Abstract
Antimicrobial resistance (AMR) is one of the major challenges of the century and should be addressed with a One Health approach. This study aimed to develop a tool that can provide a better understanding of AMR patterns and improve management practices in swine production systems to reduce its spread between farms. We generated similarity networks based on the phenotypic AMR pattern for each farm with information on important bacterial pathogens for swine farming based on the Euclidean distance. We included seven pathogens: Actinobacillus suis, Bordetella bronchiseptica, Escherichia coli, Glaesserella parasuis, Pasteurella multocida, Salmonella spp., and Streptococcus suis; and up to seventeen antibiotics from ten classes. A threshold criterion was developed to reduce the density of the networks and generate communities based on their AMR profiles. A total of 479 farms were included in the study although not all bacteria information was available on each farm. We observed significant differences in the morphology, number of nodes and characteristics of pathogen networks, as well as in the number of communities and susceptibility profiles of the pathogens to different antimicrobial drugs. The methodology presented here could be a useful tool to improve health management, biosecurity measures and prioritize interventions to reduce AMR spread in swine farming.
Similar content being viewed by others
Introduction
Antimicrobial resistance (AMR) refers to the ability of bacteria, viruses, fungi, and parasites to grow and reproduce in the presence of a given antimicrobial1. Antimicrobial resistance in bacteria is a great health and economic concern and one of the biggest challenges of the twenty-first century as demonstrated by the Global Action Plan on Antimicrobial Resistance supported by the World Health Organization (WHO), World Organization for Animal Health (WOAH), and the Food and Agriculture Organization of the United Nations (FAO)2,3. From the human perspective, 700,000 deaths every year are attributed to AMR, and only in the United States of America, at least 35,900 deaths were accredited to antibiotic-resistant bacteria and fungi each year4,5. From the animal perspective, loss of antimicrobial effectiveness in livestock leads to a reduction in animal welfare and production indices, as well as an increase in production and treatment costs6,7. This can lead to an inevitable rise in the final prices of animal products that has a negative impact on their affordability for the general public.
AMR is a complex problem, impacting human, animal and environmental health since resistance genes can disseminate to different habitats. Hence, it needs to be addressed using holistic, multi-scale methods (i.e., bacteria-, animal-, farm-, system-level) and with a One Health approach1,8,9. In this study, we focused on AMR in bacteria causing disease in the swine industry. Bacteria can be resistant to an antimicrobial because of an intrinsic characteristic of the species (innate resistance), or the resistance could be acquired. It is widely accepted that one main driver of acquired resistance is the selective pressure derived from antimicrobial usage, although the AMR problem is a complex one and many factors are involved. Excessive use and misuse of antibiotics either in humans or livestock could increase AMR in bacteria2,7. Swine production is generally characterized by being an intensive production system with high densities of animals, therefore, infectious diseases are more challenging to control even under high biosecurity conditions in case of presence or introduction6.
Here, we designed a framework to detect and assess similarities in phenotypic antimicrobial resistance status of farms for different pathogens based on the generation of monoplex (or single-layer) and multiplex (or multilayer) similarity networks. The overarching aim of this work is to generate a tool that could help improve management and clinical practices in large production systems to reduce the impact of AMR and minimize its spread within and between farms. In that sense, we propose some theoretical interventions for when complete data is available.
Methods
Data
Data from a large swine production system in the US was obtained with information on 710 animal holdings that included sow farms and finishers, although there was no information about the type of farm production for all of them. Sensitive information, such as farm names, owners, and flow was anonymized to preserve confidentiality. At each farm, we gathered information about the isolated bacterial species tested (henceforth pathogen/s). For this study, we used a total of seven pathogens: Actinobacillus suis, Bordetella bronchiseptica, Escherichia coli, Glaesserella parasuis, Pasteurella multocida, Salmonella spp., and Streptococcus suis. For each pathogen, we obtained the panel of antimicrobials tested by the laboratories, with its corresponding minimum inhibitory concentrations (MIC) values for the phenotypic AMR. The interpretation of the MIC value into “susceptible”, “intermediate” or “resistant” categories were primarily based on the clinical breakpoints provided by the Clinical and Laboratory Standards Institute´s (CLSI) veterinary guidelines, which are based on worldwide expert consensus10. When clinical breakpoints were not available by the CLSI guidelines, we used the harmonized laboratory criterion which is based on their professional experience. When interpretation for the results was missing for a given pathogen, the antimicrobial was excluded for further analysis. Category data were binarized considering resistant and intermediate as a resistant status (i.e. “1”) and susceptible as “0”. Antibiotics that were tested in less than half of the farms for a given pathogen were also excluded for subsequent analysis (Supplementary Fig. S1). In general, ten types of antibiotic classes were tested for phenotypic resistance: penicillins (ampicillin and penicillin), lincosamides (clindamycin), aminoglycosides (gentamicin, neomycin), amphenicols (florfenicol), sulfonamides (sulphadimethoxine, trimethoprim/sulphamethoxazole), fluoroquinolones (enrofloxacin), cephalosporins (ceftiofur), pleuromutilin (tiamulin), macrolides (tilmicosin, tylosin tartrate, tulathromycin), and tetracyclines (tetracycline, chlortetracycline, oxytetracycline).
The date of the reception by the laboratory was used for temporal assignment. Data ranged from the last quarter of 2018 to the end of 2021. For a given pathogen, the majority of farms had only information about one isolate for the entire study period, hence not enough information was available to conduct the analyses by year.
Generation of monoplex networks
Monoplex networks, also known as single-layer networks, can be mathematically defined as a graph G = (V, E), where |V| is the vertex of the network, and |E| the edges or links representing the connections between nodes11,12. In this case, the nodes represented the farms where the phenotypic resistance for a set of antimicrobials of a given pathogen was tested. Edges were defined as the similarity between two farms based on the resistance pattern of a pathogen. The resistance pattern of a bacteria in a farm F can be defined as a vector: F = [f1, f2, …, fn], where f is the susceptibility status of the farm to a given antimicrobial, and n is the number of tested antimicrobials for a given pathogen. From this resistance pattern, we obtained the overall resistance status of a farm F: RSF (Eq. 1). For farms with more than one isolate per pathogen, the mean susceptibility was used in the analysis.
However, for a given pathogen, the resistance pattern was defined as a matrix: P = (m x n), where m is the number of farms tested for AMR for a given pathogen, and n the number of tested antimicrobials. To obtain the similarity of resistance patterns between farms, we first calculated the pairwise Euclidean distance13 (Eq. 2). The distance measure was subsequently scaled to a maximum of 1 by dividing by the maximum value, and the similarity was obtained by subtracting the distance from 1. This similarity ranged from 0 to 1, being 1 identical resistance pattern, and 0 complete dissimilarity between two given vector farms (F and G).
To overcome the problem of missing data in the resistance pattern of farms and similarity measures, we used missing data imputation14. Missing values for the resistance to an antibiotic were replaced by the average of the available information in other farms as a proxy of the probability of resistance in a farm with missing information.
Given the definition of the edges based on similarity, monoplex networks were fully connected by edges with different weights. The application of a cut-off value or threshold is a common practice to reduce the density of networks and find node clusters15. Thus, we selected a threshold (th) and applied it to the adjacency matrix (A = (aij)); values in the matrix below a given threshold were removed and a new unweighted matrix (B) was generated (Eq. 3).
Given the possible differences in the similarity between farm AMR patterns, applying the same value for th would not be appropriate for each pathogen. Therefore, we assumed that the network given th should retain a sufficient number of edges, so there was no connected component larger than 3% of Fn disconnected from the main connected network. We iterated through the number of connected components based on the defined criterion and chose the highest value of th with only one connected component. This criterion allowed us to obtain centrality measures based on average path length, and to identify hub nodes connecting different communities16. The resultant network was undirected and unweighted.
For each node in the monoplex network for the included pathogens, degree, eigenvector and betweenness centrality measures were computed. These centrality measures are commonly used in network analysis and are thoroughly reviewed elsewhere17. Briefly, in an undirected network, the degree centrality for a node is the number of edges that connect a node with other nodes. The eigenvector centrality is based on the number of connections of a given node, as does the degree, but also takes into account the importance of its neighbors; connections to high-scoring nodes contribute more to the score of the given node12. Therefore, in the context of undirected similarity networks, a high degree or eigenvector score point to nodes that possess high similarity with other nodes in the network. Betweenness centrality quantifies the number of shortest paths between all node pairs passing through that node18.
Another important aspect to consider is the identification of communities or groups of nodes in the network and the intrinsic characteristics that differentiate them from the others. Here, we adopted the understanding of the term “community” as a subset of nodes in a network with more connections among themselves than with the rest of the nodes15,19. We used the Louvain algorithm to identify communities in the pathogen’s monoplex networks20. For each community, we computed the antimicrobial resistance profile by averaging the given resistance pattern of every farm in the community. Identifying communities in those networks is essential for identifying and analyzing differences in their antimicrobial resistance profiles.
Differences in the antimicrobial resistance profiles and the identification of hubs in the network based on centrality measures provide epidemiologic context for management in specific farms and suggest flows of pigs that may minimize AMR spread. The process mentioned above was developed in R version 3.6.321, using the following packages: “igraph”22, “tidyverse”23, “mice”24, and “reshape2”25.
Generation of multiplex networks
Similar to monoplex networks, multilayer networks are composed of nodes and edges, but they also add layers to their structure. Multiplex networks, also known as edge-colored networks, are a special type of multilayer network, in which the nodes are the same or similar in each layer, and are connected to their counterparts on other layers by coupling edges. As in monoplex networks, intra-layer edges connect nodes in the same layer11,26. Multiplex networks are valuable for studying different interactions and relationships between nodes27. In our study, the goal of using a multiplex network was to identify similarities among farms with information for different pathogens. Each layer would be every pathogen’s monoplex network included in the study. To assess how many nodes overlapped between layers, we compute the pairwise Jaccard similarity between nodes of the layers19,28. This index is equal to the number of common elements (intersection) of two sets of nodes (Li, Lj) divided by the total size of the compared sets29 (Eq. 4). The analysis was performed using the “multinet” R package30.
Results
Monoplex networks
Due to the nature of the dataset, not all the tested antimicrobials were included in the laboratory panel or had consistent susceptibility interpretation criteria for each pathogen (Supplementary Fig. S1). Therefore, the size of the vector F varied, ranging between 12 for E. coli and 15 for B. bronchiseptica and P. multocida.
Overall missing information about the AMR status (susceptible or resistant) for antibiotics in the panel of a pathogen varied from 1.77% to 11.58%. However, missing information was concentrated in only some antimicrobials, varying from one for E. coli (tetracycline) to four for Salmonella spp. (Table 1). The majority of missing data was due to the lack of inclusion of an antimicrobial when reporting the results.
We found significant differences in terms of the overall resistance status of the farms (RSF) and the percentage of resistance between pathogens (Fig. 1). The range of RSF is determined by the number of included antimicrobials from the pathogen’s panel in the study due to the inclusion criteria. Since the total number of studied antimicrobials, although similar, was not the same for every pathogen, we rescaled the overall resistance status values to a maximum of 1, dividing by the maximal RSF possible value for each pathogen. The small variation in the number of tested antimicrobials allowed us to easily compare them once rescaled. E. coli and Salmonella spp. isolates presented the higher AMR overall resistance while A. suis and G. parasuis’ isolates the lowest, closely followed by P. multocida.
Boxplots showing the comparison of the rescaled overall resistance status for the farms (RSF) and the percentage of resistance by farm for the pathogens included in the study. The RSF was calculated using Eq. (1) and rescaled to a maximum of 1 by dividing it by the maximal RSF possible value for each pathogen. Figure generated using the “tidyverse” package in R v3.6.3.
The number of farms with a resistance pattern for a given pathogen varied, as did the threshold ranging between 0.7 and 0.8 (Table 1, Supplementary Fig. S2). Densities of the resultant networks are gathered in Table 1. They varied significantly for some pathogens, being as high as 0.375 for B. bronchiseptica and as low as 0.045 for E. coli. Differences in the morphology and density of the networks for the different pathogens studied here can be observed (Fig. 2). Thus, the number of identified communities by the Louvain algorithm also varied from highly dense networks to less dense ones. Single-member communities were considered as isolates. Networks for A. suis and B. bronchiseptica had a high density, meaning that the isolates tested in the farms had extremely similar patterns of resistance, and therefore fewer communities were identified. AMR profiles were similar among communities for these pathogens (Fig. 3). In both cases, more than 48% of the farms belonged to a single community. E. coli and Salmonella spp. are pathogens with higher overall resistance in this study (Fig. 1), and that is also reflected in their community profiles (Fig. 3).
Several centrality measures were also computed, namely degree, eigenvector and betweenness centralities (Fig. 4). These measures allowed us to identify nodes that are hubs in the network17. Higher values for degree centrality showed farms that have more similarity to other farms, while the eigenvector showed farms that are connected to other well-connected farms. Higher values of betweenness showed hub farms that connect different communities, and therefore could be individually analyzed for their implication in AMR spread. It can be seen how denser networks, such as the ones for A. suis and B. bronchiseptica, have in general greater degree and eigenvector values and fewer betweenness hubs.
Multiplex networks
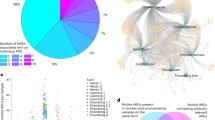
We generated a multiplex network that was comprised of 479 nodes, although not all the farms had information about every pathogen (Fig. 5). Many of them had information about only one pathogen (Fig. 5B). According to the Jaccard similarity, the overlapping of nodes between layers is low, being inferior to 0.55, and it is the lowest for A. suis with the rest of the pathogen layers (Fig. 5A).
Discussion
In this study, we propose a novel approach of using similarity networks and community detection to generate AMR profiles as a quick and easy visualization tool to support decision-making and improve antimicrobial stewardship for swine production systems. The pathogens studied in this work are etiological agents of important systemic, enteric, and respiratory diseases in swine, which create a significant impact on the health and well-being of pigs, as well as an important economic loss for the swine industry. Furthermore, these agents are primary drivers of antimicrobial use on farms. There is also a public health concern about some of these agents, such as E. coli, Pasteurella multocida, Salmonella spp., S. suis and B. bronchiseptica, due to their zoonotic nature32. Additional information on their epidemiology, pathogenesis and control can be found elsewhere32.
Here, farms have been grouped in terms of their phenotypic AMR similarity. It is worth noting that in vitro characterization of the susceptibility of an isolate does not necessarily imply the success or failure of the in vivo clinical outcome33. There are some uncertainties of clinical breakpoints in veterinary medicine for many antimicrobials since they are host-specific and even vary for different target organs34,35. In this study, we have used the currently approved approach for assigning the phenotypic AMR status based on CLSI veterinary guidelines and laboratory expertise that are therapeutically relevant10. Moreover, these clinical breakpoints were used to show the AMR profile of the bacterial population of the farm and are epidemiological relevant for the current study regardless of the sampled organ. The phenotypic AMR similarity, along with the AMR profile of the identified communities, can help improve the therapeutic management of farms, as well as guide animal movements between them to reduce the spread of resistant clones. For example, in the case of S. suis, the most frequent antimicrobial treatments are beta-lactams and fluoroquinolones36, so resistance to one of these antimicrobials would be highly relevant for infection management, and its spread could be mitigated by large swine production systems and veterinarian practitioners with the information provided by the methodology proposed here. In the S. suis community profile (Fig. 3), it can be seen that the major communities are highly susceptible to those antimicrobials. Still, if susceptibility to one of them decreases over time, specific and localized management measures could be taken to hamper the spread of resistance.
For some farms, only information about the susceptibility profile of one isolate was available. Thus, farm patterns should not be considered as the AMR frequency of the entire farm, since only one isolate per farm was used for the generation of the network in those cases. Therefore, for those farms, the information presented here should be considered as a relative risk of resistance and not as the overall resistance of the farm. In addition, there might be a possible bias in the data collection since, in some cases, sampling was clinically based and did not proceed from random sampling. This sampling criteria along with the consideration of the AMR’s categories of “resistance” and “intermediate” status as resistant isolates, might increase the percentage of resistance found for each pathogen for some antimicrobials that will be discussed below. For future studies and to minimize the former limitation, sampling protocols could be implemented. This protocol should include how to calculate sample size so it is representative of the farm’s population in terms of age, sex and number of animals, as well as to obtain an appropriate estimate of the prevalence of AMR37,38,39. Sampling should be carried out preferably randomly in healthy animals whenever possible, although the inclusion of samples from sick animals could be accepted in an admissible proportion in the shake of cost-effectiveness. Moreover, guidelines can be provided as to the target organs and type of sample admissible to ensure comparable results37,38. Even considering these limitations, the methodology is valuable from a practical point of view for identifying similarity patterns between farms affected by important pathogens of animal and public health concern.
One example of applicability is minimizing animal flows from farms belonging to other communities whenever possible; and even more so from communities with higher resistance status or where a pathogen is resistant to a therapeutically crucial antimicrobial. This could be even more beneficial when resistance genes are evaluated. The animal source may be outside the network, whether a farm that has not been tested or a farm for which no information is available for a given pathogen. In that case, an analysis of the AMR profiles of recipient farms can be conducted to find similarities between them, which could include genetic factors as well. Unfortunately, we could not assess shared features between farms, such as the geographical location and type of farm, flow or origin of pigs, farm size, treatment, or biosecurity protocols, among other factors. For example, the type of farm could be associated with different management practices7. This information would be highly beneficial to identify whether there are common features that might be contributing to increasing AMR, facilitating the benchmarking of farms and supporting the implementation of mitigation strategies. Furthermore, having information about the community’s profile can enhance the application of an effective clinical treatment in the early stages of an outbreak, and, therefore, better inform antimicrobial use and avoid the excessive use or misuse of antibiotics that could contribute to AMR. Therefore, this tool should be considered as a preliminary approach to finding meaningful associations between antimicrobial patterns of farms that can help tackle the AMR problem.
Regarding methodological aspects, missing values are a frequent problem in any analysis14. The primary source of missing data was for tetracyclines, since some laboratories reported only tetracycline, while others included chlortetracycline and oxytetracycline, but not the three of them combined. Hence, information about the resistance profile of certain antimicrobials should be evaluated taking into consideration the need for imputation for certain data. However, the imputation of missing data allows the use and analysis of valuable partial information that would not be included otherwise14. In this study, we kept phenotypic AMR data in the farm’s antimicrobial pattern even with < 50% of missing information for some pathogens, so that we could compare such resistance profiles with other pathogens, and also as a way to highlight the importance of having complete information about the same AMR panel. Community profiles for A. suis and P. multocida are a prime example of the importance of keeping phenotypic AMR for an antimicrobial with missing information, since for several of them, significant differences in community profiles could be revealed. In both cases, we identified communities that were highly susceptible to chlortetracycline: community 2 for A. suis and communities 3 and 6 for P. multocida. Something similar occurred with S. suis network and clindamycin susceptibility for the different communities. Community 25 had 1% resistant bacteria while the rest of the large communities exceeded 80% of resistance.
The reason for using the average of the available information of other farms for the same pathogen is that it shows the probability of resistance in a farm with missing information. Although it is a limitation of the study, we believe this is a simple but effective approach for this situation because it maintains the characteristics of the dataset and prevents the loss of data. Moreover, other imputation methods, such as predictive mean matching would assign a resistant or susceptible status24, directly distorting the similarity networks by potentially grouping farms with missing data with resistant or susceptible groups, when their real status is unknown for a given antimicrobial.
The use of a threshold is essential when working with fully connected networks to reduce their density and properly analyze meaningful node relationships. In many studies facing the same issue, researchers defined this threshold by trial and error40, although some threshold selection methods have been described in the literature41. We have developed a general threshold selection method that allows us to automatically create a similarity network that retains valuable information about path-length centrality measures, such as betweenness. As our results showed (Table 1), giving the same threshold for every pathogen would be arbitrary and unappropriated due to the specific intrinsic characteristics of each phenotypic AMR similarity network.
The multiplex network could not be adequately generated due to the lack of critical farm’ information about the AMR patterns for some pathogens (Fig. 5). Considering the small overlapping of nodes observed between layers, centrality measures would have just highlighted farms that appeared in more layers, and not necessarily the most important ones. A multiplex approach could be applied when more AMR data is available in all or at least in most farms. This would allow for the identification of farms that share the same AMR similarity pattern for the different pathogens. Similarity networks can be combined with other networks, such as networks representing the movement of animals between farms or other features of interest. Another application would be the generation of temporal networks to analyze the progress and evolution of relationships of AMR for a particular pathogen in a specified period11. However, temporal data about AMR in each farm is required, and this information was limited in this study (i.e., the range of farms with a single isolate ranged from 56.63% for S. suis to 78.1% for A. suis).
Although not the aim of the study, we discuss below the AMR frequency for the different pathogens and its impact based on the most relevant antimicrobials used for the clinical treatment of each pathogen. It should be noted that AMR frequency among pathogens is difficult to compare between studies due to differences in study design, the target population, as well as resistance criteria and clinical breakpoint35.
Therapeutic options to treat the disease caused by the infection of A. suis include amoxicillin, ampicillin, penicillin, tiamulin, ceftiofur, gentamicin and trimethoprim/sulfadiazine42. Taking this into account, A. suis isolates from the major communities of this study are susceptible to many of the therapeutic tools against this pathogen (Figs. 2 and 3).
Similar to previous reports, almost all isolates of B. bronchiseptica were resistant to beta-lactams (99% here vs. 100%)43, which are commonly used for the treatment of swine respiratory disease44. The resistance to beta-lactams is explained by a species-specific beta-lactamase gene described for the bacteria, as well as low membrane permeability45. Almost all isolates were susceptible to tulathromycin, an antibiotic used for the treatment of swine respiratory disease that comprises Actinobacillus pleuropneumoniae, P. multocida, Mycoplasma hyopneumoniae, G. parasuis and B. bronchiseptica46. B. bronchiseptica isolates were largely susceptible to enrofloxacin44, although community 3 was phenotypically resistant. Differences in florfenicol susceptibility have been described in several studies ranging from 10 to almost 98%45. In accordance with those results, we observed significant differences between communities (Fig. 3).
P. multocida can be found co-infecting with B. bronchiseptica in the swine upper respiratory tract, so phenotypic susceptibility of both pathogens should be taken into account in the antimicrobial treatment44,47. P. multocida isolates presented more overall susceptibility to the tested antimicrobials than other pathogens (Fig. 1), although nearly all isolates were phenotypically resistant to clindamycin and tylosin-tartrate. This elevated susceptibility status for most isolates is in agreement with the majority of published work35. Drugs commonly used to treat the disease caused by this bacteria are ampicillin, ceftiofur, enrofloxacin, and tulathromycin47, for which the majority of isolates were susceptible (Fig. 3).
There are no international criteria for G. parasuis clinical breakpoints, so the comparison between studies is challenging35. In Dayao et al., isolates of G. parasuis had higher MIC values for ampicillin, penicillin and tetracycline48. Similarly, in this study, the majority of large communities showed elevated resistance to penicillin.
The phenotypic resistance results obtained here for S. suis are in agreement with other studies. In general, low resistance was observed for beta-lactams, and high resistance was observed for tetracyclines35,36,49. In several countries, high levels of resistance to macrolides and lincosamides have been reported49, although, in this study, some communities were highly susceptible to these drugs. Moreover, S. suis was highly susceptible to florfenicol in the study farms, an observation also reported in European and North American countries35.
Escherichia coli was recently considered, alongside Brachyspira hyodysenteriae, to be the most critical antimicrobial-resistant bacteria in the European Union for swine35. In a temporal study conducted in the USA, they found high levels of resistance to tetracyclines and ampicillin, as well as moderate levels of resistance to gentamicin, neomycin, and sulfonamides50. Similar to the present study, Jiang et al. reported elevated AMR in E. coli isolates from the intestinal contents or fecal samples of diarrheic piglets from 15 states of the United States, where all isolates were found to be resistant to clindamycin, penicillin, tiamulin, and tilmicosin. In addition, they observed high resistance to ampicillin, chlortetracycline, oxytetracycline and sulphadimethoxine51. It is also worth noting that Gram-negative bacteria, for instance, E. coli, possess relative intrinsic resistance to some antimicrobial classes such as macrolides52, which is consistent with our data.
In vitro resistance to multiple drugs is often observed for species of the genus Salmonella. Therefore, the assessment of AMR prior to the onset of a therapeutic solution is essential53. Multidrug resistance bacteria constitute a problem for effective clinical treatment of infections. Therefore, for pathogens that have developed resistance to several drugs, knowing the historical resistance of a population can serve as a useful guide to assessing therapeutic options in the early stages of an outbreak53,54.
Given the importance of the AMR problem, developing new strategies is paramount to properly address and mitigate it in livestock husbandry in the context of the One Health approach. The methodology, analysis, and visualization presented here are highly relevant, not only to guide more effective clinical actions in a given farm, but to enhance antimicrobial stewardship and decrease the spread of resistance of key antimicrobials across farms, as exemplified in the discussion. This approach can be easily expanded and enhanced to incorporate animal, farm and system features to better understand their impact on AMR patterns and further support the implementation of preventive measures to reduce AMR spread. We have also highlighted the potential key role of high-betweenness farms in the different networks that connect communities. The characteristics of these farms should be analyzed in depth for their implication in AMR spread.
Data availability
The datasets analyzed during the current study are not publicly available due to confidentiality reasons and restrictions on the availability of these data, but are available from the corresponding author on reasonable request and with permission of the data provider.
References
Bright-Ponte, S. J. et al. One Health and antimicrobial resistance, a United States perspective. Rev. Sci. Tech. 38, 173–184. https://doi.org/10.20506/rst.38.1.2951 (2019).
OIE. The OIE Strategy on Antimicrobial Resistance and the Prudent Use of Antimicrobials, https://www.oie.int/app/uploads/2021/03/en-oie-amrstrategy.pdf (2016).
WHO. Global Action Plan On Antimicrobial Resistance, https://ahpsr.who.int/publications/i/item/global-action-plan-on-antimicrobial-resistance (2015)
CDC. Antibiotic Resistance Threats In The United States, https://www.cdc.gov/drugresistance/pdf/threats-report/2019-ar-threats-report-508.pdf (2019).
Shallcross, L. J., Howard, S. J., Fowler, T. & Davies, S. C. Tackling the threat of antimicrobial resistance: From policy to sustainable action. Philos. Trans. R Soc. Lond. B Biol. Sci. 370, 20140082–20140082. https://doi.org/10.1098/rstb.2014.0082 (2015).
Bengtsson, B. & Greko, C. Antibiotic resistance—Consequences for animal health, welfare, and food production. Ups. J. Med. Sci. 119, 96–102. https://doi.org/10.3109/03009734.2014.901445 (2014).
Wall, B. et al. Drivers, dynamics and epidemiology of antimicrobial resistance in animal production (Food and Agriculture Organization of the United Nations, 2016).
Hernando-Amado, S., Coque, T. M., Baquero, F. & Martínez, J. L. Defining and combating antibiotic resistance from One Health and Global Health perspectives. Nat. Microbiol. 4, 1432–1442. https://doi.org/10.1038/s41564-019-0503-9 (2019).
Pehrsson, E. C. et al. Interconnected microbiomes and resistomes in low-income human habitats. Nature 533, 212–216. https://doi.org/10.1038/nature17672 (2016).
CLSI. Performance Standards for Antimicrobial Disk and Dilution Susceptibility Tests for Bacteria Isolated From Animals. 5th edn, (Clinical and Laboratory Standards Institute, 2020).
Kivelä, M. et al. Multilayer networks. J. Complex Netw. 2, 203–271. https://doi.org/10.1093/comnet/cnu016 (2014).
Ruhnau, B. Eigenvector-centrality—A node-centrality?. Soc. Netw. 22, 357–365. https://doi.org/10.1016/S0378-8733(00)00031-9 (2000).
Tabak, J. Differential geometry in Geometry: The Language of Space and Form 248 (Facts On File, Inc, 2014).
Graham, J. W., Cumsille, P. E. & Shevock, A. E. Methods for Handling Missing Data. in Handbook of Psychology: Research methods in psychology (ed. Schinka, J. A., W. F. Velicer, & I. B. Weiner) 109–141 (John Wiley & Sons, Inc, 2013).
Fornito, A., Zalesky, A. & Bullmore, E. Fundamentals of brain network analysis (Elsevier Academic Press, 2016).
Valavanis, I., Spyrou, G. & Nikita, K. A similarity network approach for the analysis and comparison of protein sequence/structure sets. J. Biomed. Inform. 43, 257–267. https://doi.org/10.1016/j.jbi.2010.01.005 (2010).
Martínez-López, B., Perez, A. M. & Sánchez-Vizcaíno, J. M. Social network analysis. Review of general concepts and use in preventive veterinary medicine. Transbound. Emerg. Dis. 56, 109–120. https://doi.org/10.1111/j.1865-1682.2009.01073.x (2009).
Freeman, L. C. A set of measures of centrality based on betweenness. Sociometry 40, 35–41. https://doi.org/10.2307/3033543 (1977).
Magnani, M., Rossi, L. & Vega, D. Analysis of multiplex social networks with R. J. Stat. Softw. https://doi.org/10.18637/jss.v098.i08 (2021).
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. Theory Exp. 2008, P10008. https://doi.org/10.1088/1742-5468/2008/10/p10008 (2008).
R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria (2020).
Csardi, G. & Nepusz, T. The igraph software package for complex network research, https://igraph.org. (2006).
Wickham, H. et al. Welcome to the Tidyverse. J. Open Source Softw. 4, 1686. https://doi.org/10.21105/joss.01686 (2019).
van Buuren, S. & Groothuis-Oudshoorn, K. mice: Multivariate imputation by chained equations in R. J. Stat. Softw. 45, 1–67. https://doi.org/10.18637/jss.v045.i03 (2011).
Wickham, H. Reshaping data with the reshape package. J. Stat. Softw. 21, 1–20. https://doi.org/10.18637/jss.v021.i12 (2007).
De Domenico, M. et al. Mathematical formulation of multilayer networks. Phys. Rev. X 3, 041022. https://doi.org/10.1103/PhysRevX.3.041022 (2013).
Finn, K. R., Silk, M. J., Porter, M. A. & Pinter-Wollman, N. The use of multilayer network analysis in animal behaviour. Anim. Behav. 149, 7–22. https://doi.org/10.1016/j.anbehav.2018.12.016 (2019).
Bródka, P., Chmiel, A., Magnani, M. & Ragozini, G. Quantifying layer similarity in multiplex networks: A systematic study. R. Soc. Open. Sci. 5, 171747. https://doi.org/10.1098/rsos.171747 (2018).
Jaccard, P. The distribution of the flora of the alpine zone. New Phytol. 11, 37–50. https://doi.org/10.1111/j.1469-8137.1912.tb05611.x (1912).
Magnani, M., Rossi, L., Hanteer, O., Vega, D. & Dubik, M. multinet: Analysis and Mining of Multilayer Social Networks, https://CRAN.R-project.org/package=multinet (2021).
Bastian, M., Heymann, S. & Jacomy, M. Gephi: an open source software for exploring and manipulating networks. in 3rd International AAAI Conference on Weblogs and Social Media. (2009).
Zimmerman, J. et al. Diseases of Swine 11th edn. (Wiley, 2019).
Leekha, S., Terrell, C. L. & Edson, R. S. General principles of antimicrobial therapy. Mayo Clin. Proc. 86, 156–167. https://doi.org/10.4065/mcp.2010.0639 (2011).
Aarestrup, F. M., Oliver Duran, C. & Burch, D. G. Antimicrobial resistance in swine production. Anim. Health Res. Rev. 9, 135–148. https://doi.org/10.1017/s1466252308001503 (2008).
EFSA Panel on Animal Health and Welfare (AHAW) et al. Scientific Opinion on the assessment of animal diseases caused by bacteria resistant to antimicrobials: Swine. EFSA J. 19, 7114 (2021).
Seitz, M., Valentin-Weigand, P. & Willenborg, J. Use of antibiotics and antimicrobial resistance in veterinary medicine as exemplified by the swine pathogen Streptococcus suis. Curr. Top Microbiol. Immunol. 398, 103–121. https://doi.org/10.1007/82_2016_506 (2016).
WHO. Integrated surveillance of antimicrobial resistance in foodborne bacteria: application of a one health approach: guidance from the WHO Advisory Group on Integrated Surveillanec of Antimicrobial Resistance (AGISAR). 88 p. (World Health Organization, 2017).
WOA H. Chapter 6.7: Harmonisation of national antimicrobial resistance surveillance and monitoring programmes. in Terrestrial Animal Health Code (ed World Organisation for Animal Health) (2022).
Cochran, W. G. Sampling Techniques 3d edn. (Wiley, 1977).
Catanese, H. N., Brayton, K. A. & Gebremedhin, A. H. A nearest-neighbors network model for sequence data reveals new insight into genotype distribution of a pathogen. BMC Bioinform. 19, 475. https://doi.org/10.1186/s12859-018-2453-2 (2018).
Apeltsin, L., Morris, J. H., Babbitt, P. C. & Ferrin, T. E. Improving the quality of protein similarity network clustering algorithms using the network edge weight distribution. Bioinformatics 27, 326–333 (2011).
Gottschalk, M. & Broes, A. Actinobacillosis. in Diseases of Swine (ed. Zimmerman, J. J. et al.) 749–766 (2019).
Dayao, D. A., Gibson, J. S., Blackall, P. J. & Turni, C. Antimicrobial resistance in bacteria associated with porcine respiratory disease in Australia. Vet. Microbiol. 171, 232–235. https://doi.org/10.1016/j.vetmic.2014.03.014 (2014).
Brockmeier, S. L., Register, K. B., Nicholson, T. L. & Loving, C. L. Bordetellosis. in Diseases of Swine (ed. Zimmerman, J. J. et al.) 767–777 (2019).
Kadlec, K. & Schwarz, S. Antimicrobial Resistance in Bordetella bronchiseptica. Microbiol. Spectr. https://doi.org/10.1128/microbiolspec.ARBA-0024-2017 (2018).
EMA. EPAR summary for the public: Draxxin, https://www.ema.europa.eu/en/documents/overview/draxxin-epar-summary-public_en.pdf (2016).
Register, K. B. & Brockmeier, S. L. Pasteurellosis. in Diseases of Swine (ed. Zimmerman, J. J. et al.) 884–897 (2019).
Dayao, D., Gibson, J. S., Blackall, P. J. & Turni, C. Antimicrobial resistance genes in Actinobacillus pleuropneumoniae, Haemophilus parasuis and Pasteurella multocida isolated from Australian pigs. Aust. Vet. J. 94, 227–231. https://doi.org/10.1111/avj.12458 (2016).
Varela, N. P. et al. Antimicrobial resistance and prudent drug use for Streptococcus suis. Anim. Health Res. Rev. 14, 68–77. https://doi.org/10.1017/s1466252313000029 (2013).
Malik, Y. S., Chander, Y., Olsen, K. & Goyal, S. M. Antimicrobial resistance in enteric pathogens isolated from Minnesota pigs from 1995 to 2004. Can. J. Vet. Res. 75, 117–121 (2011).
Jiang, F. et al. Genotypes and antimicrobial susceptibility profiles of hemolytic escherichia coli from diarrheic piglets. Foodborne Pathog. Dis. 16, 94–103. https://doi.org/10.1089/fpd.2018.2480 (2019).
Cox, G. & Wright, G. D. Intrinsic antibiotic resistance: Mechanisms, origins, challenges and solutions. Int. J. Med. Microbiol. 303, 287–292. https://doi.org/10.1016/j.ijmm.2013.02.009 (2013).
Griffith, R. W., Carlson, S. A. & Krull, A. C. Salmonellosis. in Diseases of Swine (ed. Zimmerman, J. J. et al.) 912–925 (2019).
Fairbrother, J. M. & Nadeau, É. Colibacillosis. in Diseases of Swine (ed. Zimmerman, J. J. et al.) 807–834 (2019).
Acknowledgements
Authors would like to thank the contributing swine production system for sharing their valuable antimicrobial susceptibility data. This project was supported by NSF-BIGDATA Award #: 1838207, NSF-Convergence Accelerator Phase I Award #: 2040680 and Phase II Award #: 2134901 and USDA-NIFA Award #: 2021-68014-34143. CAV was the recipient of PhD Fellowship for the Training of Future Scholars (FPU) as well as a short-term scholarship under the FPU program, both awarded by the Spanish Ministry of Science, Innovation and Universities.
This work was completed while Caterina Scoglio was visiting CADMS during her sabbatical year.
Author information
Authors and Affiliations
Contributions
C.A.V., C.S. and B.M.L. conceived the studies, C.A.V. developed and performed the analyses, C.A.V., C.S., B.M.L., X.L., M.J.C., R.R. and L.K. reviewed the methods and analyzed the results, C.A.V. and C.S. made the figures, C.A.V. wrote the manuscript, which all authors reviewed, edited, and approved.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Aguilar-Vega, C., Scoglio, C., Clavijo, M.J. et al. A tool to enhance antimicrobial stewardship using similarity networks to identify antimicrobial resistance patterns across farms. Sci Rep 13, 2931 (2023). https://doi.org/10.1038/s41598-023-29980-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-29980-4
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.