. 3a). Although two short sequence alignments do appear in this region, we note that the corresponding mouse sequences are much further away from the mouse Gpr43 gene (0.6 Mbp and 1.7 Mbp, respectively), suggesting a lack of involvement in mouse Gpr43 transcriptional control. Deletion from positions −451 to −182 resulted in a marked drop (p-value = 0.07) in activity while further deletions downstream of position −182 resulted in significant reductions in activity (p values < 0.005), suggesting that the region spanning −451 to −58 might contain crucial activator elements. This is consistent with the observed interspecies sequence conservation (Fig. 3a). Strikingly, the near abrogation of promoter activity upon deletion of −58 to −33 suggests that the core promoter element(s) lies within this 25 bp region.
Deletions define a 519-bp putative GPR43 promoter containing the core and proximal promoter.
(a) ECR Browser plot spanning −4560 bp upstream of the transcription start site (+1), the entire GPR43 gene and 2000 bp after the gene. The plot represents sequence homology of the mouse chromosome 7 (Ref Seq ID: NC_000073.6 ) region flanking the murine Gpr43 (Entrez Gene ID: 233079) in comparison to the corresponding region on the human chromosome 19 (RefSeq ID: NC_000019.9; positions: 35934599–35942669), which is acting as the baseline. Sequence lengths of >100 bp with 70% sequence conservation are shown as peaks. (b) Luciferase reporter activities of 5′ deletion constructs from a 4628 bp putative GPR43 promoter in differentiated U937 cells 22 h after transfection. Analysis of the homology with mouse sequence was used to approximate the deletion sites. Results represent the average Firefly luciferase read-outs of three independent transfections (n = 3) normalized to Renilla luciferase activity and relative to the basic (empty) luciferase vector, arbitrarily set as 1. Error bars represent the mean ± s.d.. The data shown are representative of three independent experiments. Two tailed Students' T-test was used to determine the statistical significance of the difference between promoter constructs and is annotated as: * <0.05, ** <0.01 and *** <0.001.
XBP1 is part of the core promoter while CREB, CHOP, NFAT and STAT5 act as enhancers
By in silico predictions using the MatInspector program23, we identified 8 transcription factor (TF) recognition sites with high matrix similarity of ≥90% within the 519 bp putative promoter (Fig. 4a). The null mutations of Tax/CREB, CHOP, NFAT and STAT5/5B resulted in significant losses in activity compared to the wild type (WT) promoter (Fig. 4b), indicating that these TFs may be important enhancers for the control of GPR43 expression. Conversely, no significant loss in activity was found for mutations in the IRF4, PU.1 and c-Rel recognition sites. Mutation of the XBP1 recognition site resulted in near abrogation of promoter activity (Fig. 4b), hence identifying it as a crucial core cis element. Notably, this XBP1 recognition site resides within the 25 bp region that was deleted from the 126 bp promoter (Fig. 3b). XBP1 binding also appears to be required for the low level GPR43 expression observed in A549 adenocarcinomic human alveolar basal epithelial cells (Supplementary Fig. S1) since the XBP1 null mutation also led to near abrogation in promoter activity in the A549 cells.
In silico predictions, null mutations of the 519 bp promoter region and signalling pathway modulation identify transcription factors (TFs) and pathways regulating GPR43 expression.
(a) Predicted TF binding sites by MatInspector software with corresponding WT and mutant sequences and their relative positions indicated above each TF binding site. The core recognition sequences are underlined while mutated bases are indicated in red. (b) Relative luciferase reporter activities of 519-bp promoter mutated at indicated sites were measured in differentiated U937 monocytes. The multiple binding sites of PU.1 or NFAT residing within the 519 bp promoter were simultaneously mutated in their respective constructs. Wild type (WT) promoter is arbitrarily set to 1 luciferase unit. (c) Quantitative PCR analysis of GPR43 mRNA levels upon modulation of signalling pathways after 1 h pre-treatment with activators/inhibitors followed by immune challenge for 3 h with 100 ng/mL LPS. (d) Quantitative PCR analysis of GPR43 mRNA levels upon 4 h treatment of monocytes with inhibitor/activator or 12 h treatment with inhibitor. (c) and (d) Inhibitor/Activator + (Targeted signalling proteins) are shown: SB203580, 10 μM —| p38; U73122, 5 μM —| phospholipase C (PLC); MG132, 10 μM —| Proteasome; Wortmannin, 2 μM —| PI3kinase (PI3K); BisI, 4 μM —| protein kinase C (PKC); Forskolin, 20 μM → Adenylyl cyclase (AC). Expression levels of two reference genes, beta-2-microglobulin (B2M) and cyclophilin B (CYPB) were also measured and presented as Supplementary Figure S2. All measurements were standardized to the RPL27 as the reference gene. Experiments were performed in triplicate transfections or treatments, where error bars represent the mean ± s.d. In (b), Dunnett's test (α = 0.05) was performed while in (c) and (d), the two tailed Students' T-test was used to determine statistical significance. p-value * <0.05, ** <0.01 and *** <0.001. The data shown are representative of three independent experiments.
The activities of p38 and PLC are required for GPR43 transcription
A number of the identified TFs are known to be regulated by MAPK and PLC/PKC pathways24,25,26. To determine if these pathways are involved in the modulation of GPR43 expression, we blocked the putative pathways with small molecule inhibitors (Fig. 4c, 4d). The relative expression of housekeeping genes was not markedly affected (Supp. Fig. 2). Both SB203580 and U73122, inhibitors of p38 and PLC respectively, attenuated GPR43 transcription in LPS-challenged monocytes while only SB203580 attenuated GPR43 transcription in unchallenged monocytes (Fig. 4c, 4d). This suggests that the p38 pathway is required for basal transcription of GPR43 which is active with or without LPS-challenge while the PLC-PKC pathway is important only during LPS-mediated up-regulation of GPR43. This may be due to p38 pathway activation of XBP126, which is part of the core promoter regulating basal transcription (Fig. 4b); while the PLC-PKC pathway activates NFAT25, which we find to act as an enhancer (Fig. 4b). The p38 pathway also activates CHOP24. Separately, the proteasome inhibitor, MG132, reduced GPR43 mRNA levels with or without LPS treatment while Wortmannin, an AKT inhibitor, increased GPR43 transcript levels. Overall, our findings indicate that activation of the GPR43 expression is mediated by the p38 and PLC/PKC pathways and transcriptional regulation may possibly also involve the proteasome degradation and PI3/Akt pathways.
ChIP analysis confirmed XBP1 binding to GPR43 promoter in vivo
Since the XBP1 binding sites were found to be necessary for promoter activity, we investigated XBP1 binding to the endogenous GPR43 promoter. Chromatin immunoprecipitation (ChIP) was performed with antibodies against XBP1. The GPR43 promoter XBP1 binding site was significantly enriched in the resulting immunoprecipitate, consistent with strong association of XBP1 to that site (Fig. 5). The pull down appears specific as no significant enrichment was detected for the negative control genomic regions and no GPR43 promoter enrichment was observed when the isotype control antibody was used.
XBP1 binds to the GPR43 promoter in vivo.
(a) Schematic showing the relative regions of GPR43 promoter amplified by quantitative PCR following ChIP assay with anti-XBP1 antibody. P1–P6 spans a region of ~1200 bp surrounding the XBP1 binding site on the GPR43 promoter. Only promoter regions, P3 and P4 contain the XBP1 binding site. The GPR43 enhancer region and coding sequence, located more than 3 kb and 2.5 kb up- and downstream of the XBP1 binding site respectively, were also amplified as negative controls. (b) Quantitative PCR analysis of GPR43 promoter regions following ChIP assay on U937 cells using anti-XBP1 antibody or IgG isotype control antibody. Results are expressed as fold enrichment relative to GPR43 coding sequence (GPR43_CDS) negative control region and error bars represent the mean ± s.d. of three independent pull-down. p-value * <0.05, ** <0.01 and *** <0.001. The data shown are representative of two independent experiments.
XBP1-siRNA knockdown or overexpression alters GPR43 expression
Next, we sought to confirm that XBP1 activity is required for the endogenous transcription of GPR43. Stable knockdown of XBP1 in U937 cells led to a reduction in GPR43 expression levels relative to the control siRNA (Fig. 6a, b). Conversely, in the A549 adenocarcinomic human alveolar basal epithelial cell line, which expresses low levels of XBP1-dependent GPR43 mRNA (Fig. 1 and Supplementary Fig. S1), the overexpression of the active spliced form of XBP1 (XBP1s) raised GPR43 expression levels (Fig. 6c, d). Thus, we have confirmed that XBP1 modulates endogenous GPR43 expression.
XBP1 is critical for the expression of GPR43.
(a) Quantitative PCR analysis of GPR43 and spliced XBP1 (XBP1s) mRNA in XBP1-stable knockdown U937 cells relative to the negative control (non-targeting) siRNA expression plasmid, arbitrarily set as 1. (b) Western blot analysis of unspliced XBP1 (XBP1u) protein levels in XBP1-stable knockdown U937 cells. XBP1s levels were undetectable. (c) Quantitative PCR analysis of GPR43 and XBP1 mRNA after 48 h overexpression of XBP1s in A549 cells, relative to the pcDNA empty vector which is set arbitrarily as 1. (d) Western blot analysis of XBP1s protein levels after 48 h overexpression of XBP1s in A549 cells. (b) and (c) Measurements were standardized to RPL27 as the reference gene. (b) to (d) Results shown are the average of three independent transductions (for stable knockdown) or transfections (for overexpression), with error bars representing the mean ± s.d.. Two tailed Students' T-test was used to calculate statistical significance. p-value * <0.05, *** <0.001.
GPR43 expression is up-regulated by XBP1 activators
The XBP1-mediated transactivation of GPR43 prompted us to examine whether GPR43 expression is increased by known activators of XBP1. LPS is a known activator of XBP127 while GM-CSF is a known positive regulator of dendritic cell development28, a process which requires XBP129. We found that LPS treatment increased the GPR43 transcription by at least 7-fold while GM-CSF raised transcription by 5-fold (Fig. 7a). Both stimuli were previously reported to induce GPR43 expression via an unknown mechanism17. However, our results now revealed this to be attributable to XBP1 activation. Besides LPS, treatment with TNFα, another known activator of XBP130, also up-regulated GPR43 transcription by 5-fold. Thus, we have demonstrated that pathways leading to the activation of XBP1 consistently up-regulate GPR43 transcription.
Human monocyte GPR43 expression is up-regulated by LPS, TNFα and GM-CSF treatment.
(a) Quantitative PCR analysis of GPR43 mRNA and (b) spliced XBP1 (XBP1s) mRNA levels under 3 h treatment with either 100 ng/mL LPS; 10 ng/mL TNFα; 100 ng/mL GM-CSF or 10 mM Acetate (Ac). (a) and (b) The results represent average fold change of the treated samples relative to the non-treated control (NT) and error bars represent the mean ± s.d. of three independent treatment wells. Measurements were standardized to RPL27 as the reference gene. Two tailed Students' T-test was used to determine the statistical significance of the difference between treated and NT samples. p-value * <0.05, ** <0.01 and *** <0.001. The data shown are representative of at least two independent experiments.
Discussion
In this study, we identified and characterized the GPR43 promoter, revealing that XBP1 acts as a core cis promoter element while CREB, CHOP, NFAT and STAT5 act as enhancers (Fig. 2, 4a, 4b). Based on its known ability to activate XBP1, we accurately predicted that TNFα up-regulates GPR43 expression (Fig. 7b). This suggests that other regulators of XBP1 activity may likewise be involved in the regulation of GPR43 expression. We also showed that GPR43 expression was altered by inhibition of pathways involved in the activation of CHOP and NFAT (Fig. 4c, 4d). Our identification of the pathways known to activate these GPR43 promoter elements would facilitate the application of stimuli and conditions to alter GPR43 expression and its subsequent functions, with a view to therapeutic developments.
The indispensability of XBP1 for GPR43 promoter activity has important implications since XBP1 is involved in a number of physiological processes and diseases which include B cell development and the unfolded protein response (UPR)31. It now appears likely that these physiological processes may engage the expression and function of GPR43. Notably, mutations in the XBP1 gene locus of human patients has been associated with increased risk of inflammatory bowel disease (IBD), Crohn's disease and ulcerative colitis, while Xbp1−/− mice have increased susceptibility to colitis32. Xbp1−/− mice also display impaired glucose and insulin tolerance upon high fat diet-induced obesity33. The association of XBP1 with gut inflammation and obesity may be partially attributable to the disrupted expression of GPR43, since Gpr43−/− mice are similarly more susceptible to colitis1,2,3,4 as well as exhibiting impaired glucose tolerance and increased obesity7,8. Considering the implications of XBP1 and GPR43 in inflammation and obesity, it might be pertinent to further investigate this hypothetical link.
Besides XBP1, the GPR43 promoter was also found to be up-regulated by Tax/CREB, CHOP, NFAT and STAT5 (Fig. 4). Tax/CREB is a heterodimer consisting of viral oncoprotein, Tax and the host derived CREB34. NFAT and STAT5 have been extensively implicated in immunity and cancer, through regulating cell development, growth and cytokine production35,36,37,38. CHOP plays a role in ER stress-induced apoptosis and cytokine production39. However, in comparison with XBP1, mutations of the promoter binding sites for the aforementioned four TFs only resulted in relatively modest decrease of ~30–40% in the promoter activity (Fig. 4b). It is hence likely that these TFs are involved in fine-tuning the expression levels of GPR43 in monocytes, or, they may be more active in up-regulating the gene under the appropriate external stimuli (e.g. cytokine stimulation, ER stress or viral infection). Following the same line of argument, we cannot rule out the potential involvement of other predicted transcription factors, e.g. IRF4, c-Rel (component of NF-κB) and Pu.1 (Fig. 4b), in the expression of GPR43 under the appropriate immune contexts. This will be a subject of future investigation.
The ability of inflammatory stimuli such as LPS, TNFα and GM-CSF (Fig. 7a) to up-regulate GPR43 expression infers a number of interesting possibilities. Our findings imply that the positive correlation between GPR43 expression and inflammation at the fetal cell membranes40 or TNFα levels in adipocytes41, is likely due to the up-regulation of GPR43 by the inflammatory stimuli present. Notably, GM-CSF has been found to be beneficial for the treatment of IBD42,43. Since GPR43 is also implicated in the etiology of IBD1,2,3,4, it would be interesting to investigate whether GM-CSF exerts any beneficial therapeutic effects through modulating GPR43 expression. In view of the low plasma levels of SCFA (<200 μM)9,10 and low potencies of the ligands in activating GPR43 (generally in the mid micromolar range)14,15,16, the higher levels of expression of GPR43 in the peripheral blood monocytes might thus enhance the potency of SCFAs on the GPR43-mediated inflammation.
The increase in GPR43 expression observed in phosphoinositide-3 kinase (PI3K) inhibition under basal or immune-challenged conditions (Fig. 4c, 4d) suggests that there was a relief of suppression that is mediated by the PI3K/Akt pathway. However, the current information is insufficient to deduce whether this is a direct or indirect effect from the PI3K/Akt signalling. Nevertheless, it should be noted that activation of the PI3K/Akt pathway may lead to the inhibition of p3844. Hence, the increase may be due to the recovery of p38, causing an indirect activation of GPR43 expression.
In addition to characterizing the promoter, we provide the first description of the 5′-UTR of the human GPR43 gene, identifying an intron between the 5′-UTR and the start codon. Our findings present a new perspective on the human GPR43 gene, which was initially described as being intronless, downstream of the ATG start codon45. Notably, this gene organization is also seen in the bovine GPR43 gene, the only other GPR43 homolog which has its 5′-UTR sequenced46. Such cross-species similarities in the gene organization and high sequence conservation of the entire core promoter across human and mouse (Fig. 2a) suggests that the same regulatory elements are conserved in the bovine and murine promoters. Indeed, bovine and murine tissue specific expression of GPR43 have been found to be highly similar to that of the human counterpart14,15,16,46. A recent study reported transcriptional activity from a luciferase assay of a 500 bp sequence immediately upstream of the human GPR43 start codon in mouse RAW264.7 macrophages47. Interestingly, these researchers predicted an NF-κB binding site within this 500-bp sequence, although the mutation was reported to cause only 10% reduction in the promoter activity. Perhaps, inclusion of the GPR43 TSS (located more than 1 kb upstream of the predicted NF-κB binding site as reported by our study) would offer a more complete analysis of the effect of this putative enhancer.
While the findings on the GPR43 promoter are mostly obtained from human monocytes in this study, it would be interesting to explore if the promoter elements identified, regulate GPR43 in other cell types. Our findings on A549 cells suggest that this may indeed be the case. In A549 cells, the 519 bp GPR43 promoter construct up-regulates luciferase expression; an up-regulation that is abrogated by XBP1 null mutation (Supp. Fig. S1). These findings mirror those of U937 cells (Fig. 4b) although differences are also noted. Notably, while a 50% knockdown of XBP1 mRNA led to an 80% drop in expression of GPR43 in the U937 cell, a 100-fold increase in XBP1 mRNA only resulted in a 2-fold increase in A549 expression of GPR43. This discrepancy in effect may be due to a variety of factors. XBP1 activity in A549 may already be near saturation and hence further increases in XBP1 expression will not proportionally increase GPR43 expression. Another possibility is that XBP1 may require further activation, such as by the p38 pathway26, to become fully activated. Such pathways may be more active in monocytes. The GPR43 promoter in A549 cells may also be epigenetically silenced through DNA methylation or chromatin remodeling, such that further increases in XBP1 expression may not fully overcome this silencing. Despite these differences, our findings confirm that XBP1 can serve as a regulator involved in controlling GPR43 expression in other cell types. Notably, both GPR4315,41,48 and XBP149 are found to be up-regulated in adipocytes and adipocyte-derived cell lines, suggesting that XBP1 may also be involved in regulating the expression of GPR43 in adipocytes, an interaction that may have implications in adipocyte function.
In conclusion, we present the first full characterization of the human GPR43 promoter, revealing that GPR43 expression is regulated by XBP1 as a core cis element while CREB, CHOP, NFAT and STAT5 act as enhancers. We show that by distinguishing pathways known to activate these GPR43 promoter elements, it is possible to predict stimuli and conditions that may alter GPR43 expression and function, thus providing novel drug targets for the treatment of GPR43-associated diseases such as obesity, colitis, asthma and arthritis.
Methods
Isolation of peripheral blood monocytes, cell culture and differentiation of U937 cells
Peripheral blood mononuclear cells were isolated from the buffy coat of healthy adult donors (National University of Singapore Blood Donation Centre). Ficoll-Paque PREMIUM (GE Healthcare) gradient centrifugation was performed according to the manufacturer's instructions. Briefly, the mononuclear cell layer was isolated and washed with PBS supplemented with 2% FBS (GE healthcare) and 1 mM EDTA, to remove platelets. Monocytes were subsequently purified by negative selection using the Human Monocyte Enrichment Kit (StemCell Technologies) according to the manufacturer's instructions.
Primary monocytes and U937 cell line were cultured in RPMI (Life Technologies) while A549 cells were cultured in DMEM (Life Technologies) supplemented with 10% FBS and 1% (v/v) penicillin and streptomycin (Life Technologies). The cells were grown at 37°C and 5% CO2. For differentiation of U937 into monocytes, 5 × 105 cells/mL of pre-differentiated cells were induced with 30 ng/mL phorbol 12-myristate 13-acetate (PMA) (Sigma-Aldrich) for 24 h, before changing to fresh media. The cells were cultured for another 48 h to allow for full differentiation and thereafter used for downstream assays.
The primary neonatal human fibroblasts (Life Technologies) were routinely grown in medium 106 (Life Technologies) before RNA extraction.
5′-Rapid Amplification of cDNA Ends (5′ RACE)
Amplification of 5′ cDNA ends of mature GPR43 was performed with the GeneRacer™ kit (Life Technologies, Cat. #L1-502-02) following manufacturer's instructions. Briefly, 5 μg of U937 total RNA was dephosphorylated with Calf Intestinal Phosphatase (CIP) and ligated with a sequence specified RNA oligonucleotide at 37°C for 1 h. Following ligation, the 5′ ends of GPR43 mRNA were reverse transcribed and amplified using nested PCR primers (Supplementary Table S1). The PCR product was then analyzed on a 1.5% agarose gel. The sequence of the GPR43 5′ end was confirmed using Big Dye Terminator cycle sequencing kit and ABI 3100 Genetic Analyser (Life Technologies).
Construction of GPR43 promoter and deletion mutants
To generate promoter deletion constructs, primary monocyte genomic DNA was extracted from the interphase and phenol layer using TRIzol reagent (Life Technologies), according to the manufacturer's instructions. To obtain 5′ end promoter deletion products, primers (Supplementary Table S1) flanking the desired promoter regions were used for PCR amplification, using the purified genomic DNA as template. PCR was performed using iProof High Fidelity DNA Polymerase (Bio-rad) under the following parameters: initial denaturation at 98°C; 40 cycles of amplification, denaturation at 98°C for 10 s, annealing at Tm of primer pair +3°C for 30 s, elongation at 72°C for 2.5 min; followed by a final extension at 72°C.
The isolated promoter lengths were separately cloned into pGL4.20 vector (Promega) using standard molecular cloning techniques. The restriction enzymes used were XhoI and HindIII (Thermo Fisher Scientific). T4 DNA ligase was from Roche. For small-scale purification of plasmids, AxyPrep Plasmid Miniprep kit (Axygen Biosciences) was used. Large-scale purification of plasmids intended for transfection was carried out using PureLink HiPure Plasmid Filter Purification kit (Invitrogen). The full-length sequence of each promoter construct was confirmed by sequencing using Big Dye Terminator cycle sequencing kit and ABI Prism 3100 Genetic Analyzer (Life Technologies).
In silico predictions of transcription factor binding sites and site-directed mutagenesis
Transcription factor binding sites (TFBS) along the delineated promoter regions were identified using the MatInspector programme available online at http://www.genomatix.de/23. Analysis of the sequences was performed using the MatInspector TFBS, weight matrix library version 9.0. Only predicted TFBS with a core matrix similarity of ≥0.95 and an overall matrix similarity of ≥0.90, were considered significant.
Promoters containing the specific mutated TFBS were synthesized by Integrated DNA technologies. The mutated promoters were cloned into pGL4.20 vector (Promega) and confirmed by sequencing as described above.
Transient transfection and luciferase reporter assay
The full-length promoter, deletion and mutant constructs were transiently transfected into differentiated U937 cells at 48 h after removal of PMA, with X-tremeGENE HP DNA Transfection Reagent (Roche) according to the manufacturer's guidelines. Promoter-pGL4.20 vector constructs and pRL-CMV control Renilla luciferase vector (Promega) were transfected in the molar ratio of 10:1.
Renilla luciferase reporter activities were assessed using the Dual Luciferase Reporter Assay System (Promega) 22 h after transfection. Luminescence was detected using the Glomax 20/20 Luminometer (Promega) and the resulting measurements from the Firefly luciferase were normalized to the Renilla luciferase. Relative light units were calculated using readouts from the pGL4.20-Basic (promoter-less) vector or the wild type promoter as baseline for deletion and mutation constructs, respectively.
Treatment of cells with inflammatory stimuli and pathway activation/inhibition
For stimulation of cells, Escherichia coli 055:B5 LPS and sodium acetate were purchased from Sigma-Aldrich while GM-CSF and TNFα were from Life Technologies. Purified human monocytes were seeded at a density of 4 × 106 cells/mL in 24-well plates and cultured in the presence of either LPS (100 ng/mL), PMA (30 ng/mL), TNFα (10 ng/mL), GM-CSF (100 ng/mL) or sodium acetate (10 mM) for 3 h before RNA extraction with TRIzol reagent (Life Technologies).
For modulation of signalling pathways, p38 inhibitor SB203580, PKC inhibitor Bisindolylmaleimide I (BisI) (all from Cell Signalling Technology), PLC inhibitor U73122, adenylate cyclase activator Forskolin (Tocris Biosciences), proteasome inhibitor MG132 and PI3K inhibitor Wortmannin (Sigma) were obtained.
Purified human monocytes were pre-treated for 1 h in either 20 μM DMSO (Merck) as vehicle control, SB203580 (10 μM), U73122 (5 μM), MG132 (10 μM), Wortmannin (2 μM) or BisI (4 μM). Cells were then challenged in 100 ng/mL LPS for 3 h before mRNA was extracted for analysis.
To identify regulatory pathways involved in the basal regulation of gene expression, the same conditions above were applied except that replicate wells of cells treated with MG132 (10 μM) and Forskolin (20 μM) were lysed and analyzed for mRNA expression after 4 h without LPS treatment. For treatment with other inhibitors, monocytes were lysed for RNA extraction and analyzed after 12 h in culture.
Quantitative PCR (qPCR)
Total RNA was isolated using the TRIzol reagent according to the manufacturer's instructions and the procedure was repeated to ensure thorough removal of genomic DNA. cDNA synthesis of the extracted RNA was performed using Superscript III First Strand Synthesis kit (Life Technologies). Quantitative PCR was performed using GoTaq qPCR Master Mix (Promega) and forward and reverse primers (Supplementary Table S1) with LightCycler 480 system (Roche). Spliced XBP1 was detected using the Taqman gene expression assay (Life Technologies, Hs03929085_g1) and Light Cycler Probes Master (Roche). To obtain relative target mRNA folds, cycle thresholds (Ct) were normalized to the Ct of Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) or Ribosomal protein L27 (RPL27) reference genes and expressed as fold-change by the 2−▵▵Ct method50.
Western Blotting
Cell lysates were separated by SDS-PAGE and transferred onto PVDF membranes (Bio-rad). The blots were then probed with antibodies against XBP1 (Abcam, ab37152), or β-actin (Sigma, a2066) followed by the corresponding secondary antibodies according to respective manufacturer's instructions. Following incubation with WesternBright ECL chemiluminescent substrate (Advansta), chemiluminescent signals was detected with an ImageQuant™ LAS 4000 mini (GE Healthcare).
ChIP assay
ChIP assay was performed as described previously51. Differentiated U937 cells were cross-linked with 1% formaldehyde for 10 min at room temperature and then quenched with 0.2 M of glycine. Chromatin extracts from lysed cells were sonicated to an average DNA fragment length of 350 bp and immunoprecipitated with Santa Cruz Biotechnology antibodies anti-XBP1 (sc-7160) or IgG isotype control (sc-2027). Following de-crosslinking, quantitative PCR was performed with multiple ChIP primers (Supplementary Table S1) to quantitate the relative occupancy of the GPR43 promoter region. Fold enrichment was obtained by normalizing the amount of precipitated DNA to that of the input sample and then expressed relative to the GPR43 negative control coding region.
siRNA knockdown and XBP1s overexpression
For stable knockdown of XBP1, siRNA sequences Sense: 5′-AAAGACAGCAAGTGGTAGATTTA-3′; Anti-sense: 5′-AAAATAAATCTACCACTTGCTGT-3′52 was cloned into pFIV-H1/U6 vector (System Biosciences), while a non-targeting sequence was cloned as the negative control vector. The resulting siRNA vector construct or pFIV-H1/U6-copGFP vector (System Biosystems), which contains CopepodGFP coding sequence in replacement of the puromycin selection marker, was co-transfected with lentiviral packaging vectors pFIV-34N and pVSV-G into HEK-293T cells with TurboFect transfection reagent (Thermo Scientific). Culture supernatant was collected after 48 h and used to resuspend U937 cells, with the addition of polybrene to a final concentration of 8 μg/mL. The resulting suspensions were then centrifuged at 1000 g for 2 h at 37°C, after which the transduced cells were re-suspended in 2 mL of fresh complete RPMI media and seeded into 6 well plates. Successfully transduced cells were selected with 1 μg/mL puromycin (Life technologies) in culture medium 48 h after transduction, for 1.5 weeks.
For XBP1s overexpression, XBP1s coding sequence (NM_001079539.1) was cloned into pcDNA3.1A (Invitrogen) and transfected into A549 cells using X-tremeGENE HP DNA Transfection Reagent (Roche) according to the manufacturer's instructions.
References
Maslowski, K. M. et al. Regulation of inflammatory responses by gut microbiota and chemoattractant receptor GPR43. Nature 461, 1282–1286 (2009).
Sina, C. et al. G protein-coupled receptor 43 is essential for neutrophil recruitment during intestinal inflammation. J. Immunol. Baltim. Md 1950 183, 7514–7522 (2009).
Kim, M. H., Kang, S. G., Park, J. H., Yanagisawa, M. & Kim, C. H. Short-chain fatty acids activate GPR41 and GPR43 on intestinal epithelial cells to promote inflammatory responses in mice. Gastroenterology 145, 396–406. e1–10 (2013).
Smith, P. M. et al. The Microbial Metabolites, Short-Chain Fatty Acids, Regulate Colonic Treg Cell Homeostasis. Science 341, 569–573 (2013).
Masui, R. et al. G protein-coupled receptor 43 moderates gut inflammation through cytokine regulation from mononuclear cells. Inflamm. Bowel Dis. 19, 2848–2856 (2013).
Bjursell, M. et al. Improved glucose control and reduced body fat mass in free fatty acid receptor 2-deficient mice fed a high-fat diet. Am. J. Physiol. Endocrinol. Metab. 300, E211–220 (2011).
Tolhurst, G. et al. Short-chain fatty acids stimulate glucagon-like peptide-1 secretion via the G-protein-coupled receptor FFAR2. Diabetes 61, 364–371 (2012).
Kimura, I. et al. The gut microbiota suppresses insulin-mediated fat accumulation via the short-chain fatty acid receptor GPR43. Nat. Commun. 4, 1829 (2013).
Cummings, J. H., Pomare, E. W., Branch, W. J., Naylor, C. P. & Macfarlane, G. T. Short chain fatty acids in human large intestine, portal, hepatic and venous blood. Gut 28, 1221–1227 (1987).
Bloemen, J. G. et al. Short chain fatty acids exchange across the gut and liver in humans measured at surgery. Clin. Nutr. Edinb. Scotl. 28, 657–661 (2009).
Wong, J. M. W., de Souza, R., Kendall, C. W. C., Emam, A. & Jenkins, D. J. A. Colonic health: fermentation and short chain fatty acids. J. Clin. Gastroenterol. 40, 235–243 (2006).
Roy, C. C., Kien, C. L., Bouthillier, L. & Levy, E. Short-chain fatty acids: ready for prime time? Nutr. Clin. Pract. Off. Publ. Am. Soc. Parenter. Enter. Nutr. 21, 351–366 (2006).
Vinolo, M. A. R., Rodrigues, H. G., Nachbar, R. T. & Curi, R. Regulation of inflammation by short chain fatty acids. Nutrients 3, 858–876 (2011).
Brown, A. J. et al. The Orphan G protein-coupled receptors GPR41 and GPR43 are activated by propionate and other short chain carboxylic acids. J. Biol. Chem. 278, 11312–11319 (2003).
Le Poul, E. et al. Functional characterization of human receptors for short chain fatty acids and their role in polymorphonuclear cell activation. J. Biol. Chem. 278, 25481–25489 (2003).
Nilsson, N. E., Kotarsky, K., Owman, C. & Olde, B. Identification of a free fatty acid receptor, FFA2R, expressed on leukocytes and activated by short-chain fatty acids. Biochem. Biophys. Res. Commun. 303, 1047–1052 (2003).
Senga, T. et al. LSSIG is a novel murine leukocyte-specific GPCR that is induced by the activation of STAT3. Blood 101, 1185–1187 (2003).
Harris, P. & Ralph, P. Human leukemic models of myelomonocytic development: a review of the HL-60 and U937 cell lines. J. Leukoc. Biol. 37, 407–422 (1985).
Hass, R. et al. TPA-induced differentiation and adhesion of U937 cells: changes in ultrastructure, cytoskeletal organization and expression of cell surface antigens. Eur. J. Cell Biol. 48, 282–293 (1989).
Chen, M. & Manley, J. L. Mechanisms of alternative splicing regulation: insights from molecular and genomics approaches. Nat. Rev. Mol. Cell Biol. 10, 741–754 (2009).
Ayoubi, T. A. & Van De Ven, W. J. Regulation of gene expression by alternative promoters. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 10, 453–460 (1996).
Pennacchio, L. A. & Rubin, E. M. Genomic strategies to identify mammalian regulatory sequences. Nat. Rev. Genet. 2, 100–109 (2001).
Cartharius, K. et al. MatInspector and beyond: promoter analysis based on transcription factor binding sites. Bioinforma. Oxf. Engl. 21, 2933–2942 (2005).
Wang, X. Z. & Ron, D. Stress-induced phosphorylation and activation of the transcription factor CHOP (GADD153) by p38 MAP Kinase. Science 272, 1347–1349 (1996).
Groth, R. D. & Mermelstein, P. G. Brain-derived neurotrophic factor activation of NFAT (nuclear factor of activated T-cells)-dependent transcription: a role for the transcription factor NFATc4 in neurotrophin-mediated gene expression. J. Neurosci. Off. J. Soc. Neurosci. 23, 8125–8134 (2003).
Lee, J. et al. p38 MAPK-mediated regulation of Xbp1s is crucial for glucose homeostasis. Nat. Med. 17, 1251–1260 (2011).
Martinon, F., Chen, X., Lee, A.-H. & Glimcher, L. H. TLR activation of the transcription factor XBP1 regulates innate immune responses in macrophages. Nat. Immunol. 11, 411–418 (2010).
Caux, C. et al. CD34+ hematopoietic progenitors from human cord blood differentiate along two independent dendritic cell pathways in response to GM-CSF+TNF alpha. J. Exp. Med. 184, 695–706 (1996).
Iwakoshi, N. N., Pypaert, M. & Glimcher, L. H. The transcription factor XBP-1 is essential for the development and survival of dendritic cells. J. Exp. Med. 204, 2267–2275 (2007).
Xue, X. et al. Tumor necrosis factor alpha (TNFalpha) induces the unfolded protein response (UPR) in a reactive oxygen species (ROS)-dependent fashion and the UPR counteracts ROS accumulation by TNFalpha. J. Biol. Chem. 280, 33917–33925 (2005).
Glimcher, L. H. XBP1: the last two decades. Ann. Rheum. Dis. 69 Suppl 1 i67–71 (2010).
Kaser, A. et al. XBP1 links ER stress to intestinal inflammation and confers genetic risk for human inflammatory bowel disease. Cell 134, 743–756 (2008).
Ozcan, U. et al. Endoplasmic reticulum stress links obesity, insulin action and type 2 diabetes. Science 306, 457–461 (2004).
Kim, Y.-M. et al. Molecular characterization of the Tax-containing HTLV-1 enhancer complex reveals a prominent role for CREB phosphorylation in Tax transactivation. J. Biol. Chem. 282, 18750–18757 (2007).
Fric, J. et al. NFAT control of innate immunity. Blood 120, 1380–1389 (2012).
Ihle, J. N. The Stat family in cytokine signaling. Curr. Opin. Cell Biol. 13, 211–217 (2001).
Müller, M. R. & Rao, A. NFAT, immunity and cancer: a transcription factor comes of age. Nat. Rev. Immunol. 10, 645–656 (2010).
Yu, H., Pardoll, D. & Jove, R. STATs in cancer inflammation and immunity: a leading role for STAT3. Nat. Rev. Cancer 9, 798–809 (2009).
Nishitoh, H. CHOP is a multifunctional transcription factor in the ER stress response. J. Biochem. (Tokyo) 151, 217–219 (2012).
Voltolini, C. et al. A novel antiinflammatory role for the short-chain fatty acids in human labor. Endocrinology 153, 395–403 (2012).
Dewulf, E. M. et al. Evaluation of the relationship between GPR43 and adiposity in human. Nutr. Metab. 10, 11 (2013).
Korzenik, J. R. et al. Sargramostim for active Crohn's disease. N. Engl. J. Med. 352, 2193–2201 (2005).
Xu, Y., Hunt, N. H. & Bao, S. The role of granulocyte macrophage-colony-stimulating factor in acute intestinal inflammation. Cell Res. 18, 1220–1229 (2008).
Fukao, T. & Koyasu, S. PI3K and negative regulation of TLR signaling. Trends Immunol. 24, 358–363 (2003).
Sawzdargo, M. et al. A cluster of four novel human G protein-coupled receptor genes occurring in close proximity to CD22 gene on chromosome 19q13.1. Biochem. Biophys. Res. Commun. 239, 543–547 (1997).
Wang, A., Gu, Z., Heid, B., Akers, R. M. & Jiang, H. Identification and characterization of the bovine G protein-coupled receptor GPR41 and GPR43 genes. J. Dairy Sci. 92, 2696–2705 (2009).
Shi, G. et al. FFAR2, a candidate target for T1D, induces cell apoptosis through ERK signaling. J. Mol. Endocrinol. (2014) 10.1530/JME-14-0065.
Hong, Y.-H. et al. Acetate and propionate short chain fatty acids stimulate adipogenesis via GPCR43. Endocrinology 146, 5092–5099 (2005).
Sha, H. et al. The IRE1alpha-XBP1 pathway of the unfolded protein response is required for adipogenesis. Cell Metab. 9, 556–564 (2009).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 3, 1101–1108 (2008).
Chew, J.-L. et al. Reciprocal transcriptional regulation of Pou5f1 and Sox2 via the Oct4/Sox2 complex in embryonic stem cells. Mol. Cell. Biol. 25, 6031–6046 (2005).
Pyrko, P., Schönthal, A. H., Hofman, F. M., Chen, T. C. & Lee, A. S. The unfolded protein response regulator GRP78/BiP as a novel target for increasing chemosensitivity in malignant gliomas. Cancer Res. 67, 9809–9816 (2007).
Ge, H. et al. Activation of G protein-coupled receptor 43 in adipocytes leads to inhibition of lipolysis and suppression of plasma free fatty acids. Endocrinology 149, 4519–4526 (2008).
Acknowledgements
This work was funded by a grant from the National Medical Research Council (NMRC/CBRG/0055/2014). We thank Dr. Huck Hui Ng and Dr. Jia Hui Ng (Genome Institute of Singapore) for providing valuable guidance on the ChIP experiments.
Author information
Authors and Affiliations
Contributions
A.Z.W. and E.J.Z. conceived, designed and performed the experiments and analyzed the data with intellectual input from J.L.D. J.L.D. provided overall coordination and supervision of the study. A.Z.W., E.J.Z. and J.L.D. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ang, Z., Er, J. & Ding, J. The short-chain fatty acid receptor GPR43 is transcriptionally regulated by XBP1 in human monocytes. Sci Rep 5, 8134 (2015). https://doi.org/10.1038/srep08134
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep08134
This article is cited by
-
Ffar2 expression regulates leukaemic cell growth in vivo
British Journal of Cancer (2017)
-
Human and mouse monocytes display distinct signalling and cytokine profiles upon stimulation with FFAR2/FFAR3 short-chain fatty acid receptor agonists
Scientific Reports (2016)
-
New Horizons for the Study of Dietary Fiber and Health: A Review
Plant Foods for Human Nutrition (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.