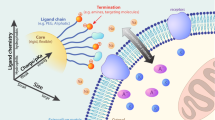
Abstract
Nanotechnology brings benefits in fields such as biomedicine but nanoparticles (NPs) may also have adverse health effects. The effects of surface-modified NPs at the cellular level have major implications for both medicine and toxicology. Semi-empirical and mechanism-based models aid to understand the cellular transport of various NPs and its implications for quantitatively biological exposure while avoiding large-scale experiments. We hypothesized relationships between NPs-cellular elimination, surface functionality and elimination pathways by cells. Surface free energy components were used to characterize the transport of NPs onto membranes and with lipid vesicles, covering both influences by size and hydrophobicity of NPs. The model was built based on properties of neutral NPs and cells, defining Van de Waals forces, electrostatic forces and Lewis acid–base (polar) interactions between NPs and vesicles as well as between vesicles and cell membranes. We yielded a generic model for estimating exocytosis rate constants of various neutral NPs by cells based on the vesicle-transported exocytosis pathways. Our results indicate that most models are well fitted (R2 ranging from 0.61 to 0.98) and may provide good predictions of exocytosis rate constants for NPs with differing surface functionalities (prediction errors are within 2 times for macrophages). Exocytosis rates differ between cancerous cells with metastatic potential and non-cancerous cells. Our model provides a reference for cellular elimination of NPs, and intends for medical applications and risk assessment.
Similar content being viewed by others
Introduction
Nanoparticles (NPs) have been extensively studied due to their artificial functionalization for biomedical development. Mediated by size and surface charge, NPs carry molecules across cell membranes to reach target organs1, cells or organelles. Size, surface charge and hydrophobicity have effect on the rates of transmembrane (cell membrane, mitochondrial membrane, etc.) transport efficiency2,3. Hydrophobicity of NPs affects bioavailability4 of NPs (hence, drugs) and increase the half-life of NPs circulating in the body by avoiding recognition by the reticuloendothelial system (RES)5. Quantifying the effects of NP surface properties on cellular elimination is of critical importance in biomedicine. NPs must be non-toxic and biocompatible to be used in biomedical applications6. However, the multiple pathways of NPs exposure to humans and low targeting efficiency easily lead to unexpected cytotoxicity in vitro and in vivo7,8. Quantifying the transport of NPs with different properties in cells can provide a reference for the assessment of toxicological risks9 of NPs.
Probabilistic methods, such as artificial neural network (ANN)10 methods, can predict dynamic equilibria cellular uptake and exocytosis rate constants of NPs disregarding mechanisms. Mechanistic models11,12 focus on known transport pathways of NPs transportation in organisms and can be extrapolated to predictions of other NPs but require experimental confirmation. Biological exposure and transportation of NPs relate to factors, such as charge density, surface charge of NPs13, and oil–water partitioning14,15. The octanol–water partition coefficient (Kow) is traditionally used to predict the accumulation/transformation of small organic compounds in environments16 and organisms17. However, NPs interact with bio-membranes in an energy-dependent manner18, preventing NPs to disperse in a thermodynamically stable way19,20. Consequently, the Kow is inadequate to describe biological exposure of NPs. Mechanism-based modelling and appropriate descriptors are needed to describe NP-biological interactions.
Here, we explore exocytosis rate constants of NPs based on cellular transport pathways21. Potential intracellular transport pathways of NPs are summarized into three categories (Fig. 1). First, NPs either encapsulated by vesicles or not, diffuse through the cytoplasm and exit cells through the membranes22,23, i.e., a non-endocytic pathway (e.g. red blood cells24). Second, NPs may be transported to lysosomes, degraded by acid (pH ~ 4–5) or not25,26, and finally be excreted out of cells27. Smaller NPs can even aggregate into a cluster and be transported by a vesicle28. Other pathways, include adhesion to the endoplasmic reticulum29, nucleus30 or golgi apparatus31, and subsequent transported by the aforementioned pathways.
Potential elimination pathways of NPs in a cell. Pathways 1–2 (single NPs) and 3–4 (aggregated NPs) are vesicle-transported exocytosis pathways, 5 and 6 are vesicle-free exocytosis pathways, 7 and 8 are degradation pathways for lysosomes and autophagosomes, respectively. Our modelling is based on vesicle-transported exocytosis pathways for unaggregated NPs (pathways 1 and 2).
NP properties affect their transport pathway and, hence, their exocytosis rates32. According to the conservation law, the apparent (experimental) elimination rate constant (keli) is a combination (summation) of different processes. The relevance of the different pathways relates to the intrinsic rate constants (k) of different pathways and the corresponding NPs concentrations in the respective cellular compartments:
Or
where [NP]vesicle and [NP]total are the concentration of vesicle-transported NPs and total NPs inside of one cell, respectively. [NP]vesicle/[NP]total is the fraction of NPs encapsulated by vesicles. Degradation (including klysosome29 and kautophagosome33) are part of elimination pathways in cells (e.g., dissolution of NPs), depending on enzyme properties34, NP properties, such as coating stability35, surface hydrophobicity36 and charge37, and inner core materials29. It is difficult to qualify the degradation of NPs directly (pathways 7 and 8). Lysosomes ‘modification’38 or aggregation19 of NPs can render the (new) surface charge and structure of NPs uncertain, especially for charged NPs.
Hypothesis/scope
Figure 1 depicts the possible cellular elimination and transport pathways of vesicle-transported NPs and vesicle-free NPs inside or outside cells. By considering inert NPs we minimized the effects of acidification within lysosomes on surface modifications, and thereby built and parameterized a generic exocytosis model based on exocytosis pathways for single NPs (Fig. 1 pathways 1 and 2):
Figure 1 depicts six exocytosis pathways (1–6). It is difficult to define the proportion of vesicle-transported NPs ([NP]vesicle/[NP]total) for our generic modelling due to rather limited data. Previous studies tried to predict transportation of NPs based on the deformation39 or combination40 energies, which relates to NPs being encapsulated by vesicles to cross cell membranes. Correspondingly, deformation energies during cell membranes and vesicles recombination drive exocytosis of NPs41, but predicting probability/frequency of vesicle-transported NPs42 remains cumbersome.
In this paper, we developed a generic model to predict the exocytosis rate constants (kexo) by calculating the interaction between vesicle-transported NPs and cell membranes based on properties of NPs and traits of cells. We considered the fraction or frequency of NPs encapsulated by vesicles to predict exocytosis of NPs by cells. We focused on Van de Waals, electrostatic and Lewis acid–base (polar) forces during the interaction of vesicle-transported NPs with cell membranes. We applied our modeling framework to inert neutral particles to avoid the influence caused by acidic environments in lysosomes. The experimental data of 64 types of NPs and six types of cell lines were used as training and test data to predict the exocytosis rate constants.
Results
Interaction energies between vesicle-transported NPs and cell membranes
Figure S1 depicts the total energies changes over the distance among NPs, vesicles and cells. Table 1 shows all interaction energies between NPs and vesicles (ΔGnp/v) and energy barrier heights between vesicles and membranes (ΔG‡v/m). All the interaction energies and barrier energies are positive values. The ΔGnp/v between different sized vesicle-transported NPs (size from 14 to 100 nm) and vesicles range from 350 to 3400 kJ/mol. The ΔG‡v/m between vesicles (size from 14 to 100 nm) and cell membranes range from 600 to 4600 kJ/mol, which is always higher than the ΔGnp/v between NPs and vesicles, except for citrate- and PEG-coated NPs.
The prediction of exocytosis rate constant kexo
The energy changes (ΔG(d), Eq. 6) and experimental rate constant ln(kexo) for NPs exocytosis by different cells are in Fig. 2. There are explicit positive correlations between ΔG(d) (Eq. 6) and experimental exocytosis rate constant ln(kexo) for non-cancerous cells (C166 cell, Fig. 2; R2 = 0.61; and p = 0.002), and explicit negative correlations between ΔG(d) and ln(kexo) for non-cancerous cells (U937 and STO cells). Interestingly, cancerous cells (Hela and SNB19 cells) gave negative correlations (with R2 and p 0.90;0.031 and 0.97;0.003, respectively), but their slopes (β, Eq. 4) are lower than non-cancerous cells (U937 and STO cells), i.e. βSTO > βU937 > βHela > βSNB19, corresponding to − 0.0010(± 0.0001) > − 0.0014(± 0.0004) > -0.0017(± 0.0004) > − 0.0024 (± 0.0003). The correlation between ΔG(d) (Eq. 4) and ln(kexo) of RAW264.7 cells (is not significant (p = 0.46), due to one potential outlier. Figure 2 indicates that Hela cells have somewhat higher experimental exocytosis rate constants than any other five cells within the ΔG range of – 250 to 1500 kJ/mol. The frequency factors (A) of the six cells are also different, the lowest is the C166 cells (e− 12.8 (± 0.3)) and the highest is the Hela cells (e− 7.3 (±0.2)).
The exocytosis rate constant ln(kexo) (/s) vs ΔG(d) (Eq. 6, kJ/mol) for six cell lines (C166 and RAW264.7 (12 neutral NPs with similar sizes and different coatings, are denoted by ‘red circle’43. All kexo were obtained based solely on initial and final intracellular NP concentrations, Table S1); U937 (three zwitterionic and three PEG-coated NPs with different sizes, are denoted by ‘blue circle’44. All kexo were obtained based on exocytosis curves, Fig. S2); Hela (three transferrin-coated NPs with different sizes, are denoted by ‘black circle’28 and one D-penicillamine coated NP, is denoted by ‘purple circle’45. All four kexo were obtained based on exocytosis curves, Fig. S2); STO and SNB19 cells (five transferrin-coated NPs with different sizes, are denoted by ‘black circle’28. All 10 kexo were obtained based solely on initial and final intracellular NP concentrations, Table S1)). The dotted lines denote 95% conference intervals. Modelling results include linear functions, R2 and p-values.
Validation of modelling
kexo for six charged NPs and 14 neutral NPs were used to test our models for U937 cells and Hela cells. Figure 3 presents that 95% confidence intervals of predicted kexo for one cationic and two anionic NPs overlap with experimental kexo while kexo of two cationic and one anionic NPs were overestimated. The 95% observation-based confidence intervals of cationic NPs are wider than anionic NPs, and cationic NP at 48 nm has the largest confidence intervals. Our model (parametrized using exocytosis curves) overestimated 14 values for kexo of neutral NPs that were based solely on initial and final intracellular NP concentrations in Hela cells. All 13 data have similar confidence intervals. Predicted kexo values for neutral NP with size of 100 nm showed the largest confidence intervals.
Plot of ln(kexo, /s) predicted by U937 and Hela models (Fig. 2) vs. experimental ln(kexo, /s): six ln(kexo, /s) based on experimental exocytosis curves of charged NPs were denoted as ‘green circle or blue circle’ (three are cationic and three are anionic44) for U937 cells and 14 ln(kexo, /s) of neutral NPs28,43 based solely on initial and final intracellular NP concentrations for Hela cells were denoted as ‘red circle’. The 95% confidence intervals for the predicted results are also represented by different bars.
Discussion
Modeling assessment
The calculated interaction energies (ΔGnp/v) and energy barrier heights (ΔG‡v/m) in Table 1 are all positive, meaning that all interaction forces among NPs, vesicles and cell membranes would be repulsive. Earlier research reported that NPs with hydrodynamic sizes of 6–7 nm46 and AuNPs47 of 2 nm permeate and absorb through/by membranes with energy barrier heights of 50–500 kJ/mol. Larger aggregated NPs caused higher energy barrier heights (600–1700 kJ/mol)47, which are similar to the interaction energies obtained from our model. All ΔG‡v/m are larger than ΔGnp/v except citrate/PEG-coated NPs44. This means that the large repulsive forces between the hydrophobic NPs (citrate/PEG-coated NPs) and the vesicles make it difficult for the NPs to bind to the vesicles, resulted a lower frequency of NPs being encapsulated by vesicles.
Our model is able to predict the effect of coatings of NPs with similar sizes on kexo by C166 cells and effect of NP sizes with the same coating on kexo by STO and SNB 19 cells. In other words, the model describes both the role of coating and size on exocytosis. Experimental exocytosis data or five cell types fit the prediction, except for RAW 264.7 cells. Five regressions had acceptable results with p < 0.05 and R2 ranging from 0.61 to 0.98 for C166, U937, Hela, SNB 19 and STO cells. Figure 2 showed half of the data outside the 95% confidence intervals for C166 cells, which means the model based on first-order kinetics may need further consideration. Near-perfect regressions for SNB 19 and STO cells were obtained as the experimental data have a linear relationship with surface area as reported28, which were also shown by our models. The result for macrophage (RAW 264.7) cells had low statistical significance (R2 = 0.06, p = 0.462), due to one potential outlier (Fig. 2). Possibly, our generic description of cells (γ, ψ, etc.) do not apply to abnormally proliferating blood macrophages. Errors in individual data (the single datapoint for RAW cells) may also exist due to experimental quirks: for example, lysosome-related demethylation may be particularly important in macrophages48.
Frequency factors A
Most regression equations for various cells showed different slopes β and intercepts A after application of generic cellular descriptors (γLWcell, γABcell and \({\psi }_{0,cell}\)). According to our hypothesis, A will apply to a single cell type. Since we cannot obtain A values based on current data, generic A values for each type of cells were obtained from those regressions. The six A values range from 2.9 × 10–6 to 7.0 × 10–4/s. Hela cells entailed the largest frequency factor (A = 7.0 × 10–4 (e− 7.3 (± 0.2))) and higher experimental kexo than other five cells (Fig. 2). According to our assumption, a higher A means a higher frequency of combination between NPs and vesicles. Reported experiments observed various NPs transport based on vesicles28,45,49, which confirms the model's prediction that NPs have a high frequency of exiting Hela cells via vesicles. We collected the initial uptake numbers of NPs per cell surface area (SI Table S2), and found U937 cells contained the highest number of NPs (14–610 per μm2) at the start of exocytosis experiment, although we got a low frequency of combination between NPs and vesicles (A = 6.3·10–6/s) for U937 cells based on our regression. As Nuri and Park reported44, PEG-coated gold NPs can migrate in the cytoplasm in the form of individual particles without vesicles, confirming that PEG-coated NPs avoid vesicles when entering cells. The large repulsive interaction energy between strong hydrophilic NPs and vesicles also verified this. These combined notions show that our model can describe the exocytotic behavior of differently coated NPs and the possibility of vesicle transport. In addition, protein corona may influence the distribution of NPs in organisms50, influencing A. The hydrophobicity of NP coating may correlate with antibody binding51. Given that hydrophobicity has already been involved in this model, our generic model ignored effects of proteins corona on NPs during exocytosis.
Free energy slope β
Our fitted value for β involving C166 cells is positive, different from negative β involving the other four cells (U937, Hela, STO and SNB19 cells). β values for two cancerous (Hela and SNB 19) cells are negative: this may be related to the hydrophilic character of cancerous cells with metastatic potential facilitating these cells’ translocation to different tissues52. Similar to cancerous cells, STO and U937 cells have high metabolic activity and migration frequency for immune response to tissue damage53, which differs from somewhat ‘tranquil’ or ‘sluggish’ endothelial cells. The energy barriers (ΔG‡v/m) between hydrophilic vesicles and membranes for cancerous cells (Hela and SNB 19 cells) and metastatic cells (STO and U937) may therefore be underestimated (see SI methods), because we used generic cellular surface free energies (γ, ψ) for all cells to build our models. We note also that cancerous cells usually express enhanced intracellular signaling due to e.g., increased vesicle production and excretion54. This may result in unexpected exocytosis patterns as compared to non-cancerous cells, i.e., affect slopes β. Diversity of proteins in biological media55 will influence the surface properties both of NPs and their interaction with cell membranes. For example, cancerous cells experience contact inhibition of locomotion (CIL)56 behavior due to lack of cadherin, which increases repulsive forces as compared to non-cancerous cells57. This in turn increases energetic barriers between cancerous vesicles and cell membranes, affecting β. Moreover, the surface roughness or curvature of cells would influence cells surface energy58: macrophages have irregular shapes but that changes after cellular uptake of NPs, resulting in different curvature and surface free energy59. The various β values of specific cells embed these factors.
Validation of modelling
Figure 3 shows that predicted kexo of six charged NPs for U937 cells are close to experimental kexo based on exocytosis curves. The margin of prediction error is within a factor of two (the range from 8 to 184%), demonstrating that our models based on neutral data still have the potential to predict exocytosis rate constants of charged NPs. Although kexo of two cationic NPs were overestimated. Our model does not consider the aggregation of cationic NPs. However, possible aggregation of cysteamine-coated NPs was observed44, causing a larger effective permeant size and underestimation of ΔG(d). This would imply a lower predicted kexo than based on the regression between ΔG(d) and ln(kexo) (Fig. 2, U937 cells). The interaction between cationic NPs and lysosomes could change the surface energy of weakly basic NP coatings in acidic environment as well. The prediction for the smallest citrate-coated NP has the highest margin of error (33%) in comparison to the other two anionic NPs. Charge-mediated exocytosis was demonstrated in previous QSAR modelling13. Although we obtained good prediction for some charged NPs, data from another dataset or experiments are needed to update this generic model and incorporate the complex effects of charged NPs on exocytosis in the future.
For Hela cells, all 14 values for kexo based solely on initial and final intracellular NP concentrations were overestimated (Fig. 3). The prediction errors are within 1–2 orders of magnitude. kexo predicted based on kinetic exocytosis curves (Fig. S2) are far larger than ‘experimental’ kexo obtained solely using initial and final intracellular NP concentrations. Both for U937 cells and Hela cells, the test NPs with largest ΔG showed the largest 95% confidence intervals, because its ΔG has exceeded ΔG of training (modelling) data. More data and research based on exocytosis curves for different cells are needed for updating and testing our model.
The relationship between surface energies and partition coefficients of coating compounds (Kow)
In this paper, we used size and surface energies (mJ/m2) of coated NPs and cell (lipid) membranes, to calculate their interaction energies. Distinct from partition coefficients (Kow), surface energies60 describe adhesion (strength) between cells and NP materials based on surface area, which can avoid the assumed shortcomings of Kow like those for characterizing NPs aggregation19. Yet, Fig. 4 shows strong correlations between log (Kow or P) of coating compounds and surface energy components (Van der Waals free energies (γLW) and polar acid–base free energies (γAB)) of coating compounds for different coatings from four datasets, confirming mechanistic similarity61 between γ values and Kow both for representing hydrophobicity. Figure 4 also indicates that polar surface free energies (γAB) have a higher correlation (R2 = 0.88) with logKow. Our results verified that surface energy components (mJ/m2) characterize surface hydrophobicity of particles with specific size. Hence, our study recommended surface energy components as available descriptors to estimate NPs exocytosis and bio-transport.
Potential and application of model
The generic model, applying descriptors of coated NPs (size, surface energy components and surface potential) and cells (surface energy components and surface potential), predict exocytosis rate constants of various NPs by calculating interactions between NPs and membranes. The model may find use in biomedicine via cellular or mitochondrial targeting62 by NPs or evaluating their antioxidant and anti-genotoxic capacity63. Our model suggests that transport experiments of NPs in the future can provide accurate surface chemical information to provide sufficient data for prediction of NPs accumulation in cells or organs, e.g., PBPK modeling17.
Strongly hydrophilic (e.g., PEG-coated) NPs are unlikely to bind to membranes and access vesicle transport machinery inside the cell. Nonetheless, our model still predicts the exocytosis of these PEG-coated NPs well (prediction errors within a factor two) and yields lower probabilities of NP-vesicle binding, illustrating the broadness of the model in describing hydrophobicity of different particles. Models were successfully applied to gold NPs and quantum dots with coatings, and can predict the transmembrane transport efficiency of biochemically inert NPs used for delivery of (hydrophobic) drugs, radiopharmaceuticals5 and nanoantioxidants to optimize biomedical applications.
Exocytosis rate constants of transferrin-coated NPs for STO cells and SNB19 cells correlate well with the NPs size as reported28, but this may not be true for other cell types and other NPs coatings. Our results confirm that our model can describe both effects of coatings and sizes on exocytosis, predicting exocytosis rate constants of NPs for both non-cancerous cells and cancerous cells, quantifying NPs transport for medical and toxicological applications.
Our models are built using data for neutral NPs with coating surface inert to acidic lysosomes64, minimizing surface modifications by lysosomes. Figure 3 shows acceptable prediction for charged NPs and U937 cells, which supports potential applicability to charged NPs. However, changes of charged, ionizable and polarizable NPs in surface free energies and surface potential after modifications within lysosomes need to be considered. Elucidating the mechanisms and behaviors of charged NPs need more experimental data. Our generic model currently does not consider aggregation and agglomeration of NPs in cells and vesicles, and a single NP is assumed to be encapsulated by a vesicle. Aggregation and agglomeration of NPs may change the properties of the whole cluster, requiring future research.
Methods
Model outline
As obtain ligand-receptor binding and deformation energy directly is difficult, we built a model based on non-covalent energy changes ΔG as predictors during exocytosis. We attempted to build relationships between ΔG values and exocytosis rate constants (kexo, /s) via Arrhenius equation:
with β as a regression coefficient. We fitted NPs exocytosis rate constants to energies based on vesicle-transported exocytosis for obtaining ‘generic’ values β (Eq. 4) applicable to different types of NPs and cells. A is frequency factor with the same unit of kexo, and energy changes are ΔG(d)
as a function of distances d between NPs, vesicles and cell membranes. Derivations below (Eq. 6) show that the two righthand terms are equal to barrier/‘activation’ (positive) energies ΔGvesicle-cell membrane‡ (ΔGv-m‡)and binding or repulsive energies ΔGnp-vesicle (ΔGnp-v). Figure 5 shows the energy changes before and after exocytosis based on vesicle-transport pathways. Vesicle-transported NPs will overcome barriers to combine vesicles with cell membranes. As such, NPs are successfully excreted by the ‘release’ of the (repulsive) interaction energy between NPs and vesicles. Thereby, ΔG(d) are the energy changes during exocytosis process and obtained via:
Energy profiles before and after combination of vesicles with cell membranes. Before the combination of vesicle-transported NPs with cell membranes, the energy barrier (ΔGv/m‡) between vesicles and cell membranes prevent the two from combining. At the moment of combination, the interaction (repulsive) energy (ΔGnp/v) between the NPs and the vesicles promote the NPs to desorb from the membrane to detach. ΔG is the energy change at the combination moment (ΔGv/m‡ minus ΔGnp/v).
We calculated ΔG between NPs and vesicles as well as between vesicles and cell membranes as a function of d (trajectory of exocytosis) from interaction forces involving Van de Waals (ΔGLW), electrostatic (ΔGe) and Lewis acid–base (polar) forces (ΔGAB) (see SI Methods)65,66, according to
We took the maximum interaction energy between NPs and vesicles as their interaction energy (see energy profiles in Fig. S1). All kexo data were obtained from exocytosis experiments for different NPs and cells. The properties of NPs (e.g., size, surface free energies) and cells/vesicles (surface potential and surface free energies) determine the computed ΔG(d). We assumed that vesicles share the same characteristics of cell membranes because vesicles combine with cell membranes during exocytosis. Additionally, we assumed that properties of neutral NPs do not change (i.e., are inert) when vesicle-transported NPs enter the lysosome’s acidic environments. Linear regressions between ΔG(d) and lnkexo were assessed based on R2 and p values. A p-value less than 0.05 was considered to be statistically significant. The 95% confidence intervals of linear relationships (Eq. 4) were obtained via Eqs. S4–S6.
Arrhenius equations
We stipulate that exocytosis rate constants can be described via interaction energy changes ΔG and the Arrhenius equation according to
in which Avesicle is a frequency factor for vesicle-transported NPs, Avesicle has the same unit as the rate constant k (/s). kBT is the product of Boltzmann's constant kB and temperature T (310 K). ΔG(d) (Fig. 2) is the energy change before and after vesicle-transported exocytosis.
Hence, exocytosis rate constant can be derived via combining Eqs. (3) and (9):
where Avesicle is the frequency factor for the vesicle-transported exocytosis pathway.
Calculation and treatment of frequency factors A
NPs may be trafficked by cellular machinery with or without vesicles. Presumably, exocytosis rate constants are proportional to the frequency of those NPs binding to vesicles38. We assumed that a single NP binds to/within a vesicle42,67, and ignored individual reports of (intracellular) aggregation28 (pathway 3 and 4, Fig. 1).
Values of A present total frequency factors of NPs encapsulated by vesicles or vesicle-free NPs trafficked in cytoplasm. We describe frequencies as
A large A describes a high intracellular trafficking efficiency (e.g., including production) of vesicles in the cell. It is hard to find good estimates for A or Avesicle for specific NPs (coatings) from exocytosis experiments that ignores differences in transport pathways. We overcome this by fitting NPs exocytosis rate constants to energies during exocytosis for obtaining ‘generic’ values A (Eq. 4) applicable to different types of NPs and cells.
Calculation of concentrations via Boltzmann
NPs interact with cell membranes to be encapsulated and transported by vesicles and released into the cytoplasm. Energy changes e.g. \(\left( {k\, \sim \,{\text{e}}^{{ - \Delta {\text{G}}/{\text{k}}_{{\text{B}}} {\text{T}}}} } \right)\)68 can be used to predict concentrations and transport rates of NPs through membranes, where ΔG denote the free energies along the translocation pathway. We thus calculated [NP]vesicle/[NP]total as the relative concentrations of NPs across the cellular compartments (vesicles/cytoplasm) according to ΔG values:
In which ΔGnp-vesicle is the interaction energy between NP and vesicle. ΔGnp-vesicle involved in the binding of NPs and vesicles, include ligand-receptor binding69, elastic or deformation energy70, and ‘generic’ surface-based DLVO (Van de Waals, electrostatic and Lewis acid–base (polar) forces). It is cumbersome to obtain ligand-receptor binding and deformation energy directly, and experimental values for β of different cells are not available. We overcome this by fitting NPs exocytosis rate constants to ΔG(d) to obtain ‘generic’ values for β (Eq. 4) applicable to different types of NPs and cells.
Dataset
We applied 64 NPs with different surface modification from four open literature28,43,44,45 for experimental exocytosis to parametrize and build our models as training data and test data. 36 NPs from the first dataset (Au@PEG-X NPs) were selected with 20 nm AuNPs and functionalized with hydrocarbyl groups (X contains one, six, nine or 12 carbons) linked by polyethylene glycol (PEG), and with hydrodynamic diameter (HD, in water) of 25.17–28.13 nm43. The exocytosis experiments were based on 6 h intracellular uptake.
The second dataset includes four AuNPs types coated with zwitterionic cysteine, polyethylene glycol (PEG), anionic citrate and cationic cysteamine. The HD of each type NP range from 14 to 56 nm. The aggregation of NPs may happen on cationic NPs44. The exocytosis experiments were based on cellular uptake after 6 h at 37 °C.
The third dataset entailed AuNPs with five sizes range of 14–100 nm were coated by transferrin28for three types of cellular exocytosis. The exocytosis experiments were based on 10 h intracellular uptake at 37 °C. The fourth group entailed a zwitterionic D-penicillamine-coated quantum dot (QD)45 with size of 8 nm (TEM size), and exocytosis experiments were executed at 37 °C.
To ensure the accuracy of our approach, we do not mix experimental exocytosis data for NPs characterized by different techniques (i.e., TEM and HD) into the same model. For most data, molecular descriptors of the coatings viz., single molecular weight per molecular volume and topological polar surface area per volume, were used to calculate the surface free energy components (the Lifshitz-van de Waals (γLW), acid–base (γAB), electron acceptor (γ+) and donor (γ-) terms)66 of various NPs according to our previous methods66. Values for γ of PEG were obtained from measurements by Van Oss et al.71. The electrostatic energies (Eq. 7) were obtained from surface potential, which in turn were obtained from zeta potential measured for six charged NPs (see SI Methods). All coating compounds’ identifiers (i.e., SMILE strings) and NPs properties are shown in Tables S3 and S4.
Those data involved six types of cells, namely RAW 264.7 (mouse macrophage), C166 (mouses endothelial), Hela (ovarian cancer cell), U937 (human macrophage), STO (fibroblast) and SNB19 (brain tumor) cells. For the descriptors of cell (membranes) and vesicles, we implemented a surface potential \({\psi }_{0,cell}\) of − 40 mV66. The γLWcell and γABcell were defined as 39 mJ/m2 and 27 mJ/m2 for a generic cell surface, respectively. The γ+cell (2.6 mJ/m2) and γ−cell (71 mJ/m2) values are calculated based on the previous method66.
Determination of experimental exocytosis rate constant kexo
All data described exocytosis involved experiments initiated without NPs in the extracellular medium. Intracellular concentrations NPt/NP0 of only 16 NPs (12 for U937 cells, four for Hela cells) were recorded over time t. The 16 experiments for U937 and Hela cells were fitted with exocytosis curves \(\left( {\left[ {NP} \right]_{{\text{t}}} = \left[ {NP} \right]_{0} \times e^{ - kt} { + }\left[ {NP} \right]_{eq} } \right)\), presented in Fig. S2. A first order kinetic equation \(\left( {\left[ {NP} \right]_{{\text{t}}} = \left[ {NP} \right]_{0} \times e^{ - kt} } \right)\) was also used to calculate exocytosis rate constants kexo of 48 NPs, wherein we took [NP]eq = 0. The experimental durations of these first and third dataset are 24 and 8 h, respectively. All exocytosis rate constants are given in Table S1.
Validation of models
We built all models based on neutral NPs in order to reduce the error in surface property changes due to charged NPs reacting with lysosome acid environment, but there are some charged data of U937 to test the error. Six neutral NPs of U937 cells were used to build robust model and another six charged NPs data based on exocytosis experiments, i.e. kexo values, were used as test data. To test the robustness of the model based on four neutral NPs with exocytosis curves for Hela cells, 14 neutral kexo (based solely on initial and final NPs concentrations) for Hela cells were used as test data. There is not enough data to test models for RAW 264.7, C166, STO and SNB19 cells.
Data availability
All data, models, or code generated or used during the study are available from the corresponding author by request.
References
Cabuzu, D., Cirja, A., Puiu, R. & Grumezescu, A. M. Biomedical applications of gold nanoparticles. Curr. Top. Med. Chem. 15, 1605–1613 (2015).
Ding, L. et al. Size, shape, and protein corona determine cellular uptake and removal mechanisms of gold nanoparticles. Small 14, 1801451. https://doi.org/10.1002/smll.201801451 (2018).
Nelson, B. C., Johnson, M. E., Walker, M. L., Riley, K. R. & Sims, C. M. Antioxidant cerium oxide nanoparticles in biology and medicine. Antioxidants 5, 15. https://doi.org/10.3390/antiox5020015 (2016).
Venditti, I. et al. Direct conjugation of resveratrol on hydrophilic gold nanoparticles: structural and cytotoxic studies for biomedical applications. Nanomaterials 10, 1898. https://doi.org/10.3390/nano10101898 (2020).
Maccora, D. et al. Gold nanoparticles and nanorods in nuclear medicine: A mini review. Appl. Sci. Basel 9, 3232. https://doi.org/10.3390/app9163232 (2019).
Singh, A. K. & Srivastava, O. N. One-step green synthesis of gold nanoparticles using black cardamom and effect of pH on its synthesis. Nanoscale Res. Lett. 10, 5. https://doi.org/10.1186/s11671-015-1055-4 (2015).
Pan, Y. et al. Size-dependent cytotoxicity of gold nanoparticles. Small 3, 1941–1949. https://doi.org/10.1002/smll.200700378 (2007).
Cheng, Y. H., Riviere, J. E., Monteiro-Riviere, N. A. & Lin, Z. M. Probabilistic risk assessment of gold nanoparticles after intravenous administration by integrating in vitro and in vivo toxicity with physiologically based pharmacokinetic modeling. Nanotoxicology 12, 453–469. https://doi.org/10.1080/17435390.2018.1459922 (2018).
Li, J. Y. et al. Mitophagy-lysosomal pathway is involved in silver nanoparticle-induced apoptosis in A549 cells. Ecotoxicol. Environ. Saf. 208, 11463. https://doi.org/10.1016/j.ecoenv.2020.111463 (2021).
Khan, A. I., Lu, Q., Du, D., Lin, Y. H. & Dutta, P. Quantification of kinetic rate constants for transcytosis of polymeric nanoparticle through blood-brain barrier. Biochim. Biophys. Acta Gen. Subj. 2779–2787, 2018. https://doi.org/10.1016/j.bbagen.2018.08.020 (1862).
Ohta, S., Inasawa, S. & Yamaguchi, Y. Real time observation and kinetic modeling of the cellular uptake and removal of silicon quantum dots. Biomaterials 33, 4639–4645. https://doi.org/10.1016/j.biomaterials.2012.03.029 (2012).
Chen, W. B., D’Argenio, D. Z., Sipos, A., Kim, K. J. & Crandall, E. D. Biokinetic modeling of nanoparticle interactions with lung alveolar epithelial cells: uptake, intracellular processing, and egress. Am. J. Physiol. Regul. Integr. Comp. Physiol. 320, R36–R43. https://doi.org/10.1152/ajpregu.00184.2020 (2021).
Bigdeli, A., Hormozi-Nezhad, M. R. & Parastar, H. Using nano-QSAR to determine the most responsible factor(s) in gold nanoparticle exocytosis. RSC Adv. 5, 57030–57037. https://doi.org/10.1039/c5ra06198a (2015).
Bao, Q. Y. et al. The effect of oil-water partition coefficient on the distribution and cellular uptake of liposome-encapsulated gold nanoparticles. Colloids Surf. B. 146, 475–481. https://doi.org/10.1016/j.colsurfb.2016.06.046 (2016).
Gao, Y. F., Xie, Z. C., Feng, J. F., Ma, W. Q. & Zhu, L. Different factors determined the toxicokinetics of organic chemicals and nanomaterials exposure to zebrafish (Danio Rerio). Ecotoxicol. Environ. Saf. 186, 109810. https://doi.org/10.1016/j.ecoenv.2019.109810 (2019).
Wang, J. Q., Hoondert, R. P. J., Thunnissen, N. W., van de Meent, D. & Hendriks, A. J. Chemical fate of persistent organic pollutants in the arctic: Evaluation of simplebox. Sci. Total Environ. 720, 137579. https://doi.org/10.1016/j.scitotenv.2020.137579 (2020).
Wang, J. Q. et al. A generalized physiologically based kinetic model for fish for environmental risk assessment of pharmaceuticals. Environ. Sci. Technol. 56, 6500–6510. https://doi.org/10.1021/acs.est.1c08068 (2022).
Price, E. & Gesquiere, A. J. An in vitro assay and artificial intelligence approach to determine rate constants of nanomaterial-cell interactions. Sci. Rep. 9, 1. https://doi.org/10.1038/s41598-019-50208-x (2019).
Praetorius, A. et al. The road to nowhere: Equilibrium partition coefficients for nanoparticles. Environ. Sci. Nano 1, 317–323. https://doi.org/10.1039/c4en00043a (2014).
Crane, M., Handy, R. D., Garrod, J. & Owen, R. Ecotoxicity test methods and environmental hazard assessment for engineered nanoparticles. Ecotoxicology 17, 421–437. https://doi.org/10.1007/s10646-008-0215-z (2008).
Ruenraroengsak, P. et al. Respiratory epithelial cytotoxicity and membrane damage (holes) caused by amine-modified nanoparticles. Nanotoxicology 6, 94–108. https://doi.org/10.3109/17435390.2011.558643 (2012).
Nativo, P., Prior, I. A. & Brust, M. Uptake and intracellular fate of surface-modified gold nanoparticles. ACS Nano 2, 1639–1644. https://doi.org/10.1021/nn800330a (2008).
Chung, H. E., Park, D. H., Choy, J. H. & Choi, S. J. Intracellular trafficking pathway of layered double hydroxide nanoparticles in human cells: Size-dependent cellular delivery. Appl. Clay Sci. 65–66, 24–30. https://doi.org/10.1016/j.clay.2012.06.007 (2012).
Geiser, M. et al. Ultrafine particles cross cellular membranes by nonphagocytic mechanisms in lungs and in cultured cells. Environ. Health Perspect. 113, 1555–1560. https://doi.org/10.1289/ehp.8006 (2005).
Poon, W. et al. Elimination pathways of nanoparticles. ACS Nano 13, 5785–5798. https://doi.org/10.1021/acsnano.9b01383 (2019).
Fortuni, B. et al. Polymeric engineering of nanoparticles for highly efficient multifunctional drug delivery systems. Sci. Rep. 9, 1. https://doi.org/10.1038/s41598-019-39107-3 (2019).
Marchesano, V. et al. Imaging inward and outward trafficking of gold nanoparticles in whole animals. ACS Nano 7, 2431–2442. https://doi.org/10.1021/nn305747e (2013).
Chithrani, B. D. & Chan, W. C. W. Elucidating the mechanism of cellular uptake and removal of protein-coated gold nanoparticles of different sizes and shapes. Nano Lett. 7, 1542–1550. https://doi.org/10.1021/nl070363y (2007).
Frohlich, E. Cellular elimination of nanoparticles. Environ. Toxicol. Pharmacol. 46, 90–94. https://doi.org/10.1016/j.etap.2016.07.003 (2016).
Song, H. I., Hong, J. A., Lee, H. & Lim, K. I. Comparative study on the catalytic activity of Fe-doped ZrO2 nanoparticles without significant toxicity through chemical treatment under various pH conditions. Sci. Rep. 9, 1. https://doi.org/10.1038/s41598-019-47443-7 (2019).
Silva, E., Barreiros, L., Segundo, M. A., Lima, S. A. C. & Reis, S. Cellular interactions of a lipid-based nanocarrier model with human keratinocytes: Unravelling transport mechanisms. Acta Biomater. 53, 439–449. https://doi.org/10.1016/j.actbio.2017.01.057 (2017).
Wang, X. W. et al. Endocytosis and organelle targeting of nanomedicines in cancer therapy. Int. J. Nanomed. 15, 9447–9467. https://doi.org/10.2147/ijn.s274289 (2020).
Zhang, J. X. et al. Systematic investigation on the intracellular trafficking network of polymeric nanoparticles. Nanoscale 9, 3269–3282. https://doi.org/10.1039/c7nr00532f (2017).
Lunov, O. et al. Lysosomal degradation of the carboxydextran shell of coated superparamagnetic iron oxide nanoparticles and the fate of professional phagocytes. Biomaterials 31, 9015–9022. https://doi.org/10.1016/j.biomaterials.2010.08.003 (2010).
Setyawati, M. I., Yuan, X., Xie, J. P. & Leong, D. T. The influence of lysosomal stability of silver nanomaterials on their toxicity to human cells. Biomaterials 35, 6707–6715. https://doi.org/10.1016/j.biomaterials.2014.05.007 (2014).
Li, J. P. et al. Rapid pH-responsive self-disintegrating nanoassemblies balance tumor accumulation and penetration for enhanced anti-breast cancer therapy. Acta Biomater. 134, 546–558. https://doi.org/10.1016/j.actbio.2021.04.022 (2021).
Zhang, J. et al. Effects of the surface charge of polyamidoamine dendrimers on cellular exocytosis and the exocytosis mechanism in multidrug-resistant breast cancer cells. J. Nanobiotechnol. https://doi.org/10.1186/s12951-021-00881-w (2021).
Sipos, A. et al. Alveolar epithelial cell processing of nanoparticles activates autophagy and lysosomal exocytosis. Am. J. Physiol. Lung Cell. Mol. Physiol. 315, 286–300. https://doi.org/10.1152/ajplung.00108.2018 (2018).
Dey, A., Stenberg, J., Dandekar, P. & Jain, R. A combinatorial study of experimental analysis and mathematical modeling: How do chitosan nanoparticles deliver therapeutics into cells?. Carbohydr. Polym. 229, 115437. https://doi.org/10.1016/j.carbpol.2019.115437 (2020).
Zhdanov, V. P. Interpretation of amperometric kinetics of content release during contacts of vesicles with a lipid membrane. Eur. Biophys. J. Biophys. Lett. 46, 461–470. https://doi.org/10.1007/s00249-016-1189-z (2017).
Deng, H., Dutta, P. & Liu, J. Stochastic modeling of nanoparticle internalization and expulsion through receptor-mediated transcytosis. Nanoscale 11, 11227–11235. https://doi.org/10.1039/c9nr02710f (2019).
Rees, P., Wills, J. W., Brown, M. R., Barnes, C. M. & Summers, H. D. The origin of heterogeneous nanoparticle uptake by cells. Nat. Commun. 10, 1. https://doi.org/10.1038/s41467-019-10112-4 (2019).
Ho, L. W. C., Yin, B. H., Dai, G. L. & Choi, C. H. J. Effect of surface modification with hydrocarbyl groups on the exocytosis of nanoparticles. Biochemistry 60, 1019–1030. https://doi.org/10.1021/acs.biochem.0c00631 (2021).
Oh, N. & Park, J. H. Surface chemistry of gold nanoparticles mediates their exocytosis in macrophages. ACS Nano 8, 6232–6241. https://doi.org/10.1021/nn501668a (2014).
Jiang, X. E. et al. Endo- and exocytosis of zwitterionic quantum dot nanoparticles. ACS Nano 4, 6787–6797. https://doi.org/10.1021/nn101277w (2010).
Gupta, R., Badhe, Y., Mitragotri, S. & Rai, B. Permeation of nanoparticles across the intestinal lipid membrane: Dependence on shape and surface chemistry studied through molecular simulations. Nanoscale 12, 6318–6333. https://doi.org/10.1039/c9nr09947f (2020).
Lunnoo, T., Assawakhajornsak, J., Ruangchai, S. & Puangmali, T. Role of surface functionalization on cellular uptake of AuNPs characterized by computational microscopy. J. Phys. Chem. B 124, 1898–1908. https://doi.org/10.1021/acs.jpcb.9b11600 (2020).
Karpurapu, M. et al. Kruppel like factor 4 promoter undergoes active demethylation during monocyte/macrophage differentiation. PLoS ONE 9, 93362. https://doi.org/10.1371/journal.pone.0093362 (2014).
Ding, L. et al. Intracellular fate of nanoparticles with polydopamine surface engineering and a novel strategy for exocytosis-inhibiting, lysosome impairment-based cancer therapy. Nano Lett. 17, 6790–6801. https://doi.org/10.1021/acs.nanolett.7b03021 (2017).
Li, X. et al. Surface chemistry governs the sub-organ transfer, clearance and toxicity of functional gold nanoparticles in the liver and kidney. J. Nanobiotechnol. 18, 1. https://doi.org/10.1186/s12951-020-00599-1 (2020).
Wenger, Y. et al. Tissue distribution and pharmacokinetics of stable polyacrylamide nanoparticles following intravenous injection in the rat. Toxicol. Appl. Pharmacol. 251, 181–190. https://doi.org/10.1016/j.taap.2010.11.017 (2011).
Minnes, R. et al. Using attenuated total reflection-fourier transform infra-red (ATR-FTIR) spectroscopy to distinguish between melanoma cells with a different metastatic potential. Sci. Rep. 7, 1. https://doi.org/10.1038/s41598-017-04678-6 (2017).
Jeon, Y. K. et al. Mesenchymal stem cells’ interaction with skin: Wound-healing effect on fibroblast cells and skin tissue. Wound Repair Regener. 18, 655–661. https://doi.org/10.1111/j.1524-475X.2010.00636.x (2010).
Becker, A. et al. Extracellular vesicles in cancer: Cell-to-cell mediators of metastasis. Cancer Cell 30, 836–848. https://doi.org/10.1016/j.ccell.2016.10.009 (2016).
Ashby, J., Pan, S. Q. & Zhong, W. W. Size and surface functionalization of iron oxide nanoparticles influence the composition and dynamic nature of their protein corona. ACS Appl. Mater. Interfaces 6, 15412–15419. https://doi.org/10.1021/am503909q (2014).
Batson, J., MacCarthy-Morrogh, L., Archer, A., Tanton, H. & Nobes, C. D. EphA receptors regulate prostate cancer cell dissemination through Vav2-RhoA mediated cell-cell repulsion. Biol. Open 3, 453–462. https://doi.org/10.1242/bio.20146601 (2014).
Halbleib, J. M. & Nelson, W. J. Cadherins in development: cell adhesion, sorting, and tissue morphogenesis. Genes Dev. 20, 3199–3214. https://doi.org/10.1101/gad.1486806 (2006).
Ferrari, M., Cirisano, F. & Moran, M. C. Mammalian cell behavior on hydrophobic substrates: Influence of surface properties. Colloids Interfaces 3, 48. https://doi.org/10.3390/colloids3020048 (2019).
Petty, H. R., Hafeman, D. G. & McConnell, H. M. Disappearance of macrophage surface folds after antibody-dependent phagocytosis. J. Cell Biol. 89, 223–229. https://doi.org/10.1083/jcb.89.2.223 (1981).
Ponsonnet, L. et al. Relationship between surface properties (roughness, wettability) of titanium and titanium alloys and cell behaviour. Mater. Sci. Eng. C 23, 551–560. https://doi.org/10.1016/s0928-4931(03)00033-x (2003).
Xiao, Y. & Wiesner, M. R. Characterization of surface hydrophobicity of engineered nanoparticles. J. Hazard. Mater. 215, 146–151. https://doi.org/10.1016/j.jhazmat.2012.02.043 (2012).
Chodari, L. et al. Targeting mitochondrial biogenesis with polyphenol compounds. Oxid. Med. Cell. Longevity. https://doi.org/10.1155/2021/4946711 (2021).
Ahmadian, E. et al. Effects of quercetin loaded nanostructured lipid carriers on the paraquat-induced toxicity in human lymphocytes. Pestic. Biochem. Physiol. 167, 5. https://doi.org/10.1016/j.pestbp.2020.104586 (2020).
Yanes, R. E. et al. Involvement of lysosomal exocytosis in the excretion of mesoporous silica nanoparticles and enhancement of the drug delivery effect by exocytosis inhibition. Small 9, 697–704. https://doi.org/10.1002/smll.201201811 (2013).
Nolte, T. M., Kettler, K., Meesters, J. A. J., Hendriks, A. J. & van de Meent, D. A semi-empirical model for transport of inorganic nanoparticles across a lipid bilayer: Implications for uptake by living cells. Environ. Toxicol. Chem. 34, 488–496. https://doi.org/10.1002/etc.2812 (2015).
Lu, B. Q., Hendriks, A. J. & Nolte, T. M. A generic model based on the properties of nanoparticles and cells for predicting cellular uptake. Colloids Surf. B. 209, 112155. https://doi.org/10.1016/j.colsurfb.2021.112155 (2022).
Tlotleng, N. et al. Cytotoxicity, intracellular localization and exocytosis of citrate capped and PEG functionalized gold nanoparticles in human hepatocyte and kidney cells. Cell Biol. Toxicol. 32, 305–321. https://doi.org/10.1007/s10565-016-9336-y (2016).
Shi, W. D., Wang, J. Z., Fan, X. J. & Gao, H. J. Size and shape effects on diffusion and absorption of colloidal particles near a partially absorbing sphere: Implications for uptake of nanoparticles in animal cells. Phys. Rev. E 78, 061914. https://doi.org/10.1103/PhysRevE.78.061914 (2008).
Shen, Z. Q., Ye, H. L. & Li, Y. Understanding receptor-mediated endocytosis of elastic nanoparticles through coarse grained molecular dynamic simulation. PCCP 20, 16372–16385. https://doi.org/10.1039/c7cp08644j (2018).
Li, Y., Kroger, M. & Liu, W. K. Shape effect in cellular uptake of PEGylated nanoparticles: Comparison between sphere, rod, cube and disk. Nanoscale 7, 16631–16646. https://doi.org/10.1039/c5nr02970h (2015).
Vanoss, C. J., Chaudhury, M. K. & Good, R. J. Monopolar surfaces. Adv. Colloid Interface Sci. 28, 35–64. https://doi.org/10.1016/0001-8686(87)80008-8 (1987).
Acknowledgements
We would like to express our gratitude to the China Scholarship Council for the support of the first author’s study at Radboud University Nijmegen for environmental science.
Author information
Authors and Affiliations
Contributions
The data was collected by B.Q.L., and the models were built by B.Q.L., based on the supervision of A.J.H., and T.M.N., J.Q.W., P.T.J.S., A.J.H. and T.M.N. revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lu, B., Wang, J., Scheepers, P.T.J. et al. Generic prediction of exocytosis rate constants by size-based surface energies of nanoparticles and cells. Sci Rep 12, 17813 (2022). https://doi.org/10.1038/s41598-022-20761-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-20761-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.