Abstract
Genome-wide association studies have identified several genetic variants associated with nonalcoholic fatty liver disease. To emphasize metabolic abnormalities in fatty liver, metabolic (dysfunction)-associated fatty liver disease (MAFLD) has been introduced; thus, we aimed to investigate single-nucleotide polymorphisms related to MAFLD and its subtypes. A genome-wide association study was performed to identify genetic factors related to MAFLD. We used a Korean population-based sample of 2282 subjects with MAFLD and a control group of 4669. We replicated the results in a validation sample which included 639 patients with MAFLD and 1578 controls. Additionally, we categorized participants into three groups, no MAFLD, metabolic dysfunction (MD)-MAFLD, and overweight/obese-MAFLD. After adjusting for age, sex, and principal component scores, rs738409 [risk allele G] and rs3810622 [risk allele T], located in the PNPLA3 gene, showed significant associations with MAFLD (P-values, discovery set = 1.60 × 10–15 and 4.84 × 10–10; odds ratios, 1.365 and 1.284, validation set = 1.39 × 10–4, and 7.15 × 10–4, odds ratios, 1.299 and 1.264, respectively). An additional SNP rs59148799 [risk allele G] located in the GATAD2A gene showed a significant association with MAFLD (P-values, discovery set = 2.08 × 10–8 and validation set = 0.034, odds ratios, 1.387 and 1.250). rs738409 was significantly associated with MAFLD subtypes ([overweight/obese-MAFLD; odds ratio (95% confidence interval), P-values, 1.515 (1.351–1.700), 1.43 × 10–12 and MD-MAFLD: 1.300 (1.191–1.416), 2.90 × 10–9]. There was a significant relationship between rs3810622 and overweight/obese-MAFLD and MD-MAFLD [odds ratios (95% confidence interval), P-values, 1.418 (1.258, 1.600), 1.21 × 10–8 and 1.225 (1.122, 1.340), 7.06 × 10–6, respectively]; the statistical significance remained in the validation set. PNPLA3 was significantly associated with MAFLD and MAFLD subtypes in the Korean population. These results indicate that genetic factors play an important role in the pathogenesis of MAFLD.
Similar content being viewed by others
Introduction
As the most common cause of chronic liver disease worldwide, the global trend of non-alcoholic fatty liver disease (NAFLD) represents an important public health problem1,2. Both environmental and genetic factors play important roles in the development of NAFLD3,4. With recent advances in genetic research, genome-wide association studies (GWASs) have identified several genetic variants. The most replicated variants associated with NAFLD include patatin-like phospholipase domain-containing 3 (PNPLA3), transmembrane 6 superfamily member 2 (TM6SF2), hydroxysteroid 17-beta dehydrogenase 13 (HSD17B13), membrane bound O-acyltransferase domain containing 7 (MBOAT7), glucokinase regulator (GCKR) and SAMM505,6.
A new definition of metabolic (dysfunction)-associated with fatty liver disease (MAFLD) has been introduced, emphasizing the role of metabolic dysfunction in the clinical outcomes of patients with hepatic steatosis7,8. MAFLD criteria have demonstrated improved capability in identifying a subset of patients with more advanced liver disease and higher mortality compared to those diagnosed solely with NAFLD9,10,11. Moreover, in MAFLD patients, MAFLD subtypes have proven to be effective in capturing differential outcomes related to cardiovascular disease or disease-specific mortality12,13.
Although MAFLD and NAFLD overlap considerably and both disease entities have similar clinical features14, utilizations of the MAFLD criteria have identified more individuals with advanced liver disease and patients at a high risk for developing complications or other metabolic co-morbidities15,16. Therefore, investigating genetic variants associated with MAFLD can provide insight into the pathogenesis of MAFLD and help stratify high-risk patients. A recent study reported no association between a single nucleotide polymorphism (SNP) and the risk of MAFLD in a Han population17; however, studies investigating the influence of genetic susceptibility on MAFLD are limited. In this study, we investigated the association between genome-wide SNP and MAFLD and its subgroups in an asymptomatic Korean population.
Materials and methods
Study population
We analyzed the Gene-Environment Interaction and phenotype cohort data as previously described18. From January 2014 and December 2014, 9676 people donated blood samples during regular health check-ups conducted at Seoul National University Hospital Healthcare System Gangnam Center. Their blood samples were stored in a biorepository with informed consent. Participants with missing data from the total enrollment were excluded. The enrolled population was divided into two groups based on the time of enrollment. Samples from subjects enrolled between January 2014 and October 2014 were used as the discovery set, and samples from subjects enrolled in subsequent months were used as the validation set. Results using the discovery set were verified using the validation set.
The Institutional Review Board (IRB) approved the storage of biospecimens with informed consent (Seoul National University Hospital/IRB number 1103-127-357). Because this study used anonymized clinical and genetic information collected and analyzed previously, the board approved this study protocol, and the informed consent was waived (Seoul National University Hospital/IRB number 2109-012-1251). This research was performed in accordance with the Declaration of Helsinki.
Clinical and laboratory assessments
Each subject completed a medical history questionnaire and received an anthropometric evaluation. Each participant’s weight was measured using a digital scale. Body mass index (BMI) was calculated as the weight (kg) divided by height (m) squared. Waist circumference was measured at the midpoint between the lower costal margin and anterior superior iliac crest by a well-trained person. Systolic and diastolic blood pressures were measured twice, and the mean values were reported.
The laboratory and radiological tests were performed on the same day. Blood specimens were obtained from each participant after an overnight fast of ≥ 8 h. Laboratory evaluations included serum total cholesterol, triglyceride, high-density lipoprotein (HDL) cholesterol, fasting glucose, and high-sensitivity C-reactive protein. Hypertension was defined as a systolic blood pressure ≥ 140 mm Hg, diastolic blood pressure ≥ 90 mm Hg, or use of antihypertensive agents. Diabetes mellitus was defined as a fasting serum glucose level of ≥ 126 mg/dL or glycosylated hemoglobin ≥ 6.5% or the use of antidiabetic medications.
The diagnosis of fatty liver was based on ultrasonographic findings (Acusion, Sequoia 512, Siemens, Mountain View, CA, USA) of experienced radiologists who were unaware of the clinical information. Sonographic features of hepatic steatosis were classified according to previously described protocols13,14.
Definition of MAFLD and subgroups
MAFLD was defined as the presence of metabolic risk factors in the setting of hepatic steatosis based on the diagnostic criteria proposed by an international consensus in 20207. This definition does not exclude those with other concomitant liver diseases or significant alcohol consumption. MAFLD was diagnosed as the presence of hepatic steatosis with one or more of the following: (1) overweight or obese (BMI ≥ 23 kg/m2), (2) diabetes mellitus, or (3) at least two metabolic risk abnormalities. Metabolic risk abnormalities consisted of (1) waist circumference (WC) ≥ 90 cm for men and 80 ≥ cm for women, (2) blood pressure ≥ 130/85 mm Hg or specific drug treatment, (3) fasting plasma triglycerides ≥ 150 mg/dL or specific drug treatment, (4) plasma HDL-cholesterol < 40 mg/dl for men and < 50 mg/dL for women or specific drug treatment, (5) prediabetes (fasting glucose 100–125 mg/dL or hemoglobin A1c 5.7–6.4%), (6) homeostasis model assessment of insulin resistance score ≥ 2.5, (7) plasma high-sensitivity C-reactive protein level > 2 mg/dL. As a homeostasis model assessment of the insulin resistance score was not available, this criterion was not implemented in our study.
In the present study, participants with MAFLD were classified into two groups, modified from a previous study19. First, we determined MAFLD-metabolic dysfunction (MD) subgroup classification based on the presence of diabetes or at least two metabolic abnormalities regardless of the BMI; the remaining non-diabetic subjects who had fewer than two metabolic abnormalities and met only the BMI ≥ 23 kg/m2 criteria of MAFLD were categorized as the MAFLD-overweight/obese group.
Genotyping and quality control
Genomic DNA was extracted from blood samples and 200 ng of DNA from each patient was genotyped using Affymetrix Axiom® Customized Biobank Genotyping Arrays (Affymetrix, Santa Clara, CA, USA). Genotype data were produced using the Korean Chip, designed by the Center for Genome Science, Korea National Institute of Health, Korea (4845–301, 3000–3031). The quality control measures were performed using the PLINK program (version 1.07; Free Software Foundation Inc., Boston, MA, USA). Samples meeting any of the following criteria were removed: (i) gender discrepancy, (ii) call rate ≤ 97%, and (iii) related and cryptically related individuals (identical by state > 90%). SNPs were filtered if (1) the call rate was < 95%, (2) minor allele frequency was ≤ 0.05, or (3) deviation from the Hardy–Weinberg equilibrium permutation test (P < 1.00 × 10−5) was observed. SHAPEIT2 v2.r904 and IMPUTE2 version 2.3.2 were used for pre-phasing the data and genotype imputations in GWAS.20,21 As a reference panel, we used 1000 Genomes Phase 3 haplotypes. Any imputed SNPs with imputation quality scores < 0.5 were excluded. After quality control, 3,631,724 SNPs in the discovery set and 3,587,584 SNPs in the validation set remained for association analysis.
Statistical analysis
Comparisons of continuous variables in the baseline characteristics between the two groups (discovery set vs validation set and no MAFLD vs. MAFLD) were performed using Student’s t-test, and categorical variables were compared using the chi-square test or Fisher’s exact test. Binary logistic regression analysis was used to analyze the influence of SNPs on MAFLD. Ordinal and multinomial logistic regression analyses were performed to investigate the association of SNPs among MAFLD subgroups. The analysis was performed using an additive genetic model, adjusting for age, sex, and five principal component scores. The genomic inflation factor (λ) was computed to confirm that population stratification did not cause confounding. A λ close to 1 indicates the absence of genomic inflation22. SNPs with a P-value of less than 5.0 × 10–8, which is conventionally considered to represent genome-wide significance, in the discovery set were re-evaluated in the validation set. We also performed a joint analysis on the combined samples. To evaluate the independence of association signals, we also performed a conditional analysis.
The PLINK software package (version 1.07), R statistical software package (version 3.1.1, R Development Core Team; R Foundation for Statistical Computing, Vienna, Austria), and Trinculo (Bayesian and frequentist multinomial logistic regression for GWASs of multicategory phenotypes) were used to test the association between MAFLD and SNPs in the genome and draw a Manhattan plot and quantile–quantile (Q-Q) plots. We used LocusZoom and linkage disequilibrium data from East Asians in the 1000 Genomes Project to generate a regional plot of significant genetic variation23.
Results
Study population
A total of 9168 subjects were included, of whom 58.6% were male. Based on the definitions described in the Methods section, there were 6951 samples in the discovery set and 2217 samples in the validation set. Characteristics of the study population are summarized in Table 1. The subjects in the validation set were older and had higher BMI, WC, and fasting glucose and lower proportions of MAFLD (P < 0.05). Most anthropometric and laboratory variables were less metabolically favorable in subjects with MAFLD than those without MAFLD (all P < 0.001, Table 1).
GWAS of MAFLD
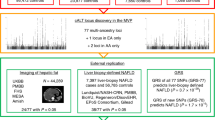
We first evaluated the genome-wide associations of MAFLD in the discovery set, with a significance threshold of P < 5.0 × 10–8, after adjusting for age, sex, and principal component scores. Figure 1 shows the Manhattan and Q-Q plots. The Q-Q plot revealed no evidence of any inflation of the test statistics (λ = 1.012, min 0.997 max 1.021). The significant SNPs in the discovery set that have been successfully replicated are listed in Table 2. In the GWAS discovery set, the two top SNPs on 22 chromosomes were significantly associated with MAFLD, which remained significant in the validation set. rs738409 and rs3810622 located in the PNPLA3 gene showed significant associations with MAFLD (P-values, discovery set = 1.60 × 10–15 and 4.84 × 10–10, odds ratio [OR], 1.365 and 1.284, respectively; validation set = 1.39 × 10–4 and 7.15 × 10–4, OR, 1.299 and 1.264, respectively). A regional plot of chromosome 22 is shown in Fig. 2. An additional SNP rs59148799, located in the GATA zinc finger domain containing 2A (GATAD2A) gene on chromosome 19, showed a significant association with MAFLD (P-values, discovery set = 2.08 × 10–8 and validation set = 0.034, Table 2). A regional plot of chromosome 19 is shown in Fig. 3. When we performed a joint analysis, these significant associations remained (Table 2).
Manhattan and quantile–quantile (Q–Q) plot for single nucleotide polymorphism in the discovery set. (A) Manhattan plot of the P-values in the genome-wide association studies for MAFLD. The horizontal dash line indicates the preset threshold of P = 5.0 × 10–8. (B) Q–Q plot showing expected versus observed (–log10 P-value). The expected line is shown in red and confidence bands are shown in gray.
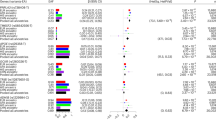
Regional association plots for chromosome 22 (A) rs738409 (B) rs3810622. Nearby SNPs are color-coded according to the level of linkage disequilibrium with the top SNP. The left y-axis represents the significance of the association based on the -log10 P-value (logistic regression in the combined samples), and the right y-axis represents the recombination rate across regions. SNP, single nucleotide polymorphism.
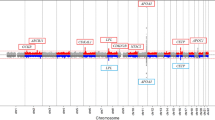
Regional association plots for chromosome 19. Nearby SNPs are color-coded according to the level of linkage disequilibrium with the top SNP. The left y-axis represents the significance of the association based on the -log10 P-value (logistic regression in the combined samples), and the right y-axis represents the recombination rate across regions. SNP, single nucleotide polymorphism.
We also performed a conditional analysis to evaluate the independence of association signals. Conditional analysis with multiple lead SNPs on same chromosome showed that the effect of rs3810622 on MAFLD disappeared, but the significance of rs738409 remained even when rs3810622 was evaluated together (Supplementary Table 1).
Impact of SNPs on MAFLD subtypes
Next, we investigated the associations between SNPs identified in the GWAS and MAFLD subtypes (no MAFLD, overweight/obese MAFLD, and MD-MAFLD). Ordinal logistic regression analysis after adjusting for age, sex, and five principal component scores showed that two SNPs, rs738409 and rs3810622, located in the PNPLA3 gene, showed significant associations with MAFLD subtypes (P-values, discovery set = 6.63 × 10–13, and 2.71 × 10–8, validation set = 5.54 × 10–4, and 0.002, Joint set = 1.64 × 10–15, and 1.75 × 10–10, respectively; Table 3). When we performed a conditional analysis, the independent association of rs738409 remained even when rs3810622 was evaluated together (Supplementary Table 2).
We investigated the ORs of each SNP of the MAFLD subtype compared to the group without MAFLD. Associations between the two SNPs and MAFLD subtypes are shown in Table 4. rs738409 was associated with a higher risk of overweight/obese MAFLD and MD-MAFLD [discovery set, OR (95% confidence interval (CI)), P-value, 1.515 (1.351–1.700), 1.43 × 10–12 and 1.300 (1.191–1.416), 2.90 × 10–9, respectively; validation set, OR (95% CI), P-value, 1.466 (1.182–1.818), 0.0005 and 1.233 (1.058–1.437), 0.0073, respectively)]. Additionally, significant associations were observed between rs3810622 and overweight/obese MAFLD and MD-MAFLD [discovery set, OR (95% CI), P-value, 1.418 (1.258, 1.600), 1.21 × 10–8 and 1.225 (1.122, 1.340), 7.06 × 10–6; validation set, OR (95% CI), P-value, 1.453 (1.164, 1.815), 9.76 × 10–4 and 1.192 (1.022, 1.389), 0.0248, respectively]. When we performed a joint analysis, these significant associations remained (Table 4). A conditional analysis with multiple lead SNPs showed that the effect of rs738409 on MAFLD remained significant when rs3810622 was evaluated together (Supplementary Table 3).
Discussion
Although the main pathogenesis is common between NAFLD and MAFLD, there have been no GWAS on MAFLD in Korea. In this study, we found a significant association between PNPLA3 and GATAD2A gene variants and MAFLD in a Korean population using GWAS. Moreover, we demonstrated that PNPLA3 rs738409 and rs3810622 were associated with MAFLD subtypes.
A recent review investigated the effects and applications of SNPs associated with MAFLD; however, the disease groups included in most of the studies were NAFLD rather than pure MAFLD24. A previous study based on the Han population found no association between SNPs and the risk of MAFLD in Western China17. Some significant associations may not be found because the sample size was not large enough (286 cases and 250 healthy controls). Recently, a systematic review reported genetic predispositions in MAFLD showing that rs641738C > T near membrane-bound O-acyltransferase domain-containing 7 is associated with increased hepatic fat, MAFLD severity, susceptibility to develop nonalcoholic steatohepatitis, advanced fibrosis, and hepatocellular carcinoma in Western populations with MAFLD25. However, these results were not based on GWAS.
In this study, three SNPs were significantly associated with MAFLD. Among them, rs738409 encodes PNPLA3 I148M, which is associated with increased hepatic fat in multiple ethnicities26,27,28,29. The PNPLA3 gene, located in hepatic lipid droplets, is involved in lipid metabolism by interfering with adipose triglyceride lipase activity and modulates hepatic triglyceride accumulation30,31. The PNPLA3 rs3810622 intron variant has been previously reported to be associated with elevated alanine aminotransferase and blood glucose32 and decreased serum triglyceride in NAFLD patients, suggesting its role in MAFLD as well as NAFLD33,34.
Additionally, rs59148799, located in GATAD2A showed a significant association with MAFLD. Although bioinformatic analysis has shown that this SNP is an intronic polymorphism that does not influence the protein structure or RNA splicing, it remains unclear whether transcription factor-binding sites are located within this SNP. In line with our results, GATAD2A rs4808199 has been reported to be associated with NAFLD in Japanese35 and U.S. multi‐ethnic populations36 although MAFLD was not considered in these studies. In contrast, GATAD2A rs4808199 showed no significant correlation with MAFLD in Western Chinese17,37. Collectively, further studies are required to confirm the association between GATAD2A and MAFLD.
PNPLA3 and SAMM50 genes were known to be significantly associated with the presence and severity of NAFLD in a Korean population in a previous study6. Compared to a previous study in Korean population6, the phenotype in this study focused on MAFLD and did not exclude heavy alcohol drinkers, or patients with concomitant liver disease. PNPLA3 variants showed a significant association with MAFLD as well as NAFLD, but no association was found between SAMM50 variants and MAFLD in this study. Other variants in the TM6SF2, HSD17B13, MBOAT7 and GCKR genes previously known to be associated with NAFLD were not found to be associated with MAFLD in this study. Ethnic differences among study populations and differences in outcome phenotypes may cause differences in results.
MAFLD affects a heterogeneous group of patients, including overweight/obese patients without metabolic abnormalities and lean patients with metabolic dysfunction. In terms of mortality, the MAFLD subgroups showed differential outcomes according to the accompanying metabolic dysfunctions13,38. On investigating the associations between SNPs and MAFLD subtypes, rs738409 and rs3810622, located in PNPLA3, showed significant associations with MAFLD subtypes. Further, there were significant relationships in both overweight/obese MAFLD and MD-MAFLD patients, suggesting possible genetic associations between PNPLA3 gene and MAFLD subtypes.
The present study had certain limitations. First, the discovery and validation sets selected were from the same population that underwent regular health checkups at our institute. Further validation studies using other populations should be conducted. Second, although ultrasonography is widely used as a first-line method to detect hepatic steatosis in clinical practice39,40, it is relatively insensitive for detecting mild steatosis41. Third, a definition of metabolic dysfunction-associated steatotic liver disease (MASLD) has been introduced recently, which overcomes the exclusive and stigmatizing nature of the NAFLD and the limitations of MAFLD42. Because this study focused on MAFLD rather than NAFLD or MASLD, we did not exclude, or distinguish concomitant liver disease or significant alcohol use according to the MAFLD definition. Therefore, samples and observations may be biased with respect to NAFLD or MASLD. Additionally, we did not consider disease activity or severity in this study. Finally, there may be selection bias because the study population may not be representative of the general population of subjects who visit a single health screening center in Korea for health checkups.
In conclusion, PNPLA3 was significantly associated with MAFLD and its subtypes in the Korean population. Our findings indicate that genetic factors play important roles in the pathogenesis of MAFLD and NAFLD.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Change history
14 May 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41598-024-61718-8
References
Paik, J. M., Golabi, P., Younossi, Y., Mishra, A. & Younossi, Z. M. Changes in the global burden of chronic liver diseases from 2012 to 2017: The growing impact of NAFLD. Hepatology 72(5), 1605–1616 (2020).
Younossi, Z. M. Non-alcoholic fatty liver disease-a global public health perspective. J. Hepatol. 70(3), 531–544 (2019).
Sookoian, S. & Pirola, C. J. Genetics in non-alcoholic fatty liver disease: The role of risk alleles through the lens of immune response. Clin. Mol. Hepat. 29(Suppl), S184-s195 (2023).
Seko, Y., Yamaguchi, K. & Itoh, Y. The genetic backgrounds in nonalcoholic fatty liver disease. Cli.n J. Gastroenterol. 11(2), 97–102 (2018).
Anstee, Q. M. et al. Genome-wide association study of non-alcoholic fatty liver and steatohepatitis in a histologically characterised cohort(☆). J. Hepatol. 73(3), 505–515 (2020).
Chung, G. E. et al. Genetic polymorphisms of PNPLA3 and SAMM50 are associated with nonalcoholic fatty liver disease in a Korean population. Gut Liver 12(3), 316–323 (2018).
Eslam, M. et al. A new definition for metabolic dysfunction-associated fatty liver disease: An international expert consensus statement. J. Hepatol. 73(1), 202–209 (2020).
Kang, S. H., Cho, Y., Jeong, S. W., Kim, S. U. & Lee, J. W. From nonalcoholic fatty liver disease to metabolic-associated fatty liver disease: Big wave or ripple?. Clin. Mol. Hepatol. 27(2), 257–269 (2021).
Yamamura, S. et al. MAFLD identifies patients with significant hepatic fibrosis better than NAFLD. Liver Int. 40(12), 3018–3030 (2020).
Nguyen, V. H., Le, M. H., Cheung, R. C. & Nguyen, M. H. Differential clinical characteristics and mortality outcomes in persons with NAFLD and/or MAFLD. Clin. Gastroenterol. Hepatol. 19, 2172 (2021).
Huang, Q., Zou, X., Wen, X., Zhou, X. & Ji, L. NAFLD or MAFLD: Which has closer association with all-cause and cause-specific mortality?-Results from NHANES III. Front. Med. (Lausanne) 8, 693507 (2021).
Lee, H., Lim, T. S., Kim, S. U. & Kim, H. C. Long-term cardiovascular outcomes differ across metabolic dysfunction-associated fatty liver disease subtypes among middle-aged population. Hepatol. Int. 16, 1308–1317 (2022).
Chung, G. E. et al. Lean or diabetic subtypes predict increased all-cause and disease-specific mortality in metabolic-associated fatty liver disease. BMC Med. 21(1), 4 (2023).
Younossi, Z. M. et al. Are there outcome differences between NAFLD and metabolic-associated fatty liver disease?. Hepatology 76, 1423–1437 (2022).
Gofton, C., Upendran, Y., Zheng, M. H. & George, J. MAFLD: How is it different from NAFLD?. Clin. Mol. Hepatol. 29(Suppl), S17-s31 (2023).
Ayada, I. et al. Systematically comparing epidemiological and clinical features of MAFLD and NAFLD by meta-analysis: Focusing on the non-overlap groups. Liver Int. 42(2), 277–287 (2022).
Liao, S. et al. Genetic variants associated with metabolic dysfunction-associated fatty liver disease in western China. J. Clin. Lab. Anal. 36(9), e24626 (2022).
Oh, S. W. et al. Genome-wide association study of metabolic syndrome in Korean populations. PLoS One 15(1), e0227357 (2020).
Sohn, W., Kwon, H. J., Chang, Y., Ryu, S. & Cho, Y. K. Liver Fibrosis in Asians with metabolic dysfunction-associated fatty liver disease. Clin. Gastroenterol. Hepatol. 20, e1135 (2021).
Delaneau, O., Zagury, J. F. & Marchini, J. Improved whole-chromosome phasing for disease and population genetic studies. Nat. Methods 10(1), 5–6 (2013).
Howie, B. N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet 5(6), e1000529 (2009).
Devlin, B., Roeder, K. & Wasserman, L. Genomic control, a new approach to genetic-based association studies. Theor. Popul. Biol. 60(3), 155–166 (2001).
Pruim, R. J. et al. LocusZoom: Regional visualization of genome-wide association scan results. Bioinformatics 26(18), 2336–2337 (2010).
Zhu, X., Xia, M. & Gao, X. Update on genetics and epigenetics in metabolic associated fatty liver disease. Ther. Adv. Endocrinol. Metab. 13, 20420188221132136 (2022).
Ismaiel, A. & Dumitrascu, D. L. Genetic predisposition in metabolic-dysfunction-associated fatty liver disease and cardiovascular outcomes-systematic review. Eur. J. Clin. Invest. 50(10), e13331 (2020).
Romeo, S. et al. Genetic variation in PNPLA3 confers susceptibility to nonalcoholic fatty liver disease. Nat. Genet. 40(12), 1461–1465 (2008).
Yuan, X. et al. Population-based genome-wide association studies reveal six loci influencing plasma levels of liver enzymes. Am. J. Hum. Genetics 83(4), 520–528 (2008).
Speliotes, E. K. et al. Genome-wide association analysis identifies variants associated with nonalcoholic fatty liver disease that have distinct effects on metabolic traits. PLoS Genet. 7(3), e1001324 (2011).
Sookoian, S. & Pirola, C. J. Meta-analysis of the influence of I148M variant of patatin-like phospholipase domain containing 3 gene (PNPLA3) on the susceptibility and histological severity of nonalcoholic fatty liver disease. Hepatology 53(6), 1883–1894 (2011).
Wang, Y., Kory, N., BasuRay, S., Cohen, J. C. & Hobbs, H. H. PNPLA3, CGI-58, and inhibition of hepatic triglyceride hydrolysis in mice. Hepatology 69(6), 2427–2441 (2019).
Carlsson, B. et al. Review article: the emerging role of genetics in precision medicine for patients with non-alcoholic steatohepatitis. Aliment. Pharmacol. Ther. 51(12), 1305–1320 (2020).
Song, G. et al. Association of patatin-like phospholipase domain-containing protein 3 gene polymorphisms with susceptibility of nonalcoholic fatty liver disease in a Han Chinese population. Med. (Baltimore) 95(33), e4569 (2016).
Larrieta-Carrasco, E. et al. Genetic variants in COL13A1, ADIPOQ and SAMM50, in addition to the PNPLA3 gene, confer susceptibility to elevated transaminase levels in an admixed Mexican population. Exp. Mol. Pathol. 104(1), 50–58 (2018).
Kitamoto, T. et al. Genome-wide scan revealed that polymorphisms in the PNPLA3, SAMM50, and PARVB genes are associated with development and progression of nonalcoholic fatty liver disease in Japan. Hum. Genet. 132(7), 783–792 (2013).
Kawaguchi, T. et al. Risk estimation model for nonalcoholic fatty liver disease in the Japanese using multiple genetic markers. PLoS One 13(1), e0185490 (2018).
Wang, J. et al. Association of genetic risk score with NAFLD in an ethnically diverse cohort. Hepatol. Commun. 5(10), 1689–1703 (2021).
Mu, T. et al. Single nucleotide polymorphism of genes associated with metabolic fatty liver disease. J. Oncol. 2022, 9282557 (2022).
Moon, J. H., Kim, W., Koo, B. K. & Cho, N. H. Metabolic dysfunction-associated fatty liver disease predicts long-term mortality and cardiovascular disease. Gut Liver 16(3), 433–442 (2022).
Korean Association for the Study of the L: KASL clinical practice guidelines: management of nonalcoholic fatty liver disease. Clin. Mol. Hepatol. 19(4), 325–348 (2013).
Chalasani, N., Younossi, Z., Lavine, JE., Diehl, A.M., Brunt, E.M., Cusi, K., Charlton, M., Sanyal, A.J., American Gastroenterological A, American Association for the Study of Liver D et al. The diagnosis and management of non-alcoholic fatty liver disease: practice guideline by the American Gastroenterological Association, American Association for the Study of Liver Diseases, and American College of Gastroenterology. Gastroenterology, 142(7), 1592–1609 (2012).
Schwenzer, N. F. et al. Non-invasive assessment and quantification of liver steatosis by ultrasound, computed tomography and magnetic resonance. J. Hepatol. 51(3), 433–445 (2009).
Rinella, M. E. et al. NAFLD Nomenclature consensus group. A multi-society Delphi consensus statement on new fatty liver disease nomenclature. Hepatology 78, 1966–1986 (2023).
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
G.E.C. designed the study. E.K.C., M.-S.K., J.I.Y., S.-W.O. and J.Y.Y. collected data. Y.L. performed analysis. E.J.C., Y.L. and G.E.C. wrote the main manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The original version of this Article contained an error in affiliation 4. In addition, an affiliation was omitted for Seung-Won Oh.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, Y., Cho, E.J., Choe, E.K. et al. Genome-wide association study of metabolic dysfunction-associated fatty liver disease in a Korean population. Sci Rep 14, 9753 (2024). https://doi.org/10.1038/s41598-024-60152-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-60152-0
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.