Abstract
The determination of trypsin in the human real sample is a routine medical investigation to assess the pancreatic disease. Herein, we fabricated an interferometric reflectance spectroscopy based biosensor for the determination trypsin. For this purpose, urease and fluorescein 5(6)-isothiocyanate (FLITC) were immobilized on the nanoporous anodic alumina (NAA). The operation principle of the proposed biosensor is based on the change in the pH of the solution during the reaction of urease and urea and therefore change in the light-absorbing ability of FLITC in the presence of trypsin. The reaction of the urease enzyme with urea increased the pH of the solution because of producing ammonia. This increase in the pH of solution increased the light-absorbing ability of the immobilized FLITC on NAA and therefore the intensity of the reflected light from the NAA to the charge-coupled device detector decreased. In the presence of trypsin, the catalytic activity of immobilized urease on NAA decreased. This decrease in the activity of urease enzyme consequent on the decrease in the amount of the generated ammonia. Therefore, the immobilized FLITC on the NAA did not absorb more light and consciously, the intensity of the light reflected light into the detector increased. The proposed biosensor exhibited a good response to the concentration of trypsin in the range of 0.25–20 μg.mL−1 with the limit of detection of 0.06 μg.mL−1.
Similar content being viewed by others
Introduction
Millions of people in the world suffer from some form of pancreatic diseases1. According to world health statistics reports, pancreatic diseases id associated with a tremendous economic burden2,3. Due to reduce the economic losses caused by this disease and also to improve the public health, the fabrication of a device to diagnose this disease is important for every government. The routine laboratory test for pancreatic disease diagnosis is the determination of trypsin the human real sample. Trypsin is a serine protease with a potential role in pancreatic cancer that can cleavage the peptides on the carbonyl side of arginine and lysine amino acids4. In the acute pancreatitis patient that is a fatal disease, the concentration of trypsin reaches up to 84.4 µg.mL−1 in the urine sample that is so higher than its normal rage (115–350 ng.mL−1)5. Therefore, it is necessary to develop a simple, selective, sensitive, and low-cost sensor for trypsin detection. However, most of them are complicated, low sensitivity, time-consuming, and expensive method6,7,8,9,10. To overcome these problems, biosensors become more interested to detect trypsin11,12.
The biosensor is an analytical device which includes a transducer and a biomolecule (such as the enzymes, antibodies, and aptamers)13,14,15. Among the various biosensors that have been reported for the sensing of bio-targets, the optical biosensors are more interested because of their high sensitivity, small and lightweight, remote sensing ability, immunity to electromagnetic interference, capability for monitoring a wide range of chemical and physical parameters, and reliable operation16,17,18,19.
Up to now, several optical biosensors have been fabricated for the determination of trypsin20,21,22,23,24. Among them, the fluorescence biosensor based on quantum dots is more interested25,26,27,28,29,30. However, the fluorescence biosensor based on quantum dots are expensive methods and can not be suited for point-of-care testing. Therefore, the fabrication of a cheap but high sensitive optical biosensor for the determination of trypsin is needed.
To the best of our knowledge, the use of the interferometric reflectance spectroscopy (IRS) method for the fabrication of biosensor to detect trypsin has not been reported yet. The IRS is a contactless optical device that can be used for the determination of the various targets19,31,32. In this method, the beams of white light interfere with a reflective thin film then the partial beams of the light reflect the charge-coupled device (CCD) detector. Macroporous silicon (MPS) and nanoporous anodic alumina (NAA) are two common reflective thin films for the fabrication of the IRS based biosensor19,33,34,35. NAA has a lot of advantages such as high surface area, biocompatibility, easy functional ability, versatile, durable low-cost, and three-dimensional nanostructure platform36,37,38,39. According to the previous report, trypsin can cleave the amino acid chain from the carboxyl side (C-terminal side) of the amino acids lysine and arginine40. Therefore, the use of an enzyme that lysine and arginine amino acids play a major role in the function and structure of the enzyme and study the effect of trypsin on the catalytic activity of this enzyme, can be a novel method for the fabrication of biosensor for trypsin detection. Urease enzyme is a metalloenzyme that can catalyze the reaction of urea molecule into carbon dioxide and ammonia selectively. The amino acids lysine and arginine are two amino acids exist on the structure of urease enzyme. One of lysine amino acid bridges two Ni2+ ions (co-factor) to each other, playing a major role in the catalytic activity of the urease enzyme41. Up to now, several optical biosensors have been reported by using urease enzyme to detect the urea in the sample42,43. In these kinds of the optical-based biosensor, a pH-sensitive optical probe molecule such as fluorescein was used to study the generated ammonia from urea. In this paper, for the first time, an IRS method was used for the fabrication of an optical biosensor to detect trypsin as a model of protease proteins. To fabricate the IRS sensor, NAA, urease enzyme and fluorescein 5(6)-isothiocyanate (FLITC) were used as a reflective thin film, an enzyme that its activity can be changed by to protease proteins and an optical probe molecule, respectively. Compare with the fluorescence biosensor based on quantum dots, the proposed IRS based biosensor exhibited high analytical performance in terms of selectivity, sensitivity, stability, linear range and limit of detection in nearly every case.
Experimental Section
Reagents and chemicals
All chemicals were of analytical reagent grade and used without further purification. Double deionized (DI) water (18.6 MΩ) was used throughout. Aluminum (Al) discs of 15 mm diameter were obtained from Goodfellow. (3-aminopropyl) trimethoxysilane (3-APTES), oxalic acid, glutaraldehyde (GLA), phosphoric acid (H3PO4), sodium acetate (NaAC), chromium (VI) oxide (H2CrO4), perchloric acid (HClO4), cysteine, dopamine, glucose, nicotinamide adenine dinucleotide trypsin, urea, urease enzyme and were obtained from Sigma-Aldrich.
Apparatus
Scanning electron microscopy (SEM) was performed with an FEI Quanta 600. Infrared spectra were obtained using a JASCO FT/IR-680 Plus Fourier transform infrared (FTIR) spectrometer. Raman scattering was performed on a Renishaw’s inVia Raman spectrometer using 514 nm laser source. The interferometric reflectance spectra were recorded using an AvaSpec-ULS3648 fiber optic spectrometer. EMITech K575X sputter coater was used to deposit 10 nm thick gold layer under vacuum at 30 mA for 1 min.
The fabrication of NAA-NH-GLA-urease-FLITC
NAA was prepared based on 2 steps electrochemical anodization process in oxalic acid solution under stirring condition9,31,44,45. The fabrication process is denoted in the electronic supporting material. The electrochemically fabricated NAA was then immersed into a 3-APTES solution (1% in ethanol/H2O 3:1) for 30 min under a nitrogen gas atmosphere to introduce amine groups inside of the pore of NAA. NAA-NH2 was then rinsed with deionized water for 1 min and dried under nitrogen gas flow. After that, the NAA-NH2 was dried under a nitrogen atmosphere at 100 °C for 2 h. After then, NAA-NH2 was immersed in a 2.5% GLA in phosphate buffer (PB) and stirred for 1 h. NAA-NH-GLA was then rinsed with water and dried under nitrogen gas flow to remove the unattached GLA. GLA is a crosslinker that can be used for the immobilization of enzymes to fabricate biosensors46,47. Subsequently, The NAA-NH-GLA was immersed in a urease solution (20 mg.mL−1, 0.1 M PB and pH 7.4) for 5 h under a stirring condition at 4 °C to immobilize urease enzyme on the NAA-NH-GLA. After rinsing with 0.1 M PB, the NAA-GLA-NH2-urease was immersed in a fluorescein iso-thiocyanate solution (1 mg.mL−1, 0.1 M PB and pH 8.4) for overnight at 4 °C to immobilize FLITC on the NAA-NH-GLA-urease. According to the previous report, the iso-thiocyanate (ITC) functional group can interact with sulfhydryl groups (–SH), amine (–NH2), and hydroxyl (–OH) functional groups48. Cysteine and lysine amino acids that have –SH and –NH2 functional groups, respectively and exist in the structure of the urease enzyme49. Therefore, the ITC function group of FLITC can interact with urease enzyme and immobilize on the NAA. After that, the NAA-GLA-NH2-urease-FLITC was rinsed with water several times to wash away any unattached FLITC. The NAA-urease-FLITC was stored at 4 °C in a refrigerator when not in use. The schematic representation for the fabrication of an IRS based biosensor is shown in Fig. 1.
The sensing process of trypsin
The determination of trypsin was done in the three steps:
(1) 3 mM urea solution (0.1 M NaAC, pH 6.4) was transferred to cell reaction by using a peristaltic pump. After 30 seconds, the pump was turned off for 4 min and the signal of the biosensor was recorded after that. Before starting the second step, the biosensor was washed by pumping NaAC solution (0.1 M, pH 6.4) for 1 min. (2) The different concentration of trypsin solution (0.1 M NaAC, pH 9.4) was pumped to the measurement cell for 45 min. During this period of time, trypsin cleaved the urease enzyme form the arginine and L-lysine sites39. Then, NAA-NH-GLA-urease-FLITC was washed by NaAC solution (0.1 M, pH 6.4) for 5 min to remove the cleaved parts of the urease enzyme. (3) The urea solution was transferred with the same condition in the first step (0.1 M NaAC, pH 6.4, 3 mM urea, reaction time 4 min). The analyzed data showed that the recorded signal of the biosensor in the third step was lower than in the first step. The reasonable explanation is that the cleaved urease by trypsin could not catalyze urea as much as the first step and then the amount of the generated ammonia was lower than before. Consequently, the immobilized FLITC could not absorb the white light in this low pH solution as much as in the first step. Therefore, the intensity of the white light was detected by the CCD detector was higher than in the first step.
Results and Discussion
The surface morphology of the prepared NAA
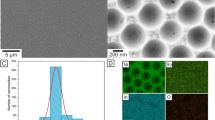
The morphological characterization of the NAA was examined by SEM (Fig. 2A–F). As shown in these figures, the surface of the NAA has a hexagonal multi-pore structure. The average pore diameter and pore length of NAA are 54 nm (with a standard deviation of 2.1 nm) and 5.17 µm (with a standard deviation of 0.07 µm), respectively. According to the previous report50, the best Fabry-Perot fringe spectrum of the NAA would be obtained as the pore length/pore diameter aspect ratio was around 90 to100. Since the aspect ratio of the fabricated NAA for the determination of trypsin was 95.7, therefore the biosensor signal would be high.
Also, the pore diameter of the fabricated NAA is bigger than trypsin enzyme (3.8 × 3.8 × 3.8 nm)51, therefore, trypsin enzyme can get in the nanopore of NAA easily.
Characterization of NAA-NH-GLA-urease-FLITC
FTIR and Raman spectroscopy are commonly characterization techniques of materials.
FTIR spectrum of NAA (Fig. 3A (curve a)) exhibited a band at 3408 cm−1 due to the O–H stretching mode, and two absorption bands at 1159 cm−1 and 955 cm−1 due to the Al-O-H and the Al-O modes of boehmite, respectively52. After the immobilization of urease-FLITC on NAA, the FTIR spectrum changed noticeably (Fig. 3A (curve b)).
As shown in Fig. 3A (curve b), an absorption band at 736 cm−1 due to the –C=S stretching of the FLITC is clearly seen. The other characteristic absorption bands a band at 1058 cm−1 due to the −Al–O–Si, a band at 1394 cm−1 due to the –C–C stretching, a band at 1991 cm−1 due the –C=C=N (from L-histidine of urease enzyme), a strong band at 2112 cm−1 due to –C=N (from L-histidine of urease enzyme), a strong band at 2324 cm−1 due to the –C–H stretching, a band 2850 cm−1 due to the −CH3, and a band at 2920 cm−1 due to the –CH2 are also clearly seen in this figure53,54.
Figure 3B shows a typical Raman spectrum of NAA (a) NAA-NH-GLA-urease-FLITC (b). As can be seen, NAA has not any band in this wavelength. However, several bands were observed in Raman spectrum of NAA-NH-GLA-urease-FLITC. The band in the range of 1140 cm−1 are attributed to the aromatic –C–H in-plane bend, a band at 1250 cm−1 due to the –C–O–C stretching mode and a band at 1414 cm−1 due to the –C–C stretching of quinone and symmetric carboxylic (–COO−) stretch mode from FLITC and a band at 1600 cm−1 due to the amide II stretching, and a band at 1723 cm−1 due to amide I of urease enzyme53,54. These results demonstrate that FLITC and urease enzyme was immobilized on the NAA, successfully.
The optical characterization of the biosensor
Figure 4A shows the typical the interference spectra of NAA-NH-GLA-urease (A), NAA-NH-GLA-urease-FLITC (B) in the absence (a) and presence (b) of 3 mM urea (0.1 M NaAC, pH 6.4). The spectrums (c) in these figures relate to the difference of spectrum (a) and (b) that obtained from the subtraction of spectrum (a) from the spectrum (b). As can be seen, the sensitivity of NAA-NH-GLA-urease to 3 mM urea is low. However, after the immobilization of FLITC as a pH photo-probe molecule, the response of NAA-NH-GLA-urease-FLITC is dramatically higher than NAA-NH-GLA-urease. This difference in the signal of the NAA-NH-GLA-urease and NAA-NH-GLA-urease-FLITC to 3 mM urea was shown in Fig. 4C. The reasonable explanation is that the reaction of the urease enzyme with urea increased the pH of the solution because of producing ammonia. This increase in the pH of solution increased the light-absorbing ability of the immobilized FLITC on NAA and therefore the intensity of the reflected light from the NAA to the charge-coupled device (CCD) detector decreased.
(A) IRS of the NAA-urease and (B) NAA-urease-FLITC in the absence (a) and presence (b) of 3.0 mM urea and (c) the difference between two interference spectrums. (C) ∆IRS of the NAA-urease (a) and NAA-urease-FLITC (b) in a solution (0.1 M NaAC, pH = 6.4) containing 3 mM urea. (D) IRS of the NAA-urease-FLITC in the absence (a) and presence (b) of 20.0 μg.mL−1 trypsin and (c) the difference between two interference spectrums.
Also, Fig. 4D shows the IRS of the NAA-urease-FLITC in the absence (a) and presence (b) of 20.0 μg.mL−1 trypsin and (c) the difference between two interference spectrums. As can be seen, the intensity of the IRS increased in the presence of trypsin due to urease enzyme digestion and deactivation, the pH of the solution during the reaction of urease and urea did not increase and then FLITC did not absorb the interfered with light. Hence, the intensity of the reflected light to the CCD detector increased.
Optimization of effective parameters on the response of the biosensor
The pH of the solution and incubation time were optimized. The respective figures are given in the electronic supporting material (Fig. S1). The following experimental conditions were found to give the best results: (a) pH of 6.4 the solution, (b) interaction time of 4 min for urea (3 mM) and (c) pH of 9.4 the solution (d) 45 min for trypsin (3 μg.mL−1). As can be seen, the optimum pH of the solution for the determination of urea is 6.4. The reasonable reason to explain this phenomenon is, the activity of urease enzyme decreased in the pH less than 6.4 and the change in the absorbance intensity of FLITC from pH 6.4 to alkaline pH is high. Therefore, the immobilized FLITC absorbed more white light and the response of NAA-NH-GLA-urease-FLITC to urea in pH 6.4 solutions is higher than the other pH solution. Also, the average pH of the urine sample is about 6. Therefore, pH 6.4 was chosen as the optimum pH of the solution. Since the proposed biosensor is a pH-based biosensor, the determination of the analyte should be done at the solution that its pH buffering capacity is lower than 6.4. Since the pH buffering capacity of phosphate solution is in the range of pH 6.2–8.2, NaAC was used as the electrolyte of solution that its pH buffering capacity is in the range of 3.8–5.8. Figure S1B shows the effect of time on the response of biosensor to urea. As can be seen, the response of biosensor to urea increased rapidly with increasing time up to 4 min and remained unchanged at longer reaction times, suggesting that the reaction has reached the saturation level. Also, Figure S1C shows the effect of pH solution to the response of the proposed biosensor in the presence of 2 μg.mL−1 trypsin. As shown, the trypsin has a high activity to cleave the urease enzyme in the pH 9.4. The result has a good agreement with the previous report55. The reasonable reason to explain this phenomenon is the structure of the enzyme is changed and the active site is distorted in the higher than 9.5 and pH solution and the activity of trypsin decreased. Figure S1D shows the effect of time on the response of biosensor to trypsin. As can be seen, the response of biosensor to trypsin increased rapidly with increasing time up to 45 min and then remained unchanged at longer reaction times, suggesting that the reaction has reached the saturation level.
Detection of trypsin
Under the optimized experimental conditions, the proposed biosensor was employed for the determination of trypsin (Fig. 5). It can be seen that the signal increase with increasing urea. As indicated in the inset of Fig. 5, a linear relationship between the logarithm of trypsin concentration (Ctryp) and the intensity of the IRS was observed in the linear dynamic range (LDR) from 0.25–20.0 µg.mL−1 with a regression equation of ∆Intensity = 115.87 log Ctryp (µg.mL−1) + 229.9 and a correlation coefficient of 0.9971. The limit of detection (LOD) was 0.06 µg.mL−1 (at 3σ/S). The LOD of the proposed method is superior to the previously reported IRS based sensor for trypsin sensing (25 µg.mL−1)56.
IRS of the NAA-urease-FLITC in a solution (0.1 M NaAC, pH = 6.4, 3 mM urea) containing different amounts of trypsin (0.25, 0.5, 1.0, 2.0, 4.0, 6.0, 10.0, and 20.0 μg.mL−1 from outer to inner). Inset: the corresponding calibration curve of the proposed IRS based sensor. Error bars represent standard deviations of four repeated experiments.
Table 1 compares the LDR and the LOD of the proposed IRS based biosensor for trypsin determination with some of the optical biosensing methods such as nanoclusters26,27 and quantum dots25,28. As can be seen, although the LOD of CuNCs-protamine is lower than the proposed IRS based biosensor, however, it´s LDR is not wider than it. In addition, the fabrication and then purification of the functionalized nanoclusters and quantum dots are a very complicated, costly and time-consuming process.
Furthermore, the proposed IRS based biosensor is not only a single-step measuring method but also is a remote sensing, lightweight, and low-cost device. Therefore, the analytical performance of the proposed optical biosensor in most cases is better than the optical biosensors, by considering all these aspects.
The reproducibility stability and selectivity of the NAA-NH-GLA-urease-FLITC
The NAA-NH-GLA-urease-FLITC exhibited good reproducibility in the detection of trypsin. The relative standard deviation (RSD) for trypsin was founded 6.9% for over 5 repeated measurements of 3.0 µg.mL−1 trypsin. When not in use, the biosensor was stored at 4 °C in a refrigerator for 3 weeks, its signal of biosensor decreased by approximately 8.1%. The effect interfering compounds were also studied on the response of biosensor (Fig. S2). No interference was observed with 0.1 mg.mL−1 of cysteine, dopamine, glucose, nicotinamide adenine dinucleotide, and common cations and anions (0.15 mg.mL−1 of Na+, Al+3, K+, Cl-, NO3). It clearly proves that the biosensor has high selectivity trypsin.
The Inhibition efficiency of the biosensor by trypsin also studied (Fig. S3). The IC50 value was obtained from the fitting curve was 12.4 µg.mL−1.
Analytical application of the biosensor
The proposed biosensor was also employed for the determination of trypsin in the urine. Since the working range of the proposed biosensor was up to 20.0 µg.mL−1 and the concentration of trypsin in the patient’s urine is high (84.4 µg.mL−1), therefore, the urine sample should be diluted to get trypsin concentration into calibration range. For this purpose, 18.2 µL of a human urine sample (urea concentration was 192.0 mM) was diluted with 981.8 µL NaAC (0.1 M, pH 6.4) to decrease the concentration of the urea to 3.5 mM. Then, 858.1 µL of the diluted human urine sample was added to 141.9 µL of trypsin (20.0 µg.mL−1, 0.1 M NaAC, pH 6.4). The concentration of urea and trypsin reached 3.0 mM and 2.0 µg.mL−1, respectively. Finally, the sample was pumped to the analytical cell to determine. The obtained recoveries for 2.0 µg.mL−1 trypsin was 86%.
Conclusion
This study introduced a biosensor based on the IRS for the determination of trypsin. To fabricate this biosensor, the urease enzyme and FLITC were immobilized on the pores of NAA as a urease enzyme and FLITC were used as an enzyme that its activity can be changed by to protease proteins and an optical probe molecule, respectively. The immobilized urease enzyme catalyzed the hydrolysis of urea to ammonia, letting to the pH-induced change in the light-absorbing ability of FLITC. The catalytic ability of the immobilized urease enzyme in the presence of trypsin as a bio-inhibitor changed. This change has a logarithmic relationship with the concentration of trypsin in the range 0.25–20.0 µg.mL−1. The proposed biosensor was also applied in the determination of trypsin in a human urine sample. The analytical performance of the proposed IRS biosensor was better than the fluorescence biosensor based on quantum dots.
References
https://www.pancan.org/news/new-global-cancer-statistics-released/.
Xiao, A. Y. et al. Global incidence and mortality of pancreatic diseases: a systematic review, meta-analysis, and meta-regression of population-based cohort studies. Lancet Gastroenterol. Hepatol. 1, 45–55 (2016).
Soreide, K., Janssen, E., Körner, H. & Baak, J. Trypsin in colorectal cancer: molecular biological mechanisms of proliferation, invasion, and metastasis. J. Pathol. 209, 147–156 (2006).
Heinrich, H. C., Gabbe, E. E. & Ičagić, F. Immunoreaktives Serum Trypsin bei Pankreaserkrankungen. Klinische Wochenschrift. 57, 1237–1238 (1979).
Zaccheo, B. A. & Crooks, R. M. Self-Powered Sensor for Naked-Eye Detection of Serum Trypsin. Anal. Chem. 83, 1185–1188 (2011).
Stoytcheva, M., Zlatev, R., Cosnier, S. & Arredondo, M. Square wave voltammetric determination of trypsin activity. Electrochim. Acta 76, 43–47 (2012).
Zhang, L., Qin, H., Cui, W., Zhou, Y. & Du, J. Label–free, turn–on fluorescent sensor for trypsin activity assay and inhibitor screening. Talanta 161, 535–540 (2016).
Santos, A., Macías, G., Ferré-Borrull, J., Pallarès, J. & Marsal, L. F. Photoluminescent Enzymatic Sensor Based on Nanoporous Anodic Alumina. ACS Appl. Mater. Interfaces 4, 3584–3588 (2012).
Neff, P. A., Serr, A., Wunderlich, B. K. & Bausch, A. R. Label-Free Electrical Determination of Trypsin Activity by a Silicon-on-Insulator Based Thin Film Resistor. Chem. Phys. Chem. 8, 2133–2137 (2007).
Dwivedi, A. K. & Iyer, P. K. A fluorescence turn on trypsin assay based on aqueous polyfluorene. J. Mater. Chem. B 1, 4005–4010 (2013).
Liu, R. et al. A real-time fluorescence turn-on assay for trypsin based on a conjugated polyelectrolyte. J. Mater. Chem. B 1, 1402–1405 (2013).
Amouzadeh Tabrizi, M., Shamsipur, M., Saber, R., Sarkar, S. & Sherkatkhameneh, N. Flow injection amperometric sandwich-type electrochemical aptasensor for the determination of adenocarcinoma gastric cancer cell using aptamer-Au@Ag nanoparticles as labeled aptamer. Electrochim. Acta 246, 1147–1154 (2017).
Amouzadeh Tabrizi, M., Shamsipur, M., Saber, R., Sarkar, S. & Zolfaghari, N. An ultrasensitive sandwich-type electrochemical immunosensor for the determination of SKBR-3 breast cancer cell using rGO-TPA/FeHCFnano labeled Anti-HCT as a signal tag. Sens. Actuator B-Chem. 243, 823–830 (2017).
Shamsipur, M. & Amouzadeh Tabrizi, M. Achieving direct electrochemistry of glucose oxidase by one step electrochemical reduction of graphene oxide and its use in glucose sensing. Mater. Sci. Eng. C. 45, 103–108 (2014).
Torrini, F. et al. Characterization of troponin T binding aptamers for an innovative enzyme-linked oligonucleotide assay (ELONA). Anal. Bioanal. Chem. 411, 7709–7716 (2019).
Allsop, T. D. P. et al. An ultra-sensitive aptasensor on optical fibre for the direct detection of bisphenol A. Biosens. Bioelectron. 135, 102–110 (2019).
Chao, J. et al. Nanostructure-based surface-enhanced Raman scattering biosensors for nucleic acids and proteins. J. Mater. Chem. B 4, 1757–1769 (2016).
Rajeev, G. et al. A label-free optical biosensor based on nanoporous anodic alumina for tumour necrosis factor-alpha detection in chronic wounds. Sens. Actuator B-Chem. 257, 116–123 (2018).
Feng, J. et al. pH-Regulated Synthesis of Trypsin-Templated Copper Nanoclusters with Blue and Yellow Fluorescent Emission. ACS Omega 2, 9109–9117 (2017).
Xue, F., Qu, F., Han, W., Xia, L. & You, J. Aggregation-induced emission enhancement of gold nanoclusters triggered by silicon nanoparticles for ratiometric detection of protamine and trypsin. Anal. Chim. Acta 1046, 170–178 (2019).
Işık, B. & Sezgintürk, M. K. Quantification of Trypsin Activity by a New Biosensing System Based on the Enzymatic Degradation and the Destructive Nature of Trypsin. Int. J. Pept. Res. Ther. 23, 313–322 (2017).
Ong, I. L. H. & Yang, K.-L. Recent developments in protease activity assays and sensors. Analyst 142, 1867–1881 (2017).
Dai, H. et al. Photoelectrochemical biosensor constructed using TiO2 mesocrystals based multipurpose matrix for trypsin detection. Biosens. Bioelectron. 92, 687–694 (2017).
Poon, C.-Y. et al. FRET-based modified graphene quantum dots for direct trypsin quantification in urine. Anal. Chim. Acta 917, 64–70 (2016).
Wang, G.-L., Jin, L.-Y., Dong, Y.-M., Wu, X.-M. & Li, Z.-J. Intrinsic enzyme mimicking activity of gold nanoclusters upon visible light triggering and its application for colorimetric trypsin detection. Biosens. Bioelectron. 64, 523–529 (2015).
Wang, L., Shi, F., Li, Y. & Su, X. An ultra-sensitive and label-free fluorescent probe for trypsin and inhibitor based on DNA hosted Cu nanoclusters. Sens. Actuator B-Chem. 222, 945–951 (2016).
Gao, X., Tang, G., Li, Y. & Su, X. A novel optical nanoprobe for trypsin detection and inhibitor screening based on Mn-doped ZnSe quantum dots. Anal. Chim. Acta 743, 131–136 (2012).
Zhu, Q., Zhan, R. & Liu, B. Homogeneous Detection of Trypsin in Protein Mixtures Based on Fluorescence Resonance Energy Transfer between Anionic Conjugated Polymer and Fluorescent Probe. Macromol. Rapid Commun. 31, 1060–1064 (2010).
Xu, K., Liu, F., Ma, J. & Tang, B. A new specific fullerene-based fluorescent probe for trypsin. Analyst 136, 1199–1203 (2011).
Amouzadeh Tabrizi, M., Ferré-Borrull, J. & Marsal, L. F. Highly sensitive aptasensor based on interferometric reflectance spectroscopy for the determination of amyloid β as an Alzheimer’s disease biomarkers using nanoporous anodic alumina. Biosens. Bioelectron. 137, 279–286 (2019).
Pol, L. et al. Real-Time Monitoring of Biotinylated Molecules Detection Dynamics in Nanoporous Anodic Alumina for Bio-Sensing. Nanomaterials 9, 478–489 (2019).
Harraz, F. A. Porous silicon chemical sensors and biosensors: A review. Sens. Actuator B-Chem. 202, 897–912 (2014).
Jani, A. M. M., Yazid, H., Habiballah, A. S., Mahmud, A. H. & Losic, D. In Nanoporous Alumina: Fabrication, Structure, Properties and Applications (eds. Dusan Losic & Abel Santos) 155–184 (Springer International Publishing, 2015).
Urmann, K., Arshavsky-Graham, S., Walter, J. G., Scheper, T. & Segal, E. Whole-cell detection of live lactobacillus acidophilus on aptamer-decorated porous silicon biosensors. Analyst 141, 5432–5440 (2016).
Ribes, À. et al. Selective and Sensitive Probe Based in Oligonucleotide-Capped Nanoporous Alumina for the Rapid Screening of Infection Produced by Candida albicans. ACS Sens. 4, 1291–1298 (2019).
Kaur, S. et al. Environmental Copper Sensor Based on Polyethylenimine-Functionalized Nanoporous Anodic Alumina Interferometers. Anal.Chem. 91, 5011–5020 (2019).
Eckstein, C. et al. Nanoporous Anodic Alumina Surface Modification by Electrostatic, Covalent, and Immune Complexation Binding Investigated by Capillary Filling. ACS Appl. Mater. Interfaces 10, 10571–10579 (2018).
Kumeria, T. et al. Advanced Structural Engineering of Nanoporous Photonic Structures: Tailoring Nanopore Architecture to Enhance Sensing Properties. ACS Photonics 1, 1298–1306 (2014).
Olsen, J. V., Ong, S.-E. & Mann, M. Trypsin Cleaves Exclusively C-terminal to Arginine and Lysine Residues. Mol. Cell. Proteom. 3, 608–614 (2004).
Benini, S., Cianci, M., Mazzei, L. & Ciurli, S. Fluoride inhibition of Sporosarcina pasteurii urease: structure and thermodynamics. J. Biol. Inorg. Chem. 19, 1243–1261 (2014).
Mishra, J., Kaur, H., Ganguli, A. K. & Kaur, N. Fluorescent chemosensor based on urea/thiourea moiety for sensing of Hg(II) ions in an aqueous medium with high sensitivity and selectivity: A comparative account on effect of molecular architecture on chemosensing. J. Mol. Struct. 1161, 34–43 (2018).
Parashar, U. K., Nirala, N. R., Upadhyay, C., Saxena, P. S. & Srivastava, A. Urease Immobilized Fluorescent Gold Nanoparticles for Urea Sensing. Appl. Biochem. Biotechnol. 176, 480–492 (2015).
Santos, A. et al. Understanding and morphology control of pore modulations in nanoporous anodic alumina by discontinuous anodization. phys. Stat. 209, 2045–2048 (2012).
Mardare, A. I., Kaltenbrunner, M., Sariciftci, N. S., Bauer, S. & Hassel, A. W. Ultra-thin anodic alumina capacitor films for plastic electronics. phys. Stat. 209, 813–818 (2012).
Devi, R. & Pundir, C. S. Construction and application of an amperometric uric acid biosensor based on covalent immobilization of uricase on iron oxide nanoparticles/chitosan-g-polyaniline composite film electrodeposited on Pt electrode. Sens. Actuator B-Chem. 193, 608–615 (2014).
Kuwabata, S., Okamoto, T., Kajiya, Y. & Yoneyama, H. Preparation and amperometric glucose sensitivity of covalently bound glucose oxidase to (2-aminoethyl)ferrocene on an Au electrode. Anal.Chem. 67, 1684–1690 (1995).
Kaschula, C. H. & Hunter, R. In Studies in Natural Products Chemistry Vol. 50 (ed Rahman Atta ur) 1–43 (Elsevier, 2016).
Caro, L. H. P., Plomp, P. J. A. M., Wolvetang, E. J., Kerkhof, C. & Meijer, A. J. 3-Methyladenine, an inhibitor of autophagy, has multiple effects on metabolism. Eur. J. Biochem. 175, 325–329 (1988).
Kumeria, T. & Losic, D. Controlling interferometric properties of nanoporous anodic aluminium oxide. Nanoscale Res. Lett. 7, 88–97 (2012).
Min, Q. et al. Size-selective proteolysis on mesoporous silica-based trypsin nanoreactor for low-MW proteome analysis. ChemComm. 46, 6144–6146 (2010).
Liu, C., Shih, K., Gao, Y., Li, F. & Wei, L. Dechlorinating transformation of propachlor through nucleophilic substitution by dithionite on the surface of alumina. J. Soil. Sedmient. 12, 724–733 (2012).
Larkin, P. In Infrared and Raman Spectroscopy (ed. Peter Larkin) 73–115 (Elsevier, 2011).
Lin-Vien, D., Colthup, N. B., Fateley, W. G. & Grasselli, J. G. The handbook of infrared and Raman characteristic frequencies of organic molecules. (Elsevier, 1991).
Seabra, I. J. & Gil, M. H. Cotton gauze bandage: a support for protease immobilization for use in biomedical applications. Rev. Bras. Cienc. Farm. 43, 535–542 (2007).
Nemati, M., Santos, A., Kumeria, T. & Losic, D. Label-Free Real-Time Quantification of Enzyme Levels by Interferometric Spectroscopy Combined with Gelatin-Modified Nanoporous Anodic Alumina Photonic Films. Anal. Chem. 87, 9016–9024 (2015).
Zhou, G. et al. Peptide-coated palladium nanoparticle for highly sensitive bioanalysis of trypsin in human urine samples. Nanomater. Nanotechno 8, 1–8 (2018).
Jia, R. P. et al. Enhanced photoluminescence properties of morin and trypsin absorbed on porous alumina films with ordered pores array. Solid. State Commun. 130, 367–372 (2004).
Acknowledgements
This research was supported by the Spanish Ministerio de Ciencia, Innovación y Universidades RTI2018-094040-B-I00 (MICINN/FERDER), by the Agency for Management of University and Research Grants (AGAUR) ref. 2017-SGR-1527, by the Catalan Institution for Research and Advanced Studies (ICREA) under the ICREA Academia Award and by the Marti-Franques II postdoctoral program under grant number 2017PMF-POST2-7. The project leading to these results has also received funding from “la Caixa” Foundation under the agreement LCF/PR/PR17/11120023.
Author information
Authors and Affiliations
Contributions
Mahmoud Amouzadeh Tabrizi: Conceptualization, Methodology, Data curation, Formal analysis, Validation, Investigation, Visualization, Writing - original draft. Josep Ferré-Borrull: Formal analysis, Validation. Lluis F. Marsal: Supervision, Writing - review & editing, Resources, Project administration, Funding acquisition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Amouzadeh Tabrizi, M., Ferré-Borrull, J. & Marsal, L.F. Remote biosensor for the determination of trypsin by using nanoporous anodic alumina as a three-dimensional nanostructured material. Sci Rep 10, 2356 (2020). https://doi.org/10.1038/s41598-020-59287-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-59287-7
This article is cited by
-
Effect of Etching Parameters on Sensing Performance of Porous Silicon Thin Film Based Optical Biosensors
Silicon (2022)
-
Solid-state nanopore systems: from materials to applications
NPG Asia Materials (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.