Abstract
In plants, microRNAs (miRNAs) play essential roles in growth, development, yield, stress response and interactions with pathogens. However no miRNA has been experimentally documented to be functionally involved in fruit ripening although many miRNAs have been profiled in fruits. Here we show that SlymiR157 and SlymiR156 differentially modulate ripening and softening in tomato (Solanum lycopersicum). SlymiR157 is expressed and developmentally regulated in normal tomato fruits and in those of the Colourless non-ripening (Cnr) epimutant. It regulates expression of the key ripening gene LeSPL-CNR in a likely dose-dependent manner through miRNA-induced mRNA degradation and translation repression. Viral delivery of either pre-SlymiR157 or mature SlymiR157 results in delayed ripening. Furthermore, qRT-PCR profiling of key ripening regulatory genes indicates that the SlymiR157-target LeSPL-CNR may affect expression of LeMADS-RIN, LeHB1, SlAP2a and SlTAGL1. However SlymiR156 does not affect the onset of ripening, but it impacts fruit softening after the red ripe stage. Our findings reveal that working together with a ripening network of transcription factors, SlymiR157 and SlymiR156 form a critical additional layer of regulatory control over the fruit ripening process in tomato.
Similar content being viewed by others

Introduction
MicroRNAs (miRNAs) are approximately 21-nucleotide (nt) regulatory RNAs that are processed by the dicer-like nuclease DCL1 from stem-loop structures of longer pre-miRNA precursors1. An increasing number of miRNAs have been identified in plants and most of them are conserved between species, although species-specific functional miRNAs exist2,3,4,5,6. MiRNAs play essential roles in diverse developmental processes and plant responses to biotic and abiotic stresses through suppression of target gene expression by miRNA-guided mRNA cleavage, translational repression or transcriptional RNA silencing2,3. One of the characteristics of plant miRNAs is that each miRNA family that is conserved among dicots and monocots has a conserved complementary target site in a mRNA, indicating that these miRNAs may have a similar function in different plants1,2,6.
Tomato (Solanum lycopersicum) is an economically important fleshy fruit and a major worldwide crop with high nutritional value to human health. However tomato becomes edible only after the fruit ripens. Current understanding of the genetic and molecular mechanisms involved in fruit ripening mainly comes from studies of various rare ripening tomato mutants including ripening inhibitor (rin) and Colourless non-ripening (Cnr)7,8. The rin locus encodes a MADS-box transcription factor (TF) LeMADS-RIN9, while the Cnr locus harbours an SBP-box gene, LeSPL-CNR, which belongs to the SQUAMOSA Promoter Binding Protein-like (SPL) TF family10. The 3′-UTR of the LeSPL-CNR mRNA possesses a potential miRNA156/7 target site, revealing the first clue that miRNAs may control the expression of a crucial ripening TF and thus have a functional role in fruit ripening in tomato. Many miRNAs and small RNAs have been profiled in fruits11,12,13,14,15,16, however no miRNA has been experimentally documented to be functionally involved in fruit ripening. Here we report differential roles of SlymiR157 and SlymiR156 in tomato fruit ripening and softening.
Results
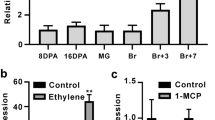
SlymiR157 and SlymiR156 are closely related, but only SlymiR157 is perfectly complementary to the potential miRNA target site in the LeSPL-CNR mRNA (Supplementary Fig. 1). We analysed the expression profiles of both miRNAs in fruits of normal Ailsa Craig (AC) and epi-mutant Cnr and found that expression of SlymiR156 was similar in AC and mutant fruits at 40 (breaker) and 54 days post anthesis (dpa). In contrast, SlymiR157 was differentially and developmentally expressed with a relatively higher level in the Cnr mutant compared to normal fruits (Fig. 1a, b). These data suggest that SlymiR157 and SlymiR156 may be involved in the regulation of the LeSPL-CNR expression at different stages of tomato development, but SlymiR157 may mainly contribute to fruit ripening. This idea is supported by previous findings that transgenic over-expression of SlymiR156 does not obviously affect fruit ripening but instead has significant impacts on fruit formation and yield17, which is incompatible with the current model generated from RNA deep sequencing and bioinformatics predictions that suggest SlymiR156 is the main small RNA effector controlling ripening in tomato12,16.
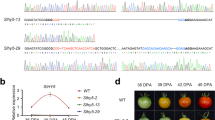
Differential and developmental regulation of SlymiR157 in tomato fruits.
(a) Transcript profiles of SlymiR156 in wild type Ailsa Craig (AC) and Cnr epimutant fruits at breaker (40 days post anthesis, dpa) and at 54 dpa (d). (b) Differential and developmental expression of SlymiR157 in AC and Cnr fruits. The amount of SlymiR157 was significantly higher at 40 dpa than 54 dpa in Cnr fruits. A similar trend of the SlymiR157 level was also found in AC and Cnr fruits, but expression of SlymiR157 decreased markedly in AC than Cnr fruits at the same stage of the ripening process. (c) Stem-loop structure of the pre-SlymiR157 RNA molecule. The genome coordinates on tomato Chromosome 3 are indicated.
We have cloned the SlyMIR157 gene through genome walking. Pre-SlymiR157 derived from SlyMIR157 was predicted to form stem-loop structures (Fig. 1c). Mining of the tomato whole genome sequence18 indicates that a single copy of SlyMIR157 is presented and located on tomato Chromosome 3 at 61,724,365–61,724,483 nt. To examine the role of SlymiR157 in ripening we exploited “miRNA-based virus induced gene silencing” (dubbed Mr VIGS), to express pre-SlymiRNA in tomato fruits. Pre-SlymiR157 was cloned into Potato virus X (PVX)- and Tobacco rattle virus (TRV)-based VIGS vectors19,20 to produce PVX/pre-SlymiR157 and TRV/pre-SlymiR157 (Fig. 2a, b). Pre-SlymiR157 was delivered into more than 100 fruits at various days post anthesis from the two different VIGS vectors and 10-20% of injected fruits developed sectors with delayed ripening (DR), whilst several dozens of fruits mock-treated with TE buffer or injected with empty PVX or TRV ripened normally (Fig. 2c, d). However the phenotypes induced by pre-SlymiR157-based Mr VIGS were considerably weaker than the non-ripening phenotypes caused by siRNA-based VIGS (dubbed Sir VIGS) of LeSPL-CNR in fruits injected with PVX/mLeSPL-CNR (Fig. 2e, f), indicating miRNA-mediated regulation of LeSPL-CNR gene expression may be a fine tuning mechanism in the control of ripening.
Pre-SlymiR157 mediated Mr VIGS inhibits fruit ripening.
(a) PVX-based vectors PVX/pre-SlymiR157 (pre157) for Mr VIGS and PVX/mLeSPL-CNR with a non-translational mutant LeSPL-CNR (mCNR) RNA for Sir VIGS. Genome organization (166K, 25K 12K, 8K and coat protein (CP)) and genomic (g), sub-genomic (sg) RNA1, gene-of-interest (GOI) sgRNA and PVX coat protein (CP) sgRNA of PVX are indicated. (b) TRV-based vector TRV/pre-SlymiR157 for Mr VIGS. Transcription of the recombinant TRV, GOI and a ribozyme (Rz) RNA is under the control the 35S promoter and NOS terminator. (c, d) Viral ectopic expression of pre-SlymiR157 from PVX (c) or TRV (d) delayed AC fruits to ripen. Ripening delayed sectors and pericarp in dissected fruits are green or yellow (arrow). Mock- or empty vectors (PVX or TRV)-treated AC fruits ripened normally. (e) Sir VIGS of LeSPL-CNR by PVX/mLeSPL-CNR developed non-ripening green sectors. 10-20% injected fruits showed the non-ripening phenotypes. (f) AC fruit injected with empty Sir VIGS vector PVX ripened normally. Photographs were taken at 5, 6, or 17 days after breaker (Br +5, Br + 6, or Br + 17).
To correlate the DR phenotype with the viral delivery of pre-SlymiR157, we performed qRT-PCR analysis on pre-SlymiR157 and mRNAs of LeSPL-CNR and several other key ripening-associated TFs including LeMADS-RIN9, LeHB121, SlAP2a22 and SlTAGL123 in three randomly collected fruits injected with PVX/pre-SlymiR157 (Fig. 3a, b, c). Viral pre-SlymiR157 accumulated to a significant high level and was much more abundant in the DR than red-ripe (RR) sectors of all fruits tested. These results suggest that PVX was able to efficiently deliver pre-SlymiR157 into fruits and that the occurrence and amount of pre-SlymiR157 were correlated with the DR phenotype. Consistent with the DR phenotype and massive accumulation of pre-SlymiR157, the level of LeSPL-CNR mRNA decreased by 22–50% in DR sectors compared to that in RR tissues of all injected fruits (Fig. 3a, b, c). Such an inverse relationship suggests that miRNA-mediated cleavage of LeSPL-CNR mRNA occurs in DR fruit tissues. A similar trend in the mRNA levels of key ripening TF genes LeHB1, SlAP2a and SlTAGL1 was also observed although LeMADS-RIN mRNA decreased significantly only in the DR sector of two out of three tested fruits (Fig. 3a, b, c). It is worthwhile noting that the four ripening TF genes contain no SlymiR157 target site18, suggesting that expression of these TF genes could not be directly regulated by SlymiR157 during fruit ripening in tomato.
Mr VIGS affects expression of LeSPL-CNR and key ripening TF genes.
(a-f) Viral delivery of pre-SlymiR157 and mature SlymiR157, as well as expression of the SlymiR157 target gene LeSPL-CNR and TF genes (LeMADS-RIN, LeHB1, SlAP2a and SlTAGL1) in delayed ripening (DR) and red-ripe (RR) sectors of three different AC fruits injected with PVX/pre-SlymiR157 (a, b, c) or PVX/SlymiR157 (d, e, f) were respectively analysed by qRT-PCR. Relative levels of RNA transcripts were obtained by normalising against the baseline expression levels of the internal control 18S rRNA. qRT-PCR of each biological sample was performed in 3-15 technical replicates. Student's t-tests were carried to evaluate the statistical significance at the level of p ≤ 0.001 (***), 0.01 (**) or 0.05 (*) for each of the specific RNA transcripts between DR and RR sectors of each different biological sample. Although there is variance, the results of these qRT-PCR assays show a similar correlation between the viral delivery of pre-SlymiR157 or mature SlymiR157 and the mRNA levels of ripening TF genes in the DR and RR sectors of fruits tested.
We also cloned the mature SlymiR157 and SlymiR156 into the PVX-based VIGS vector19 to produce PVX/SlymiR157 and PVX/SlymiR156 (Supplementary Fig. 2a). They were injected separately into the carpopodium of approximately 100 AC fruits at various stages of development on different trusses on the same plant and on different plants. About 15% of AC fruits injected with PVX/SlymiR157 produced green sectors that showed delayed ripening and later these sectors turned yellow, a phenotype characteristic of Cnr mutant (Supplementary Fig. 2b). All control fruits that were mock-injected or injected with PVX alone ripened normally. These effects mirrored those obtained using pre-SlymiR157 described above. Fruits injected with PVX/SlymiR156 ripened normally until the breaker stage, however these fruits then quickly softened and became liquefied (Supplementary Fig. 2c). These findings suggest that SlymiR157 and SlymiR156 play different roles in the modulation of ripening and softening, respectively. The softening phenotype remains to be elucidated. However, considering the high expression of SlymiR156 (Fig. 1a) and a significant decrease of LeSPL-CNR mRNA after the breaker stage10, LeSPL-CNR may also have a previously unrecognised function as a repressor of softening. Thus, SlymiR156-guided LeSPL-CNR mRNA cleavage12,16 would release such suppression and enhance fruit softening in tomato especially at the later stages of ripening.
On the other hand, viral delivery of mature SlymiR157 was readily detected in all treated fruits. Consistent with the DR phenotype, a higher level of PVX:SlymiR157 was present in the DR than RR sector even though overall levels were low (Fig. 3d, e, f). Unexpectedly, the accumulation of the transcript of the SlymiR157 target LeSPL-CNR gene, as well as of LeMADS-RIN, LeHB1, SlAP2a and SlTAGL1 mRNAs was also found to be greater in the DR than RR sectors of all fruits tested. Although miRNA-mediated posttranscriptional activation has been reported in quiescent mammalian cells and immature Xenopus laevis oocytes24,25, to our knowledge, this is the first example of miRNA-mediated up-regulation of gene expression in plants.
The increased accumulation of LeSPL-CNR by SlymiR157 is inconsistent with the phenotype in DR sectors where ripening was delayed (Supplementary Fig. 2b). To explain this discrepancy, we hypothesise that SlymiR157 may regulate tomato ripening via miRNA-mediated mRNA degradation or translational suppression through a dose-dependent mode (Fig. 3). Presence of a massive amount of SlymiR157 could effectively trigger miRNA-mediated cleavage and reduction of the LeSPL-CNR mRNA, leading to ripening-delayed phenotype as observed in DR sector of fruits injected with PVX/pre-SlymiR157 and TRV/pre-SlymiR157 (Fig. 2c, d; Fig. 3a, b, c). Conversely, the much lower amount of SlymiR157 below a threshold level would not cause degradation of LeSPL-CNR transcript but only represses translation of LeSPL-CNR protein, resulting in maintenance of fruits at breaker stage for a prolonged period during which LeSPL-CNR would be constantly transcribed10. Consequently, LeSPL-CNR mRNA accumulated to a higher amount, as found in the DR than RR sector of fruits injected with PVX/SlymiR157 (Supplementary Fig. 2b; Fig. 3d, e, f) and some of these mRNAs could remain translatable to produce an increasing amount of the LeSPL-CNR protein. The latter would result in up-regulation of the other ripening TF gene expression (Fig. 3d, e, f).
To test whether SlymiR157 could cause translation repression, we developed a “miRNA target – GFP reporter expression” assay in Nicotinana benthamiana. The SlymiR157-target site (Tar), wild-type and mutated SlymiR157 sequences were cloned into PVX/GFP19 to produce PVX/SlymiR157Tar::GFP, PVX/SlymiR157::GFP and PVX/mSlymiR157::GFP (Fig. 4a). RNA transcripts produced by in vitro transcription from the three recombinant viruses as well as from PVX/GFP were able to cause systemic infection of N. benthamiana plants. GFP protein was readily detected by direct UV visualization of green fluorescence in plants that were infected with PVX/GFP, PVX/SlymiR157::GFP or PVX/mSlymiR157::GFP. However, the insertion of the short RNA of the target site of SlymiR157 totally blocked GFP translation in PVX/SlymiR157Tar::GFP-infected plant (Fig. 4b, c, d, e). These observations are consistent with western blot detection of GFP and PVX coat protein in these plants (Fig. 4f). Importantly, semi-qRT-PCR assays showed that the levels of GFP mRNA produced from all recombinant viruses were similar (Fig. 4g). Due to the conserved and identical nature of both N. benthamiana miR157 and SlymiR157 (Supplementary Fig. 1b), we interpret these data to mean that the endogenous miR157 present in the tobacco leaf cells is down-regulating Tar::GFP protein levels through translation repression rather than via miRNA-mediated mRNA degradation. Furthermore, tomato fruits with pedicel injection of PVX/SlymiR157::GFP showed sectors of Cnr mutant phenotype whilst fruits injected with PVX/mSlymiR157::GFP, PVX/SlymiR157Tar::GFP or mock treatments ripened normally (Fig. 4h, i, j, k). Taken together, our results imply that SlymiR157 may provide an additional layer of regulation to fine-tune LeSPL-CNR expression through a combined action of miRNA-mediated degradation and translation repression in order to control tomato fruit ripening.
miRNA target – GFP reporter expression assay.
(a) PVX-based miRNA target – GFP reporter expression vectors. (b-e) Nicotinana benthamiana leaves were inoculated with PVX/GFP (GFP, b), PVX/SlymiR157Tar::GFP (miR157Tar::GFP, c), PVX/SlymiR157::GFP (miR157::GFP, d) or PVX/mSlymiR157::GFP (mmiR157::GFP, e). Photographs were taken under long-wavelength UV light at 7 days post inoculation. GFP fluorescence is green and leaf auto-fluorescence shows red. (f) Western blot detection of GFP and PVX coat protein (CP) in mock- or PVX-inoculated leaf tissues using antibody raised specifically against GFP and the viral CP. Positions of GFP and PVX CP are indicated. (g) Detection of recombinant PVX::miRNA target::GFP RNA in same leaf samples as used in Western blot by semi-qRT-PCR. The positions and sizes of amplified fragments and the number of PCR cycles are indicated. 18S rRNA was used as controls. (h-k) AC fruits were injected with PVX/GFP (h), PVX/SlymiR157::GFP (i), PVX/SlymiR157Tar::GFP (j) or PVX/mSlymiR157::GFP (k). Non-ripening sectors show green. Fruits were photographed at 7 days after breaker.
Discussion
In tomato, miRNAs are known to be involved in leaf and flower development, yield production, stress response and interactions with pathogens17,26,27,28,29. High throughput sequencing and bioinformatic analyses predict the presence of many ripening-related small RNAs and miRNAs and their targets in fruits at various developmental stages11,12,13,14,15,16. For instance, SlymiR172 is predicted to cleave SlAP2a mRNA and several miRNAs are also believed to target ripening-associated genes coding for pectate lyase (SlymiR482), endo-1,4-ß-glucanase (SlymiR395), ß-galactosidase (SlymiRZ7), ethylene-insensitive 2 (SlymiR828), serine/threonine protein kinase (SlymiR1917) and 1-aminocyclopropane-1-carboxylic acid synthase (SlymiR159)16. Nevertheless, there is no experimental evidence to demonstrate that any of these miRNAs have a functional role in tomato fruit ripening.
It has also been postulated that SlymiR156 may target and cleave LeSPL-CNR mRNA and affect ripening in tomato12,16. However transgenic over-expression of SlymiR156 in a previous report17 and viral delivery of SlymiR156 described in this study, clearly show that SlymiR156 does not significantly affect ripening per se, but that it mainly influences fruit softening (Supplementary Fig. 2) and other developmental processes. Further, a more recent study reports that miR156 regulates tomato ovary and fruit development30. Transgenic over-expression of precursor of the Arabidopsis miR156 (AtmiR156b) in tomato produced abnormal flowers and fruits with extra carpels and meristem-like structures inside the ovaries. Development of such ectopic organs is associated with miR156-mediated regulation of SPL/SBP box TF gene expression as well as with alternations of expression of TFs that are essential for meristem maintenance and reproductive development30.
However, SlymiR157 is differentially expressed in normal tomato fruits and the Cnr epi-mutant and regulates LeSPL-CNR expression. Moreover, viral delivery of either pre-SlymiR157 or its mature form by VIGS vectors can delay ripening in tomato. Thus, SlymiR157 may provide a post-transcriptional mechanism for fine-tuning LeSPL-CNR expression and protein levels through miRNA-induced mRNA cleavage or/and translation repression, possibly in a dose-dependent manner to affect LeSPL-CNR function. Furthermore, the coordinated gene expression profiles of LeSPL-CNR, LeMADS-RIN, LeHB1, SlAP2a and SlTAGL1 also suggest that LeSPL-CNR may act as a positive upstream regulator for these other four TFs (Fig. 5). This is consistent with previous findings that LeMADS-RIN protein interacts with promoters of several key ripening-associated TFs including LeHB1 and LeSPL-CNR and the latter is required for LeMADS-RIN to bind its target promoter sequence in the transcriptional control of ripening31. Thus, our findings support and greatly extend a model for the control of ripening involving a network of TFs7,8,32 and a critical extra layer of regulation involving miRNAs.
Involvement of miRNAs in modulation of tomato fruit ripening.
A transcription factor (TF) gene network including LeSPL-CNR, LeMADS-RIN, LeHB1, SlAP2a and SlTAGL1 is involved in tomato ripening. Blue arrows or “T” signs denote interactions among these TF genes that have been previously demonstrated7,8,32. Yellow arrows indicate that LeSPL-CNR may activate expression of the ripening TFs, as described in this work. SlymiR172 is proposed to down-regulate SlAP2a and affect ripening but this idea is not supported by any in planta experimental evedience7. SlymiR157 (pre-SlymiR157) may cause  miRNA-guided mRNA degradation or
miRNA-guided mRNA degradation or  translation repression to affect LeSPL-CNR function depending on whether a high above-threshold (
translation repression to affect LeSPL-CNR function depending on whether a high above-threshold ( ) or low below-threshold (
) or low below-threshold ( ) levels of SlymiR157 are present. The correlation between the high expression of SlymiR156 (Fig. 1a) and a significant decrease of LeSPL-CNR mRNA10 after the breaker stage suggests that apart from being an activator for the early ripening, LeSPL-CNR may also have a previously unrecognised function as a repressor of softening. Thus, SlymiR156-guided LeSPL-CNR mRNA cleavage12,16 would enhance fruit softening at the later stage of tomato development (
) levels of SlymiR157 are present. The correlation between the high expression of SlymiR156 (Fig. 1a) and a significant decrease of LeSPL-CNR mRNA10 after the breaker stage suggests that apart from being an activator for the early ripening, LeSPL-CNR may also have a previously unrecognised function as a repressor of softening. Thus, SlymiR156-guided LeSPL-CNR mRNA cleavage12,16 would enhance fruit softening at the later stage of tomato development ( ) in addition to its weak role in the epigenetic modulation of fruit ripening17 (
) in addition to its weak role in the epigenetic modulation of fruit ripening17 ( ).
).
Methods
Plant materials and growth conditions
Tomato (Solanum lycopersicum cultivar Ailsa Craig) and Cnr epimutant fruits were grown in a growth room with a photoperiod of 16-hr at 25°C during daytime and 20°C at night. Plants were grown to three-five trusses. Nicotiana benthamiana plants were grown under the same conditions as for tomato. Tomato fruits at various days post anthesis and N. benthamiana leaves at 7 days post-inoculation were collected and kept at −80°C until use.
Genome walking experiments
Genome walking to clone the SlyMIR157 gene was carried out as described10. The stem-loop structure of pre-SlymiR157 was predicted using an online mfold web server (mfold.rit.albany.edu).
Construction of vectors for Mr and Sir VIGS
The pre-SlymiR157 fragment was RT-PCR amplified using two sets of primers, digested with restriction enzymes and cloned into the potato virus X (PVX)- and tobacco rattle virus (TRV)-based VIGS vectors19,20 to produce PVX/pre-SlymiR157 and TRV/pre-SlymiR157, respectively (Supplementary Table 1). For construction of PVX/SlymiR157, PVX/SlymiR156, PVX/SlymiR157Tar::GFP, PVX/SlymiR157::GFP, PVX/mSlymiR157::GFP, a pair of relevant complementary primers (Supplementary Table 1) was mixed in TE (10 mM Tris-HCl (pH 8.0), 1 mM EDTA) buffer, heated at 95°C for 3 minutes, then cooled to room temperature to allow them to anneal to form double-strand forms which were cloned into either PVX or PVX/GFP vector19. The Sir VIGS vector PVX/mLeSPL-CNR carrying a mutated LeSPL-CNR gene was constructed as described10. All constructs were verified by sequencing.
In planta miRNA functional assays and in vitro molecular analysis
Mr and Sir VIGS and the miRNA target – GFP reporter expression assay were performed as described10,19,20. Absolute qRT-PCR to quantify SlymiR157, relative qRT-PCR and semi-qRT-PCR to measure viral delivery of pre- and mature miRNAs, TF and GFP reporter transcripts were performed as described10,19,32 using primers listed in Supplementary Table 2. Direct visualization of GFP green fluorescence in leaves and western blot detection of GFP and PVX coat protein were carried out as described19,32.
References
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism and function. Cell 116, 281–297 (2004).
Jones-Rhoades, M. W., Bartel, D. P. & Bartel, B. MicroRNAs and their regulatory roles in plants. Annu. Rev. Plant Biol. 57, 19–53 (2006).
Chen, X. Small RNAs and their roles in plant development. Annu. Rev. Cell Dev. Biol. 25, 21–44 (2009).
Sunkar, R., Chinnusamy, V., Zhu, J. & Zhu, J. K. Small RNAs as big players in plant abiotic stress responses and nutrient deprivation. Trends Plant Sci. 12, 301–309 (2007).
Jin, H. Endogenous small RNAs and antibacterial immunity in plants. FEBS Lett. 582, 2679–2684 (2008).
Sun, G. MicroRNAs and their diverse functions in plants. Plant Mol. Biol. 18, 17–36 (2011).
Karlova, R. et al. Transcriptional control of fleshy fruit development and ripening. J. Exp. Bot. 66, 4527–4511 (2014).
Klee, H. J. & Giovannoni, J. J. Genetics and control of tomato fruit ripening and quality attributes. Annu. Rev. Genet. 45, 41–59 (2011).
Vrebalov, J. et al. A MADS-box gene necessary for fruit ripening at the tomato ripening-inhibitor (rin) locus. Science 296, 343–346 (2002).
Manning, K. et al. A naturally occurring epigenetic mutation in a gene encoding an SBP-box transcription factor inhibits tomato fruit ripening. Nat. Genet. 38, 948–962 (2006).
Pilcher, R. L. et al. Identification of novel small RNAs in tomato (Solanumlycopersicum). Planta 226, 709–717 (2007).
Moxon, S. J. R. et al. Deep sequencing of tomato short RNAs identifies microRNAs targeting genes involved in fruit ripening. Genome Res. 18, 1602–1609 (2008).
Yin, Z. J., Li, C. H., Han, X. L. & Shen, F. F. Identification of conserved microRNAs and their target genes in tomato (Lycopersicon esculentum). Gene 414, 60–66 (2008).
Mohorianu, I. et al. Profiling of short RNAs during fleshy fruit development reveals stage-specific sRNAome expression patterns. Plant J. 67, 232–246 (2011).
Zuo, J. et al. Sculpting the maturation, softening and ethylene pathway: the influences of microRNAs on tomato fruits. Genomics 13, 7 (2012).
Karlova, R. et al. Identification of microRNA targets in tomato fruit development using high-throughput sequencing and degradome analysis. J. Exp. Bot. 64, 1863–1878 (2013).
Zhang, X. et al. Over-expression of sly-miR156a in tomato results in multiple vegetative reproductive trait alterations and partial phenocopy of the sft mutant. FEBS Lett. 585, 435–439 (2011).
Tomato Genome Consortium. The tomato genome sequence provides insight into fleshy fruit evolution. Nature 485, 635–641 (2012).
Li, C. et al. Mobile FT mRNA contribute to the system florigen signalling in floral induction. Sci. Rep. 1, 73 (2011).
Tang, Y. et al. Virus-based microRNA expression for gene functional analysis in plants. Plant Physiol. 153, 632–641 (2010).
Lin, Z. et al. A tomato HD-Zip homeobox protein, LeHB1, plays an important role in floral organogenesis and ripening. Plant J. 55, 301–310 (2008).
Chung, M. Y. et al. A tomato (Solanum lycopersicum) APETALA2/ERF gene, SlAP2a, is a negative regulator of fruit ripening. Plant J. 64, 936–947 (2010).
Vrebalov, J. et al. Fleshy fruit expansion and ripening are regulated by the tomato SHATTERPROOF gene TAGL1. Plant Cell 21, 3041–3062 (2009).
Truesdell, S. S. et al. Activation in quiescent cells and oocytes involves recruitment of a nuclear microRNP. Sci. Rep. 2, 842 (2012).
Vasudevan, S., Tong, Y. & Steitz, J. A. Switching from repression to activation: MicroRNAs can up-regulate translation. Science 318, 1931–1934 (2007).
Ori, N. et al. Regulation of LANCEOLATE by miR319 is required for compound-leaf development in tomato. Nat. Genet. 39, 787–791 (2007).
Berger, Y. et al. The NAC-domain transcription factor GOBLET specifies leaflet boundaries in compound tomato leaves. Development 136, 823–832 (2009).
Naqvi, A. R., Haq, Q. M., Mukherjee, S. K. MicroRNA profiling of tomato leaf curl new delhi virus (tolcndv) infected tomato leaves indicates that deregulation of mir159/319 and mir172 might be linked with leaf curl disease. Virol. J. 7, 281–296 (2010).
Zhang, X. H. et al. Over-expression of microRNA169 confers enhanced drought tolerance to tomato. Biotechnol. Lett. 33, 403–409 (2011).
Ferreira e Silva, G. F. et al. microRNA156-targeted SPL/SBP box transcription factors regulate tomato ovary and fruit development. Plant J. 78, 604–618 (2014).
Martel, C., Vrebalov, J., Tafelmeyer, P. & Giovannoni, J. J. The tomato MADS-box transcription factor RIPENING INHIBITOR interacts with promoters involved in numerous ripening processes in a COLORLESS NONRIPENING-dependent manner. Plant Physiol. 157, 1568–1579 (2011).
Zhou, T. et al. Virus-induced gene complementation reveals a transcription factor network in modulation of tomato fruit ripening. Sci. Rep. 2, 836 (2012).
Acknowledgements
We are grateful to D. Baulcombe for providing the original PVX vector, S. Santa-Cruz for the PVX coat protein antisera and G. B. Seymour for critical reading of this manuscript. This work was supported in part by a Pandeng Programme from Hangzhou Normal University (201108), an Innovative Grant for Science Excellence from the Hangzhou City Education Bureau China, an UK-BBSRC core grant (BBS/E/H/00YH0271) and a Research Grant from the UK Royal Society (2007/R1) to Y.H. We also thank the National Natural Science Foundation of China (31370180, 31401926, 31200913, 31201490) and the Zhejiang Provincial Natural Science Foundation (LQ13C020004, LQ13C060003, LQ12C02005, LY14C010005) for supporting our work.
Author information
Authors and Affiliations
Contributions
W.C., J.K. and T.L. designed and performed Mr and Sir VIGS and qRT-PCR; K.M. cloned and analysed SlymiR157; C.W., Y.W., C.Q., B.L., Z.Y., X.Z., M.H., P.Z. performed Mr and Sir VIGS and molecular analysis. M.G., X.Y., A.M., C.L., T.O. and N.S. performed translation repression assays; H.W., S.J., Y.L. and P.G. were involved in analysis of data and helped writing the paper. Y.H. initiated the project, analysed data and wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Chen, W., Kong, J., Lai, T. et al. Tuning LeSPL-CNR expression by SlymiR157 affects tomato fruit ripening. Sci Rep 5, 7852 (2015). https://doi.org/10.1038/srep07852
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07852
This article is cited by
-
Ectopic expression of miRNA172 in tomato (Solanum lycopersicum) reveals novel function in fruit development through regulation of an AP2 transcription factor
BMC Plant Biology (2020)
-
Noncoding RNAs: functional regulatory factors in tomato fruit ripening
Theoretical and Applied Genetics (2020)
-
Genome wide identification, characterization and validation of novel miRNA-based SSR markers in pomegranate (Punica granatum L.)
Physiology and Molecular Biology of Plants (2020)
-
Genome-wide identification of the SPL gene family in Tartary Buckwheat (Fagopyrum tataricum) and expression analysis during fruit development stages
BMC Plant Biology (2019)
-
Transcriptomics of the grape berry shrivel ripening disorder
Plant Molecular Biology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.