Abstract
Transcription activator-like effector nuclease (TALEN) and zinc finger nuclease (ZFN) genome editing technology enables site directed engineering of the genome. Here we demonstrate for the first time that both TALEN and ZFN injected directly into pig zygotes can produce live genome edited pigs. Monoallelic as well as heterozygous and homozygous biallelic events were identified, significantly broadening the use of genome editor technology in livestock by enabling gene knockout in zygotes from any chosen mating.
Similar content being viewed by others
Introduction
Gene mutation provides a powerful strategy to investigate gene function and has potential biotechnology and animal breeding applications1,2. Traditionally for livestock, specific gene knockout strategies have been based around homologous recombination in somatic cells destined for use in somatic cell nuclear transfer (SCNT) protocols3. Although significantly more efficient than when first developed, SCNT is limited by the need to isolate cells for any given genotype. The ability to perform gene knockout directly in the zygote of any chosen mating would enable engineering of any genotype.
The recent development of hybrid molecular enzymes (genome editing tools) capable of directing double-strand breaks to a specific target sequence provides a new tool for reverse genetics4,5. These targetable DNA cleavage reagents can be engineered to recognize and cleave a precise DNA sequence within a genome. The induced double stand breaks are a target for repair by the error prone non-homologous end-joining repair pathway, resulting in targeted mutation in a proportion of events. Unlike transgenesis, genome editing does not require a transferred DNA transgene and no genetic mark beyond that of the mutation is introduced into the genome.
Genome editing technology has been successfully applied to zebrafish6, rabbits7 and rodents8,9 by the direct injection of embryos, but in livestock utility has remained restricted to SCNT methodology10,11,12,13. To extend this technology beyond that of our previous work using TALEN SCNT13, we sought to establish genome editing technology in livestock by zygote injection of either TALEN or ZFN and present data on the effect of dose of genome editor on embryo development and frequency of live births. In this study we demonstrate that both TALEN and ZFN technology can be efficiently applied to engineer pig zygotes that result in gene edited live births, both mono- and bi-allelic (Table 1; Supplementary Fig. 1), significantly broadening the use of editor technology in livestock.
Results
With the aim to demonstrate the potential of this technology for animal breeding we selected RELA as the target locus. We have previously argued for a role of porcine RELA in the pathology associated with African Swine Fever Virus infection in pigs. Specifically, polymorphic sequence variation within the transactivation domains of porcine RELA parallels the differing severity associated with African Swine Fever Virus infection observed in domestic pigs in contrast to African pigs such as warthogs14. Therefore we designed genome editing tools to target mutation to this region of the porcine RELA gene (Fig. 1A). The TALENs were designed using the TALE-NT software13 and assembled to target a region located at 1458 to 1505 bp relative to the translational start site in the porcine RELA cDNA sequence (NM_001114281).
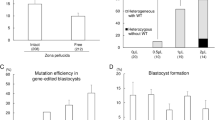
A. Location of TALEN, ZFN and PCR primers for sequencing amplicon. Porcine RELA sequence 769417-769110 accession number NW_003609513.1. B. Example Cel1 Assay. Lanes A2, A12, B6, B14 and B20 were identified as having cleavage products consistent with editing events at the target site. Direct sequencing of the same PCR products confirmed this interpretation and identified B12, B16, B17 and B18 as having only wild-type sequence.
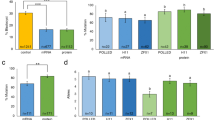
Studies in rats demonstrated that zygote cytoplasmic injection was as effective as pronuclear injection for the delivery of genome editors. Given the difficulty associated with visualization of the pronucleus in porcine zygotes, we elected to perform cytoplasmic injection. Oocytes were harvested from slaughterhouse material for in vitro studies and superovulated artificially inseminated sows were used for embryos destined for transfer into recipients. Initially 208 zygotes were subjected to a single 10 pl cytoplasmic injection of a solution composed 20 ng/μl RELA TALEN mRNA with 5 ng/μl EGFP mRNA (to enable visual identification and isolation of embryos that functionally translated the injected mRNA). After approximately 3 days of in vitro development, GFP fluorescence was detected in 36% of embryos. The presence of non-expressing embryos may reflect as yet unknown species differences in the ability to process introduced RNA since it is not generally seen in rodents nor in bovine13 but in our hands is reproducible in pigs. We conclude that mRNA injected into the cytoplasm of pig zygotes translates to functional protein in a proportion of embryos (Table 1).
GFP positive embryos were screened for editing events by Cel1 surveyor assay (Fig. 1B) and sequencing of PCR amplified fragments (Supplementary Fig. 2). We detected 16 editing events in 46 GFP-positive embryos analyzed (35%). In a second experiment we tested 34 embryos injected with 2 pl of 20 ng/μl RELA TALEN mRNA but without selection for GFP activity and detected 2 editing events (6%). In two further experiments where 2 pl of 10 ng/μl and 2 ng/μl RELA TALEN mRNA was injected, 0% and 18% editing frequency, respectively, were observed. We suspect the lack of cutting observed for the 10 ng/μl dose may simply reflect the small tested sample size (n = 3). In total we identified 21 editing events in porcine embryos in vitro (21% of tested embryos: Table 1) and a high frequency of these editing events were biallelic in nature (33% of edited embryos: Table 1). Calculating this as a frequency of tested embryos, we achieved a biallelic editing frequency of 7%. Since both the highest and lowest tested concentrations of RELA TALEN mRNA produced edited embryos in vitro we elected to transfer zygotes injected at each dosage into recipient sows and allow pregnancies to develop to term. Pregnancy rates for the higher concentrations of RELA TALEN mRNA were poor; 0 out of 2 transfers and 1 out of 2 transfers for embryos injected with 20 ng/μl or 10 ng/μl, respectively. This poor pregnancy rate reflected the visual observation that 2 ng/μl RELA TALEN mRNA injected embryos developed better in vitro than those injected with higher concentrations of TALEN mRNA; the injections carried out using lower concentrations of mRNA generally showed better development, with 7.3% developing to early morula at day three of culture following injection with 2 ng/μl, as opposed to 3.2% at 10 ng/μl and only 0.7% at 25 ng/μl. The one pregnancy from 10 ng/μl delivered 7 piglets, none of which harbored RELA editing events by direct sequencing of PCR amplified products. We did not pursue transfers of embryos with these higher TALEN mRNA concentrations any further.
In contrast, transfer of embryos injected with 2 ng/μl RELA TALEN RNA resulted in 5 pregnancies from 7 recipients. One subsequently aborted at 15 weeks of pregnancy just prior to parturition; analysis of the 9 foetuses carried revealed 3 to have editing events. In total from the remaining 4 farrowings, 39 piglets were produced of which 8 carried editing events (21%). Of the 8 edited animals, two were stillborn, another died neonatally due to being crushed by the mother; leaving 5 live piglets.
In parallel we tested a pair of ZFNs with a target location of 1496 to 1532 bp relative to the translational start site in porcine RELA cDNA sequence (NM_001114281). One transfer of embryos injected with RELA ZFN mRNA at 10 ng/μl failed to generate a pregnancy while the two transfers of embryos injected with RELA ZFN mRNA at 2 ng/μl both became pregnant resulting in the birth of 9 piglets. Of these 9 piglets, one carried an editing event at the ZFN target site (11%), comparable in frequency to our observed TALEN editing efficiency (Table 1).
Direct sequencing of PCR products revealed a variety of editing events in piglets derived from TALEN and ZFN injected embryos (Supplementary Fig. 3). Analysis of ear biopsy isolated genomic DNA identified both deletions and insertions at the target sites. Sequence data from 2 animals constituted multiple overlapping traces indicating two or more editing events; this was subsequently confirmed by sequencing of multiple cloned PCR products. Presuming that in these cases of multiple editing the frequency of events detected in ear biopsy reflects frequency in the early embryo, designer nuclease editing can remain active beyond the 2-cell stage (i.e. some events display low representation in the PCR pool and are therefore only present in a subset of cells). In total, 5 biallelic events were identified from 9 edited piglets (56%; 9% of piglets born); 4 from TALEN and 1 from ZFN mRNA injections. Of these bialleleic events 2 were homozygous, with 3 displaying different indels on each allele. While both piglets carrying homozygous biallelic event survived farrowing (milk in stomach post mortem), they were both accidentally killed by their mother within the first 24 hours of life.
Discussion
Genome editing technology offers significant advantages over the traditional methodology and is set to revolutionize the application of genetically engineered animals1. We demonstrate that this technology can be successfully applied to livestock without the need to resort to somatic cell nuclear transfer methodology10,11,12,13, significantly broadening the use of editing tools in livestock. Delivery of TALEN or ZFN RNA to the pig zygote resulted in the birth of both mono- and bi-allelic edited animals (Table 1; Fig. 1).
In summary, we observed an overall editing frequency of 2% of transferred embryos or 16% of piglets born. These figures compare favorably with that reported for zygote injection of ZFNs in rats where 2% of transferred embryos and 12% of founder animals harbored editing events8. Our editing frequencies also compare favorably with the production of monoallelic (0.1% of transferred embryos12) and biallelic (1% of transferred embryos; the greater frequency over monoallelic due to incorporation of a FACS enrichment stage prior to somatic cell nuclear transfer11) pigs using somatic cell nuclear transfer methodology.
In conclusion, we demonstrate that both TALEN and ZFN editing technology can be successfully applied to pig zygotes to produce live gene edited pigs and, contrary to previous attempts11,12 the delivery of editing tools by the direct injection into the zygote is both efficient and able to generate biallelic mutations. This novel achievement paves the way for precise genome engineering of livestock independent of SCNT (cloning) technology.
Methods
Editor design and construction
Two types of editing tool were used: TALEN and ZFN. Both were designed to target the same region of porcine RELA gene. TALEN: TALENs were designed using the TALE-NT software and assembled using methods described previously13. Briefly, intermediary arrays were produced for the porcine RELA TALEN pair for Golden Gate cloning as follows; 150 ng each pFUS_A, pFUS_B, pLR-X and pC- + 63-TAL modified vector were incubated for 10 cycles of 5 min at 37°C and 10 min at 16°C, then heated to 50°C for 5 min and then 80°C for 5 min in the presence of 50 units T4 DNA ligase (New England Biolabs), 10 units Esp3I (Fermentas), 1x T4 ligase buffer (New England Biolabs). Plasmid DNA was purified from ligated vector transformed E. coli and mRNA synthesized from SacI linearized RCIscript vectors using the mMessage Machine T3 Kit (Ambion). Transient transfection of fibroblasts showed a cutting frequency of 45%. ZFN: ZFNs targeted to porcine RELA were purchased from Sigma. The ZFN displayed 84.7% cutting in MEL1 assay (Sigma data sheet) and was supplied as mRNA.
Zygote injections
To establish the frequency of genome editing in pig embryos an in vitro embryo culture experiment was performed. Pig Ovaries were collected, washed with pre-warmed phosphate buffered saline at 38°C and follicles aspirated. Oocytes were washed in TL HEPES PVA before culturing in maturation medium for 44 hours (22 hours plus hormones and 22 hours minus hormones; 39°C, 5% CO2), followed by gentle pipetting to remove cumulus cells and IVF for 6 hours (38.5°C in 5% CO2). Zygotes were transferred to NCSU-23 HEPES base medium and subjected to a single 2–10 pl cytoplasmic injection of mRNA at 2, 10 or 20 ng/μl +/− 5 ng/μl EGFP mRNA. The 10 and 2 ng/μl mRNA samples were dilutions of the 20 ng/μl sample. Zygotes were cultured in batches for 66 hours (embryo culture medium; 5% CO2, 38.5°C), following which they were placed individually into micro-droplets of medium under oil for visual inspection or harvested for genotyping either as a total group or after manual selection of GFP fluorescing embryos.
Embryo transfers
All animal work was approved under UK Home Office licence after review by the University of Edinburgh's Animal Ethics Committee. Embryos were produced from Large-White gilts that were approximately 9 months of age and weighed at least 120 kg at time of use. Super-ovulation was achieved by feeding, between day 11 and 15 following an observed oestrus, 20 mg altrenogest (Regumate, Hoechst Roussel Vet Ltd) once daily for 4 days and 20 mg altrenogest twice on the 5th day. On the 6th day, 1500 IU of eCG (PMSG, Intervet UK Ltd) was injected at 20:00 hrs. Eighty three hours later 750 IU hCG (Chorulon, Intervet UK Ltd) was injected. Donor gilts were inseminated twice 6 hours apart after exhibiting heat generated following super-ovulation. Embryos were surgically recovered from mated donors by mid-line laparotomy under general anesthesia on day 1 following oestrus into NCSU-23 HEPES base medium. Embryos were subjected to a single 2 pl cytoplasmic injection of either ZFN or TALEN pair mRNA at 10 ng/ul or 2 ng/μl. Recipient females were treated identically to donor gilts but remained un-mated. Following TALEN or ZFN injection, fertilized embryos were transferred to recipient gilts following a mid-line laparotomy under general anaesthesia. During surgery, the reproductive tract was exposed and embryos were transferred into the oviduct of recipients using a 3.5 French gauge tomcat catheter. Litter sizes ranged from 3–17 piglets.
Genotyping
Genome editing events in porcine embryos were identified by direct sequencing of amplified, isolated DNA and through a gel electrophoresis assay. The latter identified mismatch between the two alleles through digestion by the Cel1 enzyme. Sequencing: DNA was amplified from harvested embryos using the REPLI-g® mini kit, Qiagen®. The REPLI-g® DNA sample was then used as a template for High fidelity PCR (AccuPrime™ Taq DNA Polymerase High Fidelity, Invitrogen™) using p65NJF1 5′-GCAATAACACTGACCCGACCGTG-3′ and p65NJR1 5′-GCAGGTGTCAGCCCTTTAGGAGCT-3′ as primers designed to amplify a 308 base pair region of the wild-type porcine RELA gene that overlapped the TALEN and ZFN cut sites. The PCR product was purified then sent for sequence analysis to allow identification of editing events. Alternatively, the PCR products were cloned into a plasmid and individual plasmids sequenced allowing heterozygous and mosaic editing events to be analysed separately. Cel1 assay: The presence of mutations in the RELA gene were additionally identified using a Cel1 assay (SURVEYOR® mutation detection kit, TRANSGENOMIC®). The high fidelity PCR product was denatured/re-annealed before being subjected to SURVEYOR® nuclease activity which cuts at base mismatches highlighting insertions, deletions and substitutions. The resulting fragments were subsequently separated by gel electrophoresis for analysis with size differences identifying edited events.
Additional information
DFC and SCF either have equity and/or work for Recombinetics Inc. WAR works for RoslinEmbyrology. ALM and DGM work for Genus plc. CBAW consults for Genus plc and is on the Scientific Advisory Board of Recombinetics Inc.
References
Fahrenkrug, S. C. et al. Precision genetics for complex objectives in animal agriculture. Anim. Sci. 88, 2530–2539 (2010).
Tan, W. S., Carlson, D. F., Walton, M. W., Fahrenkrug, S. C. & Hackett, P. B. Precision editing of large animal genomes. Adv. Genet. 80, 37–97 (2012).
Clark, A. J. & Whitelaw, C. B. A. A future for transgenic livestock. Nat. Rev. Genet. 4, 825–833 (2003).
Urnov, F. D., Rebar, E. J., Holmes, M. C., Zhang, H. S. & Gregory, P. D. Genome editing with engineered zinc finger nucleases. Nat. Rev. Genet. 11, 636–646 (2010).
Miller, J. C. et al. A TALE nuclease architecture for efficient genome editing. Nat. Biotechnol. 29, 143–148 (2011).
Bedell, V. M. et al. In vivo genome editing using a high-efficiency TALEN system. Nature 491, 114–118 (2012).
Flisokowska, T. et al. Efficient immunoglobulin gene disruption and targeted replacement in rabbit using zinc finger nucleases. PLoS ONE 6, e21045 (2011).
Guerts, A. M. et al. Knockout rats via embryo microinjection of zinc-finger nucleases. Science 325, 433 (2009).
Carbery, I. D. et al. Targeted genome modification in mice using zinc-finger nucleases. Genetics 186, 451–459 (2010).
Whyte, J. J. et al. Gene targeting with zinc finger nucleases to produce cloned eGFP knockout pigs. Mol. Reprod. Dev. 78, 2 (2011).
Hauschild, J. et al. Efficient generation of biallelic knockout pigs using zinc-finger nucleases. Proc. Natl. Acad. Sci. USA 108, 12013–12017 (2011).
Yang, D. et al. Generation of PPARγ mono-allelic knockout pigs via zinc-finger nucleases and nuclear transfer. Cell Res. 21, 979–982 (2011).
Carlson, D. F. et al. Efficient TALEN-mediated gene knockout in livestock. Proc. Natl. Acad. Sci. USA 109, 17382–17387 (2012).
Palgrave, C. J. et al. Species-specific variation in RELA undelies differences in NF-kB activity: a potential role in African swine fever pathogenesis. J. Virol. 85, 6008–6014 (2011).
Acknowledgements
This work was supported by Genus plc, Recombinetics Inc and through BBSRC Institute Stratregic Programme Grant; DS was supported by a Genus plc internship at The Roslin Institute; CP was supported by NSF IOS 0965413. CBAW is grateful to advice on ZFN tools provided by Fyodor Urnov.
Author information
Authors and Affiliations
Contributions
S.G.L., C.P., D.F.C., T.J.K., A.J.M., D.G.M., S.C.F. and C.B.A.W. conceived and designed the study. S.G.L., C.P., D.F.C., D.S., C.N., C.B., T.J.K., W.A.R. and W.T. performed the study. S.G.L., C.P., S.C.F. and C.B.A.W. analysed and wrote the manuscript. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Info
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Lillico, S., Proudfoot, C., Carlson, D. et al. Live pigs produced from genome edited zygotes. Sci Rep 3, 2847 (2013). https://doi.org/10.1038/srep02847
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep02847
This article is cited by
-
The transcriptomic insight into the differential susceptibility of African Swine Fever in inbred pigs
Scientific Reports (2024)
-
Versatile generation of precise gene edits in bovines using SEGCPN
BMC Biology (2023)
-
Breeding studies on red sea bream Pagrus major: mass selection to genome editing
Fisheries Science (2023)
-
Application of new biotechnologies for improvements in swine nutrition and pork production
Journal of Animal Science and Biotechnology (2019)
-
Harnessing genomic information for livestock improvement
Nature Reviews Genetics (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.