Abstract
Like all organisms, aphids, plant sap-sucking insects that house a bacterial endosymbiont called Buchnera, are members of a species interaction network. Ecological interactions across such networks can result in phenotypic change in network members mediated by molecular signals, like microRNAs. Here, we interrogated small RNA data from the aphid, Myzus persicae, to determine the source of reads that did not map to the aphid or Buchnera genomes. Our analysis revealed that the pattern was largely explained by reads that mapped to the host plant, Brassica oleracea, and a facultative symbiont, Regiella. To start elucidating the function of plant small RNA in aphid gut, we annotated 213 unique B. oleracea miRNAs; 32/213 were present in aphid gut as mature and star miRNAs. Next, we predicted targets in the B. oleracea and M. persicae genomes for these 32 plant miRNAs. We found that plant targets were enriched for genes associated with transcription, while the distribution of targets in the aphid genome was similar to the functional distribution of all genes in the aphid genome. We discuss the potential of plant miRNAs to regulate aphid gene expression and the mechanisms involved in processing, export and uptake of plant miRNAs by aphids.
Similar content being viewed by others
Introduction
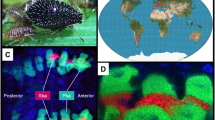
Most insects in the order Hemiptera are herbivores that feed on plant sap1. Typically, insects that feed on plant sap critically depend on bacteriome housed endosymbionts to supplement dietary shortfalls in essential amino acids and vitamins2,3,4. Many insects also host facultative symbionts that confer benefits such as resistance to attack by parasitoids and increased tolerance to heat stress5. For example, the pea aphid, Acyrthosiphon pisum, that obligately hosts the endosymbiont Buchnera aphidicola, has also been reported to host a facultative symbiont, Candidatus Hamiltonella defensa encoding a bacteriophage that provides strong protection against parasitic wasps6,7. While aphid guts contain a relatively limited microbiome8,9, aphids such as M. persicae, vector hundreds of different plant viruses10,11. Thus, the species interaction network of an aphid includes its obligate and facultative symbionts, its host plant, its gut microbiome, the microbiome of its host plant, and the viruses it vectors (Fig. 1A).
Aphid bacteriome and gut tissues were mapped against aphid, Buchnera, and host plant genomes as well as a bacterial library and viral library. (A) Myzus persicae feeding on Brassica oleracea with the aphid stylet passing through the cuticle, epidermis, and mesophyll to reach the sieve element and extract phloem. Aphid gut is colonized by a microbiome that includes facultative symbionts, gut-associated bacteria, bacterial pathogens, plant associated bacteria and viruses. The bacteriome is the aphid organ made up of bacteriocyte cells that house Buchnera and sheath cells that sometimes are infected with facultative symbionts. (B) Custom libraries: (1) Host plant, (2) Bacterial and (3) Viral.
In this paper, we study small RNA datasets from M. persicae. Preliminary data processing of the M. persicae small RNA data available in the National Center for Biotechnology Information (NCBI) data repository as BioProject PRJNA395356 identified a striking difference between gut and bacteriome tissue samples (see Fig. 1A for arrangement of tissues in aphid host) in the proportion of total reads that mapped to the M. persicae and Buchnera genomes (Fig. 2). This difference, which was not explored by Feng et al.12, motivated the current work.
More than 58% of small RNA reads from aphid gut do not map to the aphid or endosymbiont genomes. In contrast, less than 28% of small RNA reads from aphid bacteriomes do not map to the aphid or endosymbiont genome. Pie charts show the proportion of gut or bacteriome reads that aligned to the aphid and/or Buchnera genomes with less than two mismatches. Data are shown for three M. persicae lines: G006, G002, and USDA. Numbers below the pie charts refer to the total number of reads in the respective filtered dataset.
We set out to determine the source of unmapped small RNA reads in M. persicae gut samples, by mapping these reads against three genomic libraries. The first contained the genome of cabbage, Brassica oleracea13, since our aphid lines were raised on B. oleracea seedlings. The second contained genomes of both beneficial and pathogenic bacteria, that are known to be associated with aphids8,14, while the third contained genomes of aphid vectored plant viruses15,16 (Fig. 1B). Our goal was to leverage all available genomic information for members of the M. persicae species interaction network to determine the source of small reads that did not map to the M. persicae or Buchnera genomes.
Surprisingly, we found that upwards of 21% of small RNA reads in aphid gut samples mapped to the host plant genome. Motivated by recent demonstration of miRNA function across kingdoms17, we next asked if the B. oleracea small read data included any miRNAs. To address this question, we leveraged three existing B. oleracea small RNA datasets to annotate B. oleracea miRNAs using the current approaches recommended by Axtell & Meyers18. Following annotation, we found evidence of 32 B. oleracea miRNAs in aphid gut samples. Intrigued by the possibility that these B. oleracea miRNAs regulate aphid gene expression, we predicted targets in both the B. oleracea and M. persicae genomes. We found that the distribution of targets across the M. persicae genome were dissimilar to that observed in the B. oleracea genome targets that were strongly enriched for functions associated with transcription.
Results
More than one fifth of small RNA reads from aphid gut tissue samples map to the host plant genome
We analyzed M. persicae bacteriome and gut tissue small RNA data from three genetically discrete aphid lines. In bacteriomes, 78.8% ± 11.3% of small RNA reads mapped to the aphid and/or Buchnera genome. In contrast, in gut samples, only 34.8% ± 7.1% of reads mapped to the aphid and/or Buchnera genome (Fig. 2). Remarkably, in gut samples, 34.4% ± 8.6% of small RNA reads mapped to the host plant, B. oleracea, with 30.8% ± 8.2% mapping only to B. oleracea (Fig. 3, Supplementary Table S1).
More than 21% of small RNA reads in aphid gut map exclusively to the genome of the host plant, B. oleracea (green). Bars show the proportion of reads that aligned to the aphid, Buchnera, and host plant genomes for gut and bacteriome tissues in three M. persicae lines: G006, G002, and USDA. Categories <1% not displayed (see Table S1).
Small RNA reads from aphid gut tissue are of limited bacterial origin and not of viral origin
For gut samples, we mapped reads of unknown origin against two microbial libraries. The first was a bacterial library that contained the genomes of aphid facultative symbionts, aphid gut associated bacteria, and plant associated bacteria, while the second was a viral library that contained the genomes of plant viruses that aphids are known to vector (Fig. 1B, Supplementary Table S2). We found that between 2.5% and 8.6% of all small RNA reads from aphid gut tissues mapped exclusively to the bacterial library while, at most only 0.004% of all small RNA reads mapped exclusively to the viral library (Supplementary Tables S1 and S2). Most of the bacterial library mapping was attributed to Regiella insecticola with at most only 0.5% of the gut small RNA reads mapping to additional bacteria (that were not Buchnera).
Plant miRNAs are present in aphid gut and bacteriome tissue
To test our hypothesis that B. oleracea small RNAs found in M. persicae gut contain plant miRNAs, we annotated miRNAs in the B. oleracea genome using publicly available small RNA-seq data. Out of a total of 147 miRNA precursors in the B. oleracea genome 138 were homologous to known plant miRNA precursors (miRBase v22) and 9 were novel miRNA precursors (Supplementary Table S3). The 147 precursors encoded 213 miRNAs, a set that includes unique miRNAs isolated from both the 5′ and 3′ ends of the miRNA precursors. We next asked if any of these 213 miRNAs were present in our aphid gut and bacteriome datasets. In gut samples, we found perfect matches for 32 B. oleracea miRNAs that were each represented between one and 370 times (Supplementary Table S4). Seven of the 32 B. oleracea miRNAs were represented in the guts of all three M. persicae lines. These seven miRNAs map to 13 different miRNA precursors (because some miRNAs are encoded by duplicated precursors), including bol-miR395d, bol-miR_novel_104, and bol-miR168b-3p, as well as three members of the bol-miR166 family (166a-3p, 166b-3p, and 166e-3p), three members of the bol-miR159 family (159, 159a, 159c), two members of the bol-miR162 family (162-5p, 162a-5p), and two members of the bol-miR168 family (168a, 168c). In contrast, we found perfect matches for only four B. oleracea miRNAs in the aphid bacteriome tissue datasets with each miRNA represented between one and 16 times. Two of the four B. oleracea miRNAs identified in bacteriome were also represented in the gut datasets (Supplementary Table S4) that mapped to four precursors: bol-miR156-5p and three members of the bol-miR166 family (166a-3p, 166b-3p, and 166e-3p).
Plant miRNAs present in aphid gut have target sites in aphid and plant transcriptomes
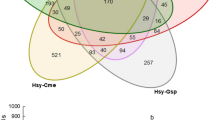
Using the intersection of three miRNA targeting programs specific to animal genomes (miRanda, PITA and RNAhybrid), we predicted 926 M. persicae 3′ UTRs as putative targets for the 32 B. oleracea miRNAs detected in M. persicae gut (Fig. 4). The 926 predicted 3′ UTR targets associated with 965 M. persicae mRNA isoforms. In contrast, in the B. oleracea genome, using the intersection of two miRNA target prediction programs specific to plant genomes (psRNAtarget and TargetFinder), we predicted 576 B. oleracea target sites for the 32 B. oleracea miRNAs detected in M. persicae gut (Fig. 4).
Prediction of targets of plant miRNAs identified 926 and 576 putative targets in the aphid and plant genomes, respectively. (A) Proportional Venn diagram of targets in aphid 3′-UTRs predicted by miRanda52, PITA53 and RNAhybrid54. (B) Proportional Venn diagram of targets in plant genome predicted by TargetFinder55 and psRNAtarget51. Proportional Venn diagrams were produced using eulerAPE v3.0.061.
Targets of plant miRNAs present in aphid gut have organism-specific COG profiles
We performed a broad functional analysis by assigning the transcripts targeted by the 32 B. oleracea miRNAs to clusters of orthologous groups (COGs)19. We used Monte Carlo simulations to test the significance of over and under-representation of each COG in our predicted target sets relative to the representation of COGs in the official gene set of the corresponding genome. In the M. persicae genome, we assigned a functional COG to 664 of the 965 predicted targets, and in the B. oleracea genome, we assigned functional COGs to 330 of the 576 predicted targets.
For the set of targets in the aphid genome, three COGs, “Carbohydrate transport and metabolism” (p = 3.5 × 10−3), “RNA processing and modification” (p = 2.2 × 10−2), and “Nuclear structure” (p = 6.2 × 10−3) were significantly overrepresented while four COGs “Amino acid transport and metabolism” (p = 2.0 × 10−4), “Cytoskeleton” (p = 1.6 × 10−2), “Replication, recombination, and repair” (p = 1.5 × 10−3), and “Translation, ribosomal structure and biogenesis” (p = 2.6 × 10−2) were significantly underrepresented (Fig. 5). Notably, the categories over and under-represented in the M. persicae genome targets differed from those over and under-represented in the B. oleracea genome.
Cluster of Orthologous Group (COG) distribution of altered plant miRNA targets over aphid genome. (Left) Percentage of targets found in each COG in increasing order. (Right) Percent difference between the distribution of COGs in the targets of the 32 B. oleracea miRNAs in the aphid genome and of the whole aphid genome. Significant categories are marked with an asterisk (p < 0.05). On the right, grey bars mark the value needed to achieve signifcant overrepresentation (positive % enrichment) or underrepresentation (negative % enrichment).
In the set of targets in the B. oleracea genome, three COGs “Transcription” (p < 1 × 10−6), “Amino acid transport and metabolism” (p = 6.0 × 10−3), and “Chromatin structure and dynamics” (p = 1.8 × 10−2) were significantly overrepresented with “Transcription” being the most significantly overrepresented as well as the COG that included the largest fraction of the B. oleracea genome targets (28%, Fig. 6). Interestingly, “Amino acid transport and metabolism” was significantly overrepresented in the B. oleracea genome targets while it was underrepresented in the M. persicae genome targets. The B. oleracea genome targets also included seven COGs that were significantly underrepresented: “Signal transduction mechanisms” (p = 2.5 × 10−5), “Intracellular trafficking, secretion, and vesicular transport” (p = 1.5 × 10−2), “Translation, ribosomal structure and biogenesis” (p = 1.6 × 10−3), “Secondary metabolites biosynthesis and catabolism” (p = 1.9 × 10−2), “Energy production and conservation” (p = 1.1 × 10−2), “Replication, recombination, and repair” (p = 4.2 × 10−2), and “Defense mechanisms” (p = 2.1 × 10−2). In both genomes, “Replication, recombination, and repair” was significantly underrepresented.
Cluster of Orthologous Group (COG) distribution of altered plant miRNA target over plant genome. (Left) Percentage of targets found in each COG in increasing order. (Right) Percent difference between the distribution of COGs in the targets of the 32 B. oleracea miRNAs in the plant genome and of the whole plant genome. Significant categories are marked with an asterisk (p < 0.05). On the right, grey bars mark the value needed to achieve signifcant overrepresentation (positive % enrichment) or underrepresentation (negative % enrichment).
Plant-derived, aphid gut small RNAs 17–35 nucleotides in length that map to plant miRNA precursors are almost exclusively miRNAs
We aligned our size-selected and filtered M. persicae gut small RNA data against full-length miRNA precursors of the 32 B. oleracea miRNAs present in our M. persicae gut datasets (Supplementary Document S1). Precursor miRNAs comprise 5′ and 3′ end regions, mature and star miRNAs, and a loop region (Fig. 7B’). None of the small RNA reads in our gut dataset aligned to 5′ or 3′ end regions while 1.8% aligned to loop regions and 91.4% of reads aligned perfectly to mature or star miRNA sequences (Fig. 7). The remaining 6.8% of reads that aligned to the 32 miRNA precursors were fragments that spanned two regions within a miRNA precursor e.g. fragments that spanned a star miRNA and a loop region (Fig. 7B), or a 5′ end fragment and a star miRNA (Fig. 7D).
Alignment of plant small RNA data against plant miRNA precursors. (A–D) Representation of M. persicae gut small RNA reads that exactly map to B. oleracea miRNA precursors. While red reads are perfect star miRNA matches green reads are perfect mature miRNA matches. X-axis indicates precursor nucleotide position, miRNA name and precursor identity in parentheses. (A) One mature miRNA sequence is shown in place of the 370 that were aligned in that position. (B′) The secondary structure of the precursor displayed in B. (E) Categorization of all reads aligned to plant miRNA precursors across all gut datasets. Reads are assigned to the mature miRNA, star miRNA, and loop sections by aligning with a maximum of one nucleotide overlap with adjacent sections. Any read that overlapped two sections by more than one nucleotide was classified as a fragment.
Discussion
We interrogated the striking difference between the proportion of total small RNA reads that mapped to the M. persicae and Buchnera genomes in M. persicae gut versus bacteriome tissues (Fig. 2). Our analyses revealed that two members of the M. persicae species interaction network, the host plant, B. oleracea, and an aphid facultative symbiont, Regiella insecticola, were the source of a large fraction of the unmapped small RNA reads in M. persicae gut libraries (Supplementary Table S1). In all three M. persicae lines, the host plant accounted for the plurality of unmapped reads while Regiella was the source of a small, yet substantial fraction of the unmapped reads in all three lines (Supplementary Table S1). While we are not the first to detect plant (diet) derived small RNA in aphids, our work is, to the best of our knowledge, the first to document that the abundance of plant small RNA is strikingly different among aphid tissues20,21. This is also not the first study to document the presence of Regiella in M. persicae as Regiella infection was previously reported in a single M. persicae line from Australia22. However, detection of Regiella in all three of the M. persicae lines used in this study was surprising as the presence of Regiella infection had not been revealed by earlier genome sequencing projects23,24. Earlier failure to detect Regiella infection coincident with previous work sequencing the genomes of Buchnera from M. persicae lines USDA and G002 likely resulted from the filtration methods used to enrich for Buchnera in genomic DNA preparations5,24. While failure to detect Regiella infection coincident with sequencing the G006 M. persicae genome is less clear, it likely resulted from low Regiella titre and bioinformatic filtering of bacterial contaminants23.
Following identification of B. oleracea small RNAs in our M. persicae gut libraries, we next asked if the B. oleracea small read data included any miRNAs. While previous studies reported annotation of 16925 (from a diversity of tissues), 15926 (from mature leaves), and 156 (from young leaves)27 miRNAs in B. oleracea, the intersection of these three sets only indicated 15 miRNAs (Supplemental Fig. S2). Further, in March 2018, the reference miRNA database, miRBase v22, contained only 10 B. oleracea unique miRNAs. Thus, we performed a new annotation of miRNAs in B. oleracea that leveraged all available datasets and followed the newly updated criteria for identification of plant miRNAs from big data RNAseq projects18. We predicted 147 miRNA precursors that produced 213 unique miRNAs (note that miRNAs can be encoded at both the 5′ and 3′ end of precursors, and two different precursors can encode identical miRNAs). Of the 213 miRNAs, 86 were previously reported in at least one of the earlier B. oleracea annotations25,26,27 (Supplemental Fig. S2), and out of the 213, 203 are members of known miRNA families (miRBase v22). The remaining 10 are novel miRNAs that lack homology to known miRNA families in miRBase v22. However, three of these 10 were previously reported by Zhang et al.27, and two were identified previously in the genus Brassica by Wei et al.28. The remaining five novel miRNAs are reported for the first time in this study (Supplementary Table S3).
Leveraging our new annotation of B. oleracea miRNAs we identified 32 B. oleracea miRNAs in our M. persicae gut data. While plant xylem sap is devoid of RNA29, plant phloem sap contains large numbers of small RNA molecules that generate a small RNA profile distinct from other plant tissues29,30,31,32. Out of the 32 B. oleracea miRNAs that we found in M. persicae gut, 16 have been found previously in phloem sap29,30,31,32, including 12 identified from Brassica napus phloem29,30 (Supplementary Table S4). Further, all four of the miRNAs found in bacteriome tissue were previously identified in B. napus phloem30. Overall, our results suggest that host plant miRNAs found in aphid tissues are members of the B. oleracea circulating miRNA profile that facilitates the systemic spread of post-transcriptional gene regulation.
An outstanding question at this time concerns function. Are the B. oleracea circulating miRNAs that we found in M. persicae gut functional? Do these B. oleracea miRNAs have targets in the M. persicae genome? Do they regulate M. persicae gene expression, or are they simply food? The ease with which it is possible to generate large amounts of small RNA and mRNA data has resulted in a recent flood of studies claiming regulation of gene expression in the genome of an organism by miRNAs encoded in the genome of its food. Unfortunately, in many cases authors have been quick to assume function in the absence of experimental validation33,34,35,36. Here, we predicted mRNA targets in the B. oleracea and M. persicae genomes for the 32 B. oleracea miRNAs found in M. persicae gut. Our analysis of the M. persicae genome failed to identify signatures of enrichment in 19 of the 22 COGs assigned to predicted targets relative to the COG composition of all genes in the M. persicae genome. That said, three M. persicae COGs were significantly enriched in targets relative to the M. persicae genome (Fig. 5); however, the degree of enrichment of these three COGs compared to the degree of enrichment of B. oleracea targets in the COG “Transcription” relative to the B. oleracea genome (Fig. 6) is much lower. While these broad-scale COG analyses revealed a lack of directed evolution in plant miRNAs targeting aphids, they also say little about the ability of individual plant miRNAs to elicit a physiological effect in aphids. However, experimental evidence from transgenic plants is compelling in arguing for the potential of B. oleracea encoded miRNAs to elicit a biological effect in M. persicae. We emphasize this point because aphids fed on transgenic plants expressing artificial miRNAs and dsRNA that target individual genes in the aphid genome, show reduced expression of the targeted gene37,38,39,40. Additionally, the cotton bollworm and the Colorado potato beetle have been shown to accumulate plant produced small RNA fragments and a corresponding reduction in expression of the targeted genes41,42. Thus, such plant-mediated RNAi experiments serve as evidence that host plant genome encoded miRNAs could indeed regulate aphid gene expression in vivo.
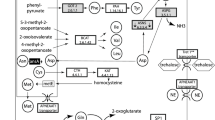
If plant circulating miRNAs do indeed function to target aphid gene expression, what processing, export, transport and uptake mechanisms may be involved? By integrating our observations with current knowledge about the production, processing, and export of miRNAs, we propose that trafficking of plant miRNA into aphid tissues is mediated by exosomes. First, B. oleracea miRNA precursors are processed yielding mature and star miRNAs within the plant cells43. Next, miRNAs and proteins that will be secreted from the plant cells are selectively loaded with B. oleracea Argonaute 1 (Ago1)44 into exosomes within multivesicular bodies43. Co-packing of the miRNAs with Ago1 facilitates stabilization and function of the secreted miRNAs when the exosome reaches its destination45. As the multivesicular bodies fuse with the plant cell membrane, the miRNA-containing exosomes are exocytosed into the circulating apoplastic fluid of B. oleracea43,46,47. The apoplast then carries the exosomes to the phloem where they can achieve long-distance transport and contact with the feeding aphids. Since aphids feed passively on moving phloem, miRNA-containing exosomes are brought into the aphid digestive tract through the aphid stylet when it is inserted into the plant sieve elements. Once in the aphid digestive tract, the host plant miRNAs are protected by their exosome membrane and move into aphid gut cells by endocytosis44. Once inside aphid cells, the B. oleracea miRNAs and associated proteins are released to function alongside the aphid’s own long-distance miRNAs37,38,39,48.
Here we document the presence of B. oleracea circulating miRNAs in gut and bacteriome tissues of the aphid M. persicae. COG analyses did not reveal strong targeting of any one functional group of aphid genes by host plant miRNAs. However, we argue that other work with transgenic plants that have been engineered to intracellularly express miRNAs and dsRNA that target aphid genes demonstrate the presence of a physiological mechanism for cross-kingdom miRNA targeting from plants to aphids. Future in vitro and in vivo studies are required to test the cross-kingdom miRNA::mRNA interactions predicted here.
Methods
We used the small RNA (sRNA) data for Myzus persicae lines G006, G002 and USDA from NCBI BioProject PRJNA395356. Sample preparation and initial data processing is described in Feng et al.12. Briefly, we processed the raw small RNA sequencing reads by first filtering out low quality reads, and then trimming sequencing adapters and primers. Mature miRNAs are typically 22 nucleotides in length, and thus following initial filtering and trimming, we retained reads that were 17–35 nucleotides long. For each sample, the set of 17–35 nucleotide long processed reads were used for genome mapping as described below. Aphid lines were raised at the University of Miami on Brassica oleracea var. Wisconsin Golden Acre.
Identification of source genome
Using Bowtie2 v2.2.6. (parameters: -L 10–very-sensitive -a) gut and bacteriome sRNA data were each aligned against the M. persicae G006 v2.0 genome23 from AphidBase (http://www.aphidbase.com) and their respective Buchnera aphidicola genomes: G006, NCBI RefSeq GCF_001939165.123; G002, NCBI RefSeq GCF_000521565.124; and USDA, NCBI RefSeq GCF_000521525.124. Following alignment, we identified reads that aligned with less than two mismatches and those that failed to align to the aphid and/or Buchnera genome; these reads were interrogated for alignment against three custom libraries. The first library contained the genome of the host plant, Brassica oleracea. The B. oleracea var. oleracea genome (NCBI RefSeq: GCF_000695525.113) was used because the genome of var. Wisconsin Golden Acre had not been sequenced. The second library contained the genomes of aphid secondary bacterial symbionts, aphid gut associated bacteria, and plant associated bacteria14 (Supplementary Table S2). The third library contained the genomes of plant viruses that aphids have been shown to vector15,16 (Supplementary Table S2). Genomes for bacteria and viruses were collected from the NCBI Assembly database. For each species or genus, preference was given to genomes with reference status and then representative status in the NCBI Assembly database. If those were not available, a complete genome was utilized. If nothing met the above criteria, manual searches were performed, using the most complete version of the genome. When only genus information was available, all species under the genus having at least a complete genome were utilized.
Annotation of B. oleracea miRNA precursors
To test our hypothesis that B. oleracea small reads from our three aphid gut libraries included plant miRNAs, we reannotated B. oleracea miRNA precursors (NCBI BioProjects: PRJNA431509, PRJNA195556, PRJNA324161, and PRJNA324144) using the recently revisited set of plant miRNA annotation criteria18. Prior to annotation we used FastQC v.0.11.4 to assess the quality of these datasets trimming sequences with a phred 33 score less than 28 and retaining sequences 17–25 nucleotides in length. For each of the four B. oleracea BioProjects, we used ShortStack v3.8.549 to identify miRNA precursors against the B. oleracea genome13. In ShortStack, hairpins were limited to 300 nucleotides, and miRNA size was limited to 20–24 nucleotides18. We set minimum coverage at 0.25 reads per million mapped and determined the placement of reads with multiple mapping locations by considering the density of all mapped reads. ShortStack predicts miRNA precursors from a user-supplied reference genome and small RNA-seq data. The small RNAs are aligned to the genome, and areas of high coverage are identified. High coverage areas with appropriate mapping are analyzed for hairpin structures and low mismatch interactions between the mature and star miRNAs. In ShortStack, mature miRNAs are those that have the higher number of mapped reads while star miRNAs have the lower number of mapped reads. For those miRNAs annotated here, mature or star miRNA designations are assigned as such. This does not reflect the 5′/3′ naming convention.
Following annotation, we collapsed miRNA precursors from each of the four BioProjects into a single data set. Duplicates across the BioProjects were removed by retaining the duplicate with the most reads mapped to the mature miRNA. Known precursors were identified as those having e-values <10−10 in local BLAST searches against miRBase v2250. Precursors with e-values ≥10−3 were identified as novel. Precursors with e-values <10−3 and ≥10−10 were further scrutinized such that precursors with query coverage ≥60% were identified as known, while precursors with query coverage <60% were subject to additional BLAST searches of their mature and star sequences using local short-BLAST against miRBase v22. Short-BLAST matches with e-values <10−10 were identified as known, and matches with e-values ≥10−10 were identified as novel. Ultimate determination of novel precursors was performed following the updated plant miRNA precursor criteria found in Axtell and Myers18.
miRNA target prediction
We predicted targets for B. oleracea miRNAs in M. persicae 3′ UTRs (AphidBase GFF) and in B. oleracea cDNA transcripts using psRNAtarget51. Potential miRNA targets in the aphid genome were predicted using three programs: miRanda v.3.3a52 (parameters: ‐sc 80 ‐go ‐8 ‐ge ‐2 ‐scale 2 ‐en 0 ‐strict ‐quiet), PITA v653 (parameters: -gxp -l 7-8 -gu 7;0,8;0 -m 7;0,8;0), and RNAhybrid v2.1.254 (parameters: -d <xi>, <theta> [from RNAcalibrate] -e 0 -p 0.05 -f 2,8). Targets in the plant genome were predicted using psRNAtarget51 and TargetFinder55. The two plant genome prediction programs were selected based on the analysis of Srivastava et al.56 using their optimized scoring parameters. All five miRNA target prediction programs use sequence complementarity in the seed region of the miRNA as well as the energetics of miRNA::mRNA binding to assess the ability of a miRNA to successfully bind to an mRNA. Predicted targets were manually filtered to include only those in which all respective target prediction programs overlapped in the miRNA seed region (Supplemental Fig. S1).
Functional enrichment analysis of miRNA targets
To determine the degree of functional enrichment in the predicted miRNA targets of the aphid and plant genomes relative to the representation of genes in each genome, targets were assigned to Clusters of Orthologous Groups (COGs) using eggnog-mapper v1.0.357 against EggNOG 4.5.1 orthology data58. 3′ UTRs targeted in the aphid genome were translated into their associated protein isoforms to be used for annotation. All isoforms sharing the 3′ UTR were included as potential targets. COG assignment was performed using Hidden Markov Models (HMMs) built from the insect (inNOG) database in EggNOG. As a comparative dataset, the entire M. persicae G006 proteome v2.0 (AphidBase) was run through a parallel COG annotation pipeline. In comparing these COG datasets, Monte Carlo simulations were performed to analyze the over/under-representation of target COGs relative to the reference proteome distribution. To generate null distributions, 926 3′ UTRs (the number of predicted miRNA targets, Fig. 4) were randomly sampled with replacement 106 times from the reference proteome. The 3′ UTRs were then expanded into their associated isoforms, and the number of proteins was added to the distribution allowing more than 926 proteins to be included in a distribution for the aphid proteome. A comparison was then made between each null COG distribution and the associated miRNA target COG value. Two-tailed p-values were calculated empirically from the null distributions to record significance in the event that the miRNA target COG category was found to be as or more extreme than the null distribution such that p < 0.05.
For the miRNA targets in the plant genome, the same functional enrichment analysis was repeated. Plant cDNA sequences were translated within EggNOG, and COG assignment was performed using HMMs built from the viridiplantae (virNOG) database in EggNOG. As a comparative dataset, the entire B. oleracea cDNA library (psRNAtarget51) was run through a parallel COG annotation pipeline. To produce Monte Carlo simulations that complement the aphid proteome analysis, 576 cDNA transcripts (the number of predicted miRNA targets, Fig. 4) were randomly sampled with replacement 106 times. Comparisons and p-values were calculated as described above.
Assessment of B. oleracea miRNA precursor processing
The secondary structure of each precursor was produced using RNAfold v2.4.459 and visualized using VARNA v3-9360 (Supplementary Document S2). These computational predictions validated the mature miRNA, star miRNA, loop, 5′ end and 3′ end regions of each miRNA precursor as previously determined through ShortStack. Next, we used Bowtie2 v2.2.6 to align our M. persicae gut small RNA data against miRNA precursors of the 32 B. oleracea miRNAs present in our gut datasets, filtering alignments to exclude any with mismatches or gaps (Supplementary Document S1). Following alignment and allowing for up to one nucleotide overlap with an adjacent region, reads were assigned to one of the following five regions: mature miRNA, star miRNA, loop, 5′ end or 3′ end. Any read that failed to meet these criteria was considered to be a “precursor fragment”.
Ethical approval and informed consent
Not required. The work did not involve vertebrates.
Data availability
All data sets supporting the conclusions are published in the article and the online supporting information. The small RNA next‐generation sequencing data sets are available at NCBI Sequence Read Archive (SRA) under BioProject PRJNA395356. The Python scripts written to perform analysis and figure production are also available on the WilsonLabMiami GitHub Repository: Thompson_etal_miRNAaphidgut (https://github.com/WilsonLabMiami/Thompson_etal_miRNAaphidgut).
References
Grimaldi, D., Engel, M. S. &. Engel, M. S. Evolution of the Insects. (Cambridge University Press, 2005).
Sudakaran, S., Kost, C. & Kaltenpoth, M. Symbiont Acquisition and Replacement as a Source of Ecological Innovation. Trends Microbiol. 25, 375–390 (2017).
Moran, N. A. Symbiosis as an adaptive process and source of phenotypic complexity. Proc. Natl. Acad. Sci. USA 104(Suppl 1), 8627–8633 (2007).
Buchner, P. Endosymbiosis of animals with plant microorganisms. (Interscience Publishers, 1965).
Moran, N. A., Russell, J. A., Koga, R. & Fukatsu, T. Evolutionary relationships of three new species of Enterobacteriaceae living as symbionts of aphids and other insects. Appl. Environ. Microbiol. 71, 3302–3310 (2005).
Oliver, K. M., Moran, N. A. & Hunter, M. S. Variation in resistance to parasitism in aphids is due to symbionts not host genotype. Proc. Natl. Acad. Sci. USA 102, 12795–12800 (2005).
Oliver, K. M., Degnan, P. H., Hunter, M. S. & Moran, N. A. Bacteriophages encode factors required for protection in a symbiotic mutualism. Science 325, 992–994 (2009).
Grigorescu, A. S. et al. Accessing the Hidden Microbial Diversity of Aphids: an Illustration of How Culture-Dependent Methods Can Be Used to Decipher the Insect Microbiota. Microb. Ecol. 75, 1035–1048 (2018).
Grenier, A. M., Nardon, C. & Rahbe, Y. Observations on the micro-organisms occurring in the gut of the pea aphid Acyrthosiphon pisum. Entomol. Exp. Appl. 70, 91–96 (1994).
Blackman, R. L. & Eastop, V. F. Aphids on the world’s crops: an identification and information guide. (John Wiley & Sons Ltd., 2000).
Hogenhout, S. A., Ammar, E.-D., Whitfield, A. E. & Redinbaugh, M. G. Insect vector interactions with persistently transmitted viruses. Annu. Rev. Phytopathol. 46, 327–359 (2008).
Feng, H., Wang, L., Wuchty, S. & Wilson, A. C. C. microRNA regulation in an ancient obligate endosymbiosis. Mol. Ecol. 27, 1777–1793 (2018).
Parkin, I. A. P. et al. Transcriptome and methylome profiling reveals relics of genome dominance in the mesopolyploid Brassica oleracea. Genome Biol. 15, R77 (2014).
Gauthier, J.-P., Outreman, Y., Mieuzet, L. & Simon, J.-C. Bacterial communities associated with host-adapted populations of pea aphids revealed by deep sequencing of 16S ribosomal DNA. PLoS One 10, e0120664 (2015).
Valenzuela, I. & Hoffmann, A. A. Effects of aphid feeding and associated virus injury on grain crops in Australia: Economic loss of grains by aphids. Austral Entomology 54, 292–305 (2015).
Ng, J. C. K. & Perry, K. L. Transmission of plant viruses by aphid vectors. Mol. Plant Pathol. 5, 505–511 (2004).
Liu, S. et al. The Host Shapes the Gut Microbiota via Fecal MicroRNA. Cell Host Microbe 19, 32–43 (2016).
Axtell, M. J. & Meyers, B. C. Revisiting Criteria for Plant MicroRNA Annotation in the Era of Big Data. Plant Cell 30, 272–284 (2018).
Galperin, M. Y., Makarova, K. S., Wolf, Y. I. & Koonin, E. V. Expanded microbial genome coverage and improved protein family annotation in the COG database. Nucleic Acids Res. 43, D261–9 (2015).
Wang, H. et al. Insect and plant-derived miRNAs in greenbug (Schizaphis graminum) and yellow sugarcane aphid (Sipha flava) revealed by deep sequencing. Gene 599, 68–77 (2017).
Sattar, S. et al. Expression of small RNA in Aphis gossypii and its potential role in the resistance interaction with melon. PLoS One 7, e48579 (2012).
von Burg, S., Ferrari, J., Müller, C. B. & Vorburger, C. Genetic variation and covariation of susceptibility to parasitoids in the aphid Myzus persicae: no evidence for trade-offs. Proc. Biol. Sci. 275, 1089–1094 (2008).
Mathers, T. C. et al. Rapid transcriptional plasticity of duplicated gene clusters enables a clonally reproducing aphid to colonise diverse plant species. Genome Biol. 18, 27 (2017).
Jiang, Z. et al. Comparative analysis of genome sequences from four strains of the Buchnera aphidicola Mp endosymbion of the green peach aphid, Myzus persicae. BMC Genomics 14, 917 (2013).
Wang, J. et al. Identification and characterization of microRNAs and their target genes in Brassica oleracea. Gene 505, 300–308 (2012).
Lukasik, A., Pietrykowska, H., Paczek, L., Szweykowska-Kulinska, Z. & Zielenkiewicz, P. High-throughput sequencing identification of novel and conserved miRNAs in the Brassica oleracea leaves. BMC Genomics 14, 801 (2013).
Zhang, J. et al. Genome-wide identification, putative functionality and interactions between lncRNAs and miRNAs in Brassica species. Sci. Rep. 8, 4960 (2018).
Wei, W. et al. Small RNA and degradome profiling involved in seed development and oil synthesis of Brassica napus. PLoS One 13, e0204998 (2018).
Buhtz, A., Springer, F., Chappell, L., Baulcombe, D. C. & Kehr, J. Identification and characterization of small RNAs from the phloem of Brassica napus. Plant J. 53, 739–749 (2008).
Buhtz, A., Pieritz, J., Springer, F. & Kehr, J. Phloem small RNAs, nutrient stress responses, and systemic mobility. BMC Plant Biol. 10, 64 (2010).
Varkonyi-Gasic, E., Gould, N., Sandanayaka, M., Sutherland, P. & MacDiarmid, R. M. Characterisation of microRNAs from apple (Malus domestica ‘Royal Gala’) vascular tissue and phloem sap. BMC Plant Biology 10, 159 (2010).
Gai, Y.-P. et al. MiRNA-seq-based profiles of miRNAs in mulberry phloem sap provide insight into the pathogenic mechanisms of mulberry yellow dwarf disease. Sci. Rep. 8, 812 (2018).
Fromm, B. et al. Plant microRNAs in human sera are likely contaminants. J. Nutr. Biochem. 65, 139–140 (2019).
Tosar, J. P., Rovira, C., Naya, H. & Cayota, A. Mining of public sequencing databases supports a non-dietary origin for putative foreign miRNAs: underestimated effects of contamination in NGS. RNA 20, 754–757 (2014).
Kang, W. et al. Survey of 800+ data sets from human tissue and body fluid reveals xenomiRs are likely artifacts. RNA 23, 433–445 (2017).
Heintz-Buschart, A. et al. Small RNA profiling of low biomass samples: identification and removal of contaminants. BMC Biol. 16, 52 (2018).
Guo, H. et al. Plant-generated artificial small RNAs mediated aphid resistance. PLoS One 9, e97410 (2014).
Bhatia, V., Bhattacharya, R., Uniyal, P. L., Singh, R. & Niranjan, R. S. Host generated siRNAs attenuate expression of serine protease gene in Myzus persicae. PLoS One 7, e46343 (2012).
Pitino, M., Coleman, A. D., Maffei, M. E., Ridout, C. J. & Hogenhout, S. A. Silencing of Aphid Genes by dsRNA Feeding from Plants. PLoS One 6, e25709 (2011).
Sun, Y. et al. Silencing an essential gene involved in infestation and digestion in grain aphid through plant-mediated RNA interference generates aphid-resistant wheat plants. Plant Biotechnol. J. 17, 852–854 (2019).
Mao, Y.-B. et al. Silencing a cotton bollworm P450 monooxygenase gene by plant-mediated RNAi impairs larval tolerance of gossypol. Nat. Biotechnol. 25, 1307–1313 (2007).
Zhang, J. et al. Pest control. Full crop protection from an insect pest by expression of long double-stranded RNAs in plastids. Science 347, 991–994 (2015).
Baldrich, P. et al. Plant Extracellular Vesicles Contain Diverse Small RNA Species and Are Enriched in 10- to 17-Nucleotide ‘Tiny’ RNAs. The Plant Cell 31, 315–324 (2019).
Buck, A. H. et al. Exosomes secreted by nematode parasites transfer small RNAs to mammalian cells and modulate innate immunity. Nat. Commun. 5, 5488 (2014).
Lv, Z. et al. Argonaute 2 in cell-secreted microvesicles guides the function of secreted miRNAs in recipient cells. PLoS One 9, e103599 (2014).
Rutter, B. D. & Innes, R. W. Extracellular Vesicles Isolated from the Leaf Apoplast Carry Stress-Response Proteins. Plant Physiol. 173, 728–741 (2017).
Regente, M. et al. Vesicular fractions of sunflower apoplastic fluids are associated with potential exosome marker proteins. FEBS Lett. 583, 3363–3366 (2009).
Cai, Q. et al. Plants send small RNAs in extracellular vesicles to fungal pathogen to silence virulence genes. Science 360, 1126–1129 (2018).
Johnson, N. R., Yeoh, J. M., Coruh, C. & Axtell, M. J. Improved Placement of Multi-mapping Small RNAs. G3 6, 2103–2111 (2016).
Kozomara, A., Birgaoanu, M. & Griffiths-Jones, S. miRBase: from microRNA sequences to function. Nucleic Acids Res. 47, D155–D162 (2019).
Dai, X., Zhuang, Z. & Zhao, P. X. psRNATarget: a plant small RNA target analysis server (2017 release). Nucleic Acids Res. 46, W49–W54 (2018).
Enright, A. J. et al. MicroRNA targets in Drosophila. Genome Biol 5, R1 (2003).
Kertesz, M., Iovino, N., Unnerstall, U., Gaul, U. & Segal, E. The role of site accessibility in microRNA target recognition. Nat. Genet. 39, 1278–1284 (2007).
Rehmsmeier, M., Steffen, P., Hochsmann, M. & Giegerich, R. Fast and effective prediction of microRNA/target duplexes. RNA 10, 1507–1517 (2004).
Allen, E., Xie, Z., Gustafson, A. M. & Carrington, J. C. microRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell 121, 207–221 (2005).
Srivastava, P. K., Moturu, T. R., Pandey, P., Baldwin, I. T. & Pandey, S. P. A comparison of performance of plant miRNA target prediction tools and the characterization of features for genome-wide target prediction. BMC Genomics 15, 348 (2014).
Huerta-Cepas, J. et al. Fast Genome-Wide Functional Annotation through Orthology Assignment by eggNOG-Mapper. Mol. Biol. Evol. 34, 2115–2122 (2017).
Huerta-Cepas, J. et al. eggNOG 4.5: a hierarchical orthology framework with improved functional annotations for eukaryotic, prokaryotic and viral sequences. Nucleic Acids Res. 44, D286–93 (2016).
Lorenz, R. et al. ViennaRNA Package 2.0. Algorithms for Molecular Biology 6 (2011).
Darty, K., Denise, A. & Ponty, Y. VARNA: Interactive drawing and editing of the RNA secondary structure. Bioinformatics 25, 1974–1975 (2009).
Micallef, L. & Rodgers, P. eulerAPE: drawing area-proportional 3-Venn diagrams using ellipses. PLoS One 9, e101717 (2014).
Acknowledgements
This work was supported by NSF Award IOS-1354154 to ACCW. MCT was supported by a University of Miami Isaac Bashevis Singer Scholarship. We thank Edward James and Celeste Banfill for useful discussions.
Author information
Authors and Affiliations
Contributions
A.C.C.W. and H.F. conceived of the study. All authors designed the research. M.C.T. performed the research. M.C.T., S.W. and A.C.C.W. analyzed the data. M.C.T. and A.C.C.W. wrote the paper. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Thompson, M.C., Feng, H., Wuchty, S. et al. The green peach aphid gut contains host plant microRNAs identified by comprehensive annotation of Brassica oleracea small RNA data. Sci Rep 9, 18904 (2019). https://doi.org/10.1038/s41598-019-54488-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-54488-1
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.