Abstract
Autoimmune hepatitis (AIH) is a chronic inflammatory liver disease characterized by an autoimmune reaction to hepatocytes. A single nucleotide polymorphism in the 3′ untranslated region of HLA-DPB1, rs9277534, is associated with HLA-DPB1 expression. rs9277534 has been linked to hepatitis B virus recovery/persistence and the risk of graft-versus-host disease with HLA-DPB1 mismatching transplantation of hematopoietic cells, but its role along with that of HLA-DP expression in AIH have not been fully clarified. We genotyped rs9277534 in 146 Japanese patients with AIH and 326 healthy subjects. HLA-DPB1 expression was determined by quantitative PCR. HLA-DPB1 expression was significantly higher for rs9277534G than for rs9277534A (P < 0.05). rs9277534 genotype was in strong linkage disequilibrium with the HLA-DPB1 allele (pairwise D′ = 0.82–1.00). Although HLA-DP alleles were not significantly associated with AIH, the frequency of the rs9277534G allele was significantly higher in AIH patients compared with healthy subjects (P = 0.002, odds ratio [OR] = 1.56). Logistic regression analysis revealed that the HLA-DRB1*04:05 allele (P < 0.001, OR = 4.61) and rs9277534 (P = 0.004, OR = 1.67) were independently associated with AIH susceptibility. rs9277534G in the HLA-DP gene is an eQTL that affects gene expression and may contribute to AIH susceptibility.
Similar content being viewed by others
Introduction
Autoimmune hepatitis (AIH) is a chronic inflammatory disease of the liver characterized by elevated transaminase, hypergammaglobulinemia, the presence of serum autoantibody, and necroinflammatory changes on hepatic histology1,2. Two types of AIH have been defined by serum autoantibody profiles3,4. Type 1 AIH is characterized by the presence of antinuclear antibodies and/or anti-smooth muscle antibodies and is the major form of AIH in Japan. Although the disease is believed to result from a combination of genetic and environmental factors, its exact etiology remains unclear. Many studies have implicated human leukocyte antigen (HLA) class II genes, especially HLA-DRB1 alleles, as major components in the genetic predisposition to AIH. For instance, the DRB1*04:04 and DRB1:04:05 alleles are linked to susceptibility in Mexican5 and Japanese6,7,8 and Korean9 populations, respectively, and studies including recent genome-wide association analyses have shown that DRB1*03:01 and DRB1*04:01 contribute to AIH susceptibility in European populations10,11,12.
HLA-DP genes have not been examined for their impact on human disease relative to HLA-DR and -DQ because they are less polymorphic and HLA-DP cell surface expression levels are believed to be lower than those of HLA-DR or -DQ13,14,15. Several genome-wide association studies have associated HLA-DP gene variations with chronic hepatitis B virus (HBV) infection in Asians16,17,18,19,20. Moreover, two single nucleotide polymorphisms (SNPs) in the 3′ untranslated region of HLA-DPB1, rs9277534 and rs9277535, are related to viral clearance in the U.S. population21 and in Asians16, respectively. Petersdorf et al. recently found that the risk of graft-versus-host disease with HLA-DPB1 mismatching transplantation of hematopoietic cells was influenced by rs927753422. Other current findings21,22 suggest that this SNP is associated with HLA-DP expression levels and may influence the strength of autoimmune responses, thus representing a new and crucial role of HLA in human disease. We therefore sought to determine whether rs9277534 and HLA-DP expression were associated with AIH in a Japanese population.
Results
HLA-DPB1 Alleles and rs9277534 in AIH Patients and Controls
The frequency of the major G allele in rs9277534 was significantly increased in AIH patients as compared with 326 healthy subjects (65.1% vs. 54.3%, P = 0.002, odds ratio [OR] = 1.56) (Table 1). Genotype frequencies also differed significantly between AIH patients and controls with a dominant model of inheritance (Table 2). The frequency of the GG or GA genotype of rs9277534 in AIH was significantly higher than that in controls (P = 0.0034, OR = 2.39).
HLA-DP allele frequency between 146 patients with AIH and 201 control subjects from our prior study23 was compared next (Table 3). DPB1*04:02 had a lower frequency in patients, but this difference was not significant after correction for multiple testing (8% vs. 13%; P = 0.023, corrected P = 0.28). In our cohort, the HLA-DRB1*04:05 allele is strongly associated with AIH susceptibility (P < 0.001), as is rs9277534 (P = 0.004). Logistic regression analysis revealed that the HLA-DRB1*04:05 allele (P < 0.001, OR = 4.61, 95% CI = 2.86–7.41) and rs9277534 (P = 0.004, OR = 1.67, 95% CI = 1.18–2.37) were independently associated with AIH susceptibility.
Association between HLA-DPB1 and rs9277534 Allele Type
The ratio of GG/GA/AA genotypes in rs9277534 was 0.301/0.485/0.215 and the respective allele frequency of G and A was 0.543 and 0.457 in 326 healthy subjects (Tables 1, 2). These were in deviation of Hardy-Weinberg equilibrium (P = 0.914). We carried out HLA-DPB1 allele typing in 146 AIH patients and analyzed for associations between HLA-DPB1 and rs9277534 allele type (Table 4). HLA-DPB1*02:01, DPB1*02:02, DPB1*04:01, DPB1*04:02, and DPB1*17:01 alleles were associated with rs9277534A, while HLA-DPB1*03:01, DPB1*05:01, DPB1*06:01, DPB1*09:01, DPB1*13:01, DPB1*14:01, and DPB1*19:01 were linked with rs9277534G. The genotype of rs9277534 was in strong linkage disequilibrium with the HLA-DPB1 allele (pairwise D′ = 0.82–1.00). The HLA-DPB1 alleles could be divided into two groups by the rs9277534 allele, which was consistent with a recent report21.
Association between rs9277534 and Clinical Findings
There were no significant differences in the serum levels of albumin, alanine aminotransferase, aspartate aminotransferase, or bilirubin, nor was there one in the frequency of antinuclear antibody positivity, between patients with rs9277534G and rs9277534A.
Correlation of HLA-DPB1 Expression Level with rs9277534 Genotypes
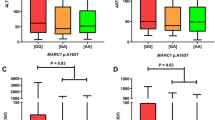
The rs9277534 genotype reportedly influences the expression level of HLA-DPB1 in European-Americans and African-Americans21,22. To validate these findings, the differences in HLA-DPB1 mRNA levels among groups of 10 healthy controls with rs9277534 AA, AG, or GG were determined by quantitative PCR. The median relative quantity of HLA-DPB1 mRNA level was significantly higher in the rs9277534 GG group than in the AA or GA groups (GG: 1.70 vs. AA: 1.45; P = 0.004, and vs. AG: 1.48; P = 0.019) (Fig. 1A). Moreover, a significant difference was detected for HLA-DPB1 mRNA between the GG + AG and AA genotypes (GG + AG: 1.66 vs. AA: 1.45; P = 0.035) (Fig. 1B). Our results confirmed that rs9277534 G-linked HLA-DPB1 mRNA levels were significantly higher than rs9277534 A-linked ones in a Japanese population21,22.
HLA-DPB1 mRNA levels correlate significantly with the rs9277534 allele in the 3′ untranslated region of HLA-DPB1. (A) HLA-DPB1 mRNA levels in individuals with rs9277534GG (n = 10) were significantly higher than in those with rs9277534AA (n = 10; P = 0.004) or rs9277534AG (n = 10; P = 0.019). (B) HLA-DPB1 mRNA levels were significantly higher for the rs9277534GG + AG genotypes than for the rs9277534AA genotype (P = 0.035).
Discussion
In the present study, the rs9277534G allele in the 3′ untranslated region of HLA-DBP1 was strongly associated with AIH compared with healthy subjects in the context of disease susceptibility, although no statistically significant DPB1 allele was detected. Logistic regression analysis showed that rs9277534G was an independent susceptibility allele to AIH in addition to the HLA-DRB1*04:05 allele. rs9277534 is an expression quantitative trait locus (eQTL) that affects gene expression. eQTL SNPs are linked to transcription factor binding, DNA methylation, alternative splicing, small RNA, and other regulatory variants. Therefore, although the functional mechanisms of rs9277434 have not been fully elucidated, this SNP could be indirectly correlated with regulatory elements such as transcription factors.
Our study also confirmed that HLA-DPB1 mRNA expression in cells from Japanese donors correlated with rs9277534A/G genotypes, as reported previously21,22. The association of HLA-DP expression with AIH and HBV infection21, both of which chronic liver diseases, is intriguing. Although HBV infection is sometimes complicated with autoimmune diseases, its precise mechanisms are unknown24. Earlier studies revealed a possible association between AIH and immune T helper (Th)1/Th2 cell balance. Exposure to interleukin (IL)-12 leads to differentiation of Th1 cells secreting interferon-γ, which in turn activates monocytes and cytotoxic CD8 T-cells and promotes natural-killer-cell functions. Interferon-γ also increases major histocompatibility class I and induces major histocompatibility class II expression by hepatocytes, further exacerbating inflammation. IL-4 exposure evokes Th2 differentiation. Th2 cells secrete IL-4, IL-10, and IL-13, all cytokines that enable B-cell maturation into plasma cells with the consequent production of autoantibodies. Ultimately, autoantibodies are involved in antibody-mediated cellular cytotoxicity and complement activation25. It is possible that high HLA-DP expression favors a Th2 response characterized by vigorous antibody production, leading to AIH. The titers of anti-liver-specific membrane lipoprotein, asialoglycoprotein receptor, and alcohol dehydrogenase are reportedly correlated with biochemical and histological indices of disease severity25. The roles of these autoantibodies in the autoimmune liver attack derive from the finding that hepatocytes are the targets of such immunoglobulins and are susceptible to cytotoxicity. Lastly, whereas Th1 responses are more often observed in HBV recovery, Th2 responses along with poor cytotoxic T-cell activity lead to chronicity26. Conversely, lower HLA-DP expression favoring Th1 might be disadvantageous in AIH but effective in HBV recovery. Taken together, higher HLA-DP expression may be deleterious for liver disease in chronic hepatitis B and AIH, while the rs9277534G allele may play an important role in the pathogenesis of chronic hepatitis. Further investigation is needed to clarify this relationship.
The mechanisms underlying the pathogenesis of AIH are not completely elucidated, but several lines of evidence support a strong influence of the large and highly polymorphic HLA region related to adaptive immunity. The most conclusive association with AIH is related to the functional role of HLA class II molecules, particularly HLA-DRB molecules, such as specific allele differences in DRB1 polymorphisms5,6,7,9,10,11 and notable amino acid sequence motifs within DRB1 polypeptides7,10,27,28. For example, lysine or arginine at position 71, valine or glycine at position 86, and histidine at position 13 reportedly contribute to AIH susceptibility in Northern European Caucasians, Argentina and Brazil, and the Japanese, respectively. Unlike HLA-DR alleles, however, HLA associations with HLA-DP alleles have been insufficiently addressed in AIH, presumably due to a smaller role of DP molecules in immune responses and relatively lower cell surface expression levels13,14,15.
In conclusion, rs9277534G in the HLA-DP gene is an eQTL affecting gene expression that may contribute to AIH susceptibility. Further large-scale studies are required to determine how varied expression of this molecule affects the immune response and development of AIH across ethnicities.
Methods
Subjects
Between January 2001 and August 2015, 146 patients with type 1 AIH were enrolled for this study and summarized in Table 5. A total of 326 previously described volunteer healthy subjects were included as well29. All controls were hospital staff whose liver function tests were within normal levels and who had indicated the absence of any major illness in a standard questionnaire. The racial background of all individuals was Japanese. All AIH patients had been diagnosed according to the scoring system of the International Autoimmune Hepatitis Group30 and were classified as type 1 AIH based on antibody profiles. Anti-nuclear antibody titers were determined by immunofluorescence using HEp-2 cells, in which a titer of ≥1:80 was considered positive31. All patients were negative for the hepatitis B surface antigen and antibodies to the hepatitis B core antigen, hepatitis C virus, and human immunodeficiency virus. Other liver diseases and AIH-primary biliary cholangitis overlap syndromes were excluded. All subjects provided written informed consent for this study, which was approved by the Institutional Review Board of Shinshu University Hospital (approval number: 527). The investigation was conducted according to the principals of the Declaration of Helsinki.
Genotyping and Linkage Analysis
Genomic DNA from patients and controls was isolated from whole blood samples using QuickGene-800 assays (Fujifilm, Tokyo, Japan). HLA typing in AIH patients was carried out using a Luminex multi-analyzer profiling system with a LAB type SSO One Lambda typing kit (One Lambda, Inc. Canoga Park, CA). HLA genotypes were determined by sequence-based typing, as earlier described32. The rs9277534 SNP in the HLA-DP locus was genotyped using a TaqMan 5′ exonuclease assay (Applied Biosystems, Tokyo, Japan). The polymerase chain reaction (PCR) was performed with the StepOne Plus Real-Time PCR System (Applied Biosystems) following the manufacturer’s instructions. Pairwise linkage disequilibrium patterns between HLA-DPB1 alleles and rs9277534 (A/G) were assessed by the Arlequin program (ver. 3.5.2.2) (http://lgb.unige.ch/arlequin/)33. The linkage of rs9277534 to HLA-DPB1 was examined in all AIH patients pursuant to a previous report23.
Measurement of HLA-DPB1 Expression
Blood for RNA analysis was collected using cellular preparation tubes (Becton Dickinson, Franklin Lakes, NJ) from groups of 10 healthy individuals with rs9277534A/A, -A/G, or -G/G genotypes and isolated using a QIAamp RNA Blood Mini Kit (Qiagen, Valencia, CA) according to the manufacturer’s instructions. Single-strand cDNA was synthesized using a high-capacity RNA-to-cDNA kit (Applied Biosystems). Relative quantities of HLA-DPB1 (LifeTechnologies assay ID: Hs00157955_m1) were determined using TaqMan® Gene Expression assays, and the expression of glucuronidase beta (LifeTechnologies assay ID: Hs99999908_m1) was used to normalize gene expression level with the ABI StepOne Plus Real Time PCR system (Applied Biosystems). Real-time data plots were presented as normalized individual data points (2−deltaCt [delta Ct = Ct gene of interest - Ct endogenous control])34.
Statistical analysis
Allele and genotype frequencies along with Hardy-Weinberg equilibrium and linkage disequilibrium were assessed using SNPStats software (Catalan Institute of Oncology, Barcelona, Spain; http://bioinfo.iconcologia.net/SNPstats)35 and Haploview 4.1 software36. The significance of an association was evaluated using chi-square analysis or Fisher’s exact test. The Mann-Whitney U-test was used to analyze continuous variables. Logistic regression was performed to investigate for associations between the HLA-DRB1*04:05 allele and rs9275534 with susceptibility to AIH. A two-sided P value of less than 0.05 was considered to be statistically significant.
References
Krawitt, E. L. Autoimmune hepatitis. N Engl J Med. 354, 54–66 (2006).
Heneghan, M. A., Yeoman, A. D., Verma, S., Smith, A. D. & Longhi, M. S. Autoimmune hepatitis. Lancet. 382, 1433–1444 (2013).
Vergani, D. et al. Liver autoimmune serology: a consensus statement from the committee for autoimmune serology of the International Autoimmune Hepatitis Group. J Hepatol. 41, 677–683 (2004).
Liberal, R., Mieli-Vergani, G. & Vergani, D. Clinical significance of autoantibodies in autoimmune hepatitis. J Autoimmun. 46, 17–24 (2013).
Vazquez-Garcia, M. N. et al. MHC class II sequences of susceptibility and protection in Mexicans with autoimmune hepatitis. J Hepatol. 28, 985–990 (1998).
Seki, T. et al. HLA class II molecules and autoimmune hepatitis susceptibility in Japanese patients. Gastroenterology. 103, 1041–1047 (1992).
Umemura, T. et al. Human leukocyte antigen class II haplotypes affect clinical characteristics and progression of type 1 autoimmune hepatitis in Japan. PLoS One. 9, e100565 (2014).
Oka, S. et al. HLA-DRB1 and DQB1 alleles in Japanese type 1 autoimmune hepatitis: The predisposing role of the DR4/DR8 heterozygous genotype. PloS one. 12, e0187325 (2017).
Lim, Y. S. et al. Susceptibility to type 1 autoimmune hepatitis is associated with shared amino acid sequences at positions 70–74 of the HLA-DRB1 molecule. J Hepatol. 48, 133–139 (2008).
Doherty, D. G. et al. Allelic sequence variation in the HLA class II genes and proteins in patients with autoimmune hepatitis. Hepatology. 19, 609–615 (1994).
Strettell, M. D. et al. Allelic basis for HLA-encoded susceptibility to type 1 autoimmune hepatitis. Gastroenterology. 112, 2028–2035 (1997).
de Boer, Y. S. et al. Genome-wide association study identifies variants associated with autoimmune hepatitis type 1. Gastroenterology. 147, 443–452 e445 (2014).
Edwards, J. A., Durant, B. M., Jones, D. B., Evans, P. R. & Smith, J. L. Differential expression of HLA class II antigens in fetal human spleen: relationship of HLA-DP, DQ, and DR to immunoglobulin expression. J Immunol. 137, 490–497 (1986).
Guardiola, J. & Maffei, A. Control of MHC class II gene expression in autoimmune, infectious, and neoplastic diseases. Crit Rev Immunol. 13, 247–268 (1993).
Hauber, I. et al. Molecular characterization of major histocompatibility complex class II gene expression and demonstration of antigen-specific T cell response indicate a new phenotype in class II-deficient patients. J Exp Med. 181, 1411–1423 (1995).
Kamatani, Y. et al. A genome-wide association study identifies variants in the HLA-DP locus associated with chronic hepatitis B in Asians. Nat Genet. 41, 591–595 (2009).
Mbarek, H. et al. A genome-wide association study of chronic hepatitis B identified novel risk locus in a Japanese population. Hum Mol Genet. 20, 3884–3892 (2011).
Nishida, N. et al. Genome-wide association study confirming association of HLA-DP with protection against chronic hepatitis B and viral clearance in Japanese and Korean. PloS one. 7, e39175 (2012).
Kim, Y. J. et al. A genome-wide association study identified new variants associated with the risk of chronic hepatitis B. Hum Mol Genet. 22, 4233–4238 (2013).
Hu, Z. et al. New loci associated with chronic hepatitis B virus infection in Han Chinese. Nat Genet. 45, 1499–1503 (2013).
Thomas, R. et al. A novel variant marking HLA-DP expression levels predicts recovery from hepatitis B virus infection. J Virol. 86, 6979–6985 (2012).
Petersdorf, E. W. et al. High HLA-DP Expression and Graft-versus-Host Disease. N Engl J Med. 373, 599–609 (2015).
Saito, S., Ota, S., Yamada, E., Inoko, H. & Ota, M. Allele frequencies and haplotypic associations defined by allelic DNA typing at HLA class I and class II loci in the Japanese population. Tissue Antigens. 56, 522–529 (2000).
Maya, R., Gershwin, M. E. & Shoenfeld, Y. Hepatitis B Virus (HBV) and Autoimmune Disease. Clinical Reviews in Allergy & Immunology. 34, 85–102 (2008).
Liberal, R. et al. Cutting edge issues in autoimmune hepatitis. Journal of Autoimmunity. 75, 6–19 (2016).
Rehermann, B. Pathogenesis of chronic viral hepatitis: differential roles of T cells and NK cells. Nat Med. 19, 859–868 (2013).
Ota, M. et al. A possible association between basic amino acids of position 13 of DRB1 chains and autoimmune hepatitis. Immunogenetics. 36, 49–55 (1992).
Donaldson, P. T. Genetics of liver disease: immunogenetics and disease pathogenesis. Gut. 53, 599–608 (2004).
Umemura, T. et al. Genetic Association of PTPN22 Polymorphisms with Autoimmune Hepatitis and Primary Biliary Cholangitis in Japan. Sci Rep. 6, 29770 (2016).
Alvarez, F. et al. International Autoimmune Hepatitis Group Report: review of criteria for diagnosis of autoimmune hepatitis. J Hepatol. 31, 929–938 (1999).
Umemura, T. et al. Immunoglobin G4-hepatopathy: association of immunoglobin G4-bearing plasma cells in liver with autoimmune pancreatitis. Hepatology. 46, 463–471 (2007).
Umemura, T. et al. Human leukocyte antigen class II molecules confer both susceptibility and progression in Japanese patients with primary biliary cirrhosis. Hepatology. 55, 506–511 (2012).
Excoffier, L. & Lischer, H. E. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour. 10, 564–567 (2010).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc. 3, 1101–1108 (2008).
Sole, X., Guino, E., Valls, J., Iniesta, R. & Moreno, V. SNPStats: a web tool for the analysis of association studies. Bioinformatics. 22, 1928–1929 (2006).
Barrett, J. C., Fry, B., Maller, J. & Daly, M. J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 21, 263–265 (2005).
Acknowledgements
The authors thank Yuki Akahane and Asami Yamazaki for their technical assistance and Trevor Ralph for his editorial assistance. This research was supported in part by a research grant from the Japan Society for the Promotion of Science KAKENHI (16K21069), and (17K09416), and the Promotion Project of Education, Research, and Medical Care from Shinshu University Hospital.
Author information
Authors and Affiliations
Contributions
T.U. and M.O. conceived and designed the study. T.Y. and S.J. performed the experiments. T.Y., T.U., S.J. and K.Y. recruited the patients and collected samples. T.Y., T.U., S.J., E.T. and M.O. analyzed the data and interpreted the experimental results. T.Y., T.U. and M.O. drafted the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yamazaki, T., Umemura, T., Joshita, S. et al. A cis-eQTL of HLA-DPB1 Affects Susceptibility to Type 1 Autoimmune Hepatitis. Sci Rep 8, 11924 (2018). https://doi.org/10.1038/s41598-018-30406-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-30406-9
This article is cited by
-
Genetic risk factors for autoimmune hepatitis: implications for phenotypic heterogeneity and biomarkers for drug response
Human Genomics (2021)
-
The role of natural killer cells in liver inflammation
Seminars in Immunopathology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.