Abstract
Acidosis causes millions of deaths each year and strategies for normalizing the blood pH in acidosis patients are greatly needed. The lactate dehydrogenase (LDH) pathway has great potential for treating acidosis due to its ability to convert protons and pyruvate into lactate and thereby raise blood pH, but has been challenging to develop into a therapy because there are no pharmaceutical-based approaches for engineering metabolic pathways in vivo. In this report we demonstrate that the metabolic flux of the LDH pathway can be engineered with the compound 5-amino-2-hydroxymethylphenyl boronic acid (ABA), which binds lactate and accelerates the consumption of protons by converting pyruvate to lactate and increasing the NAD+/NADH ratio. We demonstrate here that ABA can rescue mice from metformin induced acidosis, by binding lactate and increasing the blood pH from 6.7 to 7.2 and the blood NAD+/NADH ratio by 5 fold. ABA is the first class of molecule that can metabolically engineer the LDH pathway and has the potential to have a significant impact on medicine, given the large number of patients that suffer from acidosis.
Similar content being viewed by others
Introduction
Acidosis has a 50% mortality rate among critically ill patients. Acidosis is caused by mitochondrial dysfunction in ATP production, which leads to the accumulation of protons1. The current therapy for lactic acidosis is bicarbonate treatment, which is ineffective because of its numerous toxic side effects such as altering blood pressure, triggering apoptosis and modifying intracellular pH levels2,3,4,5. Dichloroacetate is a potential treatment for acidosis, which enhances the pyruvate dehydrogenase activity and blocks lactate production, however it is also not effective because it causes neuropathy6. Therefore, there is a great need for the development of strategies that can increase the blood pH in patients suffering from acidosis6,7,8,9.
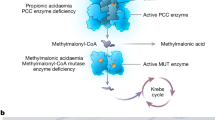
In this report we present a new strategy for treating acidosis based on the compound ABA, which binds lactate and normalizes the blood pH by increasing the consumption of protons via the LDH pathway. The mechanism by which ABA treats acidosis is shown in Figure 1. ABA forms a bivalent complex with lactate and thereby decreases the intracellular lactate concentration, shifting the equilibrium towards the production of more lactate, resulting in the consumption of protons via hydrogenation of pyruvate. Importantly, ABA is specific for lactate over other metabolites such as glucose, because of its ortho-hydroxyl, which prevents it from binding cis-diols10. In addition, ABA also increases the NAD+/NADH ratio, which also has numerous therapeutic effects, such as protection against apoptosis and suppression of inflammation.
5-amino-2-hydroxymethylphenyl boronic acid (ABA) metabolically engineers the lactate dehydrogenase pathway and is a therapy for lactic acidosis.
ABA binds lactate and normalizes the blood pH by shifting the equilibrium of the LDH pathway towards the consumption of protons (shown in red), pyruvate and the generation of NAD+.
Results
ABA was identified as a therapeutic for acidosis because of its potential ability to bind lactate. We therefore, performed experiments to determine if ABA forms a stable complex with lactate. The ABA-lactate complex was formed by mixing 100 μmoles of ABA and lactate in 1 mL of methanol for 10 min. The ABA-lactate complex was then isolated using preparative TLC, in ethyl acetate and ESI mass spectrometry was performed on the isolated ABA-lactate complex. An experimental molecular weight of 220.0786 was obtained, which correlated with the predicted m/z of 220.0786 and therefore suggests that ABA forms a stable complex with lactate.
We also investigated if ABA could bind lactate in the presence of other metabolites present in the serum, using boron nuclear magnetic resonance (NMR). ABA and sodium L-lactate were added to 500 μL of 10% fetal bovine serum in Dulbecco's Modified Eagle Medium (DMEM) and analyzed by boron NMR. Figure 2B demonstrates that ABA binds lactate in the presence of the metabolites present in fetal bovine serum and DMEM. For example, the aromatic boron of ABA in serum has an NMR peak at δ = 29 and this shifts to δ = 9 in the presence of lactate and serum, demonstrating specific complexation with lactate. The peak shifts obtained from these experiments correlated with the peak shifts obtained from the boron NMR of an ABA-lactate synthetic standard (isolated via preparative TLC). These data demonstrate that ABA specifically complexes with lactate and does not bind any of the metabolites normally present in serum and DMEM10.
ABA binds lactate, increases the NAD+/NADH ratio and the consumption of pyruvate.
(A) Boron NMR of the isolated ABA-lactate complex. Free ABA has a boron peak at δ = 29 (blue peak), which shifts to δ = 9 for the ABA-lactate complex (red peak). (B) ABA binds sodium lactate in the presence of 10% serum as determined by boron NMR. Free ABA in serum has a boron peak at δ = 29 (blue peak), which shifts to δ = 9 in the presence of equimolar lactate (red peak). (C) ABA increases the NAD+/NADH ratio by two fold in RAW 264.7 cells. Cells were incubated with 0.1 M PBS or 5 mM ABA for 5 min, the cell lysates were harvested and the NAD+/NADH ratio in cell lysates was measured. (D) ABA decreases pyruvate levels by two fold in RAW 264.7 cells. Cells were incubated with 0.1 M PBS or 5 mM ABA for 5 min, the cell lysates were harvested and the pyruvate levels in cell lysates were measured.
For ABA to effectively metabolically engineer the LDH pathway, it has to enter the cells. We therefore measured the Log D of ABA using absorbance measurements at 330 nm in an octanol/water partition assay. ABA has a Log D of 0.80 ± 0.03, which suggests that ABA should be cell membrane permeable. The ability of ABA to enhance pyruvate and NADH consumption was investigated in RAW 264.7 cells. 106 RAW 264.7 cells were incubated with 5 mM ABA, for 5 min and the cells were lysed, homogenized and separated from proteins to analyze the pyruvate, NAD+ and NADH content. Figure 2C demonstrates that ABA increases the NAD+/NADH ratio two fold and figure 2D demonstrates that the pyruvate levels are decreased by 2 fold, thereby suggesting that ABA can catalyze the detoxification of protons.
ABA is designed to deplete lactate levels during acidosis by binding lactate and thereby directing the LDH metabolic pathway to consume excess protons. We therefore investigated if ABA can bind lactate in vivo and decrease its blood concentration. 11 μmoles of ABA in 100 µL of saline were injected into mice via the tail vein and after 10 min their blood was analyzed for lactate and compared against controls. Figure 3A demonstrates that ABA can reduce blood lactate levels, for example, the blood lactate levels in healthy mice was 2 ± 0.5 mM, however, mice treated with ABA had a 3 fold reduction in blood lactate levels and was reduced to 500 ± 200 μM. ABA can decrease extracellular lactate levels by three fold within 10 min, this rapid effect is essential for treating acidosis due to the rapid time-frame in which elevated blood proton levels cause death. Importantly, at this dose ABA did not cause weight loss in mice over a three day period and we observed no changes in the brain lactate/pyruvate, NAD+/NADH levels or blood pH (figures 3B, 4C).
ABA decreases lactate levels in vivo and has minimal toxicity.
(A) ABA decreases blood lactate levels in healthy mice. 11 μmoles of ABA in PBS or PBS were injected intravenously into mice and after 10 min the plasma was isolated and lactate levels were determined. (B) ABA has minimal toxicity to mice. Mice were injected with 100 mg/kg of ABA in 100 μL PBS or 100 μL PBS, via the tail vein and their weight loss was studied over three days. Standard deviation is presented as mean ± s.d. n = 3. n.s. = not significant.
ABA rescues mice from metformin induced acidosis.
(A) ABA normalizes blood lactate levels in mice suffering from lactic acidosis. Lactic acidosis was induced in mice by treating them with 400 mg/kg of metformin. 100 mg/kg of ABA in a 100 µL of 0.1 M PBS or 100 µL of 0.1 M PBS was injected intravenously into the metformin treated mice. 15 min and 30 min after the administration of ABA or PBS the plasma was isolated from the mice and the lactate concentration was determined. (B) ABA normalizes the blood pH in mice suffering from lactic acidosis. Lactic acidosis was induced in mice by treating them with 400 mg/kg of metformin. 100 mg/kg of ABA in a 100 µL of 0.1 M PBS or 100 µL of 0.1 M PBS was injected intravenously into the metformin treated mice. 30 min post treatment the blood was isolated and the pH was measured using a pH meter. (C) ABA increases the blood NAD+/NADH ratio in mice suffering from lactic acidosis. Lactic acidosis was induced in mice by treating them with 400 mg/kg of metformin. 100 mg/kg of ABA in a 100 µL of 0.1 M PBS or 100 µL of 0.1 M PBS were injected intravenously into metformin treated mice. 30 min after administration of ABA or PBS the plasma was isolated from the mice and the NAD+/NADH ratio was determined. * p<0.05, student's t-test. Standard deviation is presented as mean ± s.d. n = 3.
ABA has numerous potential applications given its ability to decrease extracellular lactate in vivo. For example, metformin induced lactic acidosis has a 50% mortality rate due to the lack of treatments11,12. We investigated if ABA could rescue mice from metformin induced lactic acidosis. Lactic acidosis was induced in mice by an intraperitoneal injection of 400 mg/kg of metformin and blood lactate levels were monitored13,14. After the lactate levels reached 10 mM, these mice were injected with 100 mg/kg of ABA in PBS or PBS and after 30 min the blood pH, NAD+/NADH ratio and lactate levels were analyzed. Figure 4A demonstrates that ABA can rescue mice from metformin induced lactic acidosis. For example, metformin treated mice that received ABA had a 5 fold decrease in blood lactate levels, their blood lactate levels decreased from 10 ± 1.5 mM to 2 ± 0.5 mM. This decrease in lactate should induce an increase in the consumption of protons and increase the NAD+/NADH ratio. We therefore investigated if ABA caused an increase in the blood pH levels and the NAD+/NADH ratio in metformin treated mice. Figure 4B demonstrates that ABA leads to an increase in blood pH, causing the blood pH of metformin treated mice to increase from 6.7 ± 0.1 to 7.2 ± 0.1 and figure 4C demonstrates that the blood NAD+/NADH ratio increased by a factor of 5.
Discussion
The treatment of lactic acidosis is a central challenge in medicine and the medical burden caused by lactic acidosis is comparable to cancer and HIV, yet there are currently no drugs available to treat acidosis. Traditional pharmaceutical approaches for treating acidosis have focused on inhibiting the rate of lactate production, via inhibition of the Krebs cycle and have had problems with toxicity due to the central role of the Krebs cycle in metabolism. In addition, buffering of the blood pH via introduction of bicarbonates has been unsuccessful due to toxic side-effects. Therefore, the development of pharmaceutical strategies that can increase the blood pH in patients suffering from acidosis are greatly needed.
In this report we demonstrate that lactic acidosis can be treated by engineering the metabolic fluxes of protons and pyruvate, via complexation of lactate with the phenyl boronic acid derivative ABA. ABA injected intravenously was able to rescue mice from metformin induced acidosis by increasing pH levels and the blood NAD+/NADH ratio. ABA can therefore offer a potential treatment for lactic acidosis, due to its unique ability to bind lactate and restore blood pH. ABA is the first class of molecule that can metabolically engineer the LDH pathway and has the potential to have a significant impact on medicine, given the large number of patients that suffer from acidosis.
Methods
ABA binds lactate with specificity
The ability of ABA to bind lactate and make a stable complex was determined by boron NMR and mass spectrometry. 100 μmoles of ABA and sodium L-lactate were mixed in 1 mL of methanol and preparative TLC was performed in ethyl acetate. The silica band containing the ABA-lactate complex (Rf = 0.1) was removed and the complex was extracted into water and concentrated using a rotary evaporator. Electrospray ionization in a negative-ion mode was performed on the isolated complex, to obtain its mass (QB3/Chemistry Mass Spectrometry Facility at the University of California, Berkeley). We also performed boron NMR on the isolated ABA-lactate complex, in order to generate a reference boron NMR spectra (figure 2A). The preparative TLC isolated ABA-lactate complex was re-suspended in 500 μL of water. This solution was then added to a quartz NMR tube (Fisher Scientific) and boron NMR was performed using a 400 MHz Bruker NMR (n = 8 scans).
The ability of ABA to bind lactate in the presence of other metabolites was determined by boron NMR. 11 μmoles of ABA and 11 μmoles of sodium L-lactate were dissolved in 500 μL of 10% fetal bovine serum in Dulbecco's Modified Eagle Medium (DMEM). This solution was then added to a quartz NMR tube (Fisher Scientific) and boron NMR was performed using a 400 MHz Bruker NMR (n = 8 scans). As a control 11 μmoles of ABA were dissolved in 500 μL of 10% fetal bovine serum in Dulbecco's Modified Eagle Medium (DMEM) and boron NMR was performed on a 400 MHz Bruker NMR (n = 8 scans).
ABA is cell membrane permeable
The log D of ABA was measured using an octanol/water partitioning assay. 500 μL of ABA in 0.1 M PBS at 1 mg/mL concentration was added to 500 μL of octanol (Sigma Aldrich) in an eppendorf tube and sonicated in a sonicating bath for 1 h. The tube was then centrifuged for 5 min at 1000xGs and incubated for 16 h at room temperature. 100 μL of the PBS and octanol was pipetted out into separate wells of a 96 well plate (VWR), the absorbance was measured at 330 nm using a Tecan i4 plate reader and the log D was determined (n = 3).
General Cell Culture
RAW 264.7 cells were cultured in T-25 flasks in cell culture media consisting of DMEM and 10% fetal bovine serum. The cells were passaged every 3rd day by resuspending the cells using a cell scraper (Fisher Scientific) and centrifuging the cells at 300XGs for 5 min. The supernatant was removed and the cells were then resuspended in cell culture media. The cells were counted using a hemocytometer (Fisher Scientific) and utilized for further experiments described below.
ABA consumes pyruvate and increases the NAD+/NADH ratio
The ability of ABA to consume pyruvate and increase the NAD+/NADH ratio was determined by incubating ABA with RAW 264.7 cells. 5 mM of ABA in 200 μL of cell culture media was added to 106 RAW 264.7 cells and incubated for 5 min. The cells were then centrifuged at 300XGs for 5 min and the supernatant was discarded. The cells were lysed using 500 μL of cell lysis buffer provided with the NAD+/NADH quantification kit (Sigma Aldrich). The cells were then filtered through a 10 KDa centrifuge filter (Millipore) by centrifuging at 10,000XGs for 5 min. Pyruvate in the filtrate was quantified using a pyruvate quantification kit (Abcam Inc.) and the NAD+/NADH ratio was quantified using an NAD+/NADH quantification kit following the manufacturer's protocol. A Tecan i4 plate reader was utilized to measure the absorbance and fluorescence readings.
ABA decreases lactate levels in vivo
6–8 weeks old female C57BL6/j mice were utilized for all the studies in accordance with R348 animal protocol approved by the University of California. 100 μL of PBS containing 11 μmoles of ABA were injected into the mice via the tail vein. After 10 min cardiac puncture was performed on the mice and their blood was extracted and centrifuged at 10,000XGs for 10 min to isolate the plasma. A lactate assay kit (Abcam Inc.) was utilized to measure the lactate concentration in the plasma.
ABA reduces blood lactate levels in metformin induced lactic acidosis
Female 6–8 weeks old C57Bl6/j mice were injected with 400 mg/kg of metformin (in 500 μL of PBS) in the intraperitoneal cavity, the blood lactate levels of the mice were measured by performing tail snips of the mice at various time points. Once the blood lactate levels reached 10 mM, the mice were injected with either 100 mg/kg of ABA in 50 μL (pH = 7.4) via the tail vein or with 50 μL of PBS as a control. The mice were sacrificed at 0, 15 and 30 min after ABA treatment and the blood was isolated via cardiac puncture and its pH was analyzed using a pH meter (Fisher Scientific accumet). In addition a 100 μL of the blood was centrifuged at 10,000XGs for 10 min to isolate the serum and the lactate and NAD+/NADH ratio in the plasma were determined using lactate (Abcam Inc.) and NADH enzymatic kits (Sigma Aldrich) (figure 3B, D). Furthermore, liver, brain and kidneys of the mice were isolated and the NAD+/NADH and the lactate/pyruvate ratios were determined. The tissue was weighed and homogenized in the lysis buffer provided by the manufacturer (Sigma Aldrich). The tissue was then centrifuge filtered using 10 KDa centrifuge filters (Millipore) and the filtrate was then analyzed for NAD+ and NADH amounts to determine the NAD+/NADH ratio and lactate and pyruvate amounts to determine the lactate/pyruvate ratio.
Statistical Analysis
Statistical analyses were performed using a student t-test and p-values for each experiment were determined using the excel software package (Microsoft Office 2010). Statistically significant data (p<0.05) are depicted using the ‘*’ symbol.
References
Hochachka, P. W. & Mommsen, T. P. Protons and anaerobiosis. Science 219, 1391–1397 (1983).
Kozeny, G. A. et al. In vivo effects of acute changes in osmolality and sodium concentration on myocardial contractility. Am Heart J. 109, 290–296 (1985).
Cooper, D. J., Walley, K. R., Wiggs, B. R. & Russell, J. A. Bicarbonate does not improve hemodynamics in critically ill patients who have lactic acidosis. A prospective, controlled clinical study. Ann Intern Med. 112, 492–498 (1990).
Grundler, W., Weil, M. H. & Rackow, E. C. Arteriovenous carbon dioxide and pH gradients during cardiac arrest. Circulation 74, 1071–1074 (1986).
Kette, F., Weil, M. H., von Planta, M., Gazmuri, R. J. & Rackow, E. C. Buffer agents do not reverse intramyocardial acidosis during cardiac resuscitation. Circulation 81, 1660–1666 (1990).
Kaufmann, P. et al. Dichloroacetate causes toxic neuropathy in MELAS: a randomized, controlled clinical trial. Neurology 66, 324–330 (2006).
Fall, P. J. & Szerlip, H. M. Lactic acidosis: from sour milk to septic shock. J Intensive Care Med. 20, 255–271 (2005).
Luft, D., Deichsel, G., Schmulling, R. M., Stein, W. & Eggstein, M. Definition of clinically relevant lactic acidosis in patients with internal diseases. Am J Clin Pathol. 80, 484–489 (1983).
Renda, F. et al. Metformin-associated lactic acidosis requiring hospitalization. A national 10 year survey and a systematic literature review. Eur Rev Med Pharmacol Sci., 45–49 (2013).
Sartain, F. K., Yang, X. & Lowe, C. R. Complexation of L-lactate with boronic acids: a solution and holographic analysis. Chemistry 14, 4060–4067 (2008).
Peters, N. et al. Metformin-associated lactic acidosis in an intensive care unit. Crit Care 12, R149 (2008).
Misbin, R. I. et al. Lactic acidosis in patients with diabetes treated with metformin. N Engl J Med. 338, 265–266 (1998).
Snell, A. M. & Roth, G. M. The Lactic Acid of the Blood in Hepatic Disease. J Clin Invest. 11, 957–971 (1932).
Fendt, S. M. et al. Metformin decreases glucose oxidation and increases the dependency of prostate cancer cells on reductive glutamine metabolism. Cancer Res. 73, 4429–4438 (2013).
Acknowledgements
This work was supported by the following grants to NM from the NIH, 1R01HL096796-02, U01 268201000043C-0-0-1, 1R01HL090601-02, 1R21AI098799-01, R01 AI088023-03.
Author information
Authors and Affiliations
Contributions
A.P.A. performed binding studies, in vitro experiments, in vivo experiments and manuscript preparation. M.R. performed in vivo experiments. A.B.G. and E.C.W. performed binding studies. N.M. directed the research and contributed to manuscript preparation. All authors discussed the results and commented on the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. The images in this article are included in the article's Creative Commons license, unless indicated otherwise in the image credit; if the image is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the image. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Acharya, A., Rafi, M., Woods, E. et al. Metabolic engineering of lactate dehydrogenase rescues mice from acidosis. Sci Rep 4, 5189 (2014). https://doi.org/10.1038/srep05189
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep05189
This article is cited by
-
Pyruvate is a prospective alkalizer to correct hypoxic lactic acidosis
Military Medical Research (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.