Abstract
Concern about resistance to chlorhexidine has increased due to the wide use of the latter. The impact of the qacA/B and smr chlorhexidine tolerance genes on the outcome of methicillin-resistant Staphylococcus aureus (MRSA) infections is unclear. We evaluated the prevalence and clinical impact of, and microbiological risk factors for, qacA/B tolerance in MRSA bacteremia. MRSA bacteremia that occurred more than two days after intensive care unit admission between January 2009 and December 2018 was identified from a prospective cohort of S. aureus bacteremia in a tertiary-care hospital from South Korea. A total of 183 MRSA blood isolates was identified, and the major genotype found was ST5-MRSA-II (87.4%). The prevalences of qacA/B and smr were 67.2% and 3.8%, respectively. qacA/B-positive isolates were predominantly ST5-MRSA-II (96.7% [119/123]), the dominant hospital clone. In a homogenous ST5-MRSA-II background, qacA/B positivity was independently associated with septic shock (aOR, 4.85), gentamicin resistance (aOR, 74.43), and non-t002 spa type (aOR, 74.12). qacA/B positivity was found to have decreased significantly in ST5-MRSA-II in association with a decline in qacA/B-positive t2460, despite the increasing use of chlorhexidine since 2010 (P < 0.001 for trend). Continuous surveillance of the qac genes, and molecular characterization of their plasmids, are needed to understand their role in MRSA epidemiology.
Similar content being viewed by others
Introduction
Methicillin-resistant Staphylococcus aureus (MRSA) is one of the most common causes of healthcare-associated infection. It leads to increased morbidity and mortality in critically ill patients in intensive care units (ICUs). In the United States, the rate of hospital-acquired Staphylococcus aureus bacteremia was 31% and had decreased by 1.9% annually from 1995 to 2015. However, the frequency of metastatic complications of S. aureus bacteremia has increased by 0.9% per year, and the overall mortality rate is still high at 26.5%1. In South Korea, the hospital-acquired S. aureus bacteremia rate was 56% and had decreased by 2.4% annually from 2008 to 2018, and the 90-day mortality rate was 27% in 2018. The proportion of MRSA in S. aureus bacteremia was 52%, with a decrease of 1.6% annually2.
Infection control strategies to minimize MRSA infection include education about hand washing, various MRSA bundles, and chlorhexidine bathing with or without topical nasal mupirocin application3,4. Chlorhexidine bathing is estimated to reduce MRSA colonization and bacteremia in the ICU by 41% and 36%, respectively5. While there is benefit from chlorhexidine bathing and nasal mupirocin application, concern persists about the potential emergence of increased resistance due to antiseptic exposure. The qacA and qacB genes encode multidrug efflux pumps associated with chlorhexidine tolerance6,7. Several studies have shown an increase in the proportion of these genes after the widespread use of chlorhexidine in hospitals8,9,10. In addition, the presence of the qacA or qacB (qacA/B) gene has been linked to resistance to gentamicin, clindamycin, and penicillin, as well as to a high vancomycin minimal inhibitory concentration (MIC)11,12,13.
Previous studies have shown an association between qacA/B positivity and specific MRSA genotypes, including ST5- and ST239-MRSA6,7,14. ST5-MRSA is a major hospital MRSA clone in South Korea, while ST72-MRSA is a community-associated MRSA2. Because ST5-MRSA is associated with higher mortality and more accessory gene regulator (agr) dysfunctional than ST72-MRSA15,16, clinical and microbiological risk factors for qacA/B positivity might be affected by the characteristics of specific genotypes. Thus, this study aimed to evaluate the prevalence and clinical impact of, and microbiological risk factors for, qacA/B positivity in MRSA bacteremia in a homogenous genotype background after the implementation of chlorhexidine use in a tertiary hospital from South Korea.
Methods
Study population and design
A retrospective analysis of a prospective observational cohort of patients with S. aureus bacteremia was conducted at the Asan Medical Center, a 2700-bed tertiary care hospital with 118 adult ICU beds, in Seoul, South Korea. All consecutive adult patients (≥ 16 years old) who had MRSA bacteremia that developed more than two days after ICU admission between January 2009 and December 2018 were identified, and only patients whose MRSA isolates were collected and available for laboratory tests were included in the study17. Only the initial episode of MRSA bacteremia in any given patient was included. Recurrence of S. aureus bacteremia more than three months from the first bacteremia episode was regarded as a new episode and included in the analysis.
To evaluate the clinical impact of qacA/B positivity in a single genotypic background, multilocus sequence typing (MLST) of the MRSA isolates was performed. The clinical, microbiological, and genotypic characteristics of qacA/B-positive MRSA isolates, and risk factors for qacA/B positivity, were analyzed. Treatment outcome of bacteremia was evaluated based on 14-day, 30-day, and 12-week all-cause mortality. This study was approved by the Asan Medical Center Institutional Review Board (IRB number: 2008-0274). The requirement for written informed consent was waived by the Asan Medical Center Institutional Review Board, given the observational nature of the study. All methods were performed in accordance with the relevant guidelines and regulations.
Study definitions
The prognosis for the underlying disease was classified according to the system of McCabe and Jackson as rapidly fatal (when death was expected within a few months), ultimately fatal (when death was expected within 4 years), and nonfatal (when life expectancy was > 4 years)18. Charlson’s comorbidity index was used to measure comorbid burden19. The severity of bacteremia was measured by the Acute Physiology and Chronic Health Evaluation II (APACHE II) score and the Pitt bacteremia score20. Sites of infection causing MRSA bacteremia were defined according to the Centers for Disease Control and Prevention criteria21. Catheter-related infection was defined according to the Infectious Diseases Society of America guidelines22.
Microbiological data and determination of genotypes
All S. aureus isolates were identified by standard methods. Antimicrobial susceptibilities were determined with a MicroScan (Dade Behring, West Sacramento, CA, USA) and the standard criteria of the Clinical and Laboratory Standards Institute (CLSI)23. Methicillin resistance was confirmed by detection of the mecA gene by PCR. Vancomycin minimal inhibitory concentrations (MICs) were determined by the vancomycin E-test (AB Biodisk, Piscataway, NJ, USA) on Mueller–Hinton agar. Heterogeneous vancomycin-intermediate S. aureus (hVISA) was detected by population analysis profiling, as previously described24.
Staphylococcal Cassette Chromosome mec (SCCmec) type, MLST, and Staphylococcus protein A (spa) typing, as well as agr genotyping, were performed as described elsewhere25,26,27,28. MLST allele numbers and sequence types (STs) are available in the MLST database (http://www.mlst.net). agr dysfunction was detected by δ-hemolysin production29.
Chlorhexidine MIC was determined by the broth microdilution test described by CLSI30. Chlorhexidine digluconate 20% aqueous solution (Sigma-Aldrich, St. Louis, MO) was used as the starting material. The bacterial inocula (5 × 105 colony-forming unit/mL) were seeded to wells containing chlorhexidine from 0.125 to 128 mg/L and incubated at 35 ℃ for 18 h. S. aureus ATCC 29213 was used as a quality control strain in susceptibility testing. Mupirocin resistance (low and high level) was detected by the disk diffusion method31, and high-level mupirocin resistance was confirmed by PCR detection of the mupA gene. The qacA/ qacB, smr, and mupA genes were detected by PCR using primers described elsewhere32.
Infection control measures
From January 2010, our ICUs implemented a central venous catheter bundle including skin cleaning with 2% chlorhexidine gluconate in 70% alcohol. Once-daily chlorhexidine bathing was performed on patients with MRSA infections or colonizations in the surgical ICU from August 2011 and in the medical ICU from April 2013. Chlorhexidine gluconate-impregnated film dressing was used in 2016. All ICUs performed chlorhexidine bathing from August 2018. Topical nasal mupirocin has not usually been used for decolonization in ICU patients.
Statistical analysis
We analyzed categorical variables using the chi-square or Fisher’s exact test, and continuous variables using Student’s t-test or the Mann–Whitney U test. A univariate analysis of the risk factors for qacA/B-positive MRSA was performed. All variables with P values < 0.1 in the univariate analysis were included in a multiple logistic regression model to identify independent risk factors. These variables were examined for correlations before inclusion in the multivariate analysis. Categorical variables were analyzed by the chi-square test, and continuous variables over time were analyzed by linear regression for trend. All tests of significance were two-tailed, and P < 0.05 was considered statistically significant. All statistical analyses were performed with SPSS, version 24.0 K for Windows (SPSS Inc, Chicago, IL, USA) and MedCalc software (Mariakerke, Belgium).
Results
Prevalence of qacA/B in MRSA blood isolates
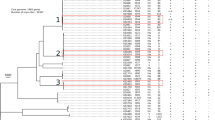
During the study period, 238 episodes of ICU-acquired MRSA bacteremia were identified. When the 183 episodes with available MRSA isolates were analyzed, the two major genotypes were ST5-MRSA-II-agr group II (87.4%), the hospital MRSA clone, and ST72-MRSA-IV-agr group I (7.1%), the community-associated MRSA clone in South Korea. Major spa types in ST5-MRSA-II were t2460 (59.4%), t9353 (14.4%), and t002 (8.1%), and those in ST72-MRSA-IV were t148 (23.7%), t664 (23.7%), and t324 (15.4%). The prevalences of the qacA/B and smr genes were 67.2% (123/183) and 3.8% (7/183), respectively. The qacA/B-positive MRSA isolates were predominantly ST5-MRSA-II (96.7% [119/123]), and all ST72-MRSA-IV isolates were qacA/B-negative. Because the clinical and microbiological characteristics of ST5- and ST72-MRSA are known to differ15, the impact of qacA/B positivity was analyzed in ST5-MRSA-II isolates only, to exclude any effect of clonal heterogeneity on qacA/B negativity.
Clinical characteristics and treatment outcomes of patients with qacA/B-positive ST5-MRSA-II bacteremia
Of the 160 ST5-MRSA-II isolates, 119 (74.4%) were qacA/B-positive. The clinical characteristics and treatment outcomes of the 160 patients with ST5-MRSA-II bacteremia according to the presence of the qacA/B gene are presented in Table 1. Median age was 64 (interquartile range [IQR], 53 to 72 years), and 133 of the patients (70.6%) were male. While there were no significant differences in clinical characteristics and outcomes between the two groups, patients with qacA/B-positive ST5-MRSA-II had a tendency to have bacteremia earlier, to have undergone recent surgery, and suffer septic shock more than those with qacA/B-negative ST5-MRSA-II.
Microbiological and genotypic characteristics of the qacA/B-positive ST5-MRSA-II isolates
The most common spa type of ST5-MRSA-II was t2460 (59.4%), followed by t9353 (14.4%) and t002 (8.1%). As shown in Table 2, t9353 was significantly associated with qacA/B positivity (18.5% vs 2.4%, P = 0.01), and t002 with qacA/B negativity (1.7% vs. 26.8%, P < 0.001). The qacA/B-positive isolates were more often resistant to fusidic acid (93.3% vs. 73.2%, P = 0.001) and gentamicin (95.8% vs. 43.9%, P < 0.001) than were the qacA/B-negative isolates. The association of gentamicin resistance with qacA/B positivity was pronounced in the t2460 isolates (100% [74/74] vs. 33.3% [7/21], P < 0.001). In addition, the qacA/B-positive isolates had significantly higher chlorhexidine MICs than the negative ones (P < 0.001) (Table 2 and Supplemental Fig. S1). The chlorhexidine MIC50 and MIC90 values were 4 mg/L and 4 mg/L in qacA/B-positive isolates, whereas those were 1 mg/L and 2 mg/L, respectively, in qacA/B-negative isolates. However, mupirocin resistance and presence of the mupA gene were not associated with qacA/B positivity. Presence of the smr gene, another chlorhexidine tolerance gene, was significantly associated with qacA/B negativity (P = 0.04).
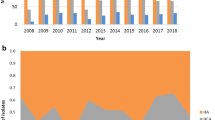
Changes in microbiological and genotypic characteristics of ST5-MRSA-II blood isolates. (a) The proportion of qacA/B-positive isolates in ST5-MRSA-II and geometric means of chlorhexidine (CHX) MICs decreased over time (P < 0.001 for trend for both). (b) Changes in spa types. The proportion of qacA/B-positivity in t2460 decreased over time (P < 0.001 for trend).
The proportion of qacA/B-positive isolates decreased from 88.1 to 50% over time (P < 0.001 for trend) (Fig. 1a), mainly due to the lowered qacA/B-positive rate in t2460 (P < 0.001 for trend) (Fig. 1b).
Risk factors for qacA/B-positive ST5-MRSA-II isolates
Univariate variables with P values of less than 0.1 in Tables 1 and 2 were included in a logistic regression model to identify independent risk factors for qacA/B-positive MRSA. Because the t9353 spa type and fusidic acid resistance were highly correlated with gentamicin resistance, we retained gentamicin resistance only. Also, we included length of ICU stay before bacteremia rather than length of hospital stay before bacteremia because the former is a more clinically meaningful variable in ICU-acquired MRSA bacteremia. Thus, included in the multivariate regression modeling were length of ICU stay before bacteremia (per 10 days), recent surgery, septic shock, gentamicin resistance, non-t002 spa type, and agr dysfunction. The independent risk factors associated with qacA/B positivity were found to be septic shock (adjusted odds ratio [aOR], 4.85; 95% confidence interval [CI], 1.09 to 21.51), gentamicin resistance (aOR, 74.43; 95% CI, 19.19 to 288.67), and non-t002 spa type (aOR, 74.12; 95% CI, 10.99 to 499.60) (Table 3).
Discussion
In this study we found that the chlorhexidine tolerance genes qacA/B and smr were mainly present in the specific hospital MRSA clone, ST5-MRSA-II, after the implementation of chlorhexidine use in a tertiary hospital from South Korea. The qacA/B gene was independently associated with septic shock, gentamicin resistance, and non-t002 spa type (especially spa type t9353) in a homogenous ST5-MRSA-II background. This implies that the qacA/B coexists well with the t9353 spa type but not the t002 spa type. During the study period, qacA/B positivity decreased significantly in ST5-MRSA-II mainly due to a reduction in qacA/B-positive t2460 isolates despite the increasing use of chlorhexidine in this hospital since 2010.
Previous clinical and microbiological studies of qacA/B and smr had been performed on strains with heterogeneous genotypic and clinical backgrounds, including both community and healthcare S. aureus strains6,13,33,34. Therefore, the proportion of qacA/B-positive isolates in the present study, which included only ICU-acquired ST5-MRSA-II bacteremia, was higher (74.4%) than in previous studies. Interestingly, the proportion of smr-positive isolates was very low (3.8%), in contrast with previous studies13,33,34. In another study conducted in South Korea6, the proportion of smr-positive MRSA was also low (2.8%), and qacA/B positivity was closely associated with ST5-MRSA, the dominant and generally agr dysfunctional hospital clone16. ST72-MRSA, a community-associated MRSA clone in South Korea that is usually agr functional16, was also significantly associated with qacA/B negativity in that study. Therefore, factors that were found to be independently associated with qacA/B positivity in that study, such as agr dysfunction, nosocomial infection, and previous antibiotic use, could have been reflections of the characteristics of the specific MLST, ST5-MRSA, because all MRSA genotypes were included. The clonal association of qacA/B with hospital MRSA clones such as ST239, ST5, and ST241 have been found in Asia14,35. Therefore, the clonal association between qacA/B and ST5-MRSA-II isolates in present study could be influenced by the dominant regional hospital MRSA clone in South Korea.
In contrast to other studies that showed the increased proportion of qacA/B positivity after the widespread use of chlorhexidine in hospitals8,9,10, the present study revealed that qacA/B positivity in ST5-MRSA-II decreased over time. This finding could be influenced by the decreased proportion of qacA/B-positive rate in t2460, which was main spa type in ST5-MRSA-II.
Several studies have shown that healthcare exposure and underlying medical condition are independently associated with qacA/B positivity in S. aureus isolates6,13,34. In the present study, we found that septic shock, gentamicin resistance, and fusidic acid resistance were also closely associated with qacA/B positivity in nosocomial ST5-MRSA-II blood isolates in a homogenous MRSA genotypic background. The qacA/B genes are usually carried by large plasmids that encode efflux pumps, and some of these plasmids contain other antibiotic resistance genes11,12,36. This fact should influence the likelihood of co-resistance to other antimicrobials such as gentamicin and fusidic acid. In addition, the presence of virulence genes on certain qac plasmids could affect the severity of the illness of patients with qacA/B-positive MRSA bacteremia37. Further detailed analysis of our qac plasmids would clarify these associations.
Interestingly, the non-t002 spa type was significantly associated with qacA/B positivity in ST5-MRSA-II isolates. t2460 and t9353 are the major spa types of MRSA in South Korea6,38. While a previous study showed that t2460 was associated with the qacA/B genes in MRSA strains as a whole6, we found that only t9353 was strongly associated with qacA/B positivity in a homogenous ST5-MRSA-II background. As shown in Fig. 1b, there has been no outbreak of qacA/B-positive t9353, and the proportion of qacA/B-positivity in t2460 strains has decreased over time (P < 0.001 for trend). Potential hypotheses for such findings are that t9353 spa type might maintain the qacA/B gene efficiently, and t2460 may have a tendency to lose a plasmid carrying qacA/B.
This study has several limitations. First, it was conducted in a single tertiary-care hospital and included exclusively MRSA bacteremia occurring in ICUs. Hence, further studies are needed to confirm our results in other hospital settings. Second, because almost all the patients had a recent history of antibiotic use, it was difficult to determine whether antibiotic use exerts selective pressure for chlorhexidine tolerance. Third, despite the proportion of available MRSA blood isolates tested was high (183/238, 77%), these results may not reflect the true prevalence of the genotypes found. Therefore, caution is needed in generalizing the results to other situations.
In conclusion, the qacA/B and smr antiseptic tolerance genes were mainly encountered in ST5-MRSA-II, the dominant hospital MRSA clone, after the implementation of chlorhexidine use in a tertiary hospital from South Korea since 2010. Septic shock, gentamicin resistance, and non-t002 spa type were independent predictors of the presence of qacA/B in ST5-MRSA-II blood isolates. Continuous surveillance for the qac genes and molecular characterization of the corresponding plasmids are needed to understand the role of these genes in MRSA epidemiology.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files. The genotypic data have been deposited to the figshare database at https://doi.org/10.6084/m9.figshare.19114976.v2.
References
Souli, M. et al. Changing characteristics of staphylococcus aureus bacteremia: results from a 21-year, prospective, longitudinal study. Clin Infect Dis 69, 1868–1877. https://doi.org/10.1093/cid/ciz112 (2019).
Choi, S. H. et al. A longitudinal study of adult patients with staphylococcus aureus bacteremia over 11 years in Korea. J Korean Med Sci 36, e104. https://doi.org/10.3346/jkms.2021.36.e104 (2021).
Huang, S. S. et al. Decolonization to reduce postdischarge infection risk among MRSA carriers. N Engl J Med 380, 638–650. https://doi.org/10.1056/NEJMoa1716771 (2019).
Jain, R. et al. Veterans AFfairs initiative to prevent methicillin-resistant Staphylococcus aureus infections. N Engl J Med 364, 1419–1430. https://doi.org/10.1056/NEJMoa1007474 (2011).
Frost, S. A. et al. Chlorhexidine bathing and health care-associated infections among adult intensive care patients: a systematic review and meta-analysis. Crit Care 20, 379–384. https://doi.org/10.1186/s13054-016-1553-5 (2016).
Hong, S. I. et al. Clinical and molecular characteristics of qacA- and qacB-positive methicillin-resistant staphylococcus aureus causing bloodstream infections. Antimicrob Agents Chemother 63, e02157-e2118. https://doi.org/10.1128/aac.02157-18 (2019).
Otter, J. A. et al. Selection for qacA carriage in CC22, but not CC30, methicillin-resistant Staphylococcus aureus bloodstream infection isolates during a successful institutional infection control programme. J Antimicrob Chemother 68, 992–999. https://doi.org/10.1093/jac/dks500 (2013).
Warren, D. K. et al. Prevalence of qacA/B genes and mupirocin resistance among methicillin-resistant Staphylococcus aureus (MRSA) isolates in the setting of chlorhexidine bathing without mupirocin. Infect Control Hosp Epidemiol 37, 590–597. https://doi.org/10.1017/ice.2016.1 (2016).
Suwantarat, N. et al. High prevalence of reduced chlorhexidine susceptibility in organisms causing central line-associated bloodstream infections. Infect Control Hosp Epidemiol 35, 1183–1186. https://doi.org/10.1086/677628 (2014).
McNeil, J. C., Hulten, K. G., Kaplan, S. L., Mahoney, D. H. & Mason, E. O. Staphylococcus aureus infections in pediatric oncology patients: high rates of antimicrobial resistance, antiseptic tolerance and complications. Pediatr Infect Dis J 32, 124–128. https://doi.org/10.1097/INF.0b013e318271c4e0 (2013).
Berg, T. et al. Complete nucleotide sequence of pSK41: evolution of staphylococcal conjugative multiresistance plasmids. J Bacteriol 180, 4350–4359. https://doi.org/10.1128/jb.180.17.4350-4359.1998 (1998).
Sidhu, M. S., Heir, E., Sørum, H. & Holck, A. Genetic linkage between resistance to quaternary ammonium compounds and beta-lactam antibiotics in food-related Staphylococcus spp. Microb Drug Resist 7, 363–371. https://doi.org/10.1089/10766290152773374 (2001).
McNeil, J. C. et al. Clinical and molecular features of decreased chlorhexidine susceptibility among nosocomial staphylococcus aureus isolates at Texas Children’s Hospital. Antimicrob Agents Chemother 60, 1121–1128. https://doi.org/10.1128/aac.02011-15 (2016).
Lu, Z. et al. Characteristics of qacA/B-positive Staphylococcus aureus isolated from patients and a hospital environment in China. J Antimicrob Chemother 70, 653–657. https://doi.org/10.1093/jac/dku456 (2015).
Park, K. H. et al. Community-associated MRSA strain ST72-SCCmecIV causing bloodstream infections: clinical outcomes and bacterial virulence factors. J Antimicrob Chemother 70, 1185–1192. https://doi.org/10.1093/jac/dku475 (2015).
Chong, Y. P. et al. Accessory gene regulator (agr) dysfunction in Staphylococcus aureus bloodstream isolates from South Korean patients. Antimicrob Agents Chemother 57, 1509–1512. https://doi.org/10.1128/aac.01260-12 (2013).
Kim, H. et al. Decreased Incidence of methicillin-resistant Staphylococcus aureus bacteremia in intensive care units: a 10-year clinical, microbiological, and genotypic analysis in a tertiary hospital. Antimicrob Agents Chemother 64, e01082-e1020. https://doi.org/10.1128/aac.01082-20 (2020).
McCabe, W. R. & Jackson, G. G. Gram-negative bacteremia: I. etiology and ecology. Arch. Internal Med. 110, 847–855. https://doi.org/10.1001/archinte.1962.03620240029006 (1962).
Charlson, M. E., Pompei, P., Ales, K. L. & MacKenzie, C. R. A new method of classifying prognostic comorbidity in longitudinal studies: development and validation. J Chronic Dis 40, 373–383. https://doi.org/10.1016/0021-9681(87)90171-8 (1987).
Chow, J. W. et al. Enterobacter bacteremia: clinical features and emergence of antibiotic resistance during therapy. Ann Intern Med 115, 585–590. https://doi.org/10.7326/0003-4819-115-8-585 (1991).
Garner, J. S., Jarvis, W. R., Emori, T. G., Horan, T. C. & Hughes, J. M. CDC definitions for nosocomial infections, 1988. Am J Infect Control 16, 128–140. https://doi.org/10.1016/0196-6553(88)90053-3 (1988).
Mermel, L. A. et al. Clinical practice guidelines for the diagnosis and management of intravascular catheter-related infection: 2009 Update by the Infectious Diseases Society of America. Clin Infect Dis 49, 1–45. https://doi.org/10.1086/599376 (2009).
Weinstein, M. P., Lewis II, J. S., Bobenchik, A. M., Campeau, S. Performance Standards for Antimicrobial Susceptibility Testing, 31st Edition, M100. (Clinical and Laboratory Standards Institute, 2021).
Wootton, M. et al. A modified population analysis profile (PAP) method to detect hetero-resistance to vancomycin in Staphylococcus aureus in a UK hospital. J Antimicrob Chemother 47, 399–403. https://doi.org/10.1093/jac/47.4.399 (2001).
Campbell, S. J. et al. Genotypic characteristics of Staphylococcus aureus isolates from a multinational trial of complicated skin and skin structure infections. J Clin Microbiol 46, 678–684. https://doi.org/10.1128/jcm.01822-07 (2008).
Enright, M. C., Day, N. P., Davies, C. E., Peacock, S. J. & Spratt, B. G. Multilocus sequence typing for characterization of methicillin-resistant and methicillin-susceptible clones of Staphylococcus aureus. J Clin Microbiol 38, 1008–1015. https://doi.org/10.1128/jcm.38.3.1008-1015.2000 (2000).
Shopsin, B. et al. Evaluation of protein A gene polymorphic region DNA sequencing for typing of Staphylococcus aureus strains. J Clin Microbiol 37, 3556–3563. https://doi.org/10.1128/jcm.37.11.3556-3563.1999 (1999).
Shopsin, B. et al. Prevalence of agr specificity groups among Staphylococcus aureus strains colonizing children and their guardians. J Clin Microbiol 41, 456–459. https://doi.org/10.1128/jcm.41.1.456-459.2003 (2003).
Traber, K. E. et al. agr function in clinical Staphylococcus aureus isolates. Microbiology (Reading) 154, 2265–2274. https://doi.org/10.1099/mic.0.2007/011874-0 (2008).
Patel, J. B., Cockerill III, F. R., Bradford, P. A. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically; approved standard, 10th edition. Clinical and Laboratory Standards Institute (2015).
Swenson, J. M. et al. Multicenter study to determine disk diffusion and broth microdilution criteria for prediction of high- and low-level mupirocin resistance in Staphylococcus aureus. J Clin Microbiol 48, 2469–2475. https://doi.org/10.1128/jcm.00340-10 (2010).
Pérez-Roth, E., Claverie-Martín, F., Villar, J. & Méndez-Alvarez, S. Multiplex PCR for simultaneous identification of Staphylococcus aureus and detection of methicillin and mupirocin resistance. J Clin Microbiol 39, 4037–4041. https://doi.org/10.1128/jcm.39.11.4037-4041.2001 (2001).
Ho, C. M. et al. High rate of qacA- and qacB-positive methicillin-resistant Staphylococcus aureus isolates from chlorhexidine-impregnated catheter-related bloodstream infections. Antimicrob Agents Chemother 56, 5693–5697. https://doi.org/10.1128/aac.00761-12 (2012).
McNeil, J. C., Hultén, K. G., Mason, E. O. & Kaplan, S. L. Impact of health care exposure on genotypic antiseptic tolerance in Staphylococcus aureus infections in a pediatric population. Antimicrob Agents Chemother 61, e00223-e217. https://doi.org/10.1128/aac.00223-17 (2017).
Sheng, W. H. et al. Epidemiology and susceptibilities of methicillin-resistant Staphylococcus aureus in Taiwan: emphasis on chlorhexidine susceptibility. Diagn Microbiol Infect Dis 63, 309–313. https://doi.org/10.1016/j.diagmicrobio.2008.11.014 (2009).
Littlejohn, T. G., DiBerardino, D., Messerotti, L. J., Spiers, S. J. & Skurray, R. A. Structure and evolution of a family of genes encoding antiseptic and disinfectant resistance in Staphylococcus aureus. Gene 101, 59–66. https://doi.org/10.1016/0378-1119(91)90224-y (1991).
McCarthy, A. J. & Lindsay, J. A. The distribution of plasmids that carry virulence and resistance genes in Staphylococcus aureus is lineage associated. BMC Microbiol 12, 104. https://doi.org/10.1186/1471-2180-12-104 (2012).
Kang, C. K. et al. agr dysfunction affects staphylococcal cassette chromosome mec type-dependent clinical outcomes in methicillin-resistant Staphylococcus aureus bacteremia. Antimicrob Agents Chemother 59, 3125–3132. https://doi.org/10.1128/aac.04962-14 (2015).
Acknowledgements
We thank Su-jin Park, Kyoung-A Jhang, and Mi Young Kim for supporting the data collection.
Funding
This work was supported by a National Research Foundation of Korea (NRF) grant funded by the Korean government (MSIT) (Grant Number: NRF-2020R1A2C1101525).
Author information
Authors and Affiliations
Contributions
Conceptualization: H.K., Y.P.C. Methodology: H.K., Y.P.C. Data curation: H.K., S.P., H.S., H.C., E.S.K. Software, formal analysis: H.K. Writing—original draft: H.K. Writing—review & editing: Y.P.C. Supervision: H.S., M.-N.K., S.B., J.J., M.J.K., S.-H.K., S.-O.L., S.-H.C., Y.S.K. Funding acquisition: Y.P.C.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kim, H., Park, S., Seo, H. et al. Clinical impact of and microbiological risk factors for qacA/B positivity in ICU-acquired ST5-methicillin-resistant SCCmec type II Staphylococcus aureus bacteremia. Sci Rep 12, 11413 (2022). https://doi.org/10.1038/s41598-022-15546-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-15546-3
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.