Abstract
The objective of this study was to correlate resistance mutations of extended spectrum beta-lactamases (ESBL) and AmpC beta-lactamases and virulence factors (VF) with 30-day mortality in patients treated with either piperacillin-tazobactam or carbapenems. A post-hoc analysis on 123 patients with ceftriaxone-resistant Escherichia coli and Klebsiella pneumoniae bacteremia treated empirically with piperacillin-tazobactam and carbapenems was performed. Beta-lactamase resistance mutations and VF were identified by whole genome sequencing (WGS). The primary endpoint was 30-day mortality. Multivariate analyses were performed using logistic regression. WGS showed diverse multilocus sequence types (MLST) in 43 K. pneumoniae strains, while ST131 predominated in E. coli strains (57/80). CTX-M was most commonly detected (76/80 [95%] of E. coli; 39/43 [91%] of K pneumoniae.), followed by OXA (53/80 [66%] of E. coli; 34/43 [79%] of K. pneumoniae). A significant correlation was found between the number of genes encoding third-generation cephalosporin-resistant beta-lactamases and 30-day mortality (p = 0.045). The positive association was not significant after controlling for empiric carbapenem, Pitt score 3 and K. pneumoniae (OR 2.43, P = 0.073). None of the VF was associated with 30-day mortality. No association was found between 30-day mortality and any ESBL and AmpC beta-lactamases or VF when piperacillin-tazobactam or carbapenems were given. No significant association between 30-day mortality and active empiric therapy was found.
Similar content being viewed by others
Introduction
Extended-spectrum and AmpC beta-lactamase Enterobacteriaceae, which lead to resistance to third generation cephalosporins, have become a global problem1,2,3. ESBL enzymes have been well studied since 1980’s, with hundreds of variants reported, including the TEM, SHV, and CTX-M families4,5,6. Multiple genetic mechanisms including plasmid transmission have facilitated the spread of CTX-M beta-lactamases7 worldwide, including in Singapore8. CTX-M genes have been commonly reported in Escherichia coli sequence type 131 (ST131)9, which was the leading cause of multidrug-resistant E. coli infections in the United States in 2011-201210. The AmpC beta-lactamases are also known to be transmissible among Enterobacteriaceae via plasmids11, and was previously reported in Singapore8. Partially due to their increased antibiotic resistance, infections by ESBL- and AmpC-producing bacteria result in increased mortality12,13,14,15.
ESBL- and AmpC-producing Enterobacteriaceae are susceptible to carbapenems, which have become the drug of choice for treatment of ESBL producing gram negative bacteremia16,17. However, the rapid rise of carbapenem-resistant Enterobacteriaceae globally18,19,20, partially driven by the increased use of carbapenems, has left little treatment options for common bacterial pathogens such as E. coli and Klebsiella pneumoniae21,22. To mitigate the rise of carbapenem resistance in Enterobacteriaceae, exploring possible effective alternative antibiotics to treat ESBL and AmpC beta-lactamase producers as a carbapenem-sparing strategy is a viable therapeutic approach.
The effectiveness of piperacillin-tazobactam against ESBL-producing bacteria has been described. The reduced efficacy in the presence of high bacterial inoculum has been described in vitro23,24,25,26. There are mixed results in clinical studies comparing efficacy of piperacillin-tazobactam and carbapenems27,28,29,30,31. An ongoing randomized trial aims to address the question of whether piperacillin-tazobactam is non-inferior to carbapenems for definitive treatment of ESBL gram negative bacteremia32. However, concerns of complex co-resistance mechanisms, including enzymes not well inhibited by tazobactam or clavulanate (e.g., plasmid-derived AmpC), may deter clinicians from using piperacillin-tazobactam for ESBL producing gram negative bacteremia33. This study aims to correlate resistance mutations and virulence factors of ESBL and AmpC beta-lactamases with 30-day mortality in patients with ceftriaxone-resistant E. coli and K. pneumoniae bacteremia when treated empirically with active piperacillin-tazobactam and carbapenems.
Method
Study setting and population
This analysis was reported according to the STROBE guidelines34. A post-hoc analysis was performed on ceftriaxone-resistant E. coli and K. pneumoniae bacteremic isolates from patients between August 2011 and May 2013 at Tan Tock Seng Hospital. The patients were included in a previously published retrospective cohort study by Ng et al.28. Briefly, patients with ceftriaxone-resistant E. coli and K. pneumoniae bacteremia were identified from electronic microbiology databases. For patients with multiple episodes of ceftriaxone-resistant E. coli or K. pneumoniae bacteremia, only the first episode was included. Patients were excluded if they had polymicrobial bacteremia, or did not receive at least 48 hours of empirical or definitive antimicrobial therapy. Other data collected included patient demographics, antibiotic susceptibility, empiric and definitive antibiotic therapy, source of bacteremia, Charlson’s co-morbidity index, Pitt bacteremia score, and 30-day mortality28. Ethics approval for this study was obtained from National Healthcare Group domain specific review boards (Approval number 2013/00083). Patient information was anonymized and de-identified prior to data collection and analysis in accordance with institutional guidelines and review board approvals, which included an approved waiver of informed consent.
In this study, we examined the effect of ESBL and AmpC resistance mutations, and virulence factors on 30-day mortality of patients with E. coli and K. pneumoniae bacteremia, treated with piperacillin-tazobactam or a carbapenem as active empiric therapy. We included Pitt bacteremia score to adjust for severity of illness.
Beta-lactamase genes that confer third-generation cephalosporin resistance according to the Bush and Jacoby functional classification were identified for further analyses35,36. AmpC genes were included as third-generation cephalosporin-resistant variants. Genes of the class SHV-OKP-LEN are closely related in sequence and grouped together in the resistance gene database we used (see below). Only some of these genes are ESBL, however, and not all alleles have been classified. We therefore performed separate analyses considering all SHV-OKP-LEN genes to be either ESBL or non-ESBL.
DNA extraction and sequencing library preparation
Each isolate from the clinical laboratory was streaked out to single colonies on LB-agar. A single colony was inoculated into Luria-Bertani broth (Gibco) and cultured overnight at 37 °C with agitation overnight. Cells from 1 ml of this culture were collected by centrifugation at 14, 000 g 1 minute. Genomic DNA was isolated from the resulting pelleted bacteria using the QIAamp DNA mini kit (Qiagen). DNA samples were quantified using a QUBIT 2.0 fluorometer (Invitrogen). Sequencing libraries were prepared with the Nextera XT Library Prep Kit (Illumina) according to the manufacturer’s instructions. The adapters were indexed using either the Nextera XT Index Kit (Illumina). Finally, 10 nM of each sample DNA sequencing library was pooled together and sequenced on a HiSeq 4000 (Illumina) with a 2 × 151 run.
Sequence analysis
Raw FASTQ reads were processed with using standard GERMS (Genome Institute of Singapore (GIS) Efficient and Rapid Microbial Sequencing) platform pipelines. Briefly, both de novo and reference-based analysis were used. De novo assemblies were performed using the Velvet assembler (version 1.2.10) with parameters optimized by Velvet Optimiser (k-mer length ranging from 81 to 127)37, scaffolded with Opera (version 1.4.1), and finished with FinIS (version 0.3)38,39. Genomes were initially annotated with Prokka (version 1.11)40. Resistance genes were called using BLASTN with a minimum identity of 90% over 90% of the gene length required for calling a gene present. MLST calls were made by using a custom BLASTN-based MLST caller.
Reference-based analyses were performed using the E. coli ST131 reference strain EC958 genome (GenBank accession NZ HG941719.1)41,42 as the reference for E. coli strains and K. pneumoniae HS11268 (GenBank accession NC 016845.1) as the reference for K. pneumoniae strains. FASTQ files were mapped using bwa (version 0.7.10)43; indel realignment and SNP (single nucleotide polymorphism) calling was performed using Lofreq* (version 2.1.2) with default parameters44.
Phylogenetic trees were made by calculating a dissimilarity matrix using SNPRelate45 and inferring a neighbor-joining tree using APE (version 3.5)46. All phylogenetic trees were visualized with GGTREE 3.247. All R packages were run in R (3.2.2). Multilocus sequence typing, resistance genes, and virulence factors were called directly from the raw FASTQ files using SRST2 (version 0.1.8)48 with default settings using the ARGAnnot database49 for resistance genes and the VFDB database50 for virulence factors. Resistance genes and MLST calls were cross-validated with the calls from the de novo assemblies, with discrepancies resolved by manual inspection of the raw data.
Novel MLST alleles and sequence types were submitted to Enterobase (for E. coli) and the Institut Pasteur MLST/cgMLST databases (for K. pneumoniae). All raw sequencing data has been deposited in GenBank under BioProject PRJNA431029.
Statistical analysis
Categorical variables were compared using the Chi-square or Fisher’s exact tests where applicable while continuous variables were compared using the Mann–Whitney U test between E. coli and K. pneumoniae bacteremia cases. A P-value of <0.05 was considered significant. The presence of beta-lactamase resistance mutations were also compared between empiric piperacillin-tazobactam and carbapenem groups using Chi-square tests. The number of third-generation cephalosporin-resistant beta-lactamase genes and mortality rate were compared using the linear-by-linear association test of Mantel-Haenszel. To assess the impact of number of resistant genes on patient mortality, a multivariate analysis was performed using binary logistic regression. Empiric carbapenem therapy, K. pneumoniae bacteremia as well as clinical (Pitt bacteremia score) and microbiological variables (Number of third-generation cephalosporin resistant genes in each isolate) with p values of <0.1 in univariate analysis were included into the model (Supplementary data); source of infection was omitted from multivariate analysis as the majority of cases (61%) were urinary. All analyses were performed using SPSS version 21 (IBM Corp, Armonk, NY, USA).
Results
The published clinical study by Ng et al. included 151 patients with ceftriaxone-resistant E. coli and K. pneumoniae bacteremia treated with active empiric monotherapy including either piperacillin-tazobactam or a carbapenem28. Only 124 bacteremic isolates were viable when revived from stocks from the microbiology department of our hospital. We included all of these 124 isolates for this study.
General features of strains
We performed whole genome sequencing on all 124 strains (80 E. coli, 44 K. pneumoniae). One of these strains was originally identified as an E. coli but, when sequenced, was very similar to K. pneumoniae reference sequences; this was excluded from further analysis. We obtained a median of 5.4 million paired end reads per strain (range 1.7–8.8 million), representing a coverage (assuming a 5Mbp genome) of 325 (range 102–526).
By multilocus sequence typing (MLST), the K pneumoniae strains were very diverse. Eight of 43 strains had novel MLST profiles. Among these 8 novel profiles, the most common profile accounted for only 3 strains. The most common MLST was ST307, accounting for 6 of 43 strains.
In contrast, by MLST the E. coli strains were dominated by ST131 (57/80 strains, including one single locus variant). The next most common sequence types, with 4 strains each, were ST38 and ST1193; neither is closely related to ST131.
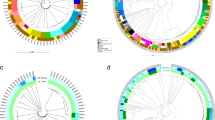
As indicated by the MLST results, the isolates obtained in this study were overall quite diverse with the exception of the ST131 strains. Separate whole genome SNP-based phylogenetic trees of the K. pneumoniae and E. coli strains are shown in Fig. 1; these demonstrate uniform clustering of related MLST types as well as the diversity of the strains across MLST types overall.
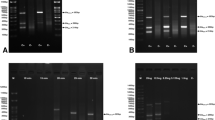
Phylogenetic trees for (a) E. coli and (b) K. pneumoniae isolates from this study. Neighbor joining trees were constructed from whole genome SNP profiles based on a reference-based analysis. MLST for each isolate is shown on the right. Black circles indicate yes (filled) or no (open) empirical carbapenem treatment and 30-day mortality, as indicated by the header. On the far right, colored squares indicate presence of a gene encoding the indicated resistance based on sequencing data.
Beta-lactamase gene analysis
The prevalence of resistance mutations in beta-lactamase genes is shown in Table 1, according to classifications on the publicly available database. The distribution of CTX-M and OXA were similar in both E. coli and K. pneumoniae groups; TEM-1D and SHV were predominantly found in K. pneumoniae samples. The frequency of AmpC was low in this population. Mortality rate and distribution of Pitt bacteremia, Charlson, and APACHE II scores in E. coli and K. pneumoniae groups were similar. 70% of K. pneumoniae strains were categorized as nosocomial compared to 26% with E. coli strains. The sources of bacteremia were similar between E. coli and K. pneumoniae groups (Table 1).
Table 2 shows the 30-day mortality of empiric piperacillin-tazobactam and carbapenem groups and presence of beta-lactamase resistance mutations. Overall there was no significant difference in 30-day mortality between empiric piperacillin-tazobactam and carbapenem groups. The difference was also not significant by the type of beta-lactamase resistance mutations present.
The correlation between 30-day mortality and number of third-generation cephalosporin-resistant beta-lactamase genes detected in the infecting strains were investigated. When considering all SHV-OKP-LEN genes to be ESBL, overall, the number of resistance genes correlated with 30-day mortality (p = 0.045) as shown in Fig. 2; this correlation was found in the empiric piperacillin-tazobactam subgroup (p = 0.040) but not in the empiric carbapenem subgroup (p = 0.504). In both E. coli and K. pneumoniae subgroups, the correlation was not significant (E. coli: p = 0.073; K. pneumoniae: p = 0.433). However, multivariate analysis showed that the number of third-generation cephalosporin-resistant genes was not a significant risk factor for 30-day mortality (odds ratio [OR] 2.43, 95% confidence interval [CI]: 0.92–6.42) after adjusting for empiric carbapenem therapy, K. pneumoniae bacteremia, and Pitt bacteremia score ≥3.
Similar results were obtained when we repeated these analyses considering all SHV-OKP-LEN genes to be non-ESBL (i.e. p = 0.054 for correlation of resistance genes with 30-day mortality; no significant association in multivariate analysis [OR 2.43, p = 0.073]).
Virulence gene analysis
An initial clustering showed that the E. coli and K. pneumoniae strains had different complements of virulence genes with clear clustering of strains by species; therefore, we analyzed the species separately.
For the 43 K. pneumoniae strains, a median of 60 virulence genes (range 27–81) was predicted in each strain; in total, 114 virulence genes were predicted in at least one of the K. pneumoniae strains. Of these 114 genes, 26 were found in all 43 strains; these were excluded from further analysis. Another 37 genes were predicted in only one of the 43 strains; these were also excluded. Among the remaining 51 virulence genes, the most prominent difference was the presence of genes for Yersiniabactin, an iron acquisition system51 present in 22/43 strains.
K. pneumoniae strains with Yersiniabactin virulence gene seemed to have higher severity of illness and 30-day mortality. (Fig. 3) However, we tested all of the 51 variable K. pneumoniae virulence genes using Fisher’s exact test for significant association with empiric carbapenem treatment or with 30-day mortality; none of these was significantly associated after a Bonferroni correction for multiple testing.
Heatmap of virulence factor calls from based on raw read mapping to VFDB for K. pneumoniae strains. Each row represents one K. pneumoniae strain, as indicated on the right of the heatmap. The strains are clustered based on their virulence factor profile, with the dendrogram displayed on the left. Each column represents one virulence factor, with selected general classes indicated below; a full list of virulence factors detected and their ordering in the heatmap can be found in Supplementary Table 1. Dark blue represents presence of a particular virulence factor in that strain; light blue indicates absence. Colored boxes on the right indicate empiric carbapenem, Pitt bacteremia score, and 30 day mortality associated with that strain.
For the 80 E. coli strains, a median of 237 virulence genes (range 164–304) was predicted in each strain; in total, 480 genes were predicted in at least one strain, 105 were present in 79 or 80 strains, and 39 were present in exactly one strain; these were excluded from further analysis. Major differences among the strains included genes encoding the ACE and SCI-I Type VI secretion systems, the ETT2 Type III secretion system, and a Type II secretion system, as well as P fimbriae and the general secretion pathway. In general, strains carried genes similar to either the ACE or the SCI-I Type VI secretion system (Fig. 4); those with the SCI-I Type VI system tended to carry P pili and to lack the ETT2 Type III system. The Type II secretion system and the general secretion pathway tended to co-occur in strains and were split among those strains carrying the ACE or the SCI-I Type VI secretion systems. The ST131 strains were those that tended to carry the SCI-I Type VI secretion system and to not carry the ETT2 Type III secretion system. We tested the association between the 336 variable genes and empiric carbapenem treatment or 30-day mortality. Again, none of these virulence factors was significantly associated with either of these clinical variables. We also tested for an association of these clinical variables with ST131; neither of these was significant. Finally, we tested for an association using only variable genes among ST131 or non-ST131 strains separately; again, none of these tests was significant. No statistically significant correlation was found between virulence factors and Pitt bacteremia score after Bonferroni p-value correction.
Heatmap of virulence factor calls from based on raw read mapping to VFDB for E. coli strains. Each row represents one E. coli strain, as indicated on the right of the heatmap. The strains are clustered based on their virulence factor profile, with the dendrogram displayed on the left. Each column represents one virulence factor, with selected general classes indicated below; a full list of virulence factors detected and their ordering in the heatmap can be found in Supplementary Table 1. Dark blue represents presence of a particular virulence factor in that strain; light blue indicates absence. Colored boxes on the right indicate empiric carbapenem, Pitt bacteremia score, and 30 day mortality associated with that strain.
Discussion
We initially found a significant correlation between increased 30-day mortality and the number of beta-lactamase genes that confer third-generation cephalosporin resistance. The correlation seemed to be present when piperacillin-tazobactam was used but not with carbapenem, suggesting that carbapenems may be more stable in presence of multiple broad spectrum beta-lactamases. However, after adjusting for empiric carbapenem therapy, K. pneumoniae bacteremia and Pitt bacteremia score 3, no significant association was found between the number of resistant genes and mortality. None of the individual beta-lactamase genes were associated with mortality according to the type of ESBL’s consistent with the findings from other studies52,53,54. Mortality rate seemed to be lower in the piperacillin-tazobactam group compared with the carbapenem group; propensity analysis in our previous publication revealed carbapenems were more likely to be used for treatment in sicker patients, explaining higher mortality in that group28.
In our multivariate analysis on 30-day mortality, we aimed to explore the effect of microbiological determinants on patient outcome in presence of clinical variables. High Pitt bacteremia score was found to be a significant risk factor for mortality (OR 3.53, 95% CI: 1.23–10.13), similar to findings in other studies27,30,31. Comorbidities were not significantly associated with patient mortality in the univariate analysis (OR 1.14, 95% CI: 0.96–1.36), and therefore was excluded from multivariate analysis. We did not look at the effect of definitive therapy (defined as antimicrobial therapy that was continued or commenced on the day that the antibiotic susceptibility results were reported to the clinicians) on mortality as most of the patients received carbapenems for definitive therapy (120/123 [98%]).
Tazobactam was reported to be active against TEM ESBL but not SHV ESBL6 although a high bacterial inoculum could confer resistance to tazobactam55. There was certainly a high prevalence of TEM and SHV in our study but we did not find that their presence affected 30-day mortality. In this study, SHV were included with OKP and LEN from the ARGAnnot database as they are relatively closely related by DNA sequence. We performed gene analyses similarly to other recent publications on determining resistance genes for ESBL organisms56,57. Among the strains positive for SHV-OKP-LEN, all predicted genes were SHV alleles. SHV-1, a commonly reported non-ESBL variant, was not found in any of our strains, although the SHV variants in our study were not all ESBL variants. We found that the ESBL classification of SHV-OKP-LEN genes did not significantly affect the correlations with mortality (p = 0.054 when SHV-OKP-LEN was omitted in correlation analyses) and multivariate analysis (OR 2.43, p = 0.073).
Tazobactam was found to be 100 time more active against CTX-M than clavulanate4, and CTX-M is the predominant ESBL in our study. However, multiple ESBL’s, hyper-production of non-ESBL beta-lactamases like TEM and SHV, and beta-lactamase production with porin loss may reduce the effectiveness of piperacillin-tazobactam4. This may explain our observation of significant association of number of resistant genes with 30-day mortality in the piperacillin-tazobactam group, which was lost after adjustment for confounders. Piperacillin-tazobactam was not considered active against AmpC beta-lactamases in vitro11, which was not very common in our study, although a meta-analysis of clinical studies did not find piperacillin-tazobactam was inferior to carbapenems in treating AmpC-producing gram negative bacteremia58.
As the E. coli and K. pneumoniae strains in our data set had very different virulence gene complements, we did not perform a combined analysis of virulence factors and mortality. This is unlike that of beta-lactamase resistance genes as both E. coli and K. pneumoniae were found to harbor similar beta-lactamases. There was no clear association between virulence factors and 30-day mortality in either E. coli or K. pneumoniae groups. This is in contrast with the findings from 3 clinical studies on E. coli, which found association between certain virulence factors and mortality; particularly papGII and hemolysin production were reported to have a protective effect on mortality52,59,60, whereas ibeA (which promotes invasion)52 and fyuA (encoding for Yersiniabactin siderophore receptor)59 increased mortality. Of note, among our E. coli strains, at least one hemolysin gene was found in all 80 strains, and 77 strains carried the fyuA gene and most of the genes annotated as required for Yersiniabactin production (all death cases were among these 77). Only two strains carried the ibeA gene; neither was from a death case. Forty five E. coli strains carried the papG gene, of which 44 were papGII and 1 was papGIII; 6 of 13 death cases were associated with strains carrying papGII. Thus none of these previously reported E. coli virulence genes were significantly associated with mortality in our data set. The role of virulence factors on mortality is still unclear in other studies60,61,62, although virulence factors and resistance genes were reported to affect each other in various ways. Specifically, a higher proportion of ESBL-producing K. pneumoniae was found to produce both type 1 and 3 fimbrial adhesins than non-ESBL strains, whereas E. coli with OXA-10 like and SFO-1 beta-lactamases were found to have a statistically significant fitness cost63. A study by Lefort et al. suggested that host factors and portal of entry outweighed bacterial determinants on mortality of patients with E. coli bacteremia64. In our multivariate analysis, only high Pitt bacteremia score was associated with mortality, suggesting that bacterial determinants may not be a significant influence when active piperacillin-tazobactam or carbapenems are given empirically.
Our study has several limitations. It was a retrospective analysis and we may not be able to control for all possible confounders. We retrieved only 124 isolates from our microbiology department out of the 151 patients included in the previous study28, and therefore we could not analyze the resistance mutations of the remaining 27 patients. We believe however that this will not affect the validity of our results. Our study sample size was low overall, so some subgroups (e.g. AmpC) likely did not have enough power to detect associations. At our current available sample size, the power to detect the same difference is 0.73. To detect a 20% difference in mortality between patients treated with piperacillin-tazobactam and carbapenem with 80% power, a sample of 160 patients would be required. Therefore, the primary value of this study in looking at effects of type of antibiotics, resistance genes, and virulence factors on mortality remains exploratory. We did not evaluate the minimum inhibitory concentration of empiric antibiotic used as it was not routinely tested in all cases, which could be a significant factor on 30-day mortality. Lastly, this study only included E. coli and K. pneumoniae samples with ESBL and/or AmpC beta-lactamases and may not be generalizable to patients with other types of infections. We did not examine the presence of efflux pumps or porin loss which may modify the efficacy of both piperacillin-tazobactam and carbapenems.
Conclusion
In conclusion, we did not find significant association between 30-day mortality and any ESBL and AmpC beta-lactamases, as well as any virulence factors when patients were given either piperacillin-tazobactam or carbapenems empirically. New studies in this area are much needed as clinical trials are underway to investigate effectiveness of piperacillin-tazobactam for treatment of ESBL-producing gram negative bacteremia33.
Data availability statement
All data generated or analysed during this study are included in this published article (and its Supplementary Information files).
References
System, N. N. I. S. National Nosocomial Infections Surveillance (NNIS) System Report, data summary from January 1992 through June 2004, issued October 2004. Am J Infect Control. 32, 470–485 (2004).
Cantoń, R. et al. Prevalence and spread of extended-spectrum β -lactamase-producing Enterobacteriaceae in Europe. Clin. Microbiol. Infect. 14, 144–153 (2008).
Sheng, W.-H., Badal, R. E. & Hsueh, P.-R. Distribution of extended-spectrum beta-lactamases, AmpC beta-lactamases, and carbapenemases among Enterobacteriaceae isolates causing intra-abdominal infections in the Asia-Pacific region: results of the study for Monitoring Antimicrobial Resistance Trend. Antimicrob. agents chemotherapy 57, 2981–2988 (2013).
Paterson, D. L. & Bonomo, R. A. Extended-Spectrum beta-Lactamases: a Clinical Update. Clin. Microbiol. Rev. 18, 657–686 (2005).
Bradford, P. Extended spectrum betalactamase in the 21 century: characterization, epidemiology, and detection of this impor- tant resistant threat. Clin. Microbiol Rev 14, 933–951 (2001).
Livermore, D. M. Beta-Lactamases in laboratory and clinical resistance. Clin. microbiology reviews 8, 557–84 (1995).
Rossolini, G. M., D’Andrea, M. M. & Mugnaioli, C. The spread of CTX-M-type extended-spectrum beta-lactamases. Clin. microbiology infection: official publication Eur. Soc. Clin. Microbiol. Infect. Dis 14(Suppl 1), 33–41 (2008).
Tan, T. Y., Ng, L. S. Y., He, J. & Hsu, L. Y. CTX-M and ampC beta-lactamases contributing to increased prevalence of ceftriaxone-resistant Escherichia coli in Changi General Hospital, Singapore. Diagn. microbiology infectious disease 66, 210–213 (2010).
Nicolas-Chanoine, M.-H., Bertrand, X. & Madec, J.-Y. Escherichia coli ST131, an intriguing clonal group. Clin. microbiology reviews 27, 543–574 (2014).
Johnson, J. R., Porter, S., Thuras, P. & Castanheira, M. The pandemic H30 subclone of sequence type 131 (ST131) as the leading cause of multidrug-resistant Escherichia coli infections in the United States (2011-2012). Open Forum Infect. Dis. 4, 1–11 (2017).
Jacoby, G. A. A. C. B.-L. Clin. Microbiol. Rev. 22, 161–182 (2009).
Tumbarello, M. et al. Costs of bloodstream infections caused by Escherichia coli and influence of extended-spectrum-beta- lactamase production and inadequate initial antibiotic therapy. Antimicrob. Agents Chemother. 54, 4085–4091 (2010).
Leistner, R. et al. Bloodstream infection due to extended-spectrum beta-lactamase (ESBL)-positive K. pneumoniae and E. coli: an analysis of the disease burden in a large cohort. Infect. 42, 991–997 (2014).
Schwaber, M. J. & Carmeli, Y. Mortality and delay in effective therapy associated with extended-spectrum β -lactamase production in Enterobacteriaceae bacteraemia: A systematic review and meta-analysis. J. Antimicrob. Chemother. 60, 913–920 (2007).
Zhang, Q. et al. Bacteraemia due to AmpC beta-lactamase-producing Escherichia coli in hospitalized cancer patients: risk factors, antibiotic therapy, and outcomes. Diagn. microbiology infectious disease 88, 247–251 (2017).
Paterson, D. L. et al. Antibiotic Therapy for Klebsiella pneumoniae Bacteremia: Implications of Production of Extended-Spectrum -Lactamases. Clin. Infect. Dis. 39, 31–37 (2004).
Harris, P. N. A. & Ferguson, J. K. Antibiotic therapy for inducible AmpC beta-lactamase-producing Gram- negative bacilli: what are the alternatives to carbapenems, quinolones and aminoglycosides? Int. J. Antimicrob. Agents 40, 297–305 (2018).
Nordmann, P., Naas, T. & Poirel, L. Global spread of Carbapenemase-producing Enterobacteriaceae. Emerg. infectious diseases 17, 1791–1798 (2011).
Kumarasamy, K. K. et al. Emergence of a new antibiotic resistance mechanism in India, Pakistan, and the UK: a molecular, biological, and epidemiological study. The Lancet. Infect. diseases 10, 597–602 (2010).
Rogers, B. A., Aminzadeh, Z., Hayashi, Y. & Paterson, D. L. Country-to-country transfer of patients and the risk of multi-resistant bacterial infection. Clin. infectious diseases: an official publication Infect. Dis. Soc. Am. 53, 49–56 (2011).
McLaughlin, M. et al. Correlations of antibiotic use and carbapenem resistance in enterobacteriaceae. Antimicrob. Agents Chemother. 57, 5131–5133 (2013).
Butler, M. S., Blaskovich, M. A. & Cooper, M. A. Antibiotics in the clinical pipeline at the end of 2015. The J. antibiotics 70, 3–24 (2017).
Thomson, K. S. & Moland, E. S. Cefepime, piperacillin-tazobactam, and the inoculum effect in tests with extended- spectrum beta-lactamase-producing Enterobacteriaceae. Antimicrob. agents chemotherapy 45, 3548–3554 (2001).
Bonfiglio, G. & Livermore, D. M. Inoculum effects on Etests and agar dilution minimum inhibitory concentrations. Piperacillin and piperacillin-tazobactam against Staphylococcus aureus. Diagn. microbiology infectious disease 19, 163–166 (1994).
Harada, Y. et al. In vitro and in vivo activities of piperacillin-tazobactam and meropenem at different inoculum sizes of ESBL-producing Klebsiella pneumoniae. Clin. Microbiol. Infect. 20, O831–O839 (2014).
Burgess, D. S. & Hall, R. Gn In vitro killing of parenteral beta-lactams against standard and high inocula of extended- spectrum beta-lactamase and non-ESBL producing Klebsiella pneumoniae. Diagn. microbiology infectious disease 49, 41–46 (2004).
Tamma, P. D. et al. Carbapenem therapy is associated with improved survival compared with piperacillin-tazobactam for patients with extended-spectrum β -lactamase bacteremia. Clin. Infect. Dis. 60, 1319–1325 (2015).
Ng, T. M. et al. Empiric piperacillin-tazobactam versus carbapenems in the treatment of bacteraemia due to extended- spectrum beta-lactamase-producing enterobacteriaceae. PLoS ONE 11, 1–11 (2016).
Ofer-Friedman, H. et al. Carbapenems Versus Piperacillin-Tazobactam for Bloodstream Infections of Nonurinary Source Caused by Extended-Spectrum Beta-Lactamase-Producing Enterobacteriaceae. Infect. Control. Hosp. Epidemiol. 36, 981–985 (2015).
Tsai, H.-Y. et al. Carbapenems and piperacillin/tazobactam for the treatment of bacteremia caused by extended- spectrum beta-lactamase-producing Proteus mirabilis. Diagn. microbiology infectious disease 80, 222–226 (2014).
Rodriguez-Bano, J., Navarro, M. D., Retamar, P., Picon, E. & Pascual, A. beta-Lactam/beta-lactam inhibitor combinations for the treatment of bacteremia due to extended-spectrum beta-lactamase-producing Escherichia coli: a post hoc analysis of prospective cohorts. Clin. infectious diseases: an official publication Infect. Dis. Soc. Am. 54, 167–174 (2012).
Harris, P. N. et al. Meropenem versus piperacillin-tazobactam for definitive treatment of bloodstream infections due to ceftriaxone non-susceptible Escherichia coli and Klebsiella spp (the MERINO trial): Study protocol for a randomised controlled trial. Trials 16, 1–8 (2015).
Harris, P. N., Tambyah, P. A. & Paterson, D. L. Beta-lactam and beta-lactamase inhibitor combinations in the treatment of extended-spectrum β -lactamase producing Enterobacteriaceae: Time for a reappraisal in the era of few antibiotic options? The Lancet Infect. Dis. 15, 475–485 (2015).
Elm, E. V. et al. guidelines for reporting observational studies Strengthening the reporting of observational studies in epidemiology (STROBE) statement: guidelines for reporting observational studies. Br. Med. J. 335, 19–22 (2007).
Bush, K. & Jacoby, G. A. Updated functional classification of beta-lactamases. Antimicrob. agents chemotherapy 54, 969–976 (2010).
Jacoby, G. A. Beta-lactamase nomenclature. Antimicrob. agents chemotherapy 50, 1123–1129 (2006).
Zerbino, D. R. Using the Velvet de novo assembler for short-read sequencing technologies. Curr. Protoc. Bioinforma. (2010).
Gao, S., Nagarajan, N. & Sung, W. K. Opera: Reconstructing optimal genomic scaffolds with high-throughput paired-end sequences. In Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), vol. 6577 LNBI, 437–451 (2011).
Gao, S., Bertrand, D. & Nagarajan, N. FinIS: Improved in silico finishing using an exact quadratic programming formulation. In Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics) 7534(LNBI), 314–325 (2012).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30(14), 2068–2069 (2014).
Forde, B. M. et al. The complete genome sequence of Escherichia coli EC958: a high quality reference sequence for the globally disseminated multidrug resistant E. coli O25b:H4-ST131 clone. PloS one 9, e104400 (2014).
Totsika, M. et al. Insights into a multidrug resistant Escherichia coli pathogen of the globally disseminated ST131 lineage: genome analysis and virulence mechanisms. PloS one 6, e26578 (2011).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinforma. (Oxford, England) 25, 1754–1760 (2009).
Wilm, A. et al. LoFreq: a sequence-quality aware, ultra-sensitive variant caller for uncovering cell-population heterogeneity from high-throughput sequencing datasets. Nucleic acids research 40, 11189–11201 (2012).
Zheng, X. et al. A high-performance computing toolset for relatedness and principal component analysis of SNP data. Bioinforma. (Oxford, England) 28, 3326–3328 (2012).
Paradis, E., Claude, J. & Strimmer, K. APE: Analyses of Phylogenetics and Evolution in R language. Bioinformatics 20(2), 289–290 (2004).
Yu, G., Smith, D. K., Zhu, H., Guan, Y. & Lam, T. T.-Y. ggtree: an r package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 8, 28–36 (2017).
Inouye, M. et al. SRST2: Rapid genomic surveillance for public health and hospital microbiology labs. Genome Medicine 6(11) (2014).
Gupta, S. K. et al. ARG-ANNOT, a new bioinformatic tool to discover antibiotic resistance genes in bacterial genomes. Antimicrob. agents chemotherapy 58, 212–220 (2014).
Chen, L., Zheng, D., Liu, B., Yang, J. & Jin, Q. VFDB 2016: hierarchical and refined dataset for big data analysis–10 years on. Nucleic acids research 44, 694–7 (2016).
Lawlor, M. S., O’connor, C. & Miller, V. L. Yersiniabactin is a virulence factor for Klebsiella pneumoniae during pulmonary infection. Infect. immunity 75, 1463–1472 (2007).
Rodríguez-Baño, J. et al. Outcome of bacteraemia due to extended-spectrum β -lactamase-producing Escherichia coli: Impact of microbiological determinants. J. Infect. 67, 27–34 (2013).
Wu, U.-I., Wang, J.-L., Chen, W.-C., Chang, S.-C. & Chen, Y.-C. Risk factors and outcomes of Escherichia coli bacteremia caused by strains that produce CTX-M or non-CTX-M extended-spectrum-beta-lactamases. Eur. journal clinical microbiology & infectious diseases: official publication Eur. Soc. Clin. Microbiol. 30, 33–39 (2011).
Miguel, G. S. et al. Clinical variables associated with the isolation of Klebsiella pneumoniae expressing different extended-spectrum beta-lactamases. Clin. microbiology infection: official publication Eur. Soc. Clin. Microbiol. Infect. Dis. 13, 532–538 (2007).
Jacoby, G. A. & Munoz-price, L. S. The New Beta-lactamases. The New Engl. J. Medicine 352, 380–391 (2005).
Holt, K. E. et al. Genomic analysis of diversity, population structure, virulence, and antimicrobial resistance in Klebsiella pneumoniae, an urgent threat to public health. Proc. Natl. Acad. Sci. United States Am. 112, 3574–81 (2015).
Long, S. W. et al. Population Genomic Analysis of 1,777 Extended-Spectrum Beta-Lactamase-Producing Kleb- siella pneumoniae Isolates, Houston, Texas: Unexpected Abundance of Clonal Group 307. mBio 8, 00489–17 (2017).
Harris, P. N. et al. Carbapenems versus alternative antibiotics for the treatment of bloodstream infections caused by Enterobacter, Citrobacter or Serratia species: A systematic review with meta-analysis. J. Antimicrob. Chemother. 71, 296–306 (2016).
Mora-Rillo, M. et al. Impact of virulence genes on sepsis severity and survival in Escherichia coli bacteremia. Virulence 6, 93–100 (2015).
Hekker, T. A. et al. Role of bacterial virulence factors and host factors in the outcome of Escherichia coli bacteraemia. Eur. journal clinical microbiology & infectious diseases: official publication Eur. Soc. Clin. Microbiol. 19, 312–316 (2000).
Bert, F. et al. Molecular epidemiology of Escherichia coli bacteremia in liver transplant recipients. Transpl. infectious disease: an official journal Transplantation Soc. 13, 359–365 (2011).
Zheng, X., Wang, J.-F., Xu, W.-L., Xu, J. & Hu, J. Clinical and molecular characteristics, risk factors and outcomes of Carbapenem-resistant Klebsiella pneumoniae bloodstream infections in the intensive care unit. Antimicrob. resistance infection control 6, 102 (2017).
Beceiro, A., Tomaś, M. & Bou, G. Antimicrobial Resistance and Virulence: a Successful or Deleterious Association in the Bacterial World? (2013).
Lefort, A. et al. Host Factors and Portal of Entry Outweigh Bacterial Determinants To Predict the Severity of Escherichia coli Bacteremia (2011).
Poirel, L., Naas, T. & Nordmann, P. Diversity, epidemiology, and genetics of class D beta-lactamases. Antimicrob. Agents Chemother. 54, 24–38 (2010).
Bethel, C. R. et al. Inhibition of OXA-1 beta-lactamase by penems. Antimicrob. Agents Chemother. 52, 3135–3143 (2008).
Acknowledgements
This study was funded by Communicable Disease Centre Centre Grant number CGSFP15002. The grant funders had no role in the design, analysis, and writing of the manuscript or the decision to publish. The sequencing was done in collaboration with the GERMS platform at Genome Institute of Singapore. We thank Balamurugan Periaswamy for help with whole genome sequencing. Sequence analysis was performed with pipelines partially supported by the National Medical Research Council, Ministry of Health, Singapore grant no. NMRC/CIRG/1357/2013. We thank the team of curators of the Institut Pasteur MLST and whole genome MLST databases for curating the data and making them publicly available at http://bigsdb.pasteur.fr.
Author information
Authors and Affiliations
Contributions
D.L. and T.M.N. were responsible for the conception and design of the study. D.L., T.M.N. and S.L.C. were responsible for the acquisition of the data. S.L.C., J.G.X.W. and S.T.H. analysed the results. All authors wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Heng, S.T., Chen, S.L., Wong, J.G.X. et al. No association between resistance mutations, empiric antibiotic, and mortality in ceftriaxone-resistant Escherichia coli and Klebsiella pneumoniae bacteremia. Sci Rep 8, 12785 (2018). https://doi.org/10.1038/s41598-018-31081-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-31081-6
This article is cited by
-
Association between the rate of third generation cephalosporin-resistant Escherichia coli and Klebsiella pneumoniae and antibiotic consumption based on 143 Chinese tertiary hospitals data in 2014
European Journal of Clinical Microbiology & Infectious Diseases (2020)
-
Prevalence of and risk factor for community-onset third-generation cephalosporin-resistant Escherichia coli bacteremia at a medical center in Taiwan
BMC Infectious Diseases (2019)
-
The higher prevalence of extended spectrum beta-lactamases among Escherichia coli ST131 in Southeast Asia is driven by expansion of a single, locally prevalent subclone
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.