Abstract
Despite plants infected by pathogens are often unable to produce offspring, it remains unclear how sterility is induced in host plants. In this study, we demonstrate that TENGU, a phytoplasmal virulence peptide known as a dwarfism inducer, acts as an inducer of sterility. Transgenic expression of TENGU induced both male and female sterility in Arabidopsis thaliana flowers similar to those observed in double knockout mutants of auxin response factor 6 (ARF6) and ARF8, which are known to regulate floral development in a jasmonic acid (JA)-dependent manner. Transcripts of ARF6 and ARF8 were significantly decreased in both tengu-transgenic and phytoplasma-infected plants. Furthermore, JA and auxin levels were actually decreased in tengu-transgenic buds, suggesting that TENGU reduces the endogenous levels of phytohormones by repressing ARF6 and ARF8, resulting in impaired flower maturation. TENGU is the first virulence factor with the effects on plant reproduction by perturbation of phytohormone signaling.
Similar content being viewed by others
Introduction
Phytoplasmas (class Mollicutes, genus Candidatus Phytoplasma) are among the smallest bacterial plant pathogens transmitted by insect vectors. Phytoplasmas induce drastic malformation of plants such as witches' broom, dwarfism, phyllody (the transformation of floral organs into leaf-like structures), virescence (the greening of floral organs) and flower sterility1,2. Because phytoplasma-infected plants are often unable to produce offspring, phytoplasmas have been described as turning plants into zombies3.
To date, four phytoplasmal effectors have been functionally characterized4,5,6,7. A virulence effector, phytoplasma tengu-su inducer (TENGU), was first identified from onion yellows phytoplasma (OY)4. The detection of TENGU in apical meristem tissue indicated that TENGU might be transported from the phloem sieve elements into neighboring tissues4. Expression of TENGU in Arabidopsis thaliana and Nicotiana benthamiana induced dwarfism and witches' broom symptoms and downregulated multiple auxin responsive genes, suggesting that disruption of the auxin signaling pathway might be the cause of these symptoms4.
In A. thaliana, the maturation of both stamens and gynoecia is regulated by AUXIN RESPONSE FACTOR (ARF) 6 and ARF88,9. Recent studies indicated that ARF6 and ARF8 act in a partially redundant manner and regulate anther dehiscence by inducing jasmonic acid (JA) production or by decreasing JA breakdown9,10. In addition, auxin has been implicated in the floral-bud development process almost through flower maturation11.
Plant sterility caused by plant pathogens is an important symptom that has a significant impact on crop production12. Plants infected by many Ca. Phytoplasma species, such as ‘Ca. Phytoplasma solani’, ‘Ca. P. oryzae’ and ‘Ca. P. asteris’ also have sterile flowers13,14,15. Some of these phytoplasmas induce virescence and phyllody in infected floral organs, which can lead to sterility. For example, Ca. P. solani-infected tomato plants exhibit drastic malformations of the stamen and carpels, which causes sterility14,16. By contrast, some phytoplasma strains belonging to ‘Ca. P. asteris’ and ‘Ca. P. oryzae’ induce sterility in both male and female flowers without floral malformations such as virescence and phyllody in particular host plants4,13,15,17. Despite these intriguing observations, how such sterility symptoms are induced in phytoplasma-infected plants remains unclear.
Here, we demonstrate that TENGU, a phytoplasmal virulence factor that causes dwarfism, also induces both male and female sterility without floral malformations in A. thaliana. TENGU significantly affects the expression of flower maturation genes and alters JA and auxin syntheses in flowers, indicating that TENGU causes developmental defects that lead to sterility through modulation of the two phytohormones.
Results
TENGU arrests flower development
We previously reported that both OY-infected and transgenic A. thaliana plants constitutively expressing TENGU exhibited dwarfism or witches' broom phenotypes4. In this study, we found that tengu-transgenic plants also showed sterility.
Whereas no difference in morphology was apparent between most wild-type Col-0 and tengu-transgenic lines during the vegetative stage (Fig. 1a), the majority of tengu-transgenic plants exhibited sterile phenotypes of varying degrees upon flowering. Approximately 37% of the tengu-transgenic plants exhibited severe sterility with closed flower buds and no seed production (Fig. 1, d and h). In the flowers of these severe-sterile lines of a 35S::tengu plant, anthers of infertile buds failed to dehisce and release pollen grains and stigmatic papillae in the carpels were somewhat shorter than those of wild-type flowers (Fig. 1, l, n and p). By contrast, ~57% of the tengu-transgenic plants showed mild sterility with many opened flowers but extremely low seed production, as reported previously4 (Fig. 1, e and i). The stigmatic papillae were as long as wild-type flowers and anthers released pollen grains (Fig. 1, n, q and m). In all OY-infected plants, flowers exhibited severe sterility with undehisced anthers and immature gynoecia, similar to severe-sterile 35S::tengu flowers (Fig. 1, c, g, k and o).
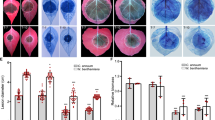
Sterility in phytoplasma-infected and tengu-transgenic plants.
(a), 27-day-old wild-type (Col-0) and 35S::tengu plants. (b–e), Inflorescence stems of Col-0 (b), onion yellows phytoplama (OY)-infected (c) and 35S::tengu plants exhibiting severe sterility (d) and mild sterility (e). (f–i), Flowers of Col-0 (f), OY-infected (g) and 35S::tengu plants exhibiting severe sterility (h) and mild sterility (i). (j–m), Anther development in mature Col-0 (j), OY-infected (k) and 35S::tengu plants exhibiting severe sterility (l) and mild sterility (m). (n–q), Scanning electron micrographs of apices of gynoecia of mature Col-0 (n), OY-infected (o), 35S::tengu plants exhibiting severe sterility (p) and mild sterility (q). Scale bars are 100 µm.
After self-crossing of 35S::tengu plants with severe sterility, the siliques of pistils of 35S::tengu did not elongate and almost all 35S::tengu fruits with severe sterility were seedless (Fig. 2a, upper panel). After self-crossing of 35S::tengu plant with mild sterility, the pistils frequently produced shorter siliques (Fig. 2a, middle panel). Among 171 fruits from 35S::tengu plants with mild sterility, 108 (63.2%) were fecund and 63 (36.8%) were infertile. In addition, tengu-transgenic pistils were pollinated with wild-type pollens using reciprocal hand-pollination (data not shown). Pollen grains from mild-sterile tengu-transgenic flowers were smaller with exine wall sculpturing (Fig. 2b), indicating that 35S::tengu plants showing mild sterility are male sterile.
Phenotypes of tengu-transgenic plants.
(a), Flowers and fruits of Col-0 and 35S::tengu plants exhibiting severe sterility and mild sterility. Flower buds and fruits from a single inflorescence are arranged from youngest to oldest. Scale bars are 1 cm. (b), Scanning electron micrographs of pollen grains of wild-type and 35S::tengu flowers exhibiting mild sterility. Scale bars are 10 µm. (c), Relative TENGU gene expression in 35S::tengu plants exhibiting severe sterility and mild sterility. The mRNA levels were determined using quantitative RT-PCR, normalized to an internal Actin2 gene control. Data are represented as the means of biological triplicates. Error bars represent the standard error of the mean. * indicates a significant difference (Student's t-test; P < 0.05) from 35S::tengu plants exhibiting severe sterility.
To evaluate the effect of TENGU expression on the severity of sterility among transformants, we examined the levels of tengu mRNA in 35S::tengu plants using quantitative reverse transcription polymerase chain reaction (qRT-PCR). The level of tengu transcripts in 35S::tengu plants with severe sterility was significantly higher than that in 35S::tengu plants with mild sterility (Fig. 2c), indicating that the severity of sterility in the transgenic plants is determined in a TENGU-dependent manner. In subsequent studies, we used the 35S::tengu plants with severe sterility to investigate TENGU function.
Altered expression of auxin response factor genes in TENGU transgenic flowers
Two paralogous ARF genes, ARF6 (At1g30330) and ARF8 (At5g37020), are known to promote flower maturation8,9,18. Developmental abnormalities in tengu-transgenic plants were very similar to those observed in double knockout mutants of ARF6 and ARF8, including the production of infertile closed buds, resulting from arrested flower development9 (Fig. 1). To determine whether ARF6 and ARF8 are involved in the severe sterility phenotypes, we examined ARF6 and ARF8 expression levels in OY-infected and severe-sterile tengu-transgenic flowers using qRT-PCR analysis. Notably, the mRNA levels of both ARF6 and ARF8 significantly decreased in both the OY-infected and tengu-transgenic flowers (Fig. 3), suggesting that TENGU represses ARF6 and ARF8 gene expression. Despite the fact that transcripts of ARF6 and ARF8 are targeted by microRNA (miR) 167 for cleavage18, the levels of miR167 were only slightly higher in the OY-infected and tengu-transgenic plants than in wild-type plants (see Supplementary Fig. S1 online), suggesting that the repression of ARF6 and ARF8 expression in the OY-infected and tengu-transgenic plants is not due solely to increased miR167 transcription.
Relative expression of ARF genes.
Relative expression of ARF6 and ARF8 genes in OY-infected and tengu-transgenic plants. The mRNA levels were determined using qRT-PCR, normalized to an internal Actin2 gene control. Data are represented as the means of biological triplicates. Error bars represent the standard error of the mean. Asterisk indicates significant differences (Student's t-test; **P < 0.01) from the wild-type (Col-0).
Altered expression of floral maturation genes in TENGU transgenic flowers
The expression of a gene encoding a putative enzyme involved in the jasmonate biosynthesis pathway, LOX2 (At3g45140), is repressed in the flowers of ARF6 or ARF8 mutant plants10. Expression of the MYB21 (At3g27180) and MYB24 (At5g40350) genes is upregulated by JA and induced downstream of ARF6 and ARF8 expression10. Therefore, we examined the expression levels of these genes in OY-infected and tengu-transgenic flowers using qRT-PCR analysis. In tengu-transgenic plants, the level of LOX2 mRNA significantly decreased (Fig. 4a). The level of LOX2 also slightly decreased in the OY-infected plants compared with wild-type plants (Fig. 4a). Levels of MYB21 and MYB24 mRNAs were significantly lower in tengu-transgenic and OY-infected plants than in wild-type plants (Fig. 4b). The transcriptional repression of JA-related genes in OY-infected and tengu-transgenic plants were consistent with the previously observed expression patterns in ARF6 and/or ARF8 mutant plants.
Relative expression of floral maturation genes.
Relative expression of LOX2 (a), MYB21 and MYB24 (b) genes in OY-infected and 35S::tengu transgenic plants. The mRNA levels were determined using qRT-PCR. Data are represented as the means of biological triplicates. Error bars represent the standard error of the mean. Asterisks indicate significant differences (Student's t-test; **P < 0.01; *P < 0.05) from the wild-type (Col-0).
Reduction of endogenous phytohormone levels in TENGU transgenic flowers
Given the very low JA concentration in flowers of the double knockout mutant of ARF6 and ARF89,19, we examined whether TENGU downregulates JA synthesis. We determined the cis-JA and active jasmonoyl-isoleucine (JA-Ile) levels in wild-type and tengu-transgenic buds. The levels of both cis-JA and JA-Ile were significantly lower in tengu-transgenic buds than in wild-type buds (Fig. 5, a and b). By contrast, the cis-JA and JA-Ile levels in seedlings were not significantly different between tengu-transgenic and wild-type plants (Fig. 5, d and e), consistent with the high expression level of ARF6 and ARF8 in flowers20. A significant decrease in the total levels of cis-JA and JA-Ile in tengu-transgenic buds indicated that TENGU expression reduced JA synthesis.
Quantification of the endogenous hormone levels.
Quantification of the endogenous phytohormone levels in buds (a–c) and in 10-day-old seedlings (d and e). (a and d), cis-JA, (b and e), JA-Ile and (c), IAA. All values represent the means of four biological replicates (n = 10 buds). Error bars represent the standard error of the mean. Asterisks indicate significant differences (Student's t-test; **P < 0.01; *P < 0.05) between the wild-type and tengu-transgenics.
TENGU was suggested previously to interfere with auxin signaling or biosynthesis pathways in A. thaliana4. Auxin presumably induces ARF6 and ARF8 activity by destabilizing auxin/indole-3-acetic acid (Aux/IAA) transcriptional repressor proteins10,21,22. To investigate whether TENGU affects the auxin production, we measured the IAA levels in wild-type and tengu-transgenic buds. The IAA levels were significantly lower in tengu-transgenic buds than in wild-type buds (Fig. 5c), suggesting that TENGU also reduced the endogenous levels of auxin.
Discussion
Our data indicate that TENGU is an effector protein with pleiotropic effects on at least two phytohormones, auxin and JA, resulting in plant sterility. In this study, we demonstrated that transgenic expression of the phytoplasmal effector TENGU induced developmental abnormalities in Arabidopsis flowers similar to those in double knockout mutants of ARF6 and ARF8 and that both ARF6 and ARF8 transcripts were significantly diminished in tengu-transgenic plants (Figs. 1,2,3). We further showed that JA and JA-Ile levels were significantly decreased in buds of tengu-transgenic plants (Fig. 5).
Since the IAA levels significantly decreased in the tengu-transgenic plants (Fig. 5c), repression of the auxin signaling pathway by TENGU may also contribute to plant sterility (Fig. 6). Understanding the roles of auxin in plant reproduction is quite limited, however, increased IAA content was shown to alter male development in tobacco11 and auxin was shown to contribute to the coordination of male and female gametophytes in orchids23. Moreover, recent studies demonstrated that some A. thaliana mutants deficient in auxin biosynthesis genes exhibited infertile phenotypes24. TENGU is thought to downregulate early auxin-responsive genes4, suggesting that repression of the auxin signaling pathway by TENGU may play a role in flower sterility.
A model for TENGU-induced reduction of the endogenous levels of JA and auxin.
Black arrows represent mRNA, protein, or phytohormone products. Blue dashed T-bars indicate the suggested molecular events leading to the repression of ARF gene expressions and auxin synthesis. Gray letters represent negative effects of TENGU on each part of the floral maturation pathway.
Based on these observations, although it remains unclear the molecular interaction between TENGU peptide and its unknown target, we can formulate some hypothesis on the targets of TENGU. TENGU may target the auxin biosynthesis and/or signaling pathways resulting in the repression of ARF6 and ARF8 expression and the reduction of JA production. Auxin is presumed to induce ARF6 and ARF8 activity10,21,22. Another interesting possibility is that TENGU might recognize or interact with an unknown regulatory factor of ARF6 and ARF8 and finally interfere with the auxin signaling pathway. Recently, one of the Arabidopsis auxin biosynthesis genes YUCCA9 was shown to be upregulated by JA25, indicating that the JA pathway is associated with auxin homeostasis. Further studies are required to establish whether TENGU directly or indirectly represses ARF6 and ARF8 expression via reduced auxin production.
We demonstrated that cis-JA and JA-Ile levels were decreased in tengu-transgenic plants (Fig. 5). Moreover, both of the phytoplasmal effectors TENGU and SAP11, a secreted protein from the aster yellows strain witches' broom (AY-WB) phytoplasma, repressed JA biosynthesis in transgenic A. thaliana26,27. In plants, the JA signaling pathway plays a central role in regulating defense responses to insects28. Yang et al. (2008) demonstrated that the tomato yellow leaf curl China virus (TYLCCNV)-encoded protein βC1 suppresses a subset of JA-responsive genes, which contribute to accelerate the increase the insect vector population in TYLCCNV-infected plants29. This indicated that the manipulation of JA levels by plant pathogens might contribute to enhanced insect vector reproduction. In fact, SAP11 leads to reduced JA biosynthesis and an increase in the colonization ability of the AY-WB insect vector27. Based on the inhibition of JA synthesis by TENGU, our results suggest that phytoplasmas reduces the endogenous JA levels via effectors to attract their insect vectors.
Note that two phytoplasmal effectors regulate JA biosynthesis in a different manner. SAP11 destabilizes the TCP transcription factors (TEOSINTEBRANCHED, CYCLOIDEA, PROLIFERATION FACTOR 1 AND 2) that lead to the repression of JA biosynthesis. By contrast, we demonstrated in this study that the TENGU-induced reduction in JA synthesis is mediated by ARF6 and ARF8, but not by TCPs. TENGU-transgenic plants did not have serrated leaves caused by the repression of TCPs in SAP11-transgenic plants27 (Fig. 1a). Moreover, ectopic expression of SAP11 in A. thaliana led to decreased JA synthesis only after wounding27. By contrast, ectopic expression of TENGU reduced the constitutive production of JA and JA-Ile even in healthy plants and is not likely to be involved in wound-induced pathways (Fig. 5). Thus, the three different effectors βC1, SAP11 and TENGU decrease the JA levels through different mechanisms.
JA and salicylic acid (SA) are important signaling molecules involved in various aspects of plant defense responses and the JA- and SA-mediated signaling pathways are mutually antagonistic30. Microarray analysis revealed that during phytoplasma infection, a marker gene of the SA signaling pathway, pathogenesis-related proteins 1 (PR-1), was strongly induced in phytoplasma-infected grapevine, suggesting that SA is involved in the response to phytoplasma infection31. However, the expression of PR-1 was not significantly induced in tengu-transgenic plants4, indicating that repression of JA synthesis in tengu-transgenic flowers is not likely the result of antagonism between SA and JA.
We demonstrated that endogenous auxin content decreased in tengu-transgenic plants (Fig. 5), consistent with previously reported microarray data4. Similarly, several effectors from the well-studied pathogenic bacterium Pseudomonas syringae pv. tomato DC3000 have been shown to modulate phytohormone levels32. P. syringae-induced miR393 negatively regulates mRNAs encoding auxin receptors thereby reducing auxin signaling, which is thought to antagonize the SA pathway33. By contrast, as discussed above, the reduction in endogenous auxin levels in tengu-transgenic plants appears to be independent of the SA signaling pathway, suggesting that TENGU may control the endogenous levels of the two phytohormones independent of SA.
Most of phytoplasma species make host plants unable to produce seeds by inducing flower sterility, phyllody and virescence. In this study, we demonstrated that phytoplasma-induced male and female sterility is caused by perturbation of the JA and auxin signaling pathways. Disruption of JA synthesis may be beneficial for phytoplasmas by attracting more insect vectors and increasing the colonization of phytoplasmas themselves. However, it can also result in plant sterility, which may be disadvantageous to the prosperity of phytoplasmas. Therefore, TENGU may balance the advantages of insect attraction and the disadvantages of plant sterility to maximize the fitness of phytoplasmas that can survive in both insects and plants to live in. Similarly, severe phyllody leads to prolonged plant life span and efficient colonization by insect vectors5,6. Therefore, sterility of flowers and phyllody appear to be common features of the manipulation of plants by phytoplasmas to attract insect vectors.
In conclusion, we demonstrated the phytoplasmal effector TENGU induces plant sterility. TENGU causes not only morphological changes during vegetative growth, but also developmental defects upon flowering by modulating of endogenous two phytohormone levels, JA and auxin.
Methods
Plant Material and Growth Conditions
A. thaliana accession Col-0 and transgenic plants were grown and maintained under a 15 h light/9 h dark photoperiod at 23°C. To inoculate A. thaliana with onion yellows phytoplasma (OY), plants with four to five rosette leaves were covered with plastic and mesh cages and seven OY-infected leafhoppers (Macrosteles striifrons) were released into each cage and then removed after seven days. Infection with OY was confirmed in all inoculated plants by PCR. For phenotypic analysis, we used multiple lines of tengu-transgenic A. thaliana exhibiting sterility. We scored fruits that elongated to greater than 6.0 mm in length as fertile because plants that produce fruits that elongate to greater than 5.5–6.0 mm in length are scored as fertile in A. thaliana8.
Scanning Electron Microscopy (SEM)
Carpels and pollen grains were fixed in 0.2 M phosphate buffered 4% glutaraldehyde at pH7.2 overnight at 4°C and then dehydrated through an ethanol series. The dehydrated specimens were immersed in isoamyl acetate. They were then critical point-dried under carbon dioxide (JCPD-5; JEOL, Tokyo, Japan), sputter-coated with platinum-palladium (Hitachi E-1030; Hitachi, Tokyo, Japan) and examined with a scanning electron microscope (SEM; Hitachi S-4000; Hitachi) at 10 kV.
RNA Extraction and Northern Blot Analysis
Total RNA was extracted from plants using ISOGEN reagent (Nippon Gene, Tokyo, Japan). Northern blot analysis of miRNAs was performed as described previously34. A primer specific to the miR167 (5′-TAGATCATGCTGGCAGCTTCA-3′) was digoxigenin (DIG)-labeled using a DIG Oligonucleotide Tailing Kit, 2nd Generation (Roche Diagnostics, Mannheim, Germany) and used as a probe for northern blot hybridization.
Gene Expression Analysis
Real-time PCR assays were performed as described previously35 with minor modifications. Fold changes were calculated using the expression of a housekeeping ACTIN2 gene as an internal control. Three biological replicates were performed for each experiment. The Actin236 mRNA levels showed only minor variations between wild-type and tengu-transgenic inflorescence stems, similar to those of two previously used housekeeping genes, U-box domain containing protein and EF-1α27,37; therefore only the data normalized against the Actin2 gene are presented in this study.
The gene-specific primer sequences (5′-3′) are as follows: qTENGU_F, TGATGATATTGAAAACGTGATAACTC; qTENGU_R, GCCCTTTTGCAATAAATCTTG; qAtARF6_F, GTGGGATCGAGGACTCCAATC; qAtARF6_R, CCCGACGTATCAAGTCTCGG; qAtARF8_F, TCAAGAACTGATTGCAAGGGATC; and qAtARF8_R, GAGATGCCGTTTGGGCTG. LOX2, MYB21 and MYB24 genes were amplified using primers described previously10.
Quantification of Phytohormones
Phytohormones were prepared and quantified according to the method of Tokuda et al. (2013)38. IAA, JA and JA-Ile were extracted and purified using solid-phase extraction. The stable isotope-labeled compounds D2-IAA (Sigma-Aldrich, St Louis, MO, USA) and D2-JA (Tokyo Kasei, Tokyo, Japan) were used as internal standards. 13C6-JA-Ile was synthesized as described previously39.
For quantification of phytohormones, we used stage 12-13 wild-type buds and tengu-transgenic infertile buds. To measure phytohormone levels simultaneously, 10 buds were lyophilized using a TissueLyser (Qiagen, Venlo, The Netherlands). Phytohormones were extracted and analyzed using a liquid chromatography-electrospray ionization-tandem mass spectrometry (LC/ESI/MS/MS; Agilent 6410; Agilent Technologies, Santa Clara, CA, USA) as described previously40 and quantified using the MassHunter vB.03.01 spectrometer software (Agilent Technologies).
References
Christensen, N. M., Axelsen, K. B., Nicolaisen, M. & Schulz, A. Phytoplasmas and their interactions with hosts. Trends Plant Sci. 10, 526–535 (2005).
Hogenhout, S. A., Oshima, K., Ammar, E. D., Kakizawa, S., Kingdom, H. N. & Namba, S. Phytoplasmas: bacteria that manipulate plants and insects. Mol. Plant Pathol. 9, 403–423 (2008).
Yong, E. Bacterial tricks for turning plants into zombies. Nature news, 10.1038/nature.2014.15011 (2014).
Hoshi, A. et al. A unique virulence factor for proliferation and dwarfism in plants identified from a phytopathogenic bacterium. Proc. Natl. Acad. Sci. USA 106, 6416–6421 (2009).
MacLean, A. M. et al. Phytoplasma effector SAP54 induces indeterminate leaf-Like flower development in Arabidopsis plants. Plant Physiol. 157, 831–841 (2011).
MacLean, A. M. et al. Phytoplasma effector SAP54 hijacks plant reproduction by degrading MADS-box proteins and promotes insect colonization in a RAD23-dependent manner. PLoS Biol. 12, e1001835 (2014).
Maejima, K. et al. Recognition of floral homeotic MADS-domain transcription factors by a phytoplasmal effector, phyllogen, induces phyllody. Plant J. 78, 541–554 (2014).
Vivian-Smith, A., Luo, M., Chaudhury, A. & Koltunow, A. Fruit development is actively restricted in the absence of fertilization in Arabidopsis. Development 128, 2321–2331 (2001).
Nagpal, P. et al. Auxin Response Factors ARF6 and ARF8 promote jasmonic acid production and flower maturation. Development 132, 4107–4118 (2005).
Reeves, P. H. et al. A regulatory network for coordinated flower maturation. PLoS Genet. 8, e1002506 (2012).
Spena, A. et al. Anther-specific expression of the rolB gene of Agrobacterium rhizogenes increases IAA content in anthers and alters anther development and whole flower growth. Theor. Appl. Genet. 84, 520–527 (1992).
Abramovitch, R. B., Anderson, J. C. & Martin, G. B. Bacterial elicitation and evasion of plant innate immunity. Nat. Rev. Mol. Cell Biol. 7, 601–611 (2006).
Jung, H. -Y. et al. ‘Candidatus Phytoplasma oryzae’, a novel phytoplasma taxon associated with rice yellow dwarf disease. Int. J. Syst. Evol. Microbiol. 53, 1925–1929 (2003).
Pracros, P., Renaudin, J., Eveillard, S., Mouras, A. & Hernould, M. Tomato flower abnormalities induced by stolbur phytoplasma infection are associated with changes of expression of floral development genes. Mol. Plant Microbe Interact. 19, 62–68 (2006).
Alcantara-Mendoza, S. et al. Detection and evaluation of Maize Bushy Stunt phytoplasma in the state of Veracruz, Mexico. Revista Mexicana de fitopatologia 28, 34–43 (2010).
Cousin, M. T. & Abadie, M. Action des mycoplasmes sur l'anthère. Etude en microscopies photonique et électronique. Rev. Cytol. Biol. Végét-Bot. 5, 41–57 (1982).
Urbanaviciene, D., Jomatiene, R., Valiunas, D. & Davis, R. E. Molecular detection of phytoplasmas in oats, barley and Triticosecale and their classification based on 16S rRNA gene polymorphisms. Zemes Ukio Mokslai 3, 15–19 (2004).
Wu, M. F., Tian, Q. T. & Reed, J. W. Arabidopsis microRNA167 controls patterns of ARF6 and ARF8 expression and regulates both female and male reproduction. Development 133, 4211–4218 (2006).
Tabata, R. et al. Arabidopsis AUXIN RESPONSE FACTOR6 and 8 regulate jasmonic acid biosynthesis and floral organ development via repression of class 1 KNOX genes. Plant Cell Physiol. 51, 164–175 (2010).
Pitaksaringkarn, W., Ishiguro, S., Asahina, M. & Satoh, S. ARF6 and ARF8 contribute to tissue reunion in incised Arabidopsis inflorescence stems. Plant Biotech. 31, 1–5 (2014).
Ulmasov, T., Hagen, G. & Guilfoyle, T. J. Activation and repression of transcription by auxin-response factors. Proc. Natl. Acad. Sci. USA 96, 5844–5849 (1999).
Tatematsu, K. et al. MASSUGU2 encodes Aux/IAA19, an auxin-regulated protein that functions together with the transcriptional activator NPH4/ARF7 to regulate differential growth responses of hypocotyl and formation of lateral roots in Arabidopsis thaliana. Plant Cell 16, 379–393 (2004).
Zhang, X. S. & O'Neill, S. D. Ovary and gametophyte development are coordinately regulated by auxin and ethylene following pollination. Plant Cell 5, 403–418 (1993).
Mashiguchi, K. et al. The main auxin biosynthesis pathway in Arabidopsis. Proc. Natl. Acad. Sci. USA 108, 18512–18517 (2011).
Hentrich, M. et al. The jasmonic acid signaling pathway is linked to auxin homeostasis through the modulation of YUCCA8 and YUCCA9 gene expression. Plant J. 74, 626–637 (2013).
Bai, X. et al. AY-WB Phytoplasma secretes a protein that targets plant cell nuclei. Mol. Plant Microbe Interact. 22, 18–30 (2009).
Sugio, A., Kingdom, H. N., MacLean, A. M., Grieve, V. M. & Hogenhout, S. A. Phytoplasma protein effector SAP11 enhances insect vector reproduction by manipulating plant development and defense hormone biosynthesis. Proc. Natl. Acad. Sci. USA 108, 1254–1263 (2011).
De Vos, M., Kim, J. H. & Jander, G. Biochemistry and molecular biology of Arabidopsis–aphid interactions. Bioessays 29, 871–883 (2007).
Yang, J. Y. et al. βC1, the pathogenicity factor of TYLCCNV, interacts with AS1 to alter leaf development and suppress selective jasmonic acid responses. Genes Dev. 22, 2564–2577 (2008).
Kunkel, B. N. & Brooks, D. M. Cross talk between signaling pathways in pathogen defense. Curr. Opin. Plant Biol. 5, 325–331 (2002).
Albertazzi, G. et al. Gene expression in grapevine cultivars in response to Bois Noir phytoplasma infection. Plant. Sci. 176, 792–804 (2009).
Cunnac, S., Lindeberg, M. & Collmer, A. Pseudomonas syringae type III secretion system effectors: repertoires in search of functions. Curr. Opin. Microbiol. 12, 53–60 (2009).
Navarro, L. et al. A plant miRNA contributes to antibacterial resistance by repressing auxin signaling. Science 312, 436–439 (2006).
Senshu, H. et al. Variability in the level of RNA silencing suppression caused by triple gene block protein 1 (TGBp1) from various potexviruses during infection. J. Gen. Virol. 90, 1014–1024 (2009).
Senshu, H. et al. A dual strategy for the suppression of host antiviral silencing: two distinct suppressors for viral replication and viral movement encoded by potato virus M. J. Virol. 85, 10269–10278 (2011).
Okano, Y. et al. In planta recognition of a double-stranded RNA synthesis protein complex by a Potexviral RNA silencing suppressor. Plant Cell 26, 2168–2183 (2014).
Curaba, J. et al. AtGA3ox2, a key gene responsible for bioactive gibberellin biosynthesis, is regulated during embryogenesis by LEAFY COTYLEDON2 and FUSCA3 in Arabidopsis. Plant J. 136, 3660–3669 (2004).
Tokuda, M. et al. Phytohormones related to host plant manipulation by a gall-inducing leafhopper. PLoS ONE 8, e62350 (2013).
Jikumaru, Y. et al. Preparation and biological activity of molecular probes to identify and analyze jasmonic acid-binding proteins. Biosci. Biotechnol. Biochem. 68, 1461–1466 (2004).
Yoshimoto, K. et al. Autophagy negatively regulates cell death by controlling NPR1-dependent salicylic acid signaling during senescence and the innate immune response in Arabidopsis. Plant Cell 21, 2914–2927 (2009).
Acknowledgements
The authors would like to thank Professor Kyoko Watanabe of Tamagawa University for her kind support in SEM photography. We are also grateful to Dr. Hiroko Senshu of our laboratory and Dr. Kyoko Sugawara of The University of Tokyo for numerous helpful discussions. This work was supported in part by the Japan Society for the Promotion of Science (JSPS) through a grants-in-aid for JSPS fellows (Grant 24-7554) and for Scientific Research (S) (Grant 25221201), the Funding program for Next Generation World-Leading Researchers (project GS005) initiated by the Council for Science and Technology Policy (CSTP) and by the Program for Promotion of Basic Research Activities for Innovative Bioscience (PROBRAIN).
Author information
Authors and Affiliations
Contributions
N.M., M.H., A.H., H.K., K.M., K.K., A.Y., Y.K. and S.N. designed research; N.M., A.H. and Y.T. performed research; N.M. and K.O. analyzed data; and N.M., M.H. and Y.Y. wrote the paper. All authors discussed the results and commented on the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Figure S1
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Minato, N., Himeno, M., Hoshi, A. et al. The phytoplasmal virulence factor TENGU causes plant sterility by downregulating of the jasmonic acid and auxin pathways. Sci Rep 4, 7399 (2014). https://doi.org/10.1038/srep07399
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07399
This article is cited by
-
The Role of Plant Defense Signaling Pathways in Phytoplasma-Infected and Uninfected Aster Leafhoppers’ Oviposition, Development, and Settling Behavior
Journal of Chemical Ecology (2024)
-
Draft genome sequence of 'Candidatus Phytoplasma asteris,’ strain SW86 associated with sandal spike disease (SSD)
3 Biotech (2024)
-
Response of Prunus species to graft-inoculation by two Iranian strains of almond witches’-broom phytoplasma
Journal of Plant Pathology (2022)
-
Biological characterization and evolution of a novel RNA silencing suppressor SWP16 from wheat blue dwarf (WBD) phytoplasma
Australasian Plant Pathology (2022)
-
Genome-wide identification and expression of the lipoxygenase gene family in jujube (Ziziphus jujuba) in response to phytoplasma infection
Journal of Plant Biochemistry and Biotechnology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.