Abstract
Electron tomography in materials science has flourished with the demand to characterize nanoscale materials in three dimensions (3D). Access to experimental data is vital for developing and validating reconstruction methods that improve resolution and reduce radiation dose requirements. This work presents five high-quality scanning transmission electron microscope (STEM) tomography datasets in order to address the critical need for open access data in this field. The datasets represent the current limits of experimental technique, are of high quality, and contain materials with structural complexity. Included are tomographic series of a hyperbranched Co2P nanocrystal, platinum nanoparticles on a carbon nanofibre imaged over the complete 180° tilt range, a platinum nanoparticle and a tungsten needle both imaged at atomic resolution by equal slope tomography, and a through-focal tilt series of PtCu nanoparticles. A volumetric reconstruction from every dataset is provided for comparison and development of post-processing and visualization techniques. Researchers interested in creating novel data processing and reconstruction algorithms will now have access to state of the art experimental test data.
Design Type(s) | reference design • nanomaterial structure generation objective |
Measurement Type(s) | 3D structure determination assay |
Technology Type(s) | electron tomography |
Factor Type(s) |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
Electron tomography attempts to reconstruct 3D objects from 2D projection images taken at different viewing angles, or tilts—producing the entire internal structure of a specimen or region of interest. Since the first 3D reconstruction from electron micrographs1, tomography with the scanning transmission electron microscope (STEM) has been widely applied to nanoscale materials2–12. Utilizing the sub-angstrom 2D resolution of modern STEM, 3D reconstructions with sub-nanometre and even atomic detail have been demonstrated13–16. In the design of advanced nanomaterials, 3D characterization of nano-scale structure offers valuable insight into a material’s macroscale function. As a result, demand for nanoscale STEM tomography is high17.
Routine tomographic methods face several challenges that reduce final quality. The geometry of most specimens and specimen holders restricts tilt range to less than roughly 140°. Commonly referred to as ‘the missing wedge’, an incomplete tilt-range limits the information available for reconstruction and manifests as an elongation in the final reconstruction18,19. Contamination and specimen radiation sensitivity limit the number of viewing angles and the signal to noise, which in turn, restricts the final resolution in 3D20. Depth-of-field limits the maximum allowable size of the object that can be reconstructed; a particular problem for aberration corrected STEM13,21,22.
Recently, efforts towards new reconstruction methods promise higher resolution reconstructions using fewer viewing angles and lower radiation doses than traditional reconstruction algorithms like Weighted Back Projection (WBP) and Simultaneous Iterative Reconstruction Technique (SIRT). New approaches such as the iterative Fourier-based equal slope tomography23 and compressed sensing inspired algorithms24,25 have demonstrated success in STEM tomography by improving reconstruction quality with reduced sampling. However, we still lack a fundamental understanding of when and how these algorithms fail. Adopting new algorithms into routine tomography requires thorough investigation17.
A lack of high-quality, open access data is impeding development and validation of new algorithms and software for 3D reconstruction, visualization, and analysis. Currently, the best tomographic datasets are harboured by a privileged few. Researchers best suited for creating novel data processing and analysis techniques do not readily have access to experimental data.
To address this deficiency, we present five datasets that have pushed the limits of electron tomography. Each dataset was acquired using a unique experimental technique, is of high quality, and contains materials with structural complexity:
Tom_1) Tomography of Hyperbranched Co2P Nanoparticle: a 150° tomographic tilt series, taken at 2° increments. The Co2P nanocrystal has a complex morphology of bundled branches that resembles a six-pointed star26. This dataset represents a tilt range and increment typical of nano-scale STEM tomography.
Tom_2) 180° Tomography of NPs on Nanofibre: a tilt series taken at 1° tilt increments over the full 180° tilt range of platinum nanoparticles on a graphitized carbon nanofibre support. This dataset provides a complete range of tilts, allowing researchers to better understand the effects of missing information. 16 fast acquisition images were acquired at each tilt, and the experimental signal-to-noise level can be adjusted by averaging different numbers of these images.
Tom_3) Atomic Resolution Tomography of Pt NP: a 145° equal slope tomography tilt series of a single platinum nanoparticle, acquired at atomic resolution, enabling reconstruction of atomic features15.
Tom_4) Atomic Resolution Tomography of Tungsten Needle: An equal slope tomography tilt series of the tip of a tungsten needle, acquired over the full 180° range, enabling atomic resolution reconstruction16.
Tom_5) Through-Focal Tomography of Pt-Cu Catalyst: a 138° through-focal tomographic tilt series acquired in an aberration-corrected microscope of Pt-Cu fuel cell catalyst nanoparticles with a complex internal pore structure on an extended carbon support at 3° increments13. This dataset overcomes the limited depth of field that accompanies high-resolution aberration corrected imaging21 by combining through-focal sectioning and tilt-series tomography to reconstruct extended objects.
The datasets include raw tilt series aligned for reconstruction and 3D reconstructions of each specimen—all in an easily readable TIF format.
Combined, these datasets provide a standard, open set of test data for the growing field of tomographic reconstruction and visualization. The datasets allow researchers to rigorously test their algorithms from alignment to reconstruction on real experimental data. The datasets will also find a use as a training tool for scientists new to tomography, a validation tool for 3D tomographic visualization, and a template to seed a future open library of tomographic data.
Methods
In ADF-STEM electron tomography, a focused electron beam with sub-nanometre diameter is rastered across a sample of interest. Electrons scattered from the sample are recorded using an annular dark field detector, which generates a 2D projection image of the sample27,28.
The viewing angle is changed by rotating the specimen and a series of projection images from different angles is acquired. For the vast majority of electron microscopes, a single axis of rotation is permitted by the stage, although more complex tilt geometries have been demonstrated17,29,30. After each successive tilt during the experiment, the specimen moves relative to the electron beam and must be re-centred. The specimen can only be re-centred approximately at the time of acquisition and so the data is said to be ‘misaligned’. The end result of a tomographic experiment is a set of ‘misaligned’ STEM images corresponding to a specific specimen tilt.
Aligning the STEM images prior to reconstruction is vital to establish a common axis of rotation (i.e., tilt-axis) in the image series. Alignment methods include the use of fiducial markers31,32, cross correlation33, and centre of mass. Once aligned, a reconstruction algorithm is used to generate a 3D reconstruction of the sample.
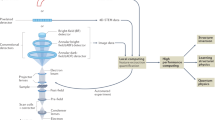
The basic steps of the tomographic method are illustrated in Fig. 1.
Series of 2D images acquired of object of unknown 3D structure at different viewing angles. Images shown are from tiltser_Co2P.tif. 2D images combined into image stack ordered by viewing angle i.e., a tilt series. Tilt series is aligned, and reconstruction algorithm is applied to produce 3D reconstruction of object. A 3D isosurface visualization of recon_Co2P.tif is shown as an example rendered using tomviz.
Tom_1: Tomography of hyperbranched Co2P nanoparticle
Sample preparation
Hyperbranched Co2P nanocrystal synthesis methods and scientific relevance are discussed in detail by Zhang et al.26 Samples were prepared for tomographic analysis by pipetting a drop of organic solution containing Co2P nanocrystals onto the surface of a copper TEM grid coated with an amorphous carbon film. Once the organic solution had dried, Co2P nanocrystals were dispersed over the grid. A drop of a solution of gold nanoparticles in water was then pipetted onto the grid, and allowed to dry. The gold nanoparticles were used as fiducial markers to align the tilt series.
Data acquisition
The tomographic tilt series of Co2P nanocrystals was acquired using an FEI Tecnai F20 scanning transmission electron microscope (STEM) at Cornell University. The microscope was operated at an accelerating voltage of 200 kV, with a convergence semi-angle of 9.6 mrad, and beam current of ~8–10 pA. This yields a nominal 2D resolution of up to 1.6 Å for STEM annular dark field (ADF) images. The tomographic tilt series was acquired over a 150° range at 2° intervals using a high angle annular dark-field (HAADF) detector. The scale in each image is ~0.71 nm per pixel.
Alignment and reconstruction
Each of the 76 projections in the tilt series, tiltser_Co2P.tif, has been aligned to a fiducial particle close to the Co2P nanocrystal using manual alignment techniques (except projections 50 and 51, which are blank to correct for 4° goniometer backlash during acquisition). Similarly, the tilt axis was determined by manually choosing the axis of rotation that minimized artifacts and maximized detail in the final reconstruction. We provide an example reconstruction, recon_Co2P.tif, produced from tiltser_Co2P.tif using the SIRT algorithm.
Tom_2: 180° Tomography of nanoparticles on nanofibre
Sample preparation
Graphitized nanofibres, loaded with platinum nanoparticles at 10 wt. % were dispersed in a methanol solution and dried onto the tip of a tungsten omniprobe needle. The needle was inserted into a Fischione 2050 On-Axis Rotation Tomography Holder for data acquisition.
Data acquisition
One 93° tilt series and one 95° tilt series were acquired with an offset of ~85.3° between the viewing angle of the first image of the first series, and the viewing angle of the first image of the second series. The two tilt series together therefore cover slightly more than the full 180° tilt range. The overlapping region between the two tilt series was used to align them together in post-processing. The angular increment for each tilt series was 1°. In order to reduce scan noise and thus improve signal to noise ratio, 16 images, each with a 1 μs per pixel dwell time, were recorded at each viewing angle and saved as an image stack to be aligned in post processing. Data was acquired using an FEI Tecnai F20 scanning transmission electron microscope (STEM) at Cornell University. The microscope was operated in low angle annular dark field (LAADF) mode an accelerating voltage of 200 kV with a probe current of approximately 5 pA. A convergence angle of ~6.9 mrad was used to optimize resolution over a large depth of field. Images were acquired with 1024×1024 pixels. Non-orthogonality in probe scan direction was observed and was corrected in all images by applying a 0.6 degree shear parallel to the tilt axis with linear interpolation. The field of view in each image was 363.52 nm.
Alignment and reconstruction
Each of the 1 μs per pixel image stacks of 16 images was aligned by cross correlation. Each stack was then summed to form a single image. The single images from both tilt series were then combined into a single 180° image stack, tiltser_180.tif, for reconstruction, with duplicate viewing angles discarded. The tilt series was aligned using a centre of mass method. A 3D reconstruction of the data, recon_180.tif, was produced using a weighted back projection algorithm. Data illustrated in Fig. 2a.
(a) Sample image from tiltser_180.tif. Mixed 3D volume/isosurface visualizations of recon_180.tif show exterior of fibre, with nanoparticles visible on exterior, and hollow interior of nanofibre, containing nanoparticles. (b) Sample image from tiltser_PtNP.tif. Mixed 3D volume/isosurface visualization of recon_PtNP.tif and volume visualization of 3D Fourier transform of recon_PtNP.tif, showing platinum reciprocal lattice spots. (c) Sample image from tiltser_W.tif. Mixed 3D volume and isosurface visualization of recon_W.tif and of the 3D Fourier transform (cropped) of recon_W.tif, showing tungsten reciprocal lattice spots. All 3D visualizations produced using tomviz.
Tom_3: Atomic resolution tomography of platinum nanoparticle
Sample preparation
Platinum nanoparticles were deposited onto a grid consisting of a 5-nm-thick silicon nitride membrane with dimensions of 100 μm×1500 μm, supported on a 100 μm-thick silicon frame designed for loading into a TEM (TEMwindows.com). High temperature coating of a 1–2 nm carbon layer was applied to mitigate charging effects due to the electron beam. The grid was then loaded onto a Fischione 2020 tomographic sample holder for data acquisition in the TEM.
Data acquisition
The tomographic tilt series of platinum nanoparticles was acquired using an uncorrected FEI Titan STEM at the University of California, Los Angeles. The microscope was operated with a beam energy of 200 keV, a 100 pA probe current, and a 10.7 mrad convergence semi-angle. A tilt series of 104 projections was acquired from a platinum nanoparticle with equal-slope increments and a tilt range of ±72.6°.
Alignment and reconstruction
The images in the tilt series, tiltser_PtNP.tif, were aligned using a centre of mass (CM) alignment method after background subtraction and removal15. We present a reconstruction of this data, recon_PtNP.tif, produced using the equal slope tomography (EST) iterative algorithm, a method described by Miao et al.23 No Fourier filters were applied to the final reconstruction. Data illustrated in Fig. 2b.
Tom_4: Atomic resolution tomography of tungsten needle
Sample preparation
A 99.95% pure tungsten wire was annealed under tension, creating a large crystalline domain with the [011] crystallographic axis aligned along the wire axis. The wire was electrochemically etched in a NaOH solution to form a sharp tip with a <10 nm diameter. The wire was plasma cleaned in an Ar/O2 gas mixture and then heated to 1,000 °C under vacuum (~10−5 Pa) to remove the oxide layer generated by the plasma cleaning. The wire was mounted in a 1 mm sample puck compatible with the TEAM microscope stage.
Data acquisition
Tomographic data was acquired using the TEAM I at the National Center for Electron Microscopy. The microscope was operated at 300 kV beam voltage in ADF-STEM mode with a convergence semi-angle of ~ 30 mrad and a ~ 70 pA beam current. The tomography rotation axis was aligned to the wire axis [011]. An equally sloped tomographic tilt series of 62 images, covering the complete angular range of ±90° was acquired from the tungsten needle sample. Two images of 1024×1024 pixels each with 6 μs per pixel dwell time and 0.405 Å pixel resolution were acquired at each angle in order to correct for drift. The TEAM stage, which is a tilt-rotate design with full 360° rotation about both axes, enabled rotation around the [011] crystalline axis.
Alignment and reconstruction
Raw experimental data can be found in tiltser_W.zip, which contains tif stacks of the two images acquired at each viewing angle, as described above. In addition, we provide an aligned tilt series, tiltser_W.tif. In the raw data, the tilt axis has a different in-plane orientation at each viewing angle, and in order to obtain an aligned tilt series, this was corrected by using Fourier methods to align the tilt direction along the image horizontal in every image. Both sample drift and scan distortion were corrected for in all images in the tilt series using Fourier techniques34. The tilt series was then aligned using a centre of mass method, with a mask applied to remove background noise. The tilt series was cropped in order to only feature the tip of the needle, which remained within the depth of focus throughout data acquisition. We present a reconstruction of the data, recon_W.tif, produced from tiltser_W.tif using the equal slope tomography iterative algorithm. The alignment and reconstruction process is explained in detail by Xu et al.16 Data illustrated in Fig. 2c.
Tom_5: Through-focal tomography of Pt-Cu catalyst
Sample preparation
The through focal tilt series was acquired on PtCu nanoparticles on a 3D Vulcan carbon support. The synthesis methods and scientific relevance of the nanoparticles as a fuel-cell electrocatalyst are discussed in detail by Wang et al.35 To prepare for observation in the electron microscope, the particles were suspended in ethanol and pipetted onto a copper TEM grid with an ultra-thin, holey carbon support film.
Data acquisition
The through-focal tomographic tilt series of de-alloyed PtCu nanoparticles on an extended 3D carbon support was acquired using TEAM I at the National Center for Electron Microscopy; a tool that provides attributes to best demonstrate the advantages of this technique. Its large convergence angle provides high lateral resolution (<0.78 Å) and a small depth-of-field (~6 nm) at 300 kV accelerating voltage. Shadowing from the TEM grid limited tilts from −68° to +71° along our chosen axis of rotation.
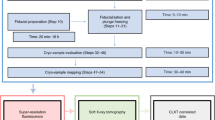
The tomographic data was acquired over a 138° tilt range using a high angle annular dark field (HAADF) detector. The 30 mrad convergence angle provided a continuum of information in the through-focal CTF that spanned a ±1.72° wedge at low and medium frequencies. A 3° tilt increment was chosen to match the convergence angle. The PtCu nanoparticles decorate a 3D Vulcan carbon support with an extended structure that far exceeds the microscope’s depth of field—making it impossible to image multiple particles in-focus within a single field of view. At every tilt a 26 image through-focal series was taken over ±250 nm defocus with 20 nm focal steps in order to ensure all objects were imaged in focus. The microscope defocus steps are calibrated from a through-focal stack (Fig. 3). Each image had a 0.38 nm per pixel lateral resolution.
A through-focal image series must be acquired at each viewing angle in through-focal tomography. Files 018.tif, 072.tif, and 120.tif are shown as examples. Through-focal tomography allows objects from an extended field of view to be reconstructed at high resolution in an aberration corrected STEM. A 3D isosurface visualization of the full view of PtCu nanoparticles on an extended carbon support in recon_ThroughFocal.tif is shown, along with high resolution 3D visualizations of individual PtCu particles in the reconstruction. All visualizations produced using tomviz.
Alignment and reconstruction
A five-dimensional alignment of the raw data in tiltser_ThroughFocal.zip was required: transverse x-y alignment, focal z-alignment, tilt axis rotation and shift. A fiduciary particle was used to align each through-focal stack in their respective x-y direction. The focal z-alignment for each focal stack was determined by identification of the best focus image to a fiduciary particle. Within each focal stack a cross-correlation alignment was used to reduce the small amounts of drift during the acquisition. After alignment, the data was reweighted in Fourier space by dividing with the microscope’s contrast transfer function (CTF) approximated by a 300 keV 30 mrad aberration-free probe plus a Wiener constant of 5 times the max CTF value. After this light deconvolution, each through-focal stack was mapped onto a universal Fourier space by bilinear extrapolation. This extrapolation distributes the complex value of an input point to its four nearest neighbors on the output Cartesian grid with a weighted average of points from all the through-focal stacks. A direct inverse 3D Fourier transform provided the final reconstruction, recon_ThroughFocal.tif. This method is described by Hovden et al.13. It should be noted that the alignment of each through-focal stack generated excess blank images for reference. Thus in the raw data provided, there are 43 images per stack; 26 images of the PtCu nanoparticles, and 17 blank reference images.
Code availability
Code equivalent to that used to reconstruct the data in Tom_1 and Tom_2 is available as part of the open source Tomviz software package at www.tomviz.org. Code used to reconstruct the data in Tom_3 is freely available online at http://www.physics.ucla.edu/research/imaging/EST/. Code used to reconstruct the data in Tom_4 is freely available online at http://www.physics.ucla.edu/research/imaging/3Datoms.
Code used to reconstruct the data in Tom_5 is available in the Supplementary Information to this paper (Supplementary File 1). Alignment tools are available as part of the open source Tomviz software package at www.tomviz.org, and the open source IMOD software package at http://bio3d.colorado.edu/imod/.
Data Records
The datasets described in this paper are available at Figshare (Data Citation 1). All files are provided in 16-bit tif image format. Table 1 describes the content of the raw tilt series datasets. Table 2 describes tilt series derived from raw data, from which reconstructions are in turn derived, and Table 3 describes the sample reconstructions derived from the raw datasets that we have provided. All tilt series have their axis of rotation along the x-axis (horizontal) of the images.
Further information on Tom_2: 180 degree tomography of NPs on nanofibre
tiltser_180.zip contains 190.tif stacks. Each stack is a series of sixteen images of one viewing angle, with each image acquired at 1 μs per pixel dwell time. The images in the each of stacks have been aligned by cross correlation. The image stacks have also been aligned with each other, allowing users to construct their own aligned tilt series from the image stacks. A full file listing for tiltser_180.zip is given below. Roman numerals indicate a stack from the first or the second tilt series. The number in the label indicates the viewing angle given by the microscope goniometer.
I_00.tif I_12.tif I_-23.tif I_35.tif I_-46.tif I_01.tif I_-12.tif I_24.tif I_-35.tif I_47.tif I_-01.tif I_13.tif I_-24.tif I_36.tif II_00.tif I_02.tif I_-13.tif I_25.tif I_-36.tif II_01.tif I_-02.tif I_14.tif I_-25.tif I_37.tif II_-01.tif I_03.tif I_-14.tif I_26.tif I_-37.tif II_02.tif I_-03.tif I_15.tif I_-26.tif I_38.tif II_-02.tif I_04.tif I_-15.tif I_27.tif I_-38.tif II_03.tif I_-04.tif I_16.tif I_-27.tif I_39.tif II_-03.tif I_05.tif I_-16.tif I_28.tif I_-39.tif II_04.tif I_-05.tif I_17.tif I_-28.tif I_40.tif II_-04.tif I_06.tif I_-17.tif I_29.tif I_-40.tif II_05.tif I_-06.tif I_18.tif I_-29.tif I_41.tif II_-05.tif I_07.tif I_-18.tif I_30.tif I_-41.tif II_06.tif I_-07.tif I_19.tif I_-30.tif I_42.tif II_-06.tif I_08.tif I_-19.tif I_31.tif I_-42.tif II_07.tif I_-08.tif I_20.tif I_-31.tif I_43.tif II_-07.tif I_09.tif I_-20.tif I_32.tif I_-43.tif II_08.tif I_-09.tif I_21.tif I_-32.tif I_44.tif II_-08.tif I_-10.tif I_22.tif I_-33.tif I_45.tif II_-09.tif I_11.tif I_-22.tif I_34.tif I_-45.tif II_10.tif I_-11.tif I_23.tif I_-34.tif I_46.tif II_-10.tif II_11.tif II_-18.tif II_26.tif II_-33.tif II_41.tif II_-11.tif II_19.tif II_-26.tif II_34.tif II_-41.tif II_12.tif II_-19.tif II_27.tif II_-34.tif II_42.tif II_-12.tif II_20.tif II_-27.tif II_35.tif II_-42.tif II_13.tif II_-20.tif II_28.tif II_-35.tif II_43.tif II_-13.tif II_21.tif II_-28.tif II_36.tif II_-43.tif II_14.tif II_-21.tif II_29.tif II_-36.tif II_44.tif II_-14.tif II_22.tif II_-29.tif II_37.tif II_-44.tif II_15.tif II_-22.tif II_30.tif II_-37.tif II_45.tif II_-15.tif II_23.tif II_-30.tif II_38.tif II_-45.tif II_16.tif II_-23.tif II_31.tif II_-38.tif II_46.tif II_-16.tif II_24.tif II_-31.tif II_39.tif II_-46.tif II_17.tif II_-24.tif II_32.tif II_-39.tif II_47.tif II_-17.tif II_25.tif II_-32.tif II_40.tif II_-47.tif II_18.tif II_-25.tif II_33.tif II_-40.tif II_48.tif
Further information on Tom_3: Atomic resolution tomography of platinum nanoparticle
The images in tiltser_PtNP.tif were acquired using the equal slope tomography method. Rather than acquiring images at a fixed angular increments as in traditional tomography, the images are acquired at viewing angles that give equal increments of the slope of the Fourier transformed image planes in Fourier space23. The viewing angles (in degrees) associated with each of the 109 images in the tif stack are listed below:
72.646 45.000 17.354 −20.556 −46.848 71.030 44.091 15.709 −22.109 −47.816 69.44 43.152 14.036 −23.629 −48.814 67.891 42.184 12.339 −25.115 −49.844 66.371 41.186 10.62 −26.565 −50.906 64.885 40.156 8.8807 −27.979 −52.001 63.435 39.094 7.125 −29.358 −53.130 62.021 37.999 5.3558 −30.700 −54.293 60.642 36.870 3.5763 −32.005 −55.491 59.300 35.707 1.7899 −33.275 −56.725 57.995 34.509 0.0000 −34.509 −57.995 56.725 33.275 −1.7899 −35.707 −59.300 55.491 32.005 −3.5763 −36.870 −60.642 54.293 30.700 −5.3558 −37.999 −62.021 53.130 29.358 −7.1250 −39.094 −63.435 52.001 27.979 −8.8807 −40.156 −64.885 50.906 26.565 −10.620 −41.186 −66.371 49.844 25.115 −12.339 −42.184 −67.891 48.814 23.629 −14.036 −43.152 −69.444 47.816 22.109 −15.709 −44.091 −71.030 46.848 20.556 −17.354 −45.000 −72.646 45.909 18.970 −18.970 −45.909
Further information on Tom_4: Atomic resolution tomography of tungsten needle
The images in tiltser_W.zip were acquired using the equal slope tomography method. Rather than acquiring images at a fixed angular increments as in traditional tomography, the images are acquired at viewing angles that give equal increments of the slope of the Fourier transformed image planes in Fourier space23. The viewing angles (in degrees) associated with each of the 63 image stacks is given in the filename of each image. A list of files in tiltser_W.zip is given below. In order to obtain a reconstruction from the raw images, they must be combined into a tilt series, and aligned (see Tom_4 Atomic Resolution Tomography of Tungsten Needle under Methods).
0.tif 39.tif 69.3.tif 116.4.tif 147.8.tif 3.5.tif 41.1.tif 72.5.tif 119.2.tif 150.5.tif 7.1.tif 43.1.tif 75.9.tif 121.9.tif 153.3.tif 10.6.tif 44.9.tif 79.3.tif 124.4.tif 156.2.tif 14.tif 46.8.tif 82.7.tif 126.7.tif 159.1.tif 17.3.tif 48.8.tif 86.3.tif 128.9.tif 159.3.tif 20.5.tif 50.8.tif 89.8.tif 131.1.tif 164.3.tif 23.6.tif 53.1.tif 93.5.tif 134.9.tif 167.8.tif 26.5.tif 55.4.tif 97.tif 136.7.tif 168.tif 29.3.tif 57.9.tif 100.4.tif 138.7.tif 175.9.tif 32.tif 60.6.tif 103.9.tif 140.8.tif 180.tif 34.5.tif 63.3.tif 107.2.tif 143.tif 36.8.tif 66.3.tif 110.4.tif 145.3.tif
Further information on Tom_5: Through-focal tomography of Pt-Cu catalyst
tiltser_ThroughFocal.zip contains 47 tif stacks. Each stack is an individual through focal series taken at a different viewing angle. Each of these tif stacks contains 43 images, 26 images of the sample, and 17 blank reference images for through-focal alignment. A reconstruction incorporating all of the information available in the data must be produced directly from the images in each of the 47 image stacks, rather than by combining the stacks into a single file tilt series as for the datasets above.
The title of each tif stack is the viewing angle in degrees that the through focal series was acquired at, measured from the first viewing angle. For example, the first through-focal tif stack, taken at 0°, is named 000.tif. The second tif stack, taken at a tilt of 3° relative to the first is labelled 003.tif. The focal increment between each image in each tif stack is 20 nm.
A full file listing for tiltser_ThroughFocal.zip is given below:
000.tif 030.tif 060.tif 090.tif 120.tif 003.tif 033.tif 063.tif 093.tif 123.tif 006.tif 036.tif 066.tif 096.tif 126.tif 009.tif 039.tif 069.tif 099.tif 129.tif 012.tif 042.tif 072.tif 102.tif 132.tif 015.tif 045.tif 075.tif 105.tif 135.tif 018.tif 048.tif 078.tif 108.tif 138.tif 021.tif 051.tif 081.tif 111.tif 024.tif 054.tif 084.tif 114.tif 027.tif 057.tif 087.tif 117.tif
Image 39 in stack 120.tif was not used as part of the direct Fourier reconstruction of the data because it was observed to contain a large scan distortion.
Technical Validation
The electron microscopes used to acquire the datasets described in this paper were professionally maintained and aligned for optimal imaging conditions prior to dataset acquisition. For Tom_1, Tom_2, Tom_3, and Tom_4 an appropriately sized C2 aperture was selected for data acquisition in order to produce a depth of field that extended over the entire height of the object (or section of the object) to be imaged and reconstructed. For Tom_5 a small depth of field enhanced the through focal technique by providing more 3D information at every tilt. The images in each tilt series presented in this paper have been aligned. The high quality sample reconstructions we provide validate the accuracy and quality of the raw data. No obvious signs of morphological distortions that would indicate poor data acquisition are visible in either the raw data or in the reconstructions themselves. Sinograms of the reconstruction were also inspected for proper alignment to ensure that reconstructions of the highest quality were obtained18.
Usage Notes
Collectively, these tomographic datasets of nanoscale materials provide a standard for the development and validation of new 3D imaging methods—from alignment, to reconstruction, to visualization and analysis. Their uses are diverse. The tilt series data can be intentionally degraded by adding misalignment or noise to explore its influence on a particular reconstruction algorithm. In Tom_2 true experimental noise can be added by discarding images within each tilt, thereby reducing the signal to noise ratio. The effects of increasing missing wedge size or tilt increment size can be explored by removing projections from the 180° tilt range in Tom_2 or Tom_4. The high resolution of data in Tom_3 and Tom_4 provide lattice peaks in the 3D Fourier transform of the final reconstruction. These lattice peaks may have appeal to understanding post processing filters. Lastly, Tom_5 provides exploration the limited depth of field that accompanies a new generation of aberration-electron microscopes and its influence on tomography. Each reconstruction provides a playground for visualization and standards for comparison. Tom_1 in particular has an intricate morphology and aesthetic beauty.
This manuscript illustrates the steps necessary to acquire, align, and reconstruct data from nanoscale specimens at the highest quality. The educational utility of the openly available datasets presented here toward training new scientists in electron tomography should not be understated.
The tilt-series data is best viewed using 2D image processing software such as ImageJ, Fiji, or Cornell Spectrum Imager36. The reconstructed datasets require 3D visualization software. The open source software tomviz (www.tomviz.org) was used to produce the data visualizations included in this paper37. Alternatives include the free to use UCSF Chimera and commercial tools. Datasets and reconstructions may be viewed on Windows, Mac OSX, and Linux operating systems. We recommend a RAM of at least twice the size of the file being viewed for best performance.
Additional information
How to cite this article: Levin, B. D. A. et al. Nanomaterial datasets to advance tomography in scanning transmission electron microscopy. Sci. Data 3:160041 doi: 10.1038/sdata.2016.41 (2016).
References
References
DeRosier, D. J. & Klug, A. Reconstruction of Three Dimensional Structures from Electron Micrographs. Nature 217, 130–134 (1968).
Weyland, M., Midgley, P. A. & Thomas, J. M. Electron Tomography of Nanoparticle Catalysts on Porous Supports: A New Technique Based on Rutherford Scattering. J. Phys. Chem. B 105, 7882–7886 (2001).
Inoue, T. et al. Electron tomography of embedded semiconductor quantum dot. Applied Physics Letters 92, 031902 (2008).
Koguchi, M. et al. Three-dimensional STEM for observing nanostructures. J. Electron Microscopy (Tokyo) 50, 235–241 (2001).
Saghi, Z., Xu, X. & Möbus, G. Three-dimensional metrology and fractal analysis of dendritic nanostructures. Physical Review B 78, 205428 (2008).
Li, H., Xin, H. L., Muller, D. A. & Estroff, L. A. Visualizing the 3D Internal Structure of Calcite Single Crystals Grown in Agarose Hydrogels. Science 326, 1244–1247 (2009).
Midgley, P. A. & Dunin-Borkowski, R. E. Electron tomography and holography in materials science. Nature Materials 8, 271–280 (2009).
Li, S. et al. Interplay of Three-Dimensional Morphologies and Photocarrier Dynamics of Polymer/TiO2 Bulk Heterojunction Solar Cells. J. Am. Chem. Soc 133, 11614–11620 (2011).
Carriazo, D. et al. Formation Mechanism of LiFePO4 Sticks Grown by a Microwave-Assisted Liquid-Phase Process. Small 8, 2231–2238 (2012).
Keller, L. M. et al. Characterization of multi-scale microstructural features in Opalinus Clay. Microporous and Mesoporous Materials 170, 83–94 (2013).
Lin, F. et al. Phase evolution for conversion reaction electrodes in lithium-ion batteries. Nature Communications 5, 3358 (2014).
Cowman, C. D. et al. Multicomponent Nanomaterials with Complex Networked Architectures from Orthogonal Degradation and Binary Metal Back filling in ABC Triblock Terpolymers. J. Am. Chem. Soc. 137, 6026–60330 (2015).
Hovden, R. et al. Breaking the Crowther Limit: Combining depth-sectioning and tilt tomography for high-resolution wide-field 3D reconstructions. Ultramicroscopy 140, 26–31 (2014).
Scott, M. C. et al. Electron tomography at 2.4-angstrom resolution. Nature 483, 444–447 (2012).
Chen, C. C. et al. Three dimensional imaging of dislocations in a nanoparticle at atomic resolution. Nature 496, 74–77 (2013).
Xu, R. et al. Three-dimensional coordinates of individual atoms in materials revealed by electron tomography. Nature Materials 14, 1099–1103 (2015).
Ercius, P., Alaidi, O., Rames, M. J. & Ren, G. Electron Tomography: A Three-Dimensional Analytic Tool for Hard and Soft Materials Research. Adv. Mater. 27, 5638–5663 (2015).
Midgley, P. A. & Weyland, M. 3D electron microscopy in the physical sciences: the development of Z-contrast and EFTEM tomography. Ultramicroscopy 96, 413–431 (2003).
Kawase, N., Kato, M., Nishioka, H. & Jinnai, H. Transmission electron microtomography without the ‘‘missing wedge’’ for quantitative structural analysis. Ultramicroscopy 107, 8–15 (2007).
Crowther, R. A., DeRosier, D. J. & Klug, A. The Reconstruction of a Three-Dimensional Structure from Projections and its Application to Electron Microscopy. Proc. R. Soc. London, Ser. A 317, 319–340 (1970).
Hovden, R., Xin, H. L. & Muller, D. A. Extended Depth of Field for High-Resolution Scanning Transmission Electron Microscopy. Microscopy & Microanalysis 17, 75–80 (2011).
Dahmen, T. et al. Combined Scanning Transmission Electron Microscopy Tilt- and Focal Series. Microscopy & Microanalysis 20, 548–560 (2014).
Miao, J., Förster, F. & Levi, O. Equally sloped tomography with oversampling reconstruction. Physical Review B 72, 052103 (2005).
Saghi, Z. et al. Three-Dimensional Morphology of Iron Oxide Nanoparticles with Reactive Concave Surfaces. A Compressed Sensing-Electron Tomography (CS-ET) Approach. Nano Letters 11, 4666–4673 (2011).
Van Aert, S., Batenburg, K. J., Rossell, M. D., Erni, R. & Van Tendeloo, G. Three-dimensional atomic imaging of crystalline nanoparticles. Nature 470, 374–377 (2011).
Zhang, H., Ha, D.-H., Hovden, R., Kourkoutis, L. F. & Robinson, R. D. Controlled Synthesis of Uniform Cobalt Phosphide Hyperbranched Nanocrystals Using Tri-n-octylphosphine Oxide as a Phosphorus Source. Nano Letters 11, 188–197 (2011).
Crewe, A. V., Wall, J. & Langmore, J. Visibility of a single atom. Science 168, 1338–1340 (1970).
Hammel, M. & Rose, H. Resolution and optimum conditions for dark-field STEM and CTEM imaging. Ultramicroscopy 49, 81–86 (1993).
Penczek, P., Marko, M., Buttle, K. & Frank, J. Double-tilt electron tomography. Ultramicroscopy 60, 393–410 (1995).
Lanzavecchia, S. et al. Conical tomography of freeze-fracture replicas: a method for the study of integral membrane proteins inserted in phospholipid bilayers. Journal of Structural Biology 149, 87–98 (2005).
Luther, P. K., Lawrence, M. C. & Crowther, R. A. A method for monitoring the collapse of plastic sections as a function of electron dose. Ultramicroscopy 24, 7–18 (1988).
Lawrence, M. C. Alignment of images for three-dimensional reconstruction of non-periodic objects. Proc. Electron Microsc. Soc. S. Afr 13, 19–20 (1983).
Guckenberger, R. Determination of a common origin in the micrographs of tilt series in three-dimensional electron microscopy. Ultramicroscopy 9, 167–173 (1982).
Larkin, K. G., Oldfield, M. A. & Klemm, H. Fast Fourier method for the accurate rotation of sampled images. Opt. Commun. 139, 99–106 (1997).
Wang, D. et al. Tuning Oxygen Reduction Reaction Activity via Controllable Dealloying: A Model Study of Ordered Cu3Pt/C Intermetallic Nanocatalysts. Nano Letters 12, 5230–5238 (2012).
Cueva, P., Hovden, R., Mundy, J. A., Xin, H. L. & Muller, D. A. Data Processing for Atomic Resolution Electron Energy Loss Spectroscopy. Microscopy and Microanalysis 18, 667–675 (2012).
Hovden, R. et al. Repeatable and Transferable Processing for Electron Tomography: An Open Platform for Visualization and Reconstruction of 3D Materials. Microscopy and Microanalysis 21, 2407–2408 (2015).
Data Citations
Levin, B. D. A. Figshare https://dx.doi.org/10.6084/m9.figshare.c.2185342 (2016)
Acknowledgements
B.L., D.M., R.H. acknowledge support from DOE Office of Science contract DE-SC0011385. E.P. acknowledges support from NSF GRFP grant number is DGE-1144153, and from General Motors (GM), and acknowledges Zhongyi (Vic) Liu of GM for providing materials. Y.J. acknowledges support from DOE grant number DE-SC0005827. J.M. acknowledges support from DOE Office of Science contract DE-SC0010378 and NSF (DMR-1437263). This work made use of the Cornell Center for Materials Research (CCMR) Facilities supported by the National Science Foundation under Award Number DMR-1120296, and the National Center for Electron Microscopy, Lawrence Berkeley National Laboratory, which is supported by the U.S. Department of Energy under Contract no. DE-AC02-05CH11231. We thank John Grazul for TEM assistance in CCMR.
Author information
Authors and Affiliations
Contributions
B.L. & R.H. wrote and prepared the manuscript. R.H., B.L. & L.K. contributed to acquiring and reconstructing data for Tom_1. E.P., B.L., D.M. & R.H. contributed to acquiring and reconstructing data for Tom_2. M.S., C.C., & J.M. contributed to acquiring and reconstructing data for Tom_3. R.X., M.S., C.C., W.T., Y. Yang, C.O., P.E. & J.M. contributed to acquiring and reconstructing data for Tom_4. R.H., P.E., Y.J. & D.M. contributed to acquiring and reconstructing data for Tom_5. H.Z., D.H., R.R & D.W, Y.Yu., H.A. synthesized particles. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
ISA-Tab metadata
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0 Metadata associated with this Data Descriptor is available at http://www.nature.com/sdata/ and is released under the CC0 waiver to maximize reuse.
About this article
Cite this article
Levin, B., Padgett, E., Chen, CC. et al. Nanomaterial datasets to advance tomography in scanning transmission electron microscopy. Sci Data 3, 160041 (2016). https://doi.org/10.1038/sdata.2016.41
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2016.41
This article is cited by
-
Imaging 3D chemistry at 1 nm resolution with fused multi-modal electron tomography
Nature Communications (2024)
-
Real-time 3D analysis during electron tomography using tomviz
Nature Communications (2022)
-
Classification and biological identity of complex nano shapes
Communications Materials (2020)
-
An open-source, end-to-end workflow for multidimensional photoemission spectroscopy
Scientific Data (2020)
-
Multiscale higher-order TV operators for L1 regularization
Advanced Structural and Chemical Imaging (2018)