Abstract
Drug-induced block of the cardiac hERG (human Ether-à-go-go-Related Gene) potassium channel delays cardiac repolarization and increases the risk of Torsade de Pointes (TdP), a potentially lethal arrhythmia. A positive hERG assay has been embraced by regulators as a non-clinical predictor of TdP despite a discordance of about 30%. To test whether assaying concomitant block of multiple ion channels (Multiple Ion Channel Effects or MICE) improves predictivity we measured the concentration-responses of hERG, Nav1.5 and Cav1.2 currents for 32 torsadogenic and 23 non-torsadogenic drugs from multiple classes. We used automated gigaseal patch clamp instruments to provide higher throughput along with accuracy and reproducibility. Logistic regression models using the MICE assay showed a significant reduction in false positives (Type 1 errors) and false negatives (Type 2 errors) when compared to the hERG assay. The best MICE model only required a comparison of the blocking potencies between hERG and Cav1.2.
Similar content being viewed by others
Introduction
The human ether-à-go-go-related gene (hERG, Kv 11.1) channel has played an especially important role in cardiac safety and the hERG patch clamp assay is a key regulatory requirement before a first in man trial1 (http://www.fda.gov/downloads/Drugs/GuidanceComplianceRegulatoryInformation/Guidances/UCM074963.pdf). HERG produces IKr the rapidly repolarizing potassium current in cardiomyocytes2. Roy et al.3 and Suessbrich et al.4 first showed that block of hERG was the molecular mechanism by which many non-antiarrhythmic drugs produced delayed repolarization, QT prolongation and torsade de pointes (TdP) a potentially lethal cardiac arrhythmia. Because TdP is rare for non-antiarrhythmic drugs, QT prolongation is used as a surrogate marker but by itself is a weak indicator of lethality; the adverse event of importance is the TdP arrhythmia5,6.
The hERG assay is interpreted by correlating in vitro concentration-responses with exposure to estimate the safety margin (SM) or ETPC Index (hERGETPC) defined as hERG IC50/ETPC where ETPC is the effective free therapeutic plasma concentration. The poster drug for the hERG assay's utility was terfenadine (Seldane) because the assay gave the clearest non-clinical signal of the delayed repolarization linked to terfenadine's TdP outcome3,4. A reported odds ratio of 1.93 for the association between anti-hERG activity and the risk of serious ventricular arrhythmias and sudden death provided further support for the hERG assay7.
To ensure safety, regulatory authorities have accorded a very high sensitivity to a positive hERG signal. The emphasis on sensitivity plus the limitations inferred from the odds ratio are consistent with the lack of specificity that has been reported6. As a result it is not known how many potentially useful drugs have been discarded. In fact, some potent blockers of hERG are already in use therapeutically and are not torsadogenic. Such an example is verapamil, a potent hERG channel blocker that is nevertheless used to treat cardiac arrhythmias, hypertension and angina pectoris8. One explanation for this discordance is that the simultaneous block of depolarizing calcium current offsets the effect of hERG block9,10,11,12. Can an understanding of multiple ion channel effects (MICE)9,10,11,12,13 result in a better discrimination between safe and unsafe drugs? We tested this hypothesis by measuring the blocking potencies of 32 torsadogenic (+TdP) and 23 non-torsadogenic (-TdP) drugs on hERG and the depolarizing Cav1.2 and Nav1.5 channel currents. We used logistic regression models to compare torsadogenic probabilities determined by block of hERG alone or block of hERG coupled with block of Cav1.2 and/or block of Nav1.5. Consistent with our hypothesis we found that models using MICE variables significantly improved TdP predictivity. Of particular interest is that our best model only required in vitro measurements of the relative blocking potencies between Cav1.2 and hERG channel currents, independent of effective therapeutic concentrations.
Results
Three major tasks in this project were to: 1) develop a reliable high throughput method to measure concentration-response curves since manual patch clamp would be impractical given the number of drugs and concentration-responses to be tested; 2) define the criteria that best assign risk to a large dataset that includes torsadogenic and non-torsadogenic drugs; and 3) find logistic regression models that best classify drugs for torsadogenic risk. The accomplishment of these tasks is addressed presently and in a Supplementary section.
Task 1: Evaluation of automated patch clamp
The manual whole cell patch clamp is the “gold standard” method in determining the effects of drugs on ion channels, but it requires a high level of technical expertise and its throughput is low. To achieve the necessary throughput for a large dataset we used the QPatch and PatchXpress automated patch clamp (APC) instruments, which can achieve gigaseals. Figure 1 plots the IC50s of 55 drugs for hERG, Cav1.2 and Nav1.5 block and the 95% confidence intervals of the data represented by the vertical lines. The IC50s for the three channels are well defined over several orders of magnitude with confidence intervals that are within the symbols. We noticed a reduced precision of Nav1.5 currents at larger values due to concentrations that approached the solubility limit, requiring that IC50 values be extrapolated assuming a Hill coefficient equal to 1. The hERG data for 29 drugs included in the dataset agree with published manual patch clamp data6,12,14,15. Where comparable protocols were used there was agreement for the Cav1.2 and Nav1.5 results12.
IC50s of the 55 drugs included in the dataset for hERG, Cav1.2 and Nav1.5.
Symbols indicate mean values and lines the 95% confidence intervals of data. Values for the three channels are plotted as a function of drugs numbered in decreased order of potency. The drug order for each channel is indicated in Supplementary Table 3.
Tasks 2 and 3: Criteria used to predict TdP outcomes and subsequently model them
The torsadogenic outcome for each drug in the dataset was determined by consulting Redfern et al.14 and the Arizona CERT database (http://www.azcert.org/), each of which assigned torsadogenic risk to large numbers of drugs. Also consulted were FDA-generated package inserts (http://dailymed.nlm.nih.gov/dailymed/). Annotation is provided in Appendix Table 1.
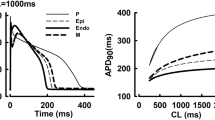
The measured IC50s recorded with the APC instruments were then correlated with the Torsade de Pointes outcomes. Figure 2a contrasts the IC50 values for Nav1.5, Cav1.2 and hERG of the +TdP compound terfenadine with those of the −TdP compound verapamil. To align these measurements with clinical exposure, the figure also shows the ETPC values. For terfenadine, the Nav1.5 and Cav1.2 IC50 values are at least one order of magnitude larger than for hERG block and the ETPC. In the case of verapamil, both Cav1.2 and hERG block are comparable and close to the ETPC. We have shown in human stem cell-derived cardiomyocytes that the pure hERG blocker terfenadine prolongs the action potential duration (APD) at 90% repolarization (APD90) whereas verapamil prolongs the APD90 but shortens the APD30, an indication of calcium channel block16.
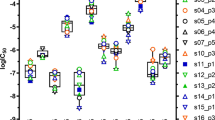
ETPC indexes of torsadogenic and non-torsadogenic drugs.
Terfenadine is concordant and verapamil is discordant with the hypothesis that potent hERG blockers are torsadogenic. (a) IC50 values for hERG, Nav1.5 and Cav1.2 and the maximum effective free plasma concentration. (b) Plot of the ETPC Indexes for hERG (indicated at the interception of blue and orange segments), Cav1.2 (point at the end of the blue segment) and Nav1.5 (point at the end of the orange segment). Dashed lines indicate the region defined by Redfern's ETPC index criteria of 30 for torsadogenic drugs14. (c) The ETPC Indexes for hERG, Cav1.2 and Nav1.5 channels of the 32 +TdP and 23 −TdP drugs included in the dataset. The profile was explained in the legend of Fig. 1. Drugs were numbered for clarity in decreased order of IC50/ETPC values. +TdP drugs included: (1) ibutilide, (2) quinidine, (3) thioridazine, (4) flecainide, (5) halofantrine, (6) droperidol, (7) terodiline, (8) bepridil, (9) procainamide, (10) terfenadine, (11) nilotinib, (12) methadone, (13) sotalol, (14) moxifloxacin, (15) cisapride, (16) paliperidone, (17) haloperidol, (18) sparfloxacin, (19) astemizole, (20) dofetilide, (21) disopyramide, (22) sertindole, (23) clozapine, (24) chlorpromazine, (25) voriconazole, (26) pimozide, (27) sunitinib, (28) cilostazol, (29) solifenacin, (30) paroxetine, (31) risperidone and (32) amiodarone. −TdP drugs included: (1) piperacillin, (2) verapamil, (3) metronidazole, (4) ceftriaxone, (5) linezolid, (6) telbivudine, (7) phenytoin, (8) ribavirin, (9) lamivudine, (10) diltiazem, (11) raltegravir, (12) saquinavir, (13) mibefradil, (14) duloxetine, (15) donepezil, (16) pentobarbital, (17) sitagliptin, (18) dasatinib, (19) diazepam, (20) mitoxantrone, (21) nifedipine, (22) nitrendipine and (23) loratadine. Dashed lines have the same meaning as before.
Figure 2b compares the relationship among the ETPC Indexes for the three channels. The hERGETPC value is the vertex of the blue and orange lines whose ends represent the CavETPC and NavETPC values respectively. Dashed lines indicate the level at which the hERG IC50 is 30-fold above the ETPC, a threshold for torsadogenic drugs14. Although the hERGETPC values for both drugs are well below 30, terfenadine's CavETPC is ten times larger than its hERGETPC value, whereas verapamil's CavETPC is lower than the hERGETPC. This profiling provides a convenient way to illustrate the MICE hypothesis: strong “V” shapes suggest +TdP compounds and “Λ” shapes suggest −TdP compounds.
Figure 2c provides separate profiles of the hERG, Cav1.2 and Nav1.5 ETPC Indexes for the +TdP and −TdP drugs. In each group, drugs were ranked from the lowest to highest hERGETPC values. The plots reveal that +TdP and −TdP drugs can be qualitatively distinguished from each other by their multiple ion channel ETPC Index profiles. In general safe drugs have either large hERGETPC values or small hERGETPC values but in the latter case, ETPC Indexes for at least one depolarizing current are within one order of magnitude of the hERGETPC value. On the other hand, +TdP drugs show small hERGETPC and large NavETPC and CavETPC values indicating very low or absent Nav1.5 and Cav1.2 block compared to hERG block.
The functional dependence between +TdP and the log transformed ETPC indices is shown in Figure 3 . The fraction of +TdP drugs per decade of ETPC indexes estimates the probability of observing a torsadogenic drug P(+TdP) in that range. Each point indicates the averaged torsadogenic assignment (+TdP = 1, −TdP = 0) and the bars indicate the standard error of the mean. Consistent with Fig. 2, high TdP risk is associated with larger −log(hERGETPC) values ( Fig. 3a ). A negative functional relationship between TdP risk and −log(CavETPC) was observed between −3 and zero values of the independent variable ( Fig. 3b ). No clear association between block of peak Nav1.5 currents and the fraction of +TdP drugs per decade was noticed ( Fig. 3c ). Figure 2 suggests that in a background of potent hERG block (hERGETPC < 30, −log(hERGETPC) > −1.477), Cav1.2 block reduces TdP risk. Thus, we explored the functional relationship between P(+TdP) and the difference between Cav1.2 and hERG blocking potencies represented by the variable CavD defined in Methods ( Fig. 3d ). The plot shows the nonlinear relationship between P(+TdP) and CavD. Note that most of the change in TdP risk occurs within one order of magnitude with a center at zero where the hERGETPC to CavETPC ratio is equal to one.
MICE models are better TdP predictors than hERG alone
What combination of the above variables results in a logistic regression model that best predicts TdP risk? To address this question we calculated the maximum log likelihood (Lmax) for each of the six models and then assessed the difference from Model 1 using the likelihood ratio test. Table 1 shows the statistical comparison between models. Note that inclusion of the −log(NavETPC) and/or −log(CavETPC) variables significantly increased the maximum log likelihood of the models and significantly decreased the probability of rejecting Model 1 when true. MICE models are significantly better than Model 1 that uses hERG block alone.
The predictive power of the Models was assessed assigning torsadogenic probabilities to each drug in the dataset using a leave-one-out (LOO) cross-validation procedure described in Methods. The resultant TdP probabilities were used to construct ROC curves to find the optimal cut-off based on the J-index where sensitivity and specificity are maximal. Figure 4a shows the torsadogenic risk of the 55 drugs included in the dataset as calculated under Models 1 (−log(hERGETPC) assay only) and 5 (CavD), the model with the smallest number of misclassified drugs. Figure 4b shows the ROC curves for both models and the J-indexes (cut-off value) indicated by arrows.
The MICE approach results in better predictive Models for TdP.
(a) Calculated torsadogenic risk under Models 1 and 5 based in cross-validated probabilities. Boxes indicated the 25, 50 and 75 percentile; whiskers indicate the 10 and 90 percentiles and filled diamond, mean values. Arrows indicate the probabilities of 0.47 and 0.46 at the J-index for Models 1 and 5 respectively. (b) ROC analysis of Model 1 and 5. The arrows indicate the cutoff point associated with the J-index. The dashed line indicates the performance of a Model that does not discriminate. (c) 2 × 2 Contingency tables for both models. Numbers in parenthesis indicate the area under the curve ± sem. Misclassified drugs are indicated in the tables at the bottom.
Note the poor separation of Model 1 probabilities compared to Model 5. Model 1 falsely identified verapamil as torsadogenic with a probability (P(+TdP) = 0.90) that was similar to terfenadine (P(+TdP) = 0.81). On the other hand, Model 5 correctly identified verapamil as non-torsadogenic (P(+TdP) = 0.31) and terfenadine as torsadogenic (P(+TdP) = 0.97). Under Model 5 only the +TdP drug voriconazole was identified as a false negative and the −TdP drugs telbivudine, loratadine, dasatinib and donepezil as false positives. The calculated torsadogenic probabilities for all Models assessed in this study are shown in Supplementary Table 2 .
Comparison of the ROC curves reveal that 96.9% sensitivity in Models 1 and 5 is achieved with 56.5% and 17.4% false positive rates, respectively. This improvement in TdP prediction is accompanied by a 16 ± 9% increase of the AUC. Both logistic models have no intrinsic classification bias since random allocation of TdP traits in the dataset resulted in ROC curves near the identity line (not shown). Figure 4c shows the 2 × 2 contingency tables at the cutoff determined by the J-index, the AUCs and the drugs misclassified by both models. Note that Model 1 misclassified 14 drugs whereas Model 5 misclassified only five drugs. Thus, compared to Model 1, false positives were halved and false negatives reduced six-fold. Table 1 shows the AUC and the change in the area with respect to Model 1 for all the models. ROC curves and contingency tables for all models are shown in Supplementary Figures 1 and 2 . Based on the D statistics ( Table 1 ), AUCs of the ROC curves ( Fig. 4 and Supplementary Figs. 1 and 2 ) and the LR+ and LR- ( Supplementary Fig. 2 ) Model 5 is the best of all the Models tested. Supplementary Fig. 3 shows the fit of Models 1 and 5 to the experimental data. From this and the previous analysis the relative position of Cav1.2 block with respect to hERG block is the major determinant of Model 5 effectiveness. By virtue of being a ratio, the CavD variable is independent of ETPC and the relative potency of hERG to Cav1.2 block is a feature of the model. The efficiency of CavD as a torsadogenic classifier as shown by the step change in TdP assignments in Figure 3d , is supported by the fact that a procedure not requiring parameter estimation to calculate torsadogenic probabilities using CavD results in a comparable classification ( Supplementary Fig. 3 ).
Discussion
This study confirms our hypothesis that Models based on MICE more accurately predict torsadogenic potential than Models based on hERG effects alone. The difference between Models 1 and 5 is statistically significant. The value of the MICE approach is clearly shown in the plots of Figure 2 , where +TdP and −TdP drugs show distinct profiles determined by the differences in blocking potencies between the depolarizing currents INa and ICa and the repolarizing current mediated by hERG. Unexpectedly, we found that a comparison of the relative blocking potencies between hERG and Cav1.2 was all that was necessary to improve predictivity of TdP for our data set of drugs; effective therapeutic plasma concentrations were not required. As a result the uncertainties surrounding measurement of clinical exposure due to protein binding of drug are removed; a comparison of potencies of block in vitro is all that is required to predict the TdP outcome.
Model 1 had fourteen falsely categorized drugs: ceftriaxone, linezolid, metronidazole, phenytoin, piperacillin, ribavirin, telbivudine and verapamil were false positives whereas amiodarone, cilostazol, paroxetine, risperidone, solifenacin and sunitinib were false negatives. Since the torsadogenic threshold for Model 1 is a hERG index value of approximately 30, it was not surprising to find this set of misclassified drugs. All false positive drugs have hERG Index values below or at the 30 threshold whereas all false negative drugs are weak hERG blockers with hERG Indexes above 90 ( Fig. 2 ).
Model 5 only had one false negative drug (voriconazole) and four false positives (dasatinib, donepezil, loratadine and telbivudine). Possible reasons for these misclassifications are as follows: voriconazole, telbivudine and loratadine displayed a large amount of variability in their Cav1.2 IC50 values ( Supplementary Table 1 ) and because Model 5 does not take into account the variability of the IC50 values, their assigned torsadogenic risk probabilities are less certain. Also, our classification of voriconazole as a torsadogenic drug may be questionable because reported incidents of TdP have been observed in patients with multiple risk co-variates. For example, TdP occurred in a patient who had undergone induction chemotherapy which may have contributed to the development of TdP17.
The reasons for false classification of dasatanib and donepezil are more difficult to explain. Both drugs have “V” shaped profiles indicating much more potent hERG block compared to Nav1.5 and Cav1.2. This is typical of many torsadogenic drugs in our training set so the model may be overly sensitive to drugs that display these characteristics. However, it should be noted that there have been isolated cases of TdP reported for donepezil and even though no cases of TdP have been reported for dasatinib it has been reported that it prolongs the QT interval. Therefore, the TdP risk of these two drugs is greater than others in the −TdP group.
The ETPC, protein binding and Cmax values were taken from multiple sources and we could not control for lab to lab variability in the data. In some cases we were limited by the solubility of the drugs in our buffer. In these cases IC50 values were estimated using a binding equation with a Hill coefficient of 1 to extrapolate from the amount of block observed at achievable concentrations. All recordings were performed at ambient temperatures. Kirsch et al15, demonstrated that erythromycin and dl-sotalol inhibit hERG current more potently at physiological temperature. Interestingly in this study, dl-sotalol is more potent than in Kirsch et al.15 (111.4 μM vs. 278 μM, respectively). Effects on ion channels, other than hERG, Nav1.5 and Cav1.2, were not considered, but block of repolarizing currents other than hERG may increase TdP risk18.
Pentamidine is a drug that does not directly inhibit hERG current but does reduce the surface expression of hERG and causes TdP19 As a result, our models may misidentify drugs that exclusively inhibit the surface expression of hERG. Trafficking data is only available for seven of the drugs in our training set. Amiodarone, thioridazine, chlorpromazine, terfenadine and mibefradil decrease surface expression of hERG but for these drugs, direct block of hERG is the predominant factor in terms of TdP risk because all hERG IC50s are < 2 μM while significant reduction of hERG surface expression occurred at 10 μM19. Cisapride and quinidine do not inhibit the surface expression of hERG19. The rest of the drugs that were used to construct our models have not been evaluated for trafficking effects so it is unknown if any of the false negatives affect ion channel trafficking.
A predictive model is normally fit to a training set of data and a test set is used to assess the strength and utility of the predictive model. The restricted size of the dataset made it impossible to have separate “training” and “test” sets. Thus, we performed a leave-one-out cross validation to calculate the torsadogenic probability of each drug. ROC curves calculated using scrambled assigned torsadogenic classifications had AUC values that were approximately 0.5 for all models demonstrating a lack of bias for particular models (not shown).
We have shown that a logistic regression Model based solely on hERG is not as good as regression Models based on MICE. In addition, the MICE regression Models are better than the conventional approach of identifying torsadogenic drugs as those that have hERG IC50 values less than 30-fold from the ETPC. Using the 30-fold cutoff value resulted in 16 misidentified drugs which is, as expected, similar to the results seen with Model 1 which resulted in 14 misidentified drugs. However, we have shown that logistic regression models that include Cav1.2 or Nav1.5 current block as a mitigating effect are more predictive of TdP risk. Other models that have been used to determine torsadogenic risk include both ex vivo and in vivo models such as the female rabbit ventricular wedge preparation20, the Langendorff-perfused female rabbit heart (Screenit Model)21, Purkinje fiber action potentials22 and AV ablated dogs23. While useful, all of these models share similar disadvantages in comparison to our ion channel approach; they are low throughput, more expensive and therefore can only be practically used late in the drug development process.
Approximately, 90% of all drugs withdrawn from the market are linked to toxicity and safety24. By far the most dramatic risk for all drugs is sudden cardiac death. TdP the outcome of present interest is associated with a death rate of about 20%. Thorough QT studies in man are meant to detect TdP risk in humans but these studies are performed after hundreds of millions of dollars have already been spent on development. For this reason drug developers give the hERG assay extraordinary sensitivity which as the present study shows is unwarranted. The MICE assay is not only better than the hERG assay but, the fact that it uses automated patch clamp instruments, provides higher throughput and faster turn-around times than the manually performed patch clamp method. This study demonstrates that TdP risk can be accurately assessed using MICE early in the development process.
Methods
Drugs
Drugs were from LGM pharma (TN), Selleck Chemicals LLC (TX), Sigma-Aldrich (MO), Synfine research (Canada) and R&D Systems (MN). Providers are indicated for each drug in Supplementary Table 1. Test article stock solutions were prepared in DMSO and stored frozen. Test article concentrations were prepared fresh daily by diluting stock solutions into an extracellular buffer (HEPES-buffered physiological saline solution, HBPS).
Cells
hERG (KCNH2 gene, Kv11.1 channel) channels were stably expressed in HEK293 cells. Nav1.5 (SCN5A gene, hNav1.5 channel) and Cav1.2 (CACNA1C/CACNB2/CACNA2D1 genes, hCav1.2/β2/α2δ channel) channels were stably expressed in CHO cells. All cell lines were from ChanTest Corporation.
Electrophysiology
The IC50 values for block of hERG, Cav1.2 and Nav1.5 channels were measured using QPatch and PatchXpress automatic patch clamp systems. All experiments were performed at ambient temperature.
Solutions
Extracellular solution
The extracellular solution for the three channels in both QPatch-HT and PatchXpress instruments was the HBPS solution containing (in mM): 137 NaCl, 4 KCl, 1.8 CaCl2, 1 MgCl2, 1; 10 HEPES, 10 Glucose, pH adjusted to 7.4 with NaOH (supplemented with 0.3% DMSO.)
Intracellular solutions
hERG currents: The intracellular solution used in QPatch-HT and PatchXpress instruments contained: 130 K-aspartate, 5 MgCl2, 5 EGTA, 4 Na2ATP, 10 HEPES, pH adjusted to 7.2 with KOH.
Cav1.2 currents: The intracellular solution for QPatch-HT experiments contained: 130 Cs-aspartate, 1 CaCl2, 5 MgCl2, 2 EDTA, 10 EGTA, 4 Na2ATP, 10 HEPES, pH adjusted to 7.2 with CsOH. The intracellular solution for PatchXpress experiments contained: 130 Cs-aspartate, 5 MgCl2, 10 EGTA, 4 Na2ATP, 0.1 GTP, 10 HEPES, pH adjusted to 7.2 with CsOH.
Nav1.5 currents: The intracellular solution for QPatch-HT experiments contained: 120 CsF, 2 MgCl2, 10 EGTA, 30 CsCl, 5 NaF, 5 HEPES, pH adjusted to 7.2 with CsOH. The intracellular solution for PatchXpress experiments contained: 130 Cs-aspartate, 5 MgCl2, 5 EGTA, 4 Na2ATP, 0.1 GTP, 10 HEPES, pH adjusted to 7.2 with CsOH.
Voltage protocols
The following voltage protocols were used to evaluate the effects of the drugs on ion channel currents.
hERG currents
hERG block was measured using a stimulus voltage pattern consisting of a 500 ms pulse to −40 mV to assess the leak current, a 2 s activating pulse to +40 mV and a 2 s test pulse to −40 mV (tail current). The pulse pattern was repeated continuously at 10 sec intervals from a holding potential of −80 mV.
Cav1.2 currents
Cav1.2 channel block was measured using a stimulus voltage pattern consisting of a 150 ms test pulse to 0 mV elicited at 5 sec intervals from a −40 mV holding potential. CdCl2 (200 μM) was added at the end of each experiment to block hCav1.2 current and calculate leak. Only one drug, pentobarbital, was evaluated in PatchXpress. In this case the holding and test potentials were −80 mV and +10 mV respectively.
Nav1.5 currents
Nav1.5 channel block was measured using a depolarizing test pulse to −10 mV from a 200 ms hyperpolarizing conditioning pre-pulse to −120 mV. The pulse pattern was repeated at 10 sec intervals from a holding potential of −80 mV. Leak current was calculated after complete inactivation of the sodium current at the test potential.
Analysis of drug effects
For ion channel testing, data acquisition and analyses were performed using the QPatch Assay or DataXpress software. When possible, dose response curves were constructed using concentrations (in half log increments) that blocked approximately 10% to 90% of the ion channel current. However, the maximal concentrations tested were limited by solubility.
Concentration-response data was fitted to an equation of the following form:

Where [Test] was the concentration of test article, IC50 was the concentration of the test article producing half-maximal inhibition, N was the Hill coefficient and % Block was the percentage of ion channel current inhibited at each concentration of the test article. Nonlinear least squares fits was solved with the Solver add-in for Excel (Microsoft, Redmond, WA). Where appropriate, data are expressed as means ± sem.
Modeling
Classification of drugs as +TdP or −TdP
We classified the torsadogenic outcome for each drug in the dataset using the five categories enunciated in Redfern et al.14, the Arizona CERT database (http://www.azcert.org/) case reports in the literature and FDA-generated package inserts (http://dailymed.nlm.nih.gov/dailymed/). The classification is annotated in Appendix Table 1.
Effective therapeutic plasma concentrations
The unbound effective therapeutic plasma concentration (ETPC) for each drug was collected from the literature14 or calculated from Cmax and percent protein binding data found in the literature. For references see Supplementary Table 1.
Statistical methods
The statistical software used was implemented by the R Development Core Team (http://www.R-project.org/).
Description of the data and models
The data set is comprised of n = 55 compounds for which both torsadogenic classification and reliable measurements of ion-channel block were available. We defined the ETPC index as the ratio of the IC50 for channel block to the effective free therapeutic plasma concentration (ETPC). For analysis we use the −log transformed ETPC indexes



Where




are the basic variables.
We also define the variables CavD as C − H and NavD as N − H. D stands for the logarithmic distance of the block potencies between the indicated channel and hERG.
The six logistic regression models for TdP under consideration are

Note that Models 5 and 6 are independent of E.
For the assumptions of logistic regression to hold, the distributions of the arguments in the two subpopulations (TdP = 0 and TdP = 1) must belong to the same one parameter exponential family. For the above six models, this requirement must hold for all linearcombinations of (H + E), (C + E) and (N + E). The simplest way for this to happen is for these three variables to have a multivariate normal distribution in each of the two subpopulations. We investigate this possibility by first checking the univariate distributions in the two subpopulations and then doing a qqplot check of multivariate normality based on Mahalanobis distances25.
Two sample comparisons
Table 2 shows the ETPC adjusted variables for the TdP = 0 and TdP = 1 classes. The only possibly non-normal distribution is that of C − E in the TdP = 0 class, which appears to be a bit left-skewed (not shown). Nevertheless, the distribution does not fail the test for univariate normality.
Check for multivariate normality
If a distribution is multivariate normal, then the squared Mahalanobis distances D2 from the points to its center follow a chi-square distribution with degrees of freedom d equal to the dimension of the data. Here d = 3. The conformity of the data with this theoretical distribution implies that the method of Logistic Regression is appropriate (not shown).
Evaluation of the LR models
The maximum log likelihood (Lmax) of each model was calculated using ad hoc programs written in Java based on the Newton-Raphson method. Differences between models were assessed using the likelihood ratio test (D test). Two times the difference of the log likelihood for each nested model was used to calculate χ2 and the probability of rejecting the model with the lower number of parameters when true26.
Receiver operating characteristic (ROC) analysis
The predictive power of the models was assessed assigning torsadogenic probabilities to each drug in the dataset using a leave-one-out (LOO) cross-validation procedure. To this end 55 datasets (training sets) were built by removing one drug at a time from the total set. For each model, its logistic equation was fitted to each training set and the fitted coefficients used to assign a torsadogenic probability P(+TdP) to the drug that was ‘left-out’. This process was repeated until all compounds had a predicted cross-validated probability assigned. Receiver Operating Characteristic (ROC) curves were used to quantify the results27. An optimal cut-off, set by the Youden's index (J)28 (defined as the threshold point where sensitivity-(1-specificity) is maximal) was chosen to build 2 × 2 contingency tables and calculate likelihood ratios (LR+ = sensitivity/(1-specificity) and LR− = (1-sensitivity)/specificity). ROC curves were compared among the different models measuring the area under the curve (AUC) and the 95% confidence intervals for the differences between AUCs29,30. ROC curves and the AUC were calculated using JMP software (Cary, NC).
References
S7B Nonclinical Evaluation of the Potential for Delayed Ventricular Repolarization (QT Interval Prolongation) by Human Pharmaceuticals (2005 October).
Sanguinetti, M. C., Jiang, C., Curran, M. E. & Keating, M. T. A mechanistic link between an inherited and an acquired cardiac arrhythmia: HERG encodes the IKr potassium channel. Cell. 81, 299–307 (1995).
Roy, M., Dumaine, R. & Brown, A. M. HERG, a primary human ventricular target of the nonsedating antihistamine terfenadine. Circulation. 94, 817–823 (1996).
Suessbrich, H., Waldegger, S., Lang, F. & Busch, A. E. Blockade of HERG channels expressed in Xenopus oocytes by the histamine receptor antagonists terfenadine and astemizole. FEBS Lett. 385, 77–80 (1996).
Sager, P. T. Key clinical considerations for demonstrating the utility of preclinical models to predict clinical drug-induced torsades de pointes. Br. J. Pharmacol. 154, 1544–1549 (2008).
Gintant, G. An evaluation of hERG current assay performance: Translating preclinical safety studies to clinical QT prolongation. Pharmacol. Ther. 129, 109–119 (2011).
De Bruin, M. L., Pettersson, M., Meyboom, R. H., Hoes, A. W. & Leufkens, H. G. Anti-HERG activity and the risk of drug-induced arrhythmias and sudden death. Eur. Heart. J. 6, 590–597 (2005).
Zhang, S., Zhou, Z., Gong, Q., Makielski, J. C. & January, C. T. Mechanism of block and identification of the verapamil binding domain to HERG potassium channels. Circ. Res. 84, 989–998 (1999).
Bril, A. et al. Combined potassium and calcium channel blocking activities as a basis for antiarrhythmic efficacy with low proarrhythmic risk: experimental profile of BRL-32872. J. Pharmacol. Exp. Ther. 276, 637–646 (1996).
Martin, R. L. et al. The utility of hERG and repolarization assays in evaluating delayed cardiac repolarization: influence of multi-channel block. J. Cardiovasc. Pharmacol. 43, 369–379 (2004).
Lacerda, A. E., Kuryshev, Y. A., Yan, G. X., Waldo, A. L. & Brown, A. M. Vanoxerine: cellular mechanism of a new antiarrhythmic. J. Cardiovasc. Electrophysiol. 21, 301–310 (2010).
Mirams, G. R. et al. Simulation of multiple ion channel block provides improved early prediction of compounds' clinical torsadogenic risk. Cardiovasc. Res. 91, 53–61 (2011).
Hanson, L. A. et al. ILSI-HESI cardiovascular safety subcommittee initiative: evaluation of three non-clinical models of QT prolongation. J. Pharmacol. Toxicol. Methods. 54, 116–129 (2006).
Redfern, W. S. et al. Relationships between preclinical cardiac electrophysiology, clinical QT interval prolongation and torsade de pointes for a broad range of drugs: evidence for a provisional safety margin in drug development. Cardiovasc. Res. 58, 32–45 (2003).
Kirsch, G. E. et al. Variability in the measurement of hERG potassium channel inhibition: effects of temperature and stimulus pattern. J. Pharmacol. Toxicol. Methods. 50, 93–101 (2004).
Peng, S., Lacerda, A. E., Kirsch, G. E., Brown, A. M. & Bruening-Wright, A. The action potential and comparative pharmacology of stem cell-derived human cardiomyocytes. J. Pharmacol. Toxicol. Methods. 61, 277–286 (2010).
Philips, J. A. et al. Torsades de pointes associated with voriconazole use. Transpl. Infect. Dis. 1, 33–36 (2007).
Towart, R. et al. Blockade of the I(Ks) potassium channel: an overlooked cardiovascular liability in drug safety screening? J. Pharmacol. Toxicol. Methods. 60, 1–10 (2009).
Kuryshev, Y. A. et al. Pentamidine-induced long QT syndrome and block of hERG trafficking. J. Pharmacol. Exp. Ther. 312, 316–323 (2005).
Liu, T. et al. Blinded validation of the isolated arterially perfused rabbit ventricular wedge in preclinical assessment of drug-induced proarrhythmias. Heart Rhythm. 3, 948–956 (2006).
Valentin, J. P., Hoffmann, P., De Clerck, F., Hammond, T. G. & Hondeghem, L. Review of the predictive value of the Langendorff heart model (Screenit system) in assessingthe proarrhythmic potential of drugs. J. Pharmacol. Toxicol. Methods. 49, 171–81 (2004).
Gintant, G. A., Limberis, J. T., McDermott, J. S., Wegner, C. D. & Cox, B. F. The canine Purkinje fiber: an in vitro model system for acquired long QT syndrome and drug-induced arrhythmogenesis. J. Cardiovasc. Pharmacol. 37, 607–618 (2001).
Thomsen, M. B. et al. Sudden cardiac death in dogs with remodeled hearts is associated with larger beat-to-beat variability of repolarization. Basic Res. Cardiol. 100, 279–287 (2005).
Schuster, D., Laggner, C. & Langer, T. Why Drugs Fail – A Study on Side Effects in New Chemical Entities. Current Pharmaceutical Design, 11, 3545–3559 (2005).
Venables, W. N. & Ripley, B. D. Modern Applied Statistics with S, Fourth Edition, Springer: New York, USA, 2002.
Hosmer, D. W. & Lemeshow, S. Applied Logistic Regression. John Wiley and Sons, Hoboken, New Jersey, USA, 2000.
Zweig, M. H. & Campbell, G. Receiver-operating characteristic (ROC) plots: a fundamental evaluation tool in clinical medicine. Clin. Chem. 39, 561–577 (1993).
Youden, W. J. Index for rating Diagnostic tests. Cancer. 3, 32–35 (1950).
DeLong, E. R., DeLong, D. M. & Clarke-Pearson, D. L. Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics. 44, 837–845 (1988).
Hanley, J. A. & Hajian-Tilaki, K. O. Sampling variability of nonparametric estimates of the areas under receiver operating characteristic curves: an update. Acad. Radiol. 4, 49–58 (1997).
Acknowledgements
Research reported in this publication was supported by the National Heart, Lung and Blood Institute of the National Institutes of Health under Award Number R44HL104948. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
A.M.B. conceived the project, J.K., C.A.O.-P., Y.K.A. and A.B.-W. designed the experiments, J.K., C.A.O.-P., Y.K.A. and A.B.-W. analyzed the data, J.K., C.A.O.-P., G.M., J.V.S. and A.M.B. wrote the manuscript, C.A.O.-P., G.M. and J.V. developed the statistical models and C.A.O.-P., G.M. and J.V.S. performed statistical modeling.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Kramer, J., Obejero-Paz, C., Myatt, G. et al. MICE Models: Superior to the HERG Model in Predicting Torsade de Pointes. Sci Rep 3, 2100 (2013). https://doi.org/10.1038/srep02100
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep02100
This article is cited by
-
Evaluation of rosuvastatin-induced QT prolongation risk using real-world data, in vitro cardiomyocyte studies, and mortality assessment
Scientific Reports (2023)
-
The atypic antipsychotic clozapine inhibits multiple cardiac ion channels
Naunyn-Schmiedeberg's Archives of Pharmacology (2023)
-
Long-term in vitro recording of cardiac action potentials on microelectrode arrays for chronic cardiotoxicity assessment
Archives of Toxicology (2023)
-
Computational analysis of arrhythmogenesis in KCNH2 T618I mutation-associated short QT syndrome and the pharmacological effects of quinidine and sotalol
npj Systems Biology and Applications (2022)
-
The grapefruit polyphenol naringenin inhibits multiple cardiac ion channels
Naunyn-Schmiedeberg's Archives of Pharmacology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.