Abstract
Researchers who aim to globally analyze the gastrointestinal immune system via flow cytometry have many protocol options to choose from, with specifics generally tied to gut wall layers of interest. To get a clearer idea of the approach we should use on full-thickness colon samples from mice, we first undertook a systematic comparison of three tissue dissociation techniques: two based on enzymatic cocktails and the other one based on manual crushing. Using flow cytometry panels of general markers of lymphoid and myeloid cells, we found that the presence of cell-surface markers and relative cell population frequencies were more stable with the mechanical method. Both enzymatic approaches were associated with a marked decrease of several cell-surface markers. Using mechanical dissociation, we then developed two minimally overlapping panels, consisting of a total of 26 antibodies, for serial profiling of lymphoid and myeloid lineages from the mouse colon in greater detail. Here, we highlight how we accurately delineate these populations by manual gating, as well as the reproducibility of our panels on mouse spleen and whole blood. As a proof-of-principle of the usefulness of our general approach, we also report segment- and life stage-specific patterns of immune cell profiles in the colon. Overall, our data indicate that mechanical dissociation is more suitable and efficient than enzymatic methods for recovering immune cells from all colon layers at once. Additionally, our panels will provide researchers with a relatively simple tool for detailed immune cell profiling in the murine gastrointestinal tract, regardless of life stage or experimental conditions.
Similar content being viewed by others
Introduction
The immune system plays a critical role in maintaining homeostasis and protecting the body against harmful pathogens. Within the intricate network of immune defenses, the gastrointestinal tract houses a complex and dynamic immune system that serves as the first line of defense against a myriad of microorganisms encountered through food and environmental exposure1,2,3,4,5,6. The gut immune system is tasked with the challenging dual role of tolerating and interacting with the vast community of commensal microorganisms residing in the intestine while swiftly responding to potential threats posed by invading pathogens7,8,9. This delicate balance is essential for preserving gut health and overall immune function.
The immune system begins its development during embryogenesis, playing a crucial role in protecting the developing fetus from neonatal infections10,11,12. Initially, the fetal immune system primarily consists of primitive immune cells, including immature T and B lymphocytes, natural killer cells, monocytes, macrophages and dendritic cells13,14,15,16,17. As embryonic development progresses, the immune system undergoes diversification and specialization, enabling it to respond to encountered antigens10,18. Yet, a pivotal milestone in the development of the gut immune system occurs at birth. During vaginal delivery, the newborn is exposed for the first time to a diverse range of commensal bacteria while passing through the vaginal canal19,20. These commensal bacteria colonize the baby’s digestive tract and thereby play a fundamental role in the post-natal maturation of the developing gastrointestinal tract, including the immune system21,22,23,24,25. Notably, some of these bacteria actively contribute to the production of short-chain fatty acids, such as butyrate, which serves as an energy source for mucosal cells while also helping to strengthen epithelial junctions, balance nitrergic and cholinergic neuron subtypes in the enteric nervous system, and regulate immune responses26,27,28,29,30,31. The immune system then continuously matures as the external environment continuously exposes the newborn to various antigens1,21,32,33. In adulthood, the mature immune system within the gut comprises approximately 70% of the total immune cell populations34,35,36,37.
Multi-parameter flow cytometry is ideally suited for quantitative and qualitative analyses of individual immune cell populations. This technique enables the comprehensive characterization of diverse immune cells from the gut wall, including their phenotypes, activation status, functional attributes, and distribution patterns. Although the intestinal mucosa/submucosa have been the primary focus of most of these studies due to the abundance of immune cells in these tissue layers5,38,39,40,41,42, the muscle layers have nonetheless been the center of interest of others43,44,45,46,47. Ideally, profiling immune cells from all layers at once would be needed for a true comprehensive understanding of immune cell profiles. However, such analysis is complicated by the different structural properties of each layer (e.g., soft mucosa vs firm muscle), implying that a tissue dissociation method that works well for one layer might not work well for another. Consequently, a wide range of methods based on enzymatic and/or mechanical processing have been developed for dissociating gastrointestinal tissues38,48,49,50. Notably, several cocktails of enzymes that include different types/concentrations of collagenase, liberase and/or dispase have been previously described, with the consequence that a standardized approach remains elusive33,38,40,41,48,51,52,53.
The main objective of the current study was to develop a standardized-like and relatively simple method for the global profiling of lymphoid and myeloid cell lineages from the whole mouse colon. To this end, we first compared the efficacy of three tissue dissociation methods (two enzymatic and one mechanical) based on total cell yield, viability, and preservation of cell-surface markers. We then addressed another gap in the field in that most studies predominantly focus on either lymphoid or myeloid cell populations, rarely both, and with a limited number of markers54,55,56. Here, we describe two antibody panels targeting a total of 26 markers that enable concomitant characterization of both innate and adaptive immune cells. Combining mechanical tissue dissociation with these antibody panels allowed us to evaluate a broad range of lymphoid and myeloid cells from all layers of the mouse colon at once. Importantly, the procedure described herein is relatively simple and expeditious, also offering the flexibility to modify cell markers based on the research questions of interest.
Methods
Animals
FVB mice initially purchased from Charles River (strain code 207) were maintained and bred in individually ventilated cages within the conventional animal facility of the Université du Québec à Montréal, under 12 h light–12 h dark cycles (7AM to 7PM) and with ad libitum access to regular chow diet (Charles River Rodent Diet #5075, Cargill Animal Nutrition). Organs and blood samples were collected from pre-weaned juvenile (postnatal day [P] 20) and weaned adult (P120) animals. Prior to anesthesia, mice were weighed and sexed based on external genitalia. Both sexes were included in the current study (n = 2 males and 1 female per experiment). Whole blood (200 µl from P20 mice and 400 µl from P120 mice) was collected via intracardiac puncture of mice kept under deep anesthesia using isoflurane. Subsequently, mice were euthanized using CO2 and transferred to the laboratory for colon and spleen collection. All experimental procedures involving mice were approved by the institutional review board and ethics committee of the Université du Québec à Montréal (CIPA protocol #959) following the biomedical research guidelines of the Canadian Council of Animal Care (CCAC). All data obtained with mice are reported in accordance with the ARRIVE guidelines 2.057.
Preparation of single-cell suspensions
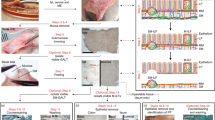
Dissected samples of whole spleen and colon were immediately placed in a sterile Petri dish containing cold RPMI 1640 1 × medium (WISENT INC, #350-000-CL) on ice. For spleen samples, single-cell suspensions were directly obtained via gentle manual grinding with the flat head of a syringe plunger on a 70 µm cell strainer (Movie S1). For colon samples, mesenteric fat was removed, and the colon was divided into two regions: proximal colon and distal colon. Each region was opened longitudinally along the mesenteric border, flushed with RPMI medium, and processed for tissue dissociation. To test the three different dissociation methods, each region was further sub-divided into three fragments of equal length. Each of the three dissociation methods was tested using a full-thickness fragment (i.e., including all unaltered tissue layers) from the proximal colon combined to a full-thickness fragment from the distal colon. Mechanical dissociation of these colon samples was performed as described above for spleen samples, but with a slightly stronger manual pressure on tissue, placing mucosal side on the cell strainer (Movie S2). Enzymatic dissociation was performed using a previously published method48,52,53. Two different enzymatic cocktails were tested, both prepared in RPMI medium supplemented with DNase I (0.1 mg/ml, Sigma-Aldrich #DN25): one cocktail contained Dispase II (1.3 mg/ml, Life Technologies #17105-041) and Collagenase I (0.4 mg/ml, Sigma-Aldrich #C2674), while the other contained Liberase TL (0.15 mg/ml, Roche #05401020001) and Collagenase V (0.4 mg/ml, Sigma-Aldrich #C9263). To compare efficiency of the mechanical approach on full-thickness colon vs mucosal/submucosal layers only, proximal and distal colon regions were further sub-divided into two fragments of equal length, with one of these further processed to peel off the mucosal/submucosal layers from the muscle/serosa layers, as previously described58,59,60. To this end, full-thickness samples were immobilized with mucosa side up by pinning onto SYLGARD-coated (Thermo Fisher Scientific, #50822180) Petri dishes and were subsequently carefully stripped under a dissecting microscope to separate the soft mucosal and associated submucosal layers from the firm muscle/serosal layers. After extensive washing in RPMI, corresponding fragments (full-thickness or mucosa/submucosa-only) from the proximal colon and distal colon were then combined for analysis.
Each resulting single-cell suspension was filtered through a 40 µm cell strainer and incubated in RPMI medium supplemented with 10% FBS for 2 h at 37 °C to promote the re-appearance of cell surface markers. After gentle centrifugation (1500 RPM during 5 min) and resuspension in 1 ml of RPMI + 2% FBS, viable cells were counted using a hemocytometer with trypan blue counterstain. For each sample, two tubes containing between 600,000 and 1,000,000 live cells in 200 µl of RPMI were prepared and processed in parallel for staining with antibody panels for lymphoid and myeloid cell lineages as described below. Additional tubes containing either non-stained cells or cells stained with viability markers only (see Tables 1, 2) were also prepared for compensation prior to data acquisition with the flow cytometer.
Staining of single-cell suspensions
Single-cell suspensions from colon, spleen, and whole blood samples were stained with cocktails of fluorescently labeled antibodies designed to mark lymphoid and myeloid cell populations as described in Tables 1 and 2, respectively. For colon and spleen samples, cells were first stained for extracellular markers during 1 h at 4 °C. These samples were then washed with 1 ml of PBS, centrifuged (1500 RPM during 5 min) and the medium was partially removed to leave cells in 200 μl. For the myeloid panel, cells were fixed using CytoFix (BD CytoFix™, #554655) according to the manufacturer’s instructions. For the lymphoid panel, cells were instead fixed and permeabilized using the Transcription Factor Buffer Set (BD Pharmingen™, #562574) according to the manufacturer’s instructions, and then processed for additional intracellular staining with FoxP3 and ROR\(\upgamma \)t antibodies (Table 1) during 1 h at 4 °C. After completion of the fixation and/or additional intracellular staining steps for both panels, cells were washed and centrifuged to leave them in 200 μl as described above. All tubes were finally kept at 4 °C protected from light until data acquisition in the flow cytometer. For blood samples, the same protocol as described above was used with an additional post-staining step to lyse red blood cells with relevant lysis buffer (BD Pharm Lyse™, # 555899) according to the manufacturer’s instructions.
Flow cytometry
All samples were analysed using a LSR Fortessa™ X-20 flow cytometer (BD Biosciences), with 3-laser configuration (Blue, Violet, Yellow-Green). To ensure accurate fluorescence compensation for each panel, a FACS tube was prepared for each antibody by adding 100 μl PBS, 1 drop of compensation beads (Invitrogen™, #01–2222-41), and 1 μl of the specific antibody. Successful compensation was achieved when there was no overlap between fluorochromes, also considering signals obtained for non-stained cells and cells stained for viability only. Data from each sample were finally acquired and recorded until reaching the desired number of events. Cell populations were manually gated using the FlowJo software V10.8.0 (BD Biosciences), which was also used to globally visualize data via 2-dimensional t-SNE analysis.
Statistical analysis
All experiments were performed using three independent biological replicates, with data expressed as the mean ± standard error of the mean (SEM). The significance of differences was determined using two-tailed Student’s t-tests in GraphPad Prism software V9.5.1. Only statistically significant differences (p < 0.05) are indicated in the figures (*, **, ***, **** represent p < 0.05, p < 0.01, p < 0.001, p < 0.0001, respectively).
Results
Mechanical dissociation best preserves cell-surface markers in immune cells from full-thickness colon samples from P20 mice
To determine the best method of colon dissociation for analyzing immune cells by flow cytometry, we tested 3 different techniques on full-thickness samples (i.e., including all unaltered tissue layers) from pre-weaned P20 mice: two enzymatic (based on either Collagenase I & Dispase II or Collagenase V & Liberase TL) and one mechanical (Fig. 1). We first analyzed the impact of these methods on the total number of recovered CD45+ hematopoietic cells, which revealed markedly greater efficiency of the mechanical approach over both enzymatic ones (Fig. 2A,B). Importantly, we also found that this gain of recovery efficiency was not made at the expense of viability, as all three methods gave similar proportions of viable CD45+ cells (Fig. 2C,D). Analysis of lymphoid and myeloid cell lineages with some general markers then revealed that the greater efficiency of the mechanical technique was in fact more specific to lymphoid cells. Frequencies of CD19+ B-cells, CD3+ T-cells, CD4+ T-cells and CD8+ T-cells were all higher with the mechanical technique (Fig. 3A–D), with an exception for the frequency of CD19+ B-cells which was equalized by the enzymatic approach based on Collagenase V & Liberase TL (Fig. 3A,B). The mechanical and the Collagenase V & Liberase TL approaches also yielded similar frequencies of F4/80+ CD11b+ macrophages (Fig. 4A,B) and CD11c+ MHCII+ dendritic cells (Fig. 4C,D), which in this case were either lower or higher than with the Collagenase I & Dispase II approach, respectively. Yet, the mechanical approach again outperformed both enzymatic techniques in the identification of Ly6C+ monocytes (Fig. 4E,F). Considering all of the above, we conclude that mechanical dissociation best balances total cell recovery yields with cell viability and presence of cell-surface markers in lymphoid and myeloid cells.
Mechanical disruption of mouse colon results in higher immune cell yields than enzymatic digestion, without impacting viability. (A,B) The mechanical technique results in increased frequencies of total CD45+ immune cells, as shown in representative contour plots (A) and associated quantitative analyses (B). (C,D) Similar percentages of live CD45+ hematopoietic cells (gated on singlets) were retrieved from the different isolation protocols, as shown in representative contour plots (C) and associated quantitative analyses (D). N = 3 biological replicates; **P ≤ 0.01 and ***P ≤ 0.001; Student’s t-test.
Mechanical disruption of mouse colon best preserves the cell surface markers of gut lymphoid cells. (A–D) Both enzymatic cocktails negatively affect the detection of the main populations of lymphoid cells (B-cells vs T-cells in panels A-B; CD4+ vs CD8+ T-cells in panels C,D), as shown in representative contour plots (A,C) and associated quantitative analyses (B,D). N = 3 biological replicates; *P ≤ 0.05 and ****P ≤ 0.0001; Student’s t-test.
Mechanical disruption of mouse colon best preserves the cell surface markers of gut myeloid cells, except for macrophages. (A,B) The Collagenase I & Dispase II cocktail is slightly better than both the Collagenase V & Liberase LT cocktail and mechanical disruption for detecting F4/80+ CD11b+ macrophages, as shown in representative contour plots (A) and associated quantitative analyses (B). (C,D) Mechanical disruption and the Collagenase V & Liberase LT cocktail are both slightly more efficient than the Collagenase I & Dispase II cocktail for detecting CD11c+ MHCII+ (hi) dendritic cells, as shown in representative contour plots (C) and associated quantitative analyses (D). (E,F) Mechanical disruption is more effective than both enzymatic cocktails for detecting Ly6C+ monocytes, as shown in representative contour plots (E) and associated quantitative analyses (F). N = 3 biological replicates; *P ≤ 0.05 and **P ≤ 0.01; Student’s t-test. MΦ, macrophages; Mo, monocytes; DC, dendritic cells.
Full-thickness colon samples enable the characterization of a diversified pool of gastrointestinal immune cells in P20 mice
To confirm that mechanical dissociation is well suited for recovering gastrointestinal immune cells from all colon layers at once, we then compared the relative abundance of the main types of lymphoid and myeloid cells in full-thickness vs mucosa/submucosa-only samples (all at P20, before weaning). Using t-SNE for global visualization in two dimensions, we notably found that mucosa/submucosa-only samples markedly overestimate the proportion CD19+ B-cells present in the colon (Fig. 5A) while the proportion of Ly6C+ monocytes is markedly underestimated (Fig. 5B). These experimental biases were found to be eliminated in full-thickness samples (Fig. 5), as also shown in representative contour plots (Fig.S1). These observations highlight the importance of including all colon layers for comprehensive characterization of the diverse populations of gastrointestinal immune cells.
Experimental biases that are present when using mucosal/submucosal layers are eliminated in full-thickness samples of mouse colon. (A,B) t-SNE maps of the main populations of lymphoid (A) and myeloid (B) cells. CD19+ B-cells and MHCII+ cells are enriched in mucosal/submucosal samples at the expense of CD4+/CD8+ T-cells and LyC6+ monocytes, respectively.
Development of antibody panels for global analysis of both innate and adaptative immune cells in the murine colon
Motivated by an interest for obtaining global snapshots of both innate and adaptative immune systems under various physiological contexts in the murine colon, we developed two antibody panels that target lymphoid and myeloid cell lineages separately. Selected markers were chosen based on existing literature50,52,56,61,62,63,64,65,66, also considering that we wanted our panels to be eventually useful in the context of inflammatory conditions as well. As detailed in Tables 1–2, these panels contain 26 antibodies in total (13 lymphoid-specific, 11 myeloid-specific, and 2 shared), mostly targeting extracellular markers (only two are targeting intracellular markers). Below is a summary of the flow cytometry gating strategies for each panel, using entire full-thickness colon samples mechanically dissociated from pre-weaned P20 mice as starting material.
With the lymphoid panel (Table 1 for antibodies list, Table 3 for gated phenotypes), single and live CD45+ cells can first be separated in B-cell and T-cell subsets (Fig. 6A; phenotypes in Table 3). From the T-cell subset, we can then evaluate CD4+ T-cells, CD8+ T-cells, and double-negative T-cells (Fig. 6B; phenotypes in Table 3). Of note, although subsequent steps shown are focused on the CD4+ T-cell subset, this is the same strategy for CD8+ and double-negative T-cell subsets (except for Th17 that is specific to the CD4+ T-cell subset). Hence, within CD4+, CD8+, and double-negative T-cells, frequencies of naïve, memory, and effector populations can be determined, along with activated cells and different populations of regulatory T-cells (Fig. 6B; phenotypes in Table 3). From the B-cell subset, we can also gate activated and effector cells (Fig. 6C; phenotypes in Table 3). Moreover, in a variant of the lymphoid panel, we replaced CD19 and CD25 markers by NK1.1 and CD103 markers (Table 1), thereby allowing to characterize natural killer cells, natural killer T-cells and ILC3-like populations, as well as to distinguish between tissue-resident and circulating memory cells (see graphs with grey axes in Fig. 6A,B,D; phenotypes in Table 3).
Gating strategy for the phenotypical characterization of lymphoid cell lineages from mechanically dissociated colon of P20 mice. (A) CD45+ hematopoietic cells are first gated for sequential exclusion of doublets/aggregated and dead cells. Single live CD45+ hematopoietic cells (orange box) are then gated to distinguish between B-cells (CD19+; blue box) and T-cells (CD3+; pink box). Alternatively, in a variant of the lymphoid panel (highlighted in grey), live CD45+ hematopoietic cells can first be gated to distinguish between natural killer T-cells (NK1.1+ CD3+; black box) and T-cells (NK1.1− CD3+; pink box). (B) As exemplified here for CD4+ T-cells (similar strategy for CD8+ T-cells and double-negative T-cells, except for T-helper 17), T-cells from the pink boxes in A can be further analyzed to evaluate the frequencies of regulatory (FoxP3+) and/or activated (CD25+ or CD69+) and/or effector (CD39+ and/or CD73+) and/or memory (CD44+ vs naïve CD62L+) subsets. ROR\(\gamma \)t further allows to identify T-helper 17 cells whereas, in the variant panel (highlighted in grey), CD103 further allows to identify tissue-resident memory cells (dashed boxes). (C) B-cells from the blue box in A can be further analyzed to evaluate the frequencies of effector (CD39+ and/or CD73+) and/or activated (CD69+) and/or memory (CD44+) subsets. (D) With the variant panel (highlighted in grey), non-T-cells (CD3− CD4−) from the orange box in A can be directly analyzed to distinguish between natural killer cells (NK1.1+) and innate lymphoid cells type 3 (ROR\(\gamma \)t+). NKT, natural killer T-cells; DN, double-negative T-cells; Th17, T-helper 17; conv, conventional; Treg, regulatory T-cells; cTreg, central Treg; eTreg, effector Treg; TRM, tissue-resident memory; NK, natural killer; ILC3, innate lymphoid cells type 3; NCR, natural cytotoxicity receptor.
With the myeloid panel (Table 2 for antibodies list, Table 4 for gated phenotypes), a first selection is made from single and live CD45+ cells using CD11b and CD11c antibodies (Fig. 7A). CD11b+ cells can then be analyzed to characterize monocytes and macrophages (Fig. 7B; phenotypes in Table 4), whereas CD11c+ cells can be analyzed to characterize dendritic cells (Fig. 7C; phenotypes in Table 4). From the LyC6+ monocyte subset, we can subsequently distinguish between classical monocytes, extravasated monocytes, non-classical monocytes, and monocyte-derived macrophages (Fig. 7B; phenotypes in Table 4). For F4 /80+ macrophages, we can further characterize tissue-resident macrophages, long-lived macrophages, sub-mucosal macrophages, muscular macrophages, lamina propria macrophages, M1-like macrophages, and M2 macrophages (Fig. 7B; phenotypes in Table 4). Finally, dendritic cell subsets can be analyzed to distinguish between monocyte-derived and conventional populations (Fig. 7C; phenotypes in Table 4). Here, it is important to bear in mind that myeloid cell populations share a lot of markers, leading to many possibilities in terms of gating strategies. What we described above is only one possible way to gate myeloid cells.
Gating strategy for the phenotypical characterization of myeloid cell lineages from mechanically dissociated colon of P20 mice. (A) CD45+ hematopoietic cells are first gated for sequential exclusion of doublets/aggregated and dead cells. Single live CD45+ hematopoietic cells are then gated to distinguish between CD11c+ (blue box) and CD11b+ (pink box) myeloid cells. (B) Panels above the dashed line show that CD11b+ cells from the pink box in A can be further analyzed to evaluate the frequencies of monocytes (LyC6+) and macrophages (LyC6-). Then, four additional markers (CCR2, CX3CR1, MHC II, CD64) allow to identify (i) classical and extravasated monocytes from the LyC6+ (hi) subset, (ii) non-classical monocytes and monocyte-derived macrophages from the LyC6+ (low) subset, and (iii) mature macrophages from the LyC6-negative subset. Panels below the dashed line show that CD11b+ F4/80+ macrophages can also be sequentially gated with MHCII/CX3CR1 and CD103/CD64 markers to evaluate the frequencies of tissue-resident macrophages, which can then be stratified for localization (lamina propria, submucosa, muscle) and inflammatory status (pro-inflammatory M1 vs anti-inflammatory M2) using three additional markers (CD163, Tim4 and CCR2). (C) Using four markers (Ly6C, CD64, CD103, CD11b), CD11c+ cells from the blue box in A can be further analyzed to distinguish between monocyte-derived dendritic cells and different subsets of conventional dendritic cells. Moreover, conventional dendritic cells type 2 can be further subdivided using Tim4/CX3CR1 markers. Mo, monocytes; MΦ, macrophages; TR, tissue-resident; LP, lamina propria; SM, submucosa; M, muscle; DC, dendritic cells; cDC, conventional DC.
In a final validation step, we sought to confirm that the very low detection of certain markers (NK1.1, CD39 in T-cells; CD115 in monocytes and macrophages) in the colon of P20 mice was biologically real and not due to an experimental artifact. To this end, we thus tested our panels on various samples from older weaned mice, including colon, whole blood and spleen. All markers that were weakly detected in the colon of P20 mice appeared markedly enriched in at least one of these additional sample types from P120 mice (Fig. S2). Therefore, we conclude that our antibody panels can efficiently provide global snapshots of both innate and adaptative immune systems, not only for colon samples but also for blood and spleen samples.
Life stage-associated changes in the immune system profile of the proximal and distal colon from mice
As a proof-of-concept of the usefulness of our approach, we conducted a systematic comparative analysis of two segments of the murine colon (full-thickness proximal vs full-thickness distal) as a function of age and weaning status corresponding to the adolescence-adulthood transition (pre-weaned P20 mice vs weaned P120 mice). With the lymphoid panel, we first evidenced that this transition impacts the frequencies of both CD19+ B-cells and CD8+ T-cells, but not overall levels of CD4+ T-cells (Fig. 8A,B). Interestingly, while the observed decline in CD19+ B-cells at P120 was found to be similar in both colon segments, the life stage-associated changes in CD8+ T-cells were instead found to be different in proximal (increased levels at P120) and distal (decreased levels at P120) segments (Fig. 8B). Upon more detailed analysis with the lymphoid panel, specific subsets of CD4+ T-cells were nonetheless found to be affected by the adolescence-adulthood transition as well. This is notably the case of CD4+ T-cell subsets also expressing the ectonucleotidases CD73 and CD39, which appeared generally increased at P120 as they are within the CD8+ T-cell population (Fig. 8C). With the myeloid panel, we detected a general decrease in the proportion of F4/80+ macrophages but not LyC6+ monocytes at P120 (Fig. 9A,B). A general adulthood-associated decrease in the frequency of CD11c+ MHC II+ dendritic cells was also noted, but without reaching statistical significance most likely because of small sample size (Fig. 9B). Upon more detailed analysis, an adulthood-associated decline in monocyte-derived macrophages (CD11b+ LyC6+ (low) CX3CR1+ CCR2+ (low) MHCII+ CD64+) was noted as well, reaching statistical significance in the distal colon only (Fig. 9C). In contrast, the frequency of tissue-resident macrophages (F4/80+ CD11b+ CX3CR1+ MHC II+ CD64+ CD103−) was found to significantly increase during the adolescence-adulthood transition in both the proximal and distal colon (Fig. 9C). Finally, a major decrease in the frequency of a specific subset of conventional dendritic cells (CD11c+ MHC II+ Ly6C− CD64− CD11b+ CD103+) was noted at P120, but only in the proximal colon. Altogether, these findings thus confirm that our combination of mechanical dissociation with both antibody panels described herein is useful for detecting physiologically relevant changes in the composition of the gastrointestinal immune system in mice.
Life stage-associated changes in the lymphoid cell profile of the proximal and distal colon of FVB mice. (A,B) Representative t-SNE maps (A) and accompanying quantitative analyses (B) of the main populations of lymphoid cells show that total levels of CD19+ B-cells are lower at P120 than at P20, in both proximal and distal colon. Total levels of CD4+ T-cells are not affected by life stage, whereas total levels of CD8+ T-cells vary differently in proximal and distal colon. (C) Quantitative analyses of lymphoid cell subsets reveal specific life stage-related changes for several CD4+ and CD8+ subsets expressing CD73 and/or CD39. Treg, regulatory T-cells. N = 3 biological replicates; *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001 and ****P ≤ 0.0001; Student’s t-test.
Life stage-associated changes in the myeloid cell profile of the proximal and distal colon of FVB mice. (A,B) Representative t-SNE maps (A) and accompanying quantitative analyses (B) of the main populations of myeloid cells show that total levels of CD11b+ F4/80+ macrophages are lower at P120 than at P20, in both proximal and distal colon. Total levels of LyC6+ monocytes are not significantly affected by life stage, whereas a general trend toward lower levels in adult mice is noted for MHC II+ CD11c+ dendritic cells. (C) Quantitative analyses of myeloid cell subsets reveal specific life stage-related changes for monocyte-derived macrophages (general decrease in adults) and tissue-resident macrophages (general increase in adults). Frequencies of CD11b+ CD103+ conventional dendritic cells are specifically lower in adult proximal colon, but not distal colon. Mo, monocytes; MΦ, macrophages; TR, tissue-resident; cDC, conventional dendritic cells. N = 3 biological replicates; *P ≤ 0.05 and ***P ≤ 0.001; Student’s t-test.
Discussion
The main goal of the current study was to identify a relatively simple method for global profiling of lymphoid and myeloid cell lineages from full-thickness mouse colon samples. Comparison of enzymatic and mechanical cell dissociation techniques via flow cytometry showed that the mechanical approach best balances total cell recovery yields with cell viability (Fig. 2) and presence of cell-surface markers in lymphoid and myeloid cells. Both tested enzyme-based techniques were found to be globally deleterious for the analysis of lymphoid lineages, presumably because of proteolytic cleavage of the cell surface markers of interest. Although the Collagenase V & Liberase LT cocktail performed better than Collagenase I & Dispase II for detecting CD19+ B-cells, both cocktails were found to be similarly much less efficient than the mechanical technique for detecting CD3+ T-cells (Fig. 3A,B). Both cocktails were also found to be similarly less efficient for detecting CD8+ T-cells, with the Collagenase I & Dispase II cocktail being in addition particularly inefficient for detecting CD4+ T-cells as well (Fig. 3C,D). Such negative impact of enzymatic cell dissociation techniques on lymphoid lineages has been previously documented in several studies38,51,52,67,68,69. This can have profound consequences when comes time to draw biologically valid conclusions. For example, based on our CD4/CD8 profiling data, both enzymatic techniques would have led to grossly erroneous conclusions about the real frequency of double negative T-cells (Fig. 3C,D). Intriguingly, enzymatic dissociation appears to be less problematic for myeloid cells. Notably, we found that both tested enzymatic cocktail were either equally efficient or even slightly more efficient than mechanical dissociation for detecting CD11b+ F4/80+ macrophages (Fig. 4A,B). However, this was not the case for CD11c+ MHCII+ dendritic cells and CD11b+ LyC6+ monocytes, with the latter being even again more represented using the mechanical approach (Fig. 4C–F). In the end, we thus think that the far superior performance of the mechanical technique with lymphoid cells in general (and monocytes) greatly outweighs the slight advantage of enzymatic-based approaches in detecting macrophages. Yet, we do recognize that mastering the mechanical technique with full-thickness mouse colon samples requires a fair amount of practice for finding the right level of manual strength to use (Movie S2). Moreover, the needed level of manual strength to use will likely be different in other segments of the gastrointestinal tract (e.g., small intestine), which is important to consider if one wants to combine multiple segments in the same analysis. When tissue crushing is too harsh (Movie S3), this leads to low cell yields and high mortality rates of immune cells, as previously reported in some studies38,70. Nonetheless, other studies have shown that even when mortality rates are high, mechanically dissociated immune cells that survive remain more immunocompetent than those obtained following collagenase digestion71.
Another key goal of this study was to develop robust antibody panels that would allow to profile a wide array of lymphoid and myeloid cells in a cost-effective manner. These panels were designed to be used on a widely used multi-color flow cytometer with minimal configuration, thereby facilitating their use by the research community. For research teams having access to a multi-color flow cytometer with cell sorting capacity, Tables S1 and S2 show that several of the cell populations that can be identified with our panels and gating strategies can also be recovered in sufficient amounts for further downstream analyses. As proof-of-concept of the usefulness of our panels for detecting physiologically relevant changes in the gastrointestinal immune system, we analyzed the impact of life stages (pre-weaned juvenile vs weaned adult mice) and anatomical region (proximal vs distal colon) on immune cell profiles. Both age32,72,73 and weaning status (via food and/or microbiome-related changes)74,75,76 are well-known determinants of immune system composition and function in the gastrointestinal tract, as we observed with both of our panels (Figs. 8–9). What was a bit more surprising was to find that some of these changes were completely different at each end of the colon, such as the adulthood-associated change in the frequency of CD8+ T-cells which is increased in proximal colon but decreased in distal colon (Fig. 8B). Differences in immune system composition and function are expected when comparing major anatomical subdivisions of the gastrointestinal tract, like the colon and the small intestine77,78. Our data now indicate that such differences can also be present in juxtaposed segments of the same major subdivision of the gastrointestinal tract. This clearly adds to the notion that care should be taken not to systematically extrapolate findings made in one region of the gastrointestinal tract to another region.
Our focus on pre-weaned P20 mice is due to our general interest for Hirschsprung disease, as most mouse models for this life-threatening condition normally succumb soon after weaning (from P21 onwards)79,80,81. As seen in human patients82,83, these mice develop Hirschsprung-associated enterocolitis (HAEC) which is then believed to progress to sepsis and thus to be the main cause of death for Hirschsprung disease84. However, how exactly HAEC develops is not well understood. Like other inflammatory conditions affecting the gastrointestinal tract, such as Crohn’s disease, an immune system aspect is presumably central, but this question has not been addressed in a global manner yet. Immune cell infiltration in mouse models of Hirschsprung disease is frequently reported using histology techniques only60,85,86, without further detailed analysis. Otherwise, the very few studies that have investigated this question in a more detailed manner were mostly restricted to Peyer’s patches87,88, based on the prior discovery that the Hirschsprung disease-associated signaling pathway GDNF-RET is key for the development of this lymphoid site in the ileum89,90. Notably, smaller Peyer’s patches in Ednrb-mutant mice have been associated with a depletion of both B-cells and immunoglobulins87,88. Another study with different mouse models of defective EDN3-EDNRB signaling reported more generalized lymphoid depletion, also impacting CD4+ /CD8+ T-cells and involving other lymphoid organs (bone marrow, spleen and thymus)91. A more recent study, again performed with an Ednrb-mutant mouse model, strongly suggests that macrophage activation in the colon locally contributes to the etiology of HAEC as well92. All these prior observations highlight the need for more comprehensive analyses of the immune system at the site of HAEC development (i.e., in the colon instead of the small intestine), ideally using various genetically distinct mouse models of Hirschsprung disease—a complex genetic disease93. Performing that kind of analysis using the method described herein will be especially important in the context of GDNF-based treatment of Hirschsprung disease58,60, as rectally administered GDNF is expected to not only locally stimulate neurogenesis but also directly influence numerous immune cell populations94,95. Such studies are currently being completed in our laboratories and will be the subject of a forthcoming publication. That said, our results already allow us to attest that our general approach works equally well with inflamed tissue, and that it can reliably detect both disease-associated and treatment-induced changes.
In conclusion, we expect that the experimentally validated approach described herein will be useful for a large number of laboratories interested in globally assessing the immune status of the mouse bowel, in a wide variety of experimental contexts. This contribution is thus an effort to democratize such analyses, including laboratories with little or no previous experience in the field.
Data availability
All relevant data are within the manuscript and its Supplementary Information files.
References
Peterson, C. T., Sharma, V., Elmen, L. & Peterson, S. N. Immune homeostasis, dysbiosis and therapeutic modulation of the gut microbiota. Clin. Exp. Immunol. 179, 363–377. https://doi.org/10.1111/cei.12474 (2015).
Owens, B. M. & Simmons, A. Intestinal stromal cells in mucosal immunity and homeostasis. Mucosal. Immunol. 6, 224–234. https://doi.org/10.1038/mi.2012.125 (2013).
Lin, S. et al. Mucosal immunity–mediated modulation of the gut microbiome by oral delivery of probiotics into Peyer’s patches. Sci. Adv. 7, 1–15 (2021).
Okumura, R. & Takeda, K. Maintenance of gut homeostasis by the mucosal immune system. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 92, 423–435. https://doi.org/10.2183/pjab.92.423 (2016).
Smith, P. D. et al. Intestinal macrophages and response to microbial encroachment. Mucosal. Immunol. 4, 31–42. https://doi.org/10.1038/mi.2010.66 (2011).
Schneider, K. M., Kim, J., Bahnsen, K., Heuckeroth, R. O. & Thaiss, C. A. Environmental perception and control of gastrointestinal immunity by the enteric nervous system. Trends Mol. Med. 28, 989–1005. https://doi.org/10.1016/j.molmed.2022.09.005 (2022).
Wu, H. J. & Wu, E. The role of gut microbiota in immune homeostasis and autoimmunity. Gut. Microbes 3, 4–14. https://doi.org/10.4161/gmic.19320 (2012).
Hine, A. M. & Loke, P. Intestinal Macrophages in Resolving Inflammation. J. Immunol. 203, 593–599. https://doi.org/10.4049/jimmunol.1900345 (2019).
Chung, H. & Kasper, D. L. Microbiota-stimulated immune mechanisms to maintain gut homeostasis. Curr. Opin. Immunol. 22, 455–460. https://doi.org/10.1016/j.coi.2010.06.008 (2010).
Landreth, K. S. Critical windows in development of the rodent immune system. HET. 21, 493–498 (2002).
Fujiwara, H. et al. Promoting roles of embryonic signals in embryo implantation and placentation in cooperation with endocrine and immune systems. Int. J. Mol. Sci. 21, 1. https://doi.org/10.3390/ijms21051885 (2020).
Fujiwara, H. Immune cells contribute to systemic cross-talk between the embryo and mother during early pregnancy in cooperation with the endocrine system. Reprod. Med. Biol. 5, 19–29. https://doi.org/10.1111/j.1447-0578.2006.00119.x (2006).
Kobayashi, M. & Yoshimoto, M. Multiple waves of fetal-derived immune cells constitute adult immune system. Immunol. Rev. 315, 11–30. https://doi.org/10.1111/imr.13192 (2023).
Mor, G. & Cardenas, I. The immune system in pregnancy: a unique complexity. Am. J. Reprod. Immunol. 63, 425–433. https://doi.org/10.1111/j.1600-0897.2010.00836.x (2010).
Morelli, S., Mandal, M., Goldsmith, L. T., Kashani, B. N. & Ponzio, N. M. The maternal immune system during pregnancy and its influence on fetal development. Res. Rep. Biol. https://doi.org/10.2147/rrb.S80652 (2015).
Hoeffel, G. et al. C-Myb(+) erythro-myeloid progenitor-derived fetal monocytes give rise to adult tissue-resident macrophages. Immunity 42, 665–678. https://doi.org/10.1016/j.immuni.2015.03.011 (2015).
Rosado, M. M. et al. From the fetal liver to spleen and gut: The highway to natural antibody. Mucosal. Immunol. 2, 351–361. https://doi.org/10.1038/mi.2009.15 (2009).
Rackaityte, E. & Halkias, J. Mechanisms of fetal T cell tolerance and immune regulation. Front. Immunol. 11, 588. https://doi.org/10.3389/fimmu.2020.00588 (2020).
Dzidic, M., Boix-Amoros, A., Selma-Royo, M., Mira, A. & Collado, M. C. Gut microbiota and mucosal immunity in the neonate. Med. Sci. (Basel) 6, 1. https://doi.org/10.3390/medsci6030056 (2018).
Geuking, M. B., Koller, Y., Rupp, S. & McCoy, K. D. The interplay between the gut microbiota and the immune system. Gut. Microbes 5, 411–418. https://doi.org/10.4161/gmic.29330 (2014).
Stras, S. F. et al. Maturation of the human intestinal immune system occurs early in fetal development. Dev. Cell 51, 357–373. https://doi.org/10.1016/j.devcel.2019.09.008 (2019).
Sanidad, K. Z. & Zeng, M. Y. Neonatal gut microbiome and immunity. Curr. Opin. Microbiol. 56, 30–37. https://doi.org/10.1016/j.mib.2020.05.011 (2020).
Tsafaras, G. P., Ntontsi, P. & Xanthou, G. Advantages and limitations of the neonatal immune system. Front. Pediatr. 8, 5. https://doi.org/10.3389/fped.2020.00005 (2020).
Battersby, A. J. & Gibbons, D. L. The gut mucosal immune system in the neonatal period. Pediatr. Allergy Immunol. 24, 414–421. https://doi.org/10.1111/pai.12079 (2013).
Schill, E. M., Floyd, A. N. & Newberry, R. D. Neonatal development of intestinal neuroimmune interactions. Trends Neurosci. 45, 928–941. https://doi.org/10.1016/j.tins.2022.10.002 (2022).
Parada Venegas, D. et al. Short chain fatty acids (SCFAs)-mediated gut epithelial and immune regulation and its relevance for inflammatory bowel diseases. Front. Immunol. 10, 277. https://doi.org/10.3389/fimmu.2019.00277 (2019).
Yao, Y. et al. The role of short-chain fatty acids in immunity, inflammation and metabolism. Crit. Rev. Food Sci. Nutr. 62, 1–12. https://doi.org/10.1080/10408398.2020.1854675 (2022).
Correa-Oliveira, R., Fachi, J. L., Vieira, A., Sato, F. T. & Vinolo, M. A. Regulation of immune cell function by short-chain fatty acids. Clin. Transl. Immunol. 5, e73. https://doi.org/10.1038/cti.2016.17 (2016).
Ratajczak, W. et al. Immunomodulatory potential of gut microbiome-derived short-chain fatty acids (SCFAs). Acta Biochim Polut. 66, 1–12. https://doi.org/10.18388/abp.2018_2648 (2019).
Soret, R. et al. Short-chain fatty acids regulate the enteric neurons and control gastrointestinal motility in rats. Gastroenterology 138, 1772–1782. https://doi.org/10.1053/j.gastro.2010.01.053 (2010).
Suply, E., de Vries, P., Soret, R., Cossais, F. & Neunlist, M. Butyrate enemas enhance both cholinergic and nitrergic phenotype of myenteric neurons and neuromuscular transmission in newborn rat colon. Am. J. Physiol. Gastrointest. Liver Physiol. 302, G1373–G1380. https://doi.org/10.1152/ajpgi.00338.2011 (2012).
Simon, A. K., Hollander, G. A. & McMichael, A. Evolution of the immune system in humans from infancy to old age. Proc. Biol. Sci. 282, 20143085. https://doi.org/10.1098/rspb.2014.3085 (2015).
Bain, C. C. et al. Constant replenishment from circulating monocytes maintains the macrophage pool in the intestine of adult mice. Nat. Immunol. 15, 929–937. https://doi.org/10.1038/ni.2967 (2014).
Wiertsema, S. P., van Bergenhenegouwen, J., Garssen, J. & Knippels, L. M. J. The interplay between the gut microbiome and the immune system in the context of infectious diseases throughout life and the role of nutrition in optimizing treatment strategies. Nutrients 13, 1. https://doi.org/10.3390/nu13030886 (2021).
Chassaing, B., Kumar, M., Baker, M. T., Singh, V. & Vijay-Kumar, M. Mammalian gut immunity. Biomed. J. 37, 246–258. https://doi.org/10.4103/2319-4170.130922 (2014).
Arya, A. K. & Hu, B. Brain-gut axis after stroke. Brain Circ. 4, 165–173. https://doi.org/10.4103/bc.bc_32_18 (2018).
Guo, Y. et al. The gut-organ-axis concept: Advances the application of gut-on-chip technology. Int. J. Mol. Sci. 24, 1. https://doi.org/10.3390/ijms24044089 (2023).
Schreurs, R. et al. Quantitative comparison of human intestinal mononuclear leukocyte isolation techniques for flow cytometric analyses. J. Immunol. Methods 445, 45–52. https://doi.org/10.1016/j.jim.2017.03.006 (2017).
Jorgensen, P. B. et al. Identification, isolation and analysis of human gut-associated lymphoid tissues. Nat. Protoc. 16, 2051–2067. https://doi.org/10.1038/s41596-020-00482-1 (2021).
Stzepourginski, I., Eberl, G. & Peduto, L. An optimized protocol for isolating lymphoid stromal cells from the intestinal lamina propria. J. Immunol. Methods 421, 14–19. https://doi.org/10.1016/j.jim.2014.11.013 (2015).
Smith, P. D. et al. Intestinal macrophages lack CD14 and CD89 and consequently are down-regulated for LPS- and IgA-mediated activities. J. Immunol. 167, 2651–2656. https://doi.org/10.4049/jimmunol.167.5.2651 (2001).
Pabst, O. & Bernhardt, G. The puzzle of intestinal lamina propria dendritic cells and macrophages. Eur. J. Immunol. 40, 2107–2111. https://doi.org/10.1002/eji.201040557 (2010).
Ozaki, H. et al. Isolation and characterization of resident macrophages from the smooth muscle layers of murine small intestine. Neurogastroenterol. Motil. 16, 39–51. https://doi.org/10.1046/j.1365-2982.2003.00461.x (2004).
Avetisyan, M. et al. Muscularis macrophage development in the absence of an enteric nervous system. Proc. Natl. Acad. Sci. USA 115, 4696–4701. https://doi.org/10.1073/pnas.1802490115 (2018).
Muller, P. A. et al. Crosstalk between muscularis macrophages and enteric neurons regulates gastrointestinal motility. Cell 158, 300–313. https://doi.org/10.1016/j.cell.2014.04.050 (2014).
Zhou, L. et al. New insights into muscularis macrophages in the gut: from their origin to therapeutic targeting. Immunol. Res. https://doi.org/10.1007/s12026-023-09397-x (2023).
Becker, L., Spear, E. T., Sinha, S. R., Haileselassie, Y. & Habtezion, A. Age-related changes in gut microbiota alter phenotype of muscularis macrophages and disrupt gastrointestinal motility. Cell. Mol. Gastroenterol. Hepatol. 7, 243–245. https://doi.org/10.1016/j.jcmgh.2018.09.001 (2019).
Singh, A., Blanco, A., Sinnott, R. & Knaus, U. Rapid isolation and flow cytometry analysis of murine intestinal immune cells after chemically induced colitis. Bio-protocol 11, 11. https://doi.org/10.21769/BioProtoc.4182 (2021).
Tamura, A. et al. Distribution of two types of lymphocytes (intraepithelial and lamina-propria-associated) in the murine small intestine. Cell Tiss. Res. 313, 47–53. https://doi.org/10.1007/s00441-003-0706-4 (2003).
Yero, A. et al. Impact of early ARV initiation on relative proportions of effector and regulatory CD8 T cell in mesenteric lymph nodes and peripheral blood during acute SIV infection of rhesus macaques. J. Virol. 96, e0025522. https://doi.org/10.1128/jvi.00255-22 (2022).
Bondonese, A. et al. Impact of enzymatic digestion on single cell suspension yield from peripheral human lung tissue. Cytometry A 103, 777–785. https://doi.org/10.1002/cyto.a.24777 (2023).
Goodyear, A. W., Kumar, A., Dow, S. & Ryan, E. P. Optimization of murine small intestine leukocyte isolation for global immune phenotype analysis. J. Immunol. Methods 405, 97–108. https://doi.org/10.1016/j.jim.2014.01.014 (2014).
Pham, T. N. Q. et al. Flt3L-mediated expansion of plasmacytoid dendritic cells suppresses HIV infection in humanized mice. Cell reports 29, 2770–2782. https://doi.org/10.1016/j.celrep.2019.10.094 (2019).
Nemoto, S., Mailloux, A. W., Kroeger, J. & Mule, J. J. OMIP-031: Immunologic checkpoint expression on murine effector and memory T-cell subsets. Cytomet. A 89, 427–429. https://doi.org/10.1002/cyto.a.22808 (2016).
DiPiazza, A. T., Hill, J. P., Graham, B. S. & Ruckwardt, T. J. OMIP-061: 20-Color flow cytometry panel for high-dimensional characterization of murine antigen-presenting cells. Cytomet. A 95, 1226–1230. https://doi.org/10.1002/cyto.a.23880 (2019).
Mincham, K. T., Young, J. D. & Strickland, D. H. OMIP 076: High-dimensional immunophenotyping of murine T-cell, B-cell, and antibody secreting cell subsets. Cytomet. A 99, 888–892. https://doi.org/10.1002/cyto.a.24474 (2021).
Percie du Sert, N. et al. The ARRIVE guidelines 2.0: Updated guidelines for reporting animal research. PLoS Biol 18, e3000410. https://doi.org/10.1371/journal.pbio.3000410 (2020).
Soret, R. et al. Genetic background influences severity of colonic aganglionosis and response to GDNF enemas in the holstein mouse model of hirschsprung disease. Int. J. Mol. Sci. 22, 1. https://doi.org/10.3390/ijms222313140 (2021).
Soret, R. et al. A collagen VI-dependent pathogenic mechanism for Hirschsprung’s disease. J. Clin. Invest. 125, 4483–4496. https://doi.org/10.1172/JCI83178 (2015).
Soret, R. et al. Glial cell derived neurotrophic factor induces enteric neurogenesis and improves colon structure and function in mouse models of Hirschsprung disease. Gastroenterology 159, 1824-1838.e1817. https://doi.org/10.1053/j.gastro.2020.07.018 (2020).
Brandi, J., Wiethe, C., Riehn, M. & Jacobs, T. OMIP-93: A 41-color high parameter panel to characterize various co-inhibitory molecules and their ligands in the lymphoid and myeloid compartment in mice. Cytomet. A 103, 624–630. https://doi.org/10.1002/cyto.a.24740 (2023).
Cossarizza, A. et al. Guidelines for the use of flow cytometry and cell sorting in immunological studies (second edition). Eur. J. Immunol. 49, 1457–1973. https://doi.org/10.1002/eji.201970107 (2019).
Jarade, A., Di Santo, J. P. & Serafini, N. Group 3 innate lymphoid cells mediate host defense against attaching and effacing pathogens. Curr. Opin. Microbiol. 63, 83–91. https://doi.org/10.1016/j.mib.2021.06.005 (2021).
Joeris, T., Muller-Luda, K., Agace, W. W. & Mowat, A. M. Diversity and functions of intestinal mononuclear phagocytes. Mucosal. Immunol. 10, 845–864. https://doi.org/10.1038/mi.2017.22 (2017).
Withers, D. R. & Hepworth, M. R. Group 3 innate lymphoid cells: Communications hubs of the intestinal immune system. Front. Immunol. 8, 1298. https://doi.org/10.3389/fimmu.2017.01298 (2017).
Yero, A. et al. Dynamics and epigenetic signature of regulatory T-cells following antiretroviral therapy initiation in acute HIV infection. EBioMedicine 71, 103570. https://doi.org/10.1016/j.ebiom.2021.103570 (2021).
Autengruber, A., Gereke, M., Hansen, G., Hennig, C. & Bruder, D. Impact of enzymatic tissue disintegration on the level of surface molecule expression and immune cell function. Eur. J. Microbiol. Immunol. (Bp) 2, 112–120. https://doi.org/10.1556/EuJMI.2.2012.2.3 (2012).
Chen, Z. et al. Collagenase digestion down-regulates the density of CD27 on lymphocytes. J. Immunol. Methods 413, 57–61. https://doi.org/10.1016/j.jim.2014.06.017 (2014).
Skulska, K., Wegrzyn, A. S., Chelmonska-Soyta, A. & Chodaczek, G. Impact of tissue enzymatic digestion on analysis of immune cells in mouse reproductive mucosa with a focus on gammadelta T cells. J. Immunol. Methods 474, 112665. https://doi.org/10.1016/j.jim.2019.112665 (2019).
Bondonese, A. et al. Impact of enzymatic digestion on single cell suspension yield from peripheral human lung tissue. Cytomet. A https://doi.org/10.1002/cyto.a.24777 (2023).
Blom, K. G. et al. Isolation of murine intrahepatic immune cells employing a modified procedure for mechanical disruption and functional characterization of the B, T and natural killer T cells obtained. Clin. Experiment. Immunol. 155, 320–329. https://doi.org/10.1111/j.1365-2249.2008.03815.x (2009).
Bosco, N. & Noti, M. The aging gut microbiome and its impact on host immunity. Genes Immun. 22, 289–303. https://doi.org/10.1038/s41435-021-00126-8 (2021).
Walrath, T. et al. Age-related changes in intestinal immunity and the microbiome. J. Leukoc Biol. 109, 1045–1061. https://doi.org/10.1002/JLB.3RI0620-405RR (2021).
Al Nabhani, Z. et al. A weaning reaction to microbiota is required for resistance to immunopathologies in the adult. Immunity 50, 1276–1288. https://doi.org/10.1016/j.immuni.2019.02.014 (2019).
Lubin, J. B. et al. Arresting microbiome development limits immune system maturation and resistance to infection in mice. Cell Host Microbe 31, 554–570. https://doi.org/10.1016/j.chom.2023.03.006 (2023).
Menezes, J. S. et al. Stimulation by food proteins plays a critical role in the maturation of the immune system. Int. Immunol. 15, 447–455. https://doi.org/10.1093/intimm/dxg043 (2003).
Agace, W. W. & McCoy, K. D. Regionalized development and maintenance of the intestinal adaptive immune landscape. Immunity 46, 532–548. https://doi.org/10.1016/j.immuni.2017.04.004 (2017).
Bowcutt, R. et al. Heterogeneity across the murine small and large intestine. World J. Gastroenterol. 20, 15216–15232. https://doi.org/10.3748/wjg.v20.i41.15216 (2014).
Bondurand, N. & Southard-Smith, E. M. Mouse models of Hirschsprung disease and other developmental disorders of the enteric nervous system: Old and new players. Dev. Biol. 417, 139–157. https://doi.org/10.1016/j.ydbio.2016.06.042 (2016).
Pilon, N. Pigmentation-based insertional mutagenesis is a simple and potent screening approach for identifying neurocristopathy-associated genes in mice. Rare Dis. 4, e1156287 (2016).
Pilon, N. Treatment and prevention of neurocristopathies. Trends Mol. Med. 27, 451–468. https://doi.org/10.1016/j.molmed.2021.01.009 (2021).
Heuckeroth, R. O. Hirschsprung disease—Integrating basic science and clinical medicine to improve outcomes. Nat. Rev. Gastroenterol. Hepatol. 15, 152–167. https://doi.org/10.1038/nrgastro.2017.149 (2018).
Montalva, L. et al. Hirschsprung disease. Nat. Rev. Dis. Primers 9, 54. https://doi.org/10.1038/s41572-023-00465-y (2023).
Li, S. et al. Update on the pathogenesis of the hirschsprung-associated enterocolitis. Int J Mol Sci 24, 1. https://doi.org/10.3390/ijms24054602 (2023).
Cheng, Z. et al. Murine model of Hirschsprung-associated enterocolitis. I: phenotypic characterization with development of a histopathologic grading system. J. Pediatr. Surg. 45, 475–482. https://doi.org/10.1016/j.jpedsurg.2009.06.009 (2010).
Porokuokka, L. L. et al. Gfra1 underexpression causes Hirschsprung’s disease and Associated enterocolitis in mice. Cell. Mol. Gastroenterol. Hepatol. 7, 655–678. https://doi.org/10.1016/j.jcmgh.2018.12.007 (2019).
Gosain, A. et al. Impaired cellular immunity in the murine neural crest conditional deletion of endothelin receptor-B model of Hirschsprung’s disease. PLoS One 10, e0128822. https://doi.org/10.1371/journal.pone.0128822 (2015).
Medrano, G. et al. B-lymphocyte-intrinsic and -extrinsic defects in secretory immunoglobulin A production in the neural crest-conditional deletion of endothelin receptor B model of Hirschsprung-associated enterocolitis. FASEB J 33, 7615–7624. https://doi.org/10.1096/fj.201801913R (2019).
Patel, A. et al. Differential RET signaling pathways drive development of the enteric lymphoid and nervous systems. Sci. Signal. 5, 55. https://doi.org/10.1126/scisignal.2002734 (2012).
Veiga-Fernandes, H. et al. Tyrosine kinase receptor RET is a key regulator of Peyer’s patch organogenesis. Nature 446, 547–551. https://doi.org/10.1038/nature05597 (2007).
Frykman, P. K., Cheng, Z., Wang, X. & Dhall, D. Enterocolitis causes profound lymphoid depletion in endothelin receptor B- and endothelin 3-null mouse models of Hirschsprung-associated enterocolitis. Eur. J. Immunol. 45, 807–817. https://doi.org/10.1002/eji.201444737 (2015).
Chen, X. et al. Intestinal proinflammatory macrophages induce a phenotypic switch in interstitial cells of Cajal. J. Clin. Invest. 130, 6443–6456. https://doi.org/10.1172/JCI126584 (2020).
Karim, A., Tang, C. S. & Tam, P. K. The emerging genetic landscape of hirschsprung disease and its potential clinical applications. Front. Pediatr. 9, 638093. https://doi.org/10.3389/fped.2021.638093 (2021).
Ibiza, S. et al. Glial-cell-derived neuroregulators control type 3 innate lymphoid cells and gut defence. Nature 535, 440–443. https://doi.org/10.1038/nature18644 (2016).
Vargas-Leal, V. et al. Expression and function of glial cell line-derived neurotrophic factor family ligands and their receptors on human immune cells. J. Immunol. 175, 2301–2308. https://doi.org/10.4049/jimmunol.175.4.2301 (2005).
Mizoguchi, A. & Bhan, A. K. A case for regulatory B cells. J. Immunol. 176, 705–710. https://doi.org/10.4049/jimmunol.176.2.705 (2006).
Vazquez, M. I., Catalan-Dibene, J. & Zlotnik, A. B cells responses and cytokine production are regulated by their immune microenvironment. Cytokine 74, 318–326. https://doi.org/10.1016/j.cyto.2015.02.007 (2015).
Li, T., Liu, M., Sun, S., Liu, X. & Liu, D. Epithelial cells orchestrate the functions of dendritic cells in intestinal homeostasis. J. Biomed. Res. Environ. Sci. 1, 343–352. https://doi.org/10.37871/jbres1165 (2020).
Shresta, S., Kyle, J. L., Robert Beatty, P. & Harris, E. Early activation of natural killer and B cells in response to primary dengue virus infection in A/J mice. Virology 319, 262–273. https://doi.org/10.1016/j.virol.2003.09.048 (2004).
Kitoh, A. et al. Indispensable role of the Runx1-Cbfbeta transcription complex for in vivo-suppressive function of FoxP3+ regulatory T cells. Immunity 31, 609–620. https://doi.org/10.1016/j.immuni.2009.09.003 (2009).
Ascon, D. B. et al. Normal mouse kidneys contain activated and CD3+CD4- CD8- double-negative T lymphocytes with a distinct TCR repertoire. J Leukoc Biol 84, 1400–1409. https://doi.org/10.1189/jlb.0907651 (2008).
Freeman, B. E., Hammarlund, E., Raue, H. P. & Slifka, M. K. Regulation of innate CD8+ T-cell activation mediated by cytokines. Proc. Natl. Acad. Sci. USA 109, 9971–9976. https://doi.org/10.1073/pnas.1203543109 (2012).
Keir, M., Yi, Y., Lu, T. & Ghilardi, N. The role of IL-22 in intestinal health and disease. J. Exp. Med. 217, e20192195. https://doi.org/10.1084/jem.20192195 (2020).
Li, H. & Tsokos, G. C. Double-negative T cells in autoimmune diseases. Curr. Opin. Rheumatol. 33, 163–172. https://doi.org/10.1097/BOR.0000000000000778 (2021).
Wu, Z. et al. CD3(+)CD4(−)CD8(−) (Double-Negative) T cells in inflammation, immune disorders and cancer. Front. Immunol. 13, 816005. https://doi.org/10.3389/fimmu.2022.816005 (2022).
Rodriguez-Galán, M. C., Bream, J. H., Farr, A. & Young, H. A. Synergistic effect of IL-2, IL-12, and IL-18 on thymocyte apoptosis and Th1/Th2 cytokine expression. J. Immunol. 174, 2796–2804. https://doi.org/10.4049/jimmunol.174.5.2796 (2005).
Parra, M. et al. Memory CD73+IgM+ B cells protect against Plasmodium yoelii infection and express Granzyme B. PLoS One 15, e0238493. https://doi.org/10.1371/journal.pone.0238493 (2020).
Nascimento, D. C. et al. Sepsis expands a CD39(+) plasmablast population that promotes immunosuppression via adenosine-mediated inhibition of macrophage antimicrobial activity. Immunity 54, 2024–2041. https://doi.org/10.1016/j.immuni.2021.08.005 (2021).
Sallusto, F. & Lanzavecchia, A. Heterogeneity of CD4+ memory T cells: Functional modules for tailored immunity. Eur. J. Immunol. 39, 2076–2082. https://doi.org/10.1002/eji.200939722 (2009).
Ivanov, I. I. et al. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 139, 485–498. https://doi.org/10.1016/j.cell.2009.09.033 (2009).
Feng, T. et al. Th17 cells induce colitis and promote Th1 cell responses through IL-17 induction of innate IL-12 and IL-23 production. J. Immunol. 186, 6313–6318. https://doi.org/10.4049/jimmunol.1001454 (2011).
Rogers, P. R., Huston, G. & Swain, S. L. High antigen density and IL-2 are required for generation of CD4 effectors secreting Th1 rather than Th0 cytokines. J. Immunol. 161, 3844–3852. https://doi.org/10.4049/jimmunol.161.8.3844 (1998).
Srivastava, R. K., Dar, H. Y. & Mishra, P. K. Immunoporosis: Immunology of osteoporosis-role of T cells. Front. Immunol. 9, 657. https://doi.org/10.3389/fimmu.2018.00657 (2018).
Shevchenko, I. et al. Enhanced expression of CD39 and CD73 on T cells in the regulation of anti-tumor immune responses. Oncoimmunology 9, 1744946. https://doi.org/10.1080/2162402X.2020.1744946 (2020).
Ono-Ohmachi, A. et al. Effector memory CD4(+)T cells in mesenteric lymph nodes mediate bone loss in food-allergic enteropathy model mice, creating IL-4 dominance. Mucosal Immunol. 14, 1335–1346. https://doi.org/10.1038/s41385-021-00434-2 (2021).
Cho, M. J. et al. Steady-state memory-phenotype conventional CD4(+) T cells exacerbate autoimmune neuroinflammation in a bystander manner via the Bhlhe40/GM-CSF axis. Exp. Mol. Med. 55, 1033–1045. https://doi.org/10.1038/s12276-023-00995-1 (2023).
Lee, H. G. et al. Pathogenic function of bystander-activated memory-like CD4(+) T cells in autoimmune encephalomyelitis. Nat. Commun. 10, 709. https://doi.org/10.1038/s41467-019-08482-w (2019).
Dobson, H. E. et al. Antigen discovery unveils resident memory and migratory cell roles in antifungal resistance. Mucosal Immunol. 13, 518–529. https://doi.org/10.1038/s41385-019-0244-3 (2020).
Wilk, M. M. et al. Lung CD4 tissue-resident memory T cells mediate adaptive immunity induced by previous infection of mice with Bordetella pertussis. J. Immunol. 199, 233–243. https://doi.org/10.4049/jimmunol.1602051 (2017).
Chen, B. et al. TIGIT deficiency protects mice from DSS-induced colitis by regulating IL-17A-producing CD4(+) tissue-resident memory T cells. Front. Immunol. 13, 931761. https://doi.org/10.3389/fimmu.2022.931761 (2022).
Maddaloni, M. et al. Milk-based nutraceutical for treating autoimmune arthritis via the stimulation of IL-10- and TGF-beta-producing CD39+ regulatory T cells. PLoS One 10, e0117825. https://doi.org/10.1371/journal.pone.0117825 (2015).
Kochetkova, I., Thornburg, T., Callis, G. & Pascual, D. W. Segregated regulatory CD39+CD4+ T cell function: TGF-beta-producing Foxp3- and IL-10-producing Foxp3+ cells are interdependent for protection against collagen-induced arthritis. J. Immunol. 187, 4654–4666. https://doi.org/10.4049/jimmunol.1100530 (2011).
Fina, D. et al. Regulation of gut inflammation and th17 cell response by interleukin-21. Gastroenterology 134, 1038–1048. https://doi.org/10.1053/j.gastro.2008.01.041 (2008).
Yu, L. et al. CD69 enhances immunosuppressive function of regulatory T-cells and attenuates colitis by prompting IL-10 production. Cell Death Dis. 9, 905. https://doi.org/10.1038/s41419-018-0927-9 (2018).
Noble, A., Giorgini, A. & Leggat, J. A. Cytokine-induced IL-10-secreting CD8 T cells represent a phenotypically distinct suppressor T-cell lineage. Blood 107, 4475–4483. https://doi.org/10.1182/blood-2005-10-3994 (2006).
Alam, M. S., Cavanaugh, C., Pereira, M., Babu, U. & Williams, K. Susceptibility of aging mice to listeriosis: Role of anti-inflammatory responses with enhanced Treg-cell expression of CD39/CD73 and Th-17 cells. Int. J. Med. Microbiol. 310, 151397. https://doi.org/10.1016/j.ijmm.2020.151397 (2020).
Ochoa-Reparaz, J. et al. Induction of gut regulatory CD39(+) T cells by teriflunomide protects against EAE. Neurol. Neuroimmunol. Neuroinflam. 3, e291. https://doi.org/10.1212/NXI.0000000000000291 (2016).
Zhou, L. et al. TGF-beta-induced Foxp3 inhibits T(H)17 cell differentiation by antagonizing RORgammat function. Nature 453, 236–240. https://doi.org/10.1038/nature06878 (2008).
Sanchez, A. M., Zhu, J., Huang, X. & Yang, Y. The development and function of memory regulatory T cells after acute viral infections. J. Immunol. 189, 2805–2814. https://doi.org/10.4049/jimmunol.1200645 (2012).
Kleinewietfeld, M. et al. CCR6 expression defines regulatory effector/memory-like cells within the CD25(+)CD4+ T-cell subset. Blood 105, 2877–2886. https://doi.org/10.1182/blood-2004-07-2505 (2005).
Anthony, S. M. et al. Protective function and durability of mouse lymph node-resident memory CD8(+) T cells. Elife 10, 1. https://doi.org/10.7554/eLife.68662 (2021).
Schneider, E. et al. CD73-mediated adenosine production by CD8 T cell-derived extracellular vesicles constitutes an intrinsic mechanism of immune suppression. Nat. Commun. 12, 5911. https://doi.org/10.1038/s41467-021-26134-w (2021).
Jiang, X. et al. The ectonucleotidases CD39 and CD73 on T cells: The new pillar of hematological malignancy. Front. Immunol. 14, 1110325. https://doi.org/10.3389/fimmu.2023.1110325 (2023).
Nath, P. R. et al. Loss of CD47 alters CD8+ T cell activation in vitro and immunodynamics in mice. Oncoimmunology 11, 2111909. https://doi.org/10.1080/2162402X.2022.2111909 (2022).
Lykhopiy, V., Malviya, V., Humblet-Baron, S. & Schlenner, S. M. IL-2 immunotherapy for targeting regulatory T cells in autoimmunity. Genes Immun. 24, 248–262. https://doi.org/10.1038/s41435-023-00221-y (2023).
Peterson, R. A. Regulatory T-cells: Diverse phenotypes integral to immune homeostasis and suppression. Toxicol. Pathol. 40, 186–204. https://doi.org/10.1177/0192623311430693 (2012).
Juvet, S. C. & Zhang, L. Double negative regulatory T cells in transplantation and autoimmunity: Recent progress and future directions. J. Mol. Cell Biol. 4, 48–58. https://doi.org/10.1093/jmcb/mjr043 (2012).
Aychek, T. et al. IL-23-mediated mononuclear phagocyte crosstalk protects mice from Citrobacter rodentium-induced colon immunopathology. Nat. Commun. 6, 6525. https://doi.org/10.1038/ncomms7525 (2015).
Holland, A. M., Bon-Frauches, A. C., Keszthelyi, D., Melotte, V. & Boesmans, W. The enteric nervous system in gastrointestinal disease etiology. Cell Mol. Life Sci. 78, 4713–4733. https://doi.org/10.1007/s00018-021-03812-y (2021).
Wu, R. Q., Zhang, D. F., Tu, E., Chen, Q. M. & Chen, W. The mucosal immune system in the oral cavity-an orchestra of T cell diversity. Int. J. Oral. Sci. 6, 125–132. https://doi.org/10.1038/ijos.2014.48 (2014).
Bain, C. C. & Mowat, A. M. Macrophages in intestinal homeostasis and inflammation. Immunol. Rev. 260, 102–117 (2014).
Viola, M. F. & Boeckxstaens, G. Intestinal resident macrophages: Multitaskers of the gut. Neurogastroenterol. Motil. 32, e13843. https://doi.org/10.1111/nmo.13843 (2020).
De Calisto, J., Villablanca, E. J. & Mora, J. R. FcgammaRI (CD64): an identity card for intestinal macrophages. Eur. J. Immunol. 42, 3136–3140. https://doi.org/10.1002/eji.201243061 (2012).
Cerovic, V., Bain, C. C., Mowat, A. M. & Milling, S. W. Intestinal macrophages and dendritic cells: what’s the difference?. Trends Immunol. 35, 270–277. https://doi.org/10.1016/j.it.2014.04.003 (2014).
Corbin, A. L. et al. IRF5 guides monocytes toward an inflammatory CD11c+ macrophage phenotype and promotes intestinal inflammation. Sci. Immunol. 5, 1–15 (2020).
Persson, E. K., Scott, C. L., Mowat, A. M. & Agace, W. W. Dendritic cell subsets in the intestinal lamina propria: Ontogeny and function. Eur. J. Immunol. 43, 3098–3107. https://doi.org/10.1002/eji.201343740 (2013).
Schridde, A. et al. Tissue-specific differentiation of colonic macrophages requires TGFbeta receptor-mediated signaling. Mucosal. Immunol. 10, 1387–1399. https://doi.org/10.1038/mi.2016.142 (2017).
Soehnlein, O., Lindbom, L. & Weber, C. Mechanisms underlying neutrophil-mediated monocyte recruitment. Blood 114, 4613–4623. https://doi.org/10.1182/blood-2009-06-221630 (2009).
Nakata, K., Yamamoto, M., Inagawa, H. & Soma, G. Effects of interactions between intestinal microbiota and intestinal macrophages on health. Anticancer Res. 33, 2849–2854 (2013).
Troy, A. E. et al. IL-27 regulates homeostasis of the intestinal CD4+ effector T cell pool and limits intestinal inflammation in a murine model of colitis. J. Immunol. 183, 2037–2044. https://doi.org/10.4049/jimmunol.0802918 (2009).
Shaw, T. N. et al. Tissue-resident macrophages in the intestine are long lived and defined by Tim-4 and CD4 expression. J. Exp. Med. 215, 1507–1518. https://doi.org/10.1084/jem.20180019 (2018).
Bain, C. C. & Schridde, A. Origin, differentiation, and function of intestinal macrophages. Front. Immunol. 9, 2733. https://doi.org/10.3389/fimmu.2018.02733 (2018).
Baghdadi, M. et al. TIM-4 glycoprotein-mediated degradation of dying tumor cells by autophagy leads to reduced antigen presentation and increased immune tolerance. Immunity 39, 1070–1081. https://doi.org/10.1016/j.immuni.2013.09.014 (2013).
Orecchioni, M., Ghosheh, Y., Pramod, A. B. & Ley, K. Macrophage polarization: Different gene signatures in M1(LPS+) vs. classically and M2(LPS-) vs. alternatively activated macrophages. Front Immunol 10, 1084. https://doi.org/10.3389/fimmu.2019.01084 (2019).
Coskun, M. Intestinal epithelium in inflammatory bowel disease. Front. Med. (Lausanne) 1, 24. https://doi.org/10.3389/fmed.2014.00024 (2014).
Schreurs, M. W. J., Eggert, A. A. O., de Boer, A. J., Figdor, C. G. & Adema, G. J. Generation and functional characterization of mouse monocyte-derived dendritic cells. Eur. J. Immunol. 29, 2835–2841. https://doi.org/10.1002/(sici)1521-4141(199909)29:09%3c2835::Aid-immu2835%3e3.0.Co;2-q (1999).
Li, N. et al. Serotonin activates dendritic cell function in the context of gut inflammation. Am. J. Pathol. 178, 662–671. https://doi.org/10.1016/j.ajpath.2010.10.028 (2011).
Bain, C. C. et al. TGFbetaR signalling controls CD103(+)CD11b(+) dendritic cell development in the intestine. Nat. Commun. 8, 620. https://doi.org/10.1038/s41467-017-00658-6 (2017).
Stagg, A. J. Intestinal dendritic cells in health and gut inflammation. Front. Immunol. 9, 2883. https://doi.org/10.3389/fimmu.2018.02883 (2018).
Li, L. J. et al. Induction of colitis in mice with food allergen-specific immune response. Sci. Rep. 6, 32765. https://doi.org/10.1038/srep32765 (2016).
Yang, P. C. et al. TIM-4 expressed by mucosal dendritic cells plays a critical role in food antigen-specific Th2 differentiation and intestinal allergy. Gastroenterology 133, 1522–1533. https://doi.org/10.1053/j.gastro.2007.08.006 (2007).
Acknowledgements
The authors would like to thank the Cellular analyses and Imaging core (CERMO-FC, UQAM) for assistance with flow cytometry data acquisition and FlowJo analyses. This work was supported by grants from the Canadian Institutes of Health Research (CIHR # PJT-180290) and the Programme de soutien aux organismes de recherche et d’innovation from the Quebec government (PSOv2b #54829) to NP. NP also holds the UQAM Research Chair on Rare Genetic Diseases, while MAJ holds the Tier 2 CIHR Canada Research Chair in Immuno-Virology. NL and AY were supported by scholarships from the CERMO-FC and the Fonds de recherche du Québec – Santé (FRQS), respectively.
Funding
RS and NP are co-founders of Neurenati Therapeutics Inc. Funding organizations had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Author information
Authors and Affiliations
Contributions
Conceived the study: NP and MAJ. Supervised the study: NP, MAJ and RS. Designed the experiments: AY (mechanical digestion, panels, and staining protocol) and NL (enzymatic digestions, lymphoid panel). Performed the experiments: NL (sample collection and digestion, cell preparation, staining and acquisition) and AY (compensation). Analyzed the data: NL (gating and t-SNE), AY (gating and t-SNE), and RS (statistical analyses). Contributed reagents/materials/analysis tools: NP and MAJ. Wrote the paper: NL (original draft), RS (review and editing), and NP (review and editing). All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lassoued, N., Yero, A., Jenabian, MA. et al. Efficient enzyme-free method to assess the development and maturation of the innate and adaptive immune systems in the mouse colon. Sci Rep 14, 11063 (2024). https://doi.org/10.1038/s41598-024-61834-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-61834-5
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.