Abstract
ABT-737 inhibits the anti-apoptotic proteins B-cell lymphoma 2 (BCL-2) and BCL-XL. Meayamycin B switches the splicing pattern of myeloid cell leukemia factor 1 (MCL1) pre-mRNA. Specifically, inhibition of splicing factor 3B subunit 1 (SF3B1) with meayamycin B promotes the generation of the proapoptotic, short splicing variant (MCL1-S) and diminishes the antiapoptotic, long variant (MCL1-L). This action was previously associated with the cytotoxicity of meayamycin B in non-small cell lung carcinoma cell lines. ABT-737 induced apoptosis in response to an ablation of MCL1-L by meayamycin B. In this study, we further exploited this synergistic combination in head and neck squamous cell carcinoma (HNSCC), up to 90% of which overexpress MCL1 and BCL-XL. In a panel of seven HNSCC cell lines, the combination of meayamycin B and ABT-737 rapidly triggered a Bax/Bak-mediated apoptosis that overcame the resistance from HPV16-positive HNSCC against each agent alone. Both RT-PCR and Western blotting showed that meayamycin B up-regulated MCL1-S and down-regulated MCL1-L. Significantly, we discovered that SF3B1 was involved in the splicing of oncogenic HPV16 E6 to produce non-oncogenic HPV16 E6*, indicating that SF3B1 may inhibit HPV16-induced tumorigenesis.
Similar content being viewed by others
Introduction
Head and neck squamous cell carcinoma (HNSCC) is the sixth most common cancer worldwide1. HNSCC mostly occurs by cigarette/alcohol consumption; however, a subgroup of HNSCC (26%) is caused by the integration of high-risk human papillomavirus (HPV), especially HPV162. It has been perplexing that only ~10% of those who were exposed to high-risk HPV16 or 18 develop cancer3. In oropharyngeal carcinoma particularly, HPV16 is detected in over 60% of the cancer tissues4. Due to their distinct mechanisms of tumorigenesis, HPV-positive and HPV-negative HNSCC should be treated separately5,6. Platinum-based chemotherapy, the most commonly used drug regardless of HPV status, is far less effective against HPV-negative HNSCC7,8. Moreover, in the past thirty years, the five-year overall survival rate of HNSCC has not improved9, warranting the discovery of new pharmacological interventions.
The discovery of novel therapeutics requires a comprehensive understanding of the disease biology. Since HPV16 is present in nearly 60% of all cervical cancers10, its expression is most widely studied in cervical cancer models. The HPV16 genome encodes six early (E) viral proteins (E1, E2, E4, E5, E6 and E7) and two late (L) viral capsid proteins (L1 and L2), accompanying the differentiation stages of the host keratinocytes11. Among these proteins, only the overexpression of E6 and E7 was correlated with malignant transformation12. At the transcription level, E6 and E7 share a mutual early p97 promoter. E6 is alternatively spliced to generate the full-length E6 and E6* lacking intron 1 (Figure 1A). Because the open reading frames of E6 and E7 are only 2 nucleotides apart, the transcription of E6 prohibits the translation of E7 protein. Otherwise, when intron 1 is excised, splicing variant E6* allows the translation of E713,14. However, there is evidence showing that the translation of E7 is independent of the splicing of E615. In response to epidermal growth factor, the activation of ERK1/2 pathway facilitates the production of E6. SiRNA knockdown experiments demonstrated that hnRNP A1 and hnRNP A2/B1 favored E6, while splicing factor Sam68 and transcription factor Brm favored E6*16. This report indicated that splicing and transcription are coupled in HPV16 E6/E7 expression.
Functionally, E6 inhibits apoptosis by triggering the degradation of p53 and E7 enhances cell proliferation by binding to another tumor suppressor protein retinoblastoma-associated protein (pRb)17,18. Whereas the function of full-length E6 is relatively well understood, the function of E6* is still elusive and available data lend controversial conclusions; through RT-PCR analysis, E6* expression was found to be significantly higher in late stage cervical lesion than early stage counterparts19,20. In contrast, E6* counteracts the anti-apoptotic action of E6 in the degradation of p53 and precaspase 821,22. Additionally, the overexpression of E6* is cytotoxic when expressed in immortalized monkey fibroblasts CV-1 cells23. In sum, these unconnected pieces of data argue that both basic and translational studies of E6 splicing are lagging, calling for further studies advanced by novel strategies.
Meayamycin B (Figure 1B), the most potent inhibitor of splicing factor 3B (SF3B), sensitized non-small cell lung carcinoma cells (A549 and H1299) to ABT-737 (Figure 1B) by modulating the splicing of Mcl-124. Similar observations were reported for the combination of spliceostatin A (structurally similar to meayamycin B) and ABT-737 in neuroblastoma25. Given the mutual risk factors, such as alcohol and cigarette consumption, shared between non-small cell lung carcinoma and HNSCC26,27, we reasoned that meayamycin B might also prime HNSCC cells for ABT-737. Therefore, we wished to examine the anticancer activity of meayamycin B, as a single-agent or in combination with ABT-737. In addition, we used meayamycin B to examine the role that SF3B subunit 1 (SF3B1) plays in the splicing of MCL1 and more importantly, HPV16 E6.
Methods
Cell lines
Four HPV16-positive HNSCC cell lines, UD-SSC2 (gift from Professor Henning Bier, University of Dusseldorf, Germany)28, UM-SCC47 (gift from Professor Thomas Carey, University of Michigan)29,30, 93-VU-147T (gift from Professor Hans Joenje, VU Medical Center, The Netherlands)31, UPCI:SCC9032 and the HPV16-negative HNSCC cell lines PCI-13 and PCI-15B (gifts from Professor Theresa Whiteside, University of Pittsburgh Cancer Institute)33 and UM-SCC22B (gift from Professor Thomas Carey, University of Michigan)34 were used throughout the study. Bax−/−/Bak−/− and wild-type mouse embryonic fibroblasts (MEFs) were gifts from the late Professor Korsmeyer (Harvard Medical School) through Professor Scott Oakes (University of California, San Francisco)35. 93-VU-147T cell line was cultured in DMEM/F12 medium (Gibco, New York) and other cell lines were cultured in Dulbecco's Modified Eagle's Medium (Gibco, New York). All cell culture media were supplemented with 10% fetal bovine serum, 1% penicillin/streptomycin and 2% L-glutamine. Cells were maintained at 37°C in air containing 5% CO2.
Pharmacodynamic interaction analysis
The cells were exposed to meayamycin B and ABT-737 at various concentrations, either as single agents or in concomitant combinations (at a constant 3:5000 ratio) for 72 h until cell viability was monitored using the MTS (([3-(4,5-dimethylthiazol-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-tetrazolium, inner salt])) method. Bliss expected effect (EEXP, expressed as cell viability) from the combination of meayamycin B and ABT-737 were calculated using EEXP = EA + EB − EA*EB, where EA and EB are effects from each single-agent treatment at a given dose. The difference between the Bliss expected and the observed effect (EOBS) of the combination is ΔE = EEXP − EOBS (Figure 2A). ΔE scores were summed across the dose range to generate a Bliss sum. Bliss sum = 0 indicates that the combination treatment is additive (as expected for independent pathway effects); Bliss sum > 0 indicates an activity greater than additive (synergy); and Bliss sum < 0 indicates the combination is less than additive (antagonism).
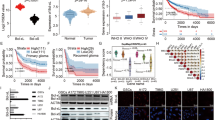
Meayamycin B (MAMB) synergized with ABT-737 in HNSCC cell lines.
(A). 72-h viability assays with MTS using meayamycin B and ABT-737 as single agents or in combination at a constant 3:5000 ratio. B–C. Evaluation of synergistic interaction between meayamycin B and ABT-737 in HNSCC cells using the Bliss independence model. (B). 3D graph of EEXP – EOBS expressed as differences in the percentage of viable cells that are treated by the combination. (C). Bliss sums demonstrate strong synergism in HNSCC cell lines (n = 3). The data represent results from three separate experiments and each data point constitutes mean ± SD.
Annexin V - fluorescein isothiocyanate (FITC) and propidium iodide (PI) staining
In all cases, cells were treated with 3 nM meayamycin B and 5 μM ABT-737 (alone or in combination), an equal volume of DMSO as a negative control, or 133 μM cisplatin as a positive control. Cells were treated for 5 h and stained with Annexin V-FITC (BD Pharmingen, CA) and PI (BD Pharmingen, CA) following the manufacturer's protocol. The cells were then analyzed by flow cytometry using a CyAn cytometer (Beckman Coulter, CA). Data were analyzed using Summit V4.3 software.
Caspase 3/7 activity assays
The cells, seeded at 1.0 × 104 per well in triplicate in white solid-bottom 96-well plates, were treated with 3 nM meayamycin B and 5 μM ABT-737 (alone or in combination) or an equal volume of DMSO as a negative control for 9 h. Then, caspase 3/7 activity was monitored using Caspase-Glo® 3/7 reagent (Promega, WI) with a Modulus II Microplate Reader (Promega, WI) following the manufacturers' protocols. The caspase 3/7 activity was expressed as the mean luminescence signal from the compound-treated wells divided by that from vehicle-treated wells.
RNA isolation and RT-PCR
Total RNA was isolated using RNeasy mini kit (QIAGEN, Maryland) and subjected to RT-PCR analysis (1 μg per reaction) using the SuperScript® II reverse transcriptase (Invitrogen, New York). The thermocycler program for the HPV-16 E6 and E7, BCL-X, MCL1 and β-actin involved an initial denaturation at 94°C for 5 min, 30 cycles at 94°C for 30 sec, 58°C for 30 sec, 72°C for 50 sec and a final elongation at 72°C for 7 min. The primer sequences are included in Table S1 (Supporting Information). The PCR products were examined on 1.5% agarose gels and imaged by a Molecular Imager Gel Doc™ XR+ (Bio-Rad, CA). The intensities of the bands stained with ethidium bromide were quantified with the Lab Imager software (Bio-Rad, CA). Of note, the sizes of the splicing variants for HPV16 E6 and Mcl-1 were not factored when calculating the relative abundance of a splicing variant of interest.
siRNA transfection
The pre-designed ON-TARGETplus modified siRNA duplexes targeting SF3B1 (020061-13) was obtained from Dharmacon (Dharmacon, MA). 93-VU-147T and UM-SCC47 cells were seeded (2.5 × 105) into 35-mm dishes and transfected with 125 nM siRNA using lipofectamine 2000 (Invitrogen, NY) for 24 h before usage. siGENOME Non-Targeting siRNA Pool #2 (Dharmacon, MA) was used as a negative control. Transfection efficiency was evaluated by western blotting of SF3B1 protein after 48–72 h post transfection.
Western blotting
Cells were harvested in a cell lysis buffer (50 mM Tris-HCl, 150 mM NaCl, 1% Triton X-100, 0.1% SDS and 1 mM PhCH2SO2F) and a cocktail of protease inhibitors (Roche, IN). Total protein (50 μg) was electrophoresed on a 12% SDS-polyacrylamide gel and transferred to polyvinylidene difluoride membranes (Millipore, MA). Membranes were blocked at room temperature for 1 h with a blocking buffer (5% non-fat dry milk in 10 mM Tris-HCl pH 7.6, 150 mM NaCl, 0.1% Tween-20) and then incubated at 4°C overnight with rabbit anti-MCL1 (5453S), anti-BCL-2 (2870S), anti-β-actin (4970S) (Cell signaling technology, MA), anti-BCL-XL/BCL-XS (sc-634) (Santa Cruz, CA), or mouse anti-SF3B1 (D221-3) (MBL, IL) followed by 1-h incubation with a horseradish peroxidase (HRP)-conjugated anti-rabbit IgG (7074S) or anti-mouse secondary antibodies (Cell signaling technology, MA). Blots were developed with ECL Plus reagents (PerkinElmer Life and Analytical Science, MA) and quantified using an Image Gauge Version 4.0 (FUJIFILM, NJ).
Statistics
Data analysis and graph plotting were carried out using a GraphPad Prism 5.0c for Mac (GraphPad Software). All of the data were presented as mean ± standard deviation and were analyzed by Student's t test or ANOVA followed by the Tukey's multiple comparison; the significance level for all analyses was 5%.
Results
Meayamycin B synergized with ABT-737 to cause apoptosis
We previously reported that meayamycin B sensitized non-small cell lung carcinoma cells (A549 and H1299) to ABT-737 by modulating the splicing of MCL124. Here, we investigated the apoptotic response of a panel of HNSCC cells to the combination of meayamycin B and ABT-737. There are two methods in common use for calculating the dose-response relationship expected from combination therapy when two drugs are assumed to have no interactions: Loewe additivity36 and Bliss independence37. Loewe additivity assumes that two inhibitors act on a target through a similar mechanism, as shown by Chou and Talalay for mutually exclusive enzyme inhibitors38. Bliss independence assumes that inhibitors can bind mutually nonexclusively through distinct mechanisms. The insufficient water solubility of ABT-737 prohibited the generation of dose response curves required for the Loewe method. Therefore, Bliss independence was used to determine drug interactions. To this end, we performed 72-h MTS-based antiproliferation assays in a panel of seven cell lines PCI-13, PCI-15B, UM-SCC22B, UPCI:SCC90, 93-VU-147T, UM-SCC47 and UD-SCC2 (GI50 data shown in Table 1). Meayamycin B only inhibited the growth of 93-VU-147T, UM-SCC47 and UD-SCC2 cells. Intriguingly, the combination of meayamycin B and ABT-737 eradicated these cells (Figure 2A). In others, especially UM-SCC22B and UPCI:SCC90, single-agent meayamycin B induced substantial cell death and the combination with ABT-737 did not enhance this effect. As an evaluation of drug interaction with Bliss independence, ΔE values – expressed as the cell viability-difference between expected and observed combination treatment – demonstrated a significantly higher cell growth inhibition by the combination (Figure 2B). In addition, Bliss sums clustered in the range of 400–600 (Figure 2C), indicating strong synergism. Meayamycin B dropped ABT-737's GI50 values by 1–2 orders of magnitude (Table 1). These data underscore the scope and generality of the efficacy for the combination of meayamycin B and ABT-737 in HNSCC cell lines.
To confirm that the meayamycin B and ABT-737 causes apoptosis, we stained treated cells with Annexin V-FITC and PI and monitored the rates of apoptosis using flow cytometry. To demonstrate an early onset of apoptosis triggered by the combination of the two drugs, we chose 4-h exposure for four HNSCC cell lines (PCI-15B and PCI-13 as HPV16-negative and UM-SCC47 and UPCI:SCC90 as HPV16-positive). After 4-h exposure, we observed 5–10% more Annexin V-positive population (Figure 3, right quadrants) in the combination-treated than in cisplatin-treated (positive control) cells, indicating the rapidity of apoptosis stimulation. Necrosis (Figure 3, upper left quadrants) was also induced as a secondary event in the apoptotic pathway39. As a downstream apoptotic event, activated caspase 3/7 (9-h exposure) experienced 7- to 15-fold increase with more evident changes observed in cells relatively resistant to each single-agent (Figure 4A). Taken together, the Annexin V-FITC/PI staining and the caspase 3/7 activity assays both suggest rapid synergistic apoptosis-stimulation by the combination of meayamycin B and ABT-737 in HNSCC cell lines.
Annexin V-FITC/PI staining.
Effects of meayamycin B and ABT-737 on the induction of early apoptosis (Annexin V-FITC staining) and late apoptosis (PI staining) as measured by flow cytometry. The relative cell populations with positive Annexin V-FITC stains were plotted in each cell line. The data show that within 4 h, the combination of 3 nM MAMB and 5 μM ABT-737 caused significantly higher population of apoptotic cells than cisplatin (n = 3).
Caspase 3/7 activity assays and viability assays.
(A) HNSCC cells exposed to 3 nM meayamycin B and 5 μM ABT-737 as single agents or in combination for 9 h. (B) Wild-type (WT) and Bax−/−/Bak−/− MEFs treated with the combinations of meayamycin B and ABT-737 for 9 h. Data representative of three separate experiments is shown. DKO: double knockout.
To further support the onset of apoptosis mediated by Bax/Bak, we exposed Bax−/−/Bak−/− MEFs and SV40-immortalized wild-type MEFs to the combinations of meayamycin B and ABT-737. We chose 9-h exposure for caspase 3/7 activity assays because it should take more than 4 h to observe the downstream effects of meayamycin B. Similarly to HNSCC, the wild-type, but not Bax−/−/Bak−/− MEFs, showed dose-dependent caspase 3/7 activation upon combination treatment (Figure 4B lower panel). Accordingly, Bax−/−/Bak−/− MEFs were significantly more resistant to the combination treatments (Figure 4B upper panel), suggesting that Bax/Bak-mediated apoptosis is a mechanism of cell killing by the combination of meayamycin B and ABT-737. Of note, other cell death pathways such as autophagy might have contributed to a ~15% decrease of proliferation in Bax−/−/Bak−/− MEFs caused by the combination, since MCL1-elimination triggers autophagic cell death when Bax is defected39,40.
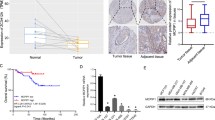
Basal expression of MCL1 correlates with meayamycin B sensitivity
In PCI-13 and 93-VU-147T cells, we previously observed a correlation between the MCL1-L level and the response to meayamycin B. To determine whether basal MCL1-L expression more broadly correlates with meayamycin B sensitivity, we assessed the basal expression of MCL1-L, BCL-X and BCL-2 using Western blotting (Figure 5A). BCL-XL was expressed in all the cell lines at levels statistically unrelated to meayamycin B response; BCL-2 was abundant in UPCI:SCC90. MCL1-L was the dominant MCL1 isoform in all the cell lines except UM-SCC22B, which exclusively expressed MCL1-S. In addition, the total levels of MCL1 in PCI-13, PCI-15B, UM-SCC22B and UPCI:SCC90 were consistently higher than those in the other cell lines (MCL1/β-actin = ~1.5 versus ≤0.8). Interestingly, three out of the four MCL1-abundant cell lines were HPV16-negative.
(A) Constitutive expression of the anti-apoptotic BCL-2 family members in HNSCC in the absence of meayamycin B. Cropped blots show the protein levels of MCL1, BCL-X, BCL-2 and β-actin. The relative expression levels of MCL1-L and BCL-XL were normalized with β-actin. (B) 72-h antiproliferation assays using varying doses of single-agent meayamycin B. Data represent the results of three independent experiments.
The efficacy of meayamycin B against the seven HNSCC cell lines was revisited. Based on their responses to the compound, they fell into two sub-groups; PCI-13, PCI-15B, UM-SCC22B and UPCI:SCC90 were nearly eradicated, while UM-SCC47, UD-SCC2 and 93-VU-147T stopped growing upon meayamycin B exposure at sub- to low-nanomolar concentrations (Figure 5B). With the exception of UPCI:SCC90, all meayamycin B-sensitive cell lines were HPV16-negative, indicating that HPV16 might desensitize cells to meayamycin B. Generally, we observed higher sensitivity toward meayamycin B in HPV16-negative, MCL1-abundant cells.
Regulation of splicing of MCL1 pre-mRNA
MCL1 pre-mRNA is spliced into antiapoptotic MCL1-L (solid line; Figure 6A) and proapoptotic MCL1-S (dotted line). We exposed the cells to meayamycin B either at 3 nM for various periods (1, 9 and 24 h) or for 9 h at various doses (0.03, 0.3 and 3 nM). At the mRNA level, meayamycin B up-regulated MCL1-S and down-regulated MCL1-L in a dose- and time-dependent manner in all tested HNSCC cell lines (Figure 6B and 6C). This reciprocal change of the two isoforms was detectable at 9 h with 0.3 nM or 1 h with 3 nM meayamycin B treatments and peaked within 9 h of 3 nM meayamycin B exposure. Meayamycin B did not generate other splicing variants of MCL1 as evidenced by results from the RT-PCR with an amplicon (for sequences, see Suppl. Table 1) spanning all three exons of MCL1. At the protein level, we observed the same pattern in all the cell lines, with UM-SCC22B as the only exception (Figure 6D and 6E); in UM-SCC22B, RT-PCR showed a switch of MCL1 alternative splicing but Western blotting showed only a dominant MCL1-S upon meayamycin B exposure. This observation indicates that MCL1 expression may be regulated at other steps, such as post-translational modifications, that are specific for this cell line. Also, in comparison to a partial obliteration of MCL1-L caused by single-agent meayamycin B, the cells exposed to the combination for the same period (9 h in PCI-15B) completely lost MCL1, demonstrating the onset of apoptotic cell death (Figure 6F).
Time- and dose-dependent regulation of MCL1 alternative splicing.
(A) Alternative splicing of MCL1. Arrows indicate the locations of the primers for RT-PCR experiments. (B)–(C) Dose-dependent regulation of MCL1 alternative splicing after 9 h meayamycin B-treatment. (D)–(E) Time-dependent regulation of MCL1 alternative splicing by 3 nM meayamycin B. Cropped blots show the relative abundance of each MCL1 splicing variant. (F) Ablation of MCL1 after 9 h of treatment by the combination in PCI-15B. Data shown represent the results of three separate experiments.
Meayamycin B also inhibited the constitutive splicing of MCL1 in UM-SCC22B, PCI-15B, PCI-13 and UM-SCC47 cell lines, as manifested by the accumulation of a transcript without exon 1 (Suppl. Figure S1). Together, these data suggest that meayamycin B decreased antiapoptotic MCL1-L levels by modulating both alternative splicing and a cell type-dependent constitutive splicing.
In parallel, we also examined BCL-X, which is also prone to regulation at the pre-mRNA splicing level in response to anticancer drugs, enhancing the production of proapoptotic BCL-XS24,41. The splicing of BCL-X did not change when the cells were incubated with 3 nM meayamycin B for 1, 9, or 24 h (Suppl. Figure S2 and S3).
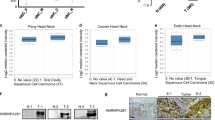
Cell type-dependent inhibition of HPV-16 E6 splicing
During the splicing process, inclusion or exclusion of intron 1 of E6 produces E6 or E6* splicing variant, respectively (Figure 1A). We assessed the levels of E6 and E7 mRNA in meayamycin B- or DMSO-treated HPV16-positive HNSCC cell lines. Figures 7A and B show that the full-length transcript was dominant for E6 in the DMSO-treated cells. In meayamycin B-treated UM-SCC47, 93-VU-147T and UPCI:SCC90 cells, E6 mRNA increased and E6* mRNA decreased in a dose- and time-dependent manner. The inhibition was observed as early as 1 h of meayamycin B-exposure as demonstrated in UPCI:SCC90 cells (Figure 7C). However, this alteration was not as prominent in cell lines constitutively expressing lower levels of E6*, such as UD-SCC2 cells. Due to the unavailability of HPV16 E6 antibody, we did not examine the changes of splicing of E6 at the protein level. In all HPV16-positive cell lines, the E7 mRNA level was unaffected, although E6 and E7 genes are co-transcribed42.
Meayamycin B regulates alternative splicing of HPV16 E6.
(A) Dose-dependent regulation of the constitutive splicing of HPV16 E6 in a cell type-dependent manner by meayamycin B. (B)–(C) Time-dependent regulation of the constitutive splicing of HPV16 E6 by meayamycin B. Data represent results of three independent experiments.
SF3B1 inhibition by siRNA and meayamycin B
We asked whether meayamycin B regulates the splicing of HPV16 E6 and MCL1 exclusively through inhibiting SF3B1. Since none of these patient-derived HNSCC cell lines have been subject to gene transfection, we used HeLa cells to test the transfection methods for SF3B1 knockdown. At 48 and 72 h post transfection, HeLa cells were directly subject to RNA extraction for the detection of MCL1 alternative splicing pattern using RT-PCR. SF3B1-siRNA remarkably switched the dominant MCL1 variant from MCL1-L to MCL1-S, indicating SF3B1 is involved in the alternative splicing of MCL1 in HeLa (Suppl. Figure S4)43.
Subsequently, we knocked down SF3B1 in HPV16-positive UM-SCC47 and 93-VU-147T cell lines. In all cases, cells were collected at 48 h post transfection, because MCL1 was regulated similarly at 48 h and 72 h post SF3B1 knockdown in HeLa cells. As shown in Figure 8A, SF3B1-siRNA effectively eliminated a majority of SF3B1 protein in both cell lines. Figure 8B shows the effect of SF3B1 knockdown and meayamycin B, as single treatment or in combination, on the splicing of HPV16 E6 and MCL1. RT-PCR was used to examine the relative abundance of each splicing variant of E6 and MCL1. Without SF3B1 inhibition, the E6/E6* ratios were 47/53 and 92/8 in UM-SCC47 and 93-VU-147T, respectively. In UM-SCC47, SF3B1 knockdown increased the E6/E6* ratio to 82/18, while non-targeting siRNAs did not alter the ratio. The effect was less evident in 93-VU-147T, probably because of the low basal level of E6*. Analogous to E6, the MCL1-L/MCL1-S ratios in UM-SCC47 and 93-VU-147T were decreased by SF3B1 knockdown from 86/14 and 81/19 to 65/35 and 48/52, respectively. These data indicate SF3B1 is involved in the regulation of the splicing of HPV16 E6 and MCL1.
Inhibition of SF3B1 by siRNA or meayamycin B.
(A) Cropped blots show the knock down of SF3B1 48 h after transfection of the siRNA. (B) RT-PCR analysis of the levels of splicing variants of HPV16 E6 and MCL1 48 h after transfection of siRNA, with or without 3 h of treatment with meayamycin B. Data represent results of three independent experiments.
Meayamycin B caused the splicing alterations to the E6 and MCL1 in HPV16-positive HNSCC cells. Hence, we asked whether such effect from meayamycin B (Figures 6 and 7) was solely via SF3B1 inhibition. We exposed UM-SCC47 and 93-VU-147T cells treated with SF3B1 siRNA or non-targeting siRNAs to 1 nM meayamycin B for 3 h. Interestingly, although meayamycin B alone only caused negligible changes to the splicing of E6 and MCL1, the drug enhanced the levels of changes caused by SF3B1 knockdown (Figure 8B). This was especially prominent in UM-SCC47 cells, with the E6/E6* ratios increased from 82/18 to 90/10 and MCL1-L/MCL1-S ratios increased from 65/35 to 23/77. The fact that short-term treatment with low-dose meayamycin B could further change the splicing pattern beyond silencing SF3B1 might be explained by the different modes of inhibition by pharmacological and biological techniques. For a protein target such as SF3B1 that interacts with RNAs and proteins, an siRNA knockdown eliminates the entire protein entity, thereby obliterating these interactions. In contract, meayamycin B might inhibit a subset of the functions of the target, causing partial disturbance to the protein scaffold that might still allow a subset of RNA-protein and/or protein-protein interactions to occur44.
Discussion
More than 90% of human genes undergo alternative splicing to generate multiple mRNA variants in a tissue-specific manner45. The corresponding protein isoforms have different, even antagonistic, properties. Their deregulation has been implicated in various disease states, including cancer46. Therefore, splicing modulators represent a promising molecular therapeutic strategy. This in vitro study in seven HNSCC cell lines revealed that meayamycin B synergized with ABT-737 to quickly trigger Bax/Bak-mediated apoptosis. Meayamycin B selectively regulated the alternative splicing of MCL1, increasing proapoptotic MCL1-S and decreasing antiapoptotic MCL1-L. Importantly, the total MCL-1 level correlated with the single-agent efficacy of meayamycin B. In selected cell lines, meayamycin B also inhibited constitutive splicing of MCL1 and HPV16 E6. In 93-VU-147T cells, knockdown of HPV16 E6 induced an increase of total MCL1 level by an unknown mechanism.
The regulation of HPV16 E6 splicing in vivo awaits full characterization. Spliceosomal U1snRNP-binding cis-acting elements have been identified in HPV16 genome47. Whether and how the U2 snRNP and its associated subcomplexes SF3a and 3b interact with HPV16 genome is still unclear. In addition, it was found that viruses can modify the expression of splicing factors of the host cells to meet their needs48. This lack of systematic understanding calls for dedicated attention to the splicing regulations of this tumorigenic virus. These observations also justify the rising necessity to treat HPV 16-positive and HPC16-negative HNSCC as two separate diseases.
In this study in HNSCC, SF3B1 inhibitor meayamycin B and SF3B1 knockdown each could lead to dominant splice variants of HPV16 E6 and MCL-1 to E6 and MCL1-S, respectively. These demonstrated that the biogenesis of E6* and Mcl-1S require SF3B1 in HNSCC cells. Interestingly, 3-h treatment of meayamycin B could significantly enhance the effect of SF3B1 knockdown, indicating that meayamycin B affected other splicing regulatory proteins involved in these splicing events. Concerning the short-term of meayamycin B exposure, it is likely resulted from a direct interaction. Possible explanations include (1) meayamycin B fits into an interface where SF3B1 interact with its binding partners, which was proposed in the target ID studies of another SF3b inhibitor pladienolide49; (2) meayamycin B has other distinct binding target.
In general, HPV16 positive cells were more resistant to meayamycin B or ABT-737 alone, which was overcome when the drugs were combined. The resistance might stem from HPV16 proteins such as E6, E7 and E2 that individually negatively regulate pro-apoptotic proteins in the host cells18,50,51.
A myriad of strategies employ genetic and pharmacologic approaches for targeting MCL152. Since MCL1 is alternatively spliced into isoforms with antagonistic functions, targeting alternative splicing is advantageous; with the obliteration of antiapoptotic MCL1-L, proapoptotic MCL1-S increases and selectively neutralizes residual MCL1-L. Here, we report the first small molecule that modulates the splicing of MCL1 in HNSCC. Combined with our previous reports in non-small cell lung carcinoma, these studies demonstrate a positive correlation between the MCL1 level and the efficacy of SF3B1 inhibitors, indicating their potential usage for cancers overexpressing MCL1.
Meayamycin B and other FR901464 analogs presumably target SF3B1 that was recently reported to regulate BCL-X alternative splicing53. Interestingly, BCL-X, also abundantly expressed in HNSCC, was not altered by meayamycin B. This was consistent with our previous findings in non-small cell lung carcinoma and Webb's findings in pediatric rhabdomyosarcoma Rh18 cells using their structurally similar analog24,54. This lack of changes in the BCL-X system may be a cell type-dependent splicing regulation for this gene. Importantly, the unresponsiveness of BCL-X with extended exposure (48-h data not shown) indicates that the rapid modulation of MCL1 splicing was a main contributor to the rapid apoptosis we observed from the combination of meayamycin B and ABT-737.
In summary, we demonstrated that the combination of SF3B1 inhibitor meayamycin B and BCL-X/BCL-2 inhibitor ABT-737 triggered apoptosis in HNSCC in vitro models. Single-agent meayamycin B elicited stronger toxicity in MCL1-abundant HPV16-negative than in MCL1-deficient HPV16-positive HNSCC, representing a potential therapy for MCL1-addicted cancer. In addition, HPV E6 is the first viral gene discovered to be regulated by SF3B1 inhibitors at a splicing level; this discovery indicates the feasibility of meayamycin B as a chemical probe to study the splicing regulation of HPV16 E6 and other viral genes.
References
Jemal, A., Siegel, R., Xu, J. & Ward, E. Cancer statistics, 2010. CA Cancer J. Clin. 60, 277–300 (2010).
Kreimer, A. R., Clifford, G. M., Boyle, P. & Franceschi, S. Human papillomavirus types in head and neck squamous cell carcinomas worldwide: A systematic review. Cancer Epidem. Biomar. 14, 467–475 (2005).
Koutsky, L. A. et al. A cohort study of the risk of cervical intraepithelial neoplasia grade 2 or 3 in relation to papillomavirus infection. N. Engl. J. Med. 327, 1272–1278 (1992).
D'souza, G. et al. Case-control study of human papillomavirus and oropharyngeal cancer. N. Engl. J. Med. 356, 1944–1956 (2007).
Chaturvedi, A. K., Engels, E. A., Anderson, W. F. & Gillison, M. L. Incidence trends for human papillomavirus-related and -unrelated oral squamous cell carcinomas in the United States. J. Clin. Oncol. 26, 612–619 (2008).
Gillison, M. L. et al. Survival outcomes by tumor human papillomavirus (HPV) status in stage III–IV oropharyngeal cancer (OPC) in RTOG 0129. J. Clin. Oncol. 27, abstr 6003 (2009).
Spanos, W. C. et al. Immune response during therapy with cisplatin or radiation for human papillomavirus-related head and neck cancer. Arch. Otolaryngol. Head Neck Surg. 135, 1137–1146 (2009).
Nagel, R. et al. Treatment response of HPV-positive and HPV-negative head and neck squamous cell carcinoma cell lines. Oral Oncol. 49, 560–566 (2013).
Colevas, A. D. Chemotherapy options for patients with metastatic or recurrent squamous cell carcinoma of the head and neck. J. Clin. Oncol. 24, 2644–2652 (2006).
Walboomers, J. M. M. et al. Human papillomavirus is a necessary cause of invasive cervical cancer worldwide. J. Pathol. 189, 12–19 (1999).
Danos, O., Katinka, M. & Yaniv, M. Human Papillomavirus-1a Complete DNA-Sequence - a Novel Type of Genome Organization among Papovaviridae. EMBO J. 1, 231–236 (1982).
Chow, L. T., Broker, T. R. & Steinberg, B. M. The natural history of human papillomavirus infections of the mucosal epithelia. Apmis 118, 422–449 (2010).
Smotkin, D., Prokoph, H. & Wettstein, F. O. Oncogenic and nononcogenic human genital papillomaviruses generate the E7 mRNA by different mechanisms. J. Virol. 63, 1441–1447 (1989).
Sedman, S. A. et al. The Full-Length E6 Protein of Human Papillomavirus Type-16 Has Transforming and Trans-Activating Activities and Cooperates with E7 to Immortalize Keratinocytes in Culture. J. Virol. 65, 4860–4866 (1991).
Stacey, S. N. et al. Translation of the human papillomavirus type 16 E7 oncoprotein from bicistronic mRNA is independent of splicing events within the E6 open reading frame. J. Virol. 69, 7023–7031 (1995).
Rosenberger, S., De-Castro Arce, J., Langbein, L., Steenbergen, R. D. & Rosl, F. Alternative splicing of human papillomavirus type-16 E6/E6* early mRNA is coupled to EGF signaling via Erk1/2 activation. Proc. Natl. Acad. Sci. U S A 107, 7006–7011 (2010).
Munger, K., Phelps, W. C., Bubb, V., Howley, P. M. & Schlegel, R. The E6 and E7 genes of the human papillomavirus type 16 together are necessary and sufficient for transformation of primary human keratinocytes. J. Virol. 63, 4417–4421 (1989).
Scheffner, M., Werness, B. A., Huibregtse, J. M., Levine, A. J. & Howley, P. M. The E6 oncoprotein encoded by human papillomavirus types 16 and 18 promotes the degradation of p53. Cell 63, 1129–1136 (1990).
Kosel, S., Burggraf, S., Engelhardt, W. & Olgemoller, B. Increased levels of HPV16 E6*I transcripts in high-grade cervical cytology and histology (CIN II+) detected by rapid real-time RT-PCR amplification. Cytopathology 18, 290–299 (2007).
Cricca, M. et al. Molecular Analysis of HPV 16 E6I/E6II Spliced mRNAs and Correlation With the Viral Physical State and the Grade of the Cervical Lesion. J. Med. Virol. 81, 1276–1282 (2009).
Pim, D., Massimi, P. & Banks, L. Alternatively spliced HPV-18 E6* protein inhibits E6 mediated degradation of p53 and suppresses transformed cell growth. Oncogene 15, 257–264 (1997).
Filippova, M. et al. The large and small isoforms of human papillomavirus type 16 E6 bind to and differentially affect procaspase 8 stability and activity. J. Virol. 81, 4116–4129 (2007).
Shirasawa, H., Tanzawa, H., Matsunaga, T. & Simizu, B. Quantitative detection of spliced E6-E7 transcripts of human papillomavirus type 16 in cervical premalignant lesions. Virology 184, 795–798 (1991).
Gao, Y. & Koide, K. Chemical Perturbation of Mcl-1 Pre-mRNA Splicing to Induce Apoptosis in Cancer Cells. Acs Chem. Biol. 8, 895–900 (2013).
Laetsch, T. W. et al. Multiple components of the spliceosome regulate Mcl1 activity in neuroblastoma. Cell Death Dis. 5, e1072 (2014).
Ahrendt, S. A. et al. Alcohol consumption and cigarette smoking increase the frequency of p53 mutations in non-small cell lung cancer. Cancer Res. 60, 3155–3159 (2000).
Hashibe, M. et al. Alcohol drinking in never users of tobacco, cigarette smoking in never drinkers and the risk of head and neck cancer: pooled analysis in the International Head and Neck Cancer Epidemiology Consortium. J. Natl. Cancer Inst. 99, 777–789 (2007).
Gwosdz, C., Balz, V., Scheckenbach, K. & Bier, H. p53, p63 and p73 expression in squamous cell carcinomas of the head and neck and their response to cisplatin exposure. Adv. Otorhinolaryngol. 62, 58–71 (2005).
Brenner, J. C. et al. Genotyping of 73 UM-SCC head and neck squamous cell carcinoma cell lines. Head Neck 32, 417–426 (2010).
Bradford, C. R. et al. P53 mutation correlates with cisplatin sensitivity in head and neck squamous cell carcinoma lines. Head Neck 25, 654–661 (2003).
Steenbergen, R. D. et al. Integrated human papillomavirus type 16 and loss of heterozygosity at 11q22 and 18q21 in an oral carcinoma and its derivative cell line. Cancer Res. 55, 5465–5471 (1995).
Ferris, R. L. et al. Human papillomavirus-16 associated squamous cell carcinoma of the head and neck (SCCHN): a natural disease model provides insights into viral carcinogenesis. Eur. J. Cancer 41, 807–815 (2005).
Chikamatsu, K. et al. Generation of anti-p53 cytotoxic T lymphocytes from human peripheral blood using autologous dendritic cells. Clin. Cancer Res. 5, 1281–1288 (1999).
Krause, C. J. et al. Human squamous cell carcinoma. Establishment and characterization of new permanent cell lines. Arch. Otolaryngology 107, 703–710 (1981).
Wei, M. C. et al. Proapoptotic BAX and BAK: a requisite gateway to mitochondrial dysfunction and death. Science 292, 727–730 (2001).
Loewe, S. The problem of synergism and antagonism of combined drugs. Arzneimittelforschung 3, 285–290 (1953).
Greco, W. R., Bravo, G. & Parsons, J. C. The search for synergy: a critical review from a response surface perspective. Pharmacol. Rev. 47, 331–385 (1995).
Chou, T. C. & Talalay, P. Generalized equations for the analysis of inhibitions of Michaelis-Menten and higher-order kinetic systems with two or more mutually exclusive and nonexclusive inhibitors. Eur. J. Biochem. 115, 207–216 (1981).
Silva, M. T. Secondary necrosis: the natural outcome of the complete apoptotic program. FEBS Lett. 584, 4491–4499 (2010).
Germain, M. et al. MCL-1 is a stress sensor that regulates autophagy in a developmentally regulated manner. EMBO J. 30, 395–407 (2011).
Shkreta, L. et al. Anticancer drugs affect the alternative splicing of Bcl-x and other human apoptotic genes. Mol. Cancer Ther. 7, 1398–1409 (2008).
Smotkin, D. & Wettstein, F. O. Transcription of human papillomavirus type 16 early genes in a cervical cancer and a cancer-derived cell line and identification of the E7 protein. Proc. Natl. Acad. Sci. U S A 83, 4680–4684 (1986).
Moore, M. J., Wang, Q. Q., Kennedy, C. J. & Silver, P. A. An Alternative Splicing Network Links Cell-Cycle Control to Apoptosis. Cell 142, 625–636 (2010).
Weiss, W. A., Taylor, S. S. & Shokat, K. M. Recognizing and exploiting differences between RNAi and small-molecule inhibitors. Nat. Chem. Biol. 3, 739–744 (2007).
Wang, E. T. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008).
Kalsotra, A. & Cooper, T. A. Functional consequences of developmentally regulated alternative splicing. Nat. Rev. Genet. 12, 715–729 (2011).
Cumming, S. A., McPhillips, M. G., Veerapraditsin, T., Milligan, S. G. & Graham, S. V. Activity of the human papillomavirus type 16 late negative regulatory element is partly due to four weak consensus 5′ splice sites that bind a U1 snRNP-like complex. J. Virol. 77, 5167–5177 (2003).
Mole, S., Milligan, S. G. & Graham, S. V. Human papillomavirus type 16 E2 protein transcriptionally activates the promoter of a key cellular splicing factor, SF2/ASF. J. Virol. 83, 357–367 (2009).
Kotake, Y. et al. Splicing factor SF3b as a target of the antitumor natural product pladienolide. Nat. Chem. Biol. 3, 570–575 (2007).
Ramirez-Salazar, E. et al. HPV16 E2 could act as down-regulator in cellular genes implicated in apoptosis, proliferation and cell differentiation. Virol. J. 8, 247–256 (2011).
Lee, J. O., Russo, A. A. & Pavletich, N. P. Structure of the retinoblastoma tumour-suppressor pocket domain bound to a peptide from HPV E7. Nature 391, 859–865 (1998).
Quinn, B. A. et al. Targeting Mcl-1 for the therapy of cancer. Expert Opin. Invest. Drugs 20, 1397–1411 (2011).
Massiello, A., Roesser, J. R. & Chalfant, C. E. SAP155 Binds to ceramide-responsive RNA cis-element 1 and regulates the alternative 5′ splice site selection of Bcl-x pre-mRNA. FASEB J. 20, 1680–1682 (2006).
Fan, L., Lagisetti, C., Edwards, C. C., Webb, T. R. & Potter, P. M. Sudemycins, novel small molecule analogues of FR901464, induce alternative gene splicing. ACS Chem. Biol. 6, 582–589 (2011).
Acknowledgements
This work was supported by the US National Cancer Institute (R01 CA120792 and 3P50 CA097190 0751) and UPCI Translational Microenvironment/Virus Oncology Center. We thank Sami Osman in our group for providing meayamycin B. This project used the UPCI flow cytometry facility that is supported in part by the US National Cancer Institute (P30CA047904). We would like to thank Dr. Joel Gillespie at the Materials Characterization Laboratory.
Author information
Authors and Affiliations
Contributions
Y.G., R.L.F. and K.K. designed experiments and wrote the manuscript. Y.G. produced the data presented in Figure 1–2 and Figure 4–8. Y.G. and S.T. produced the date presented in Figure 3. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Gao, Y., Trivedi, S., Ferris, R. et al. Regulation of HPV16 E6 and MCL1 by SF3B1 inhibitor in head and neck cancer cells. Sci Rep 4, 6098 (2014). https://doi.org/10.1038/srep06098
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06098
This article is cited by
-
Myeloid cell leukemia-1 expression in cancers of the oral cavity: a scoping review
Cancer Cell International (2022)
-
Sensitivity to splicing modulation of BCL2 family genes defines cancer therapeutic strategies for splicing modulators
Nature Communications (2019)
-
DNA methylation in silkworm genome may provide insights into epigenetic regulation of response to Bombyx mori cypovirus infection
Scientific Reports (2017)
-
Splicing modulators act at the branch point adenosine binding pocket defined by the PHF5A–SF3b complex
Nature Communications (2017)
-
The SF3B1 inhibitor spliceostatin A (SSA) elicits apoptosis in chronic lymphocytic leukaemia cells through downregulation of Mcl-1
Leukemia (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.