Abstract
Lesion mimic mutants (LMM) usually show spontaneous cell death and enhanced defence responses similar to hypersensitive response (HR) in plants. Many LMM have been reported in rice, wheat, maize, barley, Arabidopsis, etc., but little was reported in xylophyta. BpGH3.5 is an early auxin-response factor which regulates root elongation in birch. Here, we found a T-DNA insertion mutant in a BpGH3.5 transgenic line named lmd showing typical LMM characters and early leaf senescence in Betula platyphylla × B. pendula. lmd showed H2O2 accumulation, increased SA level and enhanced resistance to Alternaria alternate, compared with oe21 (another BpGH3.5 transgenic line) and NT (non-transgenic line). Cellular structure observation showed that programmed cell death occurred in lmd leaves. Stereomicroscope observation and Evans’ blue staining indicated that lmd is a member of initiation class of LMM. Transcriptome analysis indicated that defence response-related pathways were enriched. Southern-blot indicated that there were two insertion sites in lmd genome. Genome re-sequencing and thermal asymmetric interlaced PCR (TAIL-PCR) confirmed the two insertion sites, one of which is a T-DNA insertion in the promoter of BpEIL1 that may account for the lesion mimic phenotype. This study will benefit future research on programmed cell death, HR and disease resistance in woody plants.
Similar content being viewed by others
Introduction
Plants experience spontaneous cell death during development and as a means of resistance to harmful environments, which is called programmed cell death (PCD)1. PCD influences the whole cyclogeny of plant, including germination, vegetative and reproductive growth, senescence, and especially the response to biotic or abiotic stress2. When attacked by imcompatible pathogens, plants can initiate a form of partial PCD at the infected sites to limit the pathogen spread. This is an innate immune response called hypersensitive response (HR). At the same time, certain defence-related genes are activated and systemic acquired resistance (SAR) is formed to protect the plants from attacking3.
Lesion mimic mutants (LMM) are a class of mutants that show spontaneous cell death and defence response without any pathogen attack4. The lesions of LMM resulting from the altered regulation of cell death processes resemble to HR-mediated cell death. LMM have been ideal materials to study cell death and defence pathways in plants. Currently, many LMM have been isolated from Arabidopsis 5, rice6, maize7, barley8, wheat9, potato4, and cotton10, etc. The genes that have been cloned in LMM encode various proteins involved in different pathways including chloroplast activity and light energy, sphingolipids and fatty acids metabolism, signal perception, ion fluxes and reactive oxygen species (ROS) changes11, indicating a complex mechanism of LMM formation. The overexpression of certain genes can also lead to the lesion mimic phenotype12, 13. LMM are usually divided into two groups: the initiation class, which shows location lesions and is discrete in size, and the propagation class, which includes runaway cell death wherein lesions, once formed, can expand continuously to the entire tissue14. Most LMM are accompanied by molecular and cellular changes such as defence-related genes expression, callose deposition, ROS accumulation, increased salicylic acid level and activation of SAR15. Some important phytohormones, including salicylic acid (SA), jasmonic acid (JA) and ethylene, are usually involved in a complex pathogen-plant interaction16,17,18. These phytohormones also play important roles in different LMM.
Reactive oxygen species (ROS), such as superoxide anion radical and hydrogen peroxide (H2O2), are important factors that signal the onset of abiotic and biotic stress. Different damaging and protective signalling pathways are activated by ROS. Many transcription factors (AS1, MYB30, MYC2 and WRKY70), hormone regulators (AXR1, ERA1, SID2, EDS1 and SGT1b) and cell death regulators (RCD1 and DND1) are regulated in H2O2-mediated cell death19.
Plants have evolved sophisticated defence systems to protect themselves from various biotic and abiotic attacks20. Once attacked by a pathogen, the plant immune system is triggered immediately by pathogen-associated molecular patterns (PAMPs) which is called PAMP-triggered immunity (PTI). PTI constitutes a basic defence response21. Effector recognition is mediated by R protein. The R protein can activate effector-triggered immunity (ETI) during the interaction between the pathogen and plants after PTI. Most R proteins contains nucleotide binding site (NBS) and conserved leucine-rich-repeat (LRR) domain22. ETI induces SA accumulation and MAPK activation, which are important for plant disease resistance23. This is followed by activation of the SAR. SA plays a critical role in this process and induces the expression of pathogenesis-related genes (PR1, PR2, PR5)24. Furthermore, many plants including tomato, wheat, and Arabidopsis 25,26,27, etc., show increased SA level after pathogen infection. The most direct evidence of the importance of SA function is provided by transgenic tobacco plants expressing the NahG gene. NahG gene encodes salicylate hydroxylase which can convert SA to catechol. NahG overexpression in tobacco can inhibit SA accumulation and the transgenic line defective in the ability to induce SAR against tobacco mosaic virus28. Besides, the PeaT1-induced SAR pathway is also mediated by salicylic acid and NPR1 gene29.
Ethylene (ET) is an endogenous hormone involved in seed germination, organ abscission, fruit ripening, senescence, disease and stress resistance5. ET and SA usually act synergistically to confer disease resistance in plants. ET treatments on NahG transgenic tobacco can elevate SA level. Ethylene insensitive 3 (EIN3) which belongs to the EIN3 family is a positive regulator of the ET signal transduction30. The degradation of EIN3 protein is a primary means by which the sensitivity of plants to ethylene is regulated31. There are five EIN3-like proteins (EILs), namely, EIL1, EIL2, EIL3, EIL4, EIL5, among which EIL1 is the most similar and functionally redundant to EIN3 in Arabidopsis thaliana. EIN3/EIL1 can activate many genes related to ethylene response32, senescence33, flower development30, SA production34, salt stress35, pathogen invasion36, iron homeostasis37, and Fe metabolism38, indicates that EIN3/EIL1 is a junction of different pathways.
In this study, we described a T-DNA insertion mutant of Betula platyphylla × B. pendula named lmd with a typical LMM phenotype. The lmd mutant exhibited spontaneous cell death and leaf abscission both in vitro and in soil. Southern blotting showed that there were two insertion sites in the lmd genome. Genome re-sequencing and TAIL-PCR results indicated that the insertion of BpEIL1 promoter might cause the formation of the lesion mimic phenotype. The lesions formation and the enhanced defence response are related to SA accumulation. Our study characterized and identified a woody lesion mimic mutant in birch. It suggests that BpEIL1 plays a crosstalk role in the signalling cascade leading to defence response and HR cell death.
Results
Characterization of lmd mutant
The GH3 family is an important class of early auxin-response genes involved in the development of the hypocotyls and roots in Arabidopsis thaliana. BpGH3.5 is a GH3-like gene in birch, which can regulate root elongation as an early auxin-response factor. We obtained a T-DNA insertion mutant named lmd when studying the BpGH3.5 function in Betula platyphylla × B. pendula. The lmd showed necrotic spots on mature leaves with early leaf senescence. That was different from the other 20 BpGH3.5 overexpression lines. The necrotic spots appeared from the edge of the lmd leaves. The number of the lesions increased as leaf aged but the lesion size was limited (Fig. 1G and J). Interestingly, we observed the same phenotype on the seedlings growing in woody plant medium (WPM). This indicated that the necrotic lesions were not caused by microorganism but spontaneous. We judged that the lmd mutant was a member of lesion mimic mutants. Before this, few lesion mimic mutants in xylophyta were reported. Therefore, we performed a series of experiments to confirm this result.
Phenotype of the lmd mutant. (A and B) Show the ground diameter and height, respectively. (C) Net photosynthetic efficient (Pn). (D) Absorbance of NT, oe21 and lmd eluent. The absorbance of lmd is higher than that of NT and oe21which indicated that the cell death is more in lmd than in NT and oe21. (E) Comparison of MDA content in NT, oe21 and lmd. (F) Increment of NT, oe21 and lmd. (G) A branch of lmd. (H) Result of Evans’ blue staining. We can see few lesions on the leaf before staining. After staining, we can see more stained spots on the leaf. Bars = 50μm. (I) Callose deposition. (J) Lesions of the lmd leaves under a stereomicroscope. The number of lesions increased as leaf aged. Bars = 1 mm.
The lmd showed slower growth than its counterparts NT (non-transgenic line) and oe21 (another BpGH3.5 transgenic line) (Fig. 1A,B and F). The leaf net photosynthetic rate was lower in lmd (Fig. 1C), which may account for the low increment of lmd. To examine the cell death of lmd, we performed Evans’ blue staining. Evans’ blue which can enter dead cells is a histochemical indicator of cell death. The staining result showed that lmd exhibited a few deep blue spots all over the leaves, even though we could see few necrosis spots on leaves before staining, indicating that necrotic spots would appear subsequently in these positions (Fig. 1H). However, the NT and oe21 leaves did not exhibit positive Evans’ blue staining (Supplemental Fig. S1). To quantify the dead cells in lmd, Evans’ blue was eluted by SDS-methanol solution, and the absorbance of the eluent was measured in 595 nm. The absorbance of lmd eluent was much higher than that of NT and oe21, which indicated that the dead cells in lmd was much more (Fig. 1D). Malondialdehyde (MDA) content indirectly reflects the degree of cellular damage. The MDA content of lmd was significantly higher than that of NT and oe21 (Fig. 1E), which indicated that cellular damage occured in lmd. The aniline blue staining showed an obviously callose deposition in lmd (Fig. 1I).
Cellular structure observation
Programmed cell death must be accompanied by cellular structure changes. To observe if there were any cellular structure changes in lmd, we performed obversation with both light microscope and electron microscope. Light microscope observation showed that the dead cells clustered in certain areas, but the cells nearby were absolutely normal. It indicated that lmd was an initiation type of LMM (Fig. 2C). We observed cell ultrastructure of lmd leaves using a transmission electron microscope (TEM). More dead cells were observed in lmd compared with oe21 and NT. Some cells generated phagocytosis, and the organelles disappeared, leaving an empty cell wall at last where the lesions were in lmd (Fig. 2B). This result was indicative of typical programmed cell death development in plants. There were many leaf glands on the leaf hypodermis of birch, which would degenerated upon leaf maturity. Owing to lmd showed early leaf senescence, we scanned the leaf surface using a scanning electron microscope (SEM) to see if there were any changes in lmd. On the 3rd leaves, we found fewer leaf glands on lmd leaves than on NT and oe21 leaves (Fig. 2A,D). On the 4th leaves, there was no difference among NT, oe21 and lmd. The number of leaf glands showed no difference between the 3rd and the 4th leaves in lmd. SEM observation showed that the degeneration of leaf glands in lmd occurred earlier than in NT and oe21, which was evident from both the number and morphology of the leaf glands. It indicated an early maturity happened in lmd.
Observation of tissular and cellular structure. (A) SEM observation of lmd, NT and oe21. Bars = 100 μm. (B) TEM observation result showing vacuole (V), nucleus (Nu), chloroplast (Chl), mitochondria (M), cell wall (CW) and autophagosome (the arrows). Bars = 0.5 μm. (C) Paraffin section showing cell death was limited in certain area (the arrow). Bar = 100 μm. (D) Leaf gland numbers of the 3 lines.
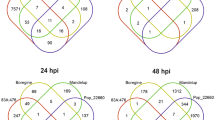
Transcriptome sequencing explore transcriptional changes in lmd
To further learn lmd at the molecular level, we used an Illumina 2500 platform to perform lmd, oe21 and NT transcriptome sequencing. After filtering the low-quality reads, we obtained 3.46 Gb clean reads of each sample on average. Q30 percentages were all above 91.20%. In total, 23,209,269 (80.40%) of the lmd, 27,280,130 (81.95%) of the oe21 and 22,584,972(82.30%) of the NT reads were mapped onto the reference genome. Statistical analysis identified 995 differentially expressed genes (DEGs) among lmd, oe21 and NT (Supplemental Fig. S2), of which 560 genes were up-regulated and 435 genes were down-regulated. Many pathogenesis-related genes, such as WRKY, glutathione-S-transferases(GST), ethylene response factor(ERF) and serine/threonine-protein kinase, which have been previously implicated in defence response39, 40, were among the significantly up-regulated fraction. Gene Ontology (GO) analysis revealed that DEGs involved in catalytic activity, electron carrier activity, antioxidant activity, immune system process and response to stimulus were enriched (Fig. 3). KEGG analysis indicated that DEGs were enriched in plant hormone signal transduction, plant-pathogen interaction and peroxisome pathways.
Enriched GO terms of biological process. Grey bars represent the percentage of genes corresponding to each GO term among genes in the cluster, whereas black bars represent the percentage of genes corresponding to these GO terms in the whole birch genome. All of these terms were significantly enriched (P values are indicated to the right of the graphs).
H2O2 accumulation in lmd
The formation of many LMM necrotic lesions is associated with ROS. To determine whether the programmed cell death of lmd was accompanied by ROS, we performed 3,3C-diaminobenzidine (DAB) and 2,7-dichlorodihydrofluorescein diacetate (DCFH-DA) staining. DAB staining showed that lots of brown spots occured on the lmd leaves (from 1st to 5th leaves, including immature and mature leaves), whereas the leaves of oe21 and NT showed a negative reaction. It indicated H2O2 accumulated in lmd (Fig. 4A). DCFH-DA can move freely through the cytomembrane. It indicates the level of intracellular ROS. DCFH-DA staining showed a higher accumulation of ROS in lmd (Fig. 4A). Peroxidases (PODs) are involved in plant resistance to biotic and abiotic stresses. The POD activity of lmd was higher than that of NT and oe21 (Fig. 4B). Superoxide dismutase (SOD) is a primary biological scavenger of free radicals. It is an important antioxidant enzyme. SOD activity in the lmd mutant was higher than that of NT but was not significantly different from that of oe21 (Fig. 4E). To confirm the results, we performed qRT-PCR to measure the relative expression level of peroxidase genes (peroxidase 15, peroxidase 21) and GLPs (GLP1 and GLP2). Our results indicated higher relative expression level of peroxidase 15, peroxidase 21 and GLPs in lmd than in NT and oe21 (Fig. 4C,D,F and G). All above indicated that H2O2 accumulated in lmd and the activities of enzymes related to ROS were increased.
H2O2 accumulation and activities of POD and SOD in NT, oe21 and lmd. (A) DAB and DCFH-DA staining for H2O2 observation. Bars = 20 μm. (B) Comparison of POD activity in NT, oe21 and lmd. (C and D) Show the relative expression level of peroxidase 15 and peroxidase 21. (E) Comparison of SOD activity in NT, oe21 and lmd. (F and G) Show the relative expression level of GLPs.
SA involvement in enhanced resistance of lmd
Disease resistance is regulated by multiple signal transduction pathways such as SA, JA and ET41. We performed gas chromatography to measure the ethylene release. The ethylene release of lmd, NT and oe21 showed no difference at 1 ppm. HPLC-MS assay for measuring SA and JA content showed that SA accumulated in lmd (Fig. 5A). The JA content in lmd was lower than that of NT but higher than that of oe21 (Fig. 5B). There are some molecular markers for each pathway. SA usually induces the expression of PRs to enhance disease resistance in plants42. PDF1.2 is a marker for the ET/JA pathway. Therefore, we performed qRT-PCR to quantify these genes. lmd showed a high expression level of PRs, including PR1, PR1-like, PR1a and PR5 (Fig. 6D). PDF1.2 expression level showed no difference compared with oe21 (Fig. 5C). All of the above results indicated that SA accumulated and the expression of PRs was induced in lmd.
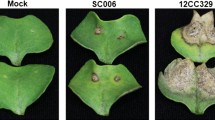
lmd is more resistant to Alternaria alternate. (A) Phenotype of NT, oe21 and lmd 12 days after spraying (DAS) with Alternaria alternate spores. (B) Morbidity of NT, oe21, and lmd observed at 7, 12, 17, 22, and 27 days after spraying. (C) Individual incidence of NT, oe21 and lmd at 7, 12, 17, 22, and 27 days after spraying.
Enhanced resistance to Alternaria alternata
Many LMM exhibite enhanced resistance to pathogens43, 44. To test whether the lmd mutant was resistant to Alternaria alternata, we sprayed fresh spore suspension onto NT, oe21 and lmd seedlings. 7 days later, NT and oe21 were heavily infected with several wilted and flag leaves, whereas few wilted leaves were observed in the lmd mutant, despite the many lesions on its leaves (Fig. 6A). 17 days after spraying (DAS), the morbidity of both NT and oe21 were above 90%, but still no lmd plants were infected. The onset time of lmd was delayed for 15 days at least, which was 22 days after spraying. At that time, the morbidity of lmd was 45.55%.The morbidities of NT and oe21 were both upto 100% (Fig. 6B). To measure the disease severity, we counted the ratio of infected leaves to all leaves on one plant (named individual incidence). We can see the the ratio of infected leaves on each plant in lmd was much lower than that of NT and oe21 (Fig. 6C). It indicated lmd’s resistance to Alternaria alternata was enhanced.
Discussion
We obtained 21 BpGH3.5 overexpression transgenic lines when studying the function of BpGH3.5 in 2012. One of them showed necrotic spots on the mature leaves. The number of the necrotic spots increased as leaf aged. The leaves of this line dropped earlier than the other 20 transgenic lines. Molecular and histochemical assays indicated that this transgenic line was a lesion mimic mutant. We named it lmd (lesion mimic and early deciduous leaf). Lesion mimic mutants have been reported in many species45, 46, but little was reported in xylophyta. The lmd mutant showed H2O2 accumulation from the 2nd leaves to the 5th leaves (the 1st leaves were too small to observe). H2O2 is reported as a cell death inducer. We examined cell death using Evans’ blue staining. Our result indicated that there was little cell death in the 2nd leaves of lmd except for few spots at the edge of the leaves. The cell death increased in the 3rd and 4th leaves in lmd. The H2O2 accumulated in the 2nd leaves, but the cell death was little. It indicated that cell death induced by H2O2 was related to the development of leaves in lmd. Plants can form protective physical barriers, such as callose, to protect themselves from invasion. Callose is a multifaceted defence response that depends on the environmental conditions and the challenging pathogen-associated molecular patterns47. Aniline blue staining showed callose deposition in lmd leaves. qRT-PCR showed that the relative expression level of β-1,3-glucanase (callose degradation)48 gene was lower in lmd, which indicated a transcriptional change of related genes (Supplemental Fig. S3).
LMM can be divided into two types: initiation mutants and propagation mutants49. Microscope observation showed the dead cells were limitied in certain areas and the other areas showed absolutely normal. This result was in line with the Evans’ blue staining result. We can conclude that lmd is an initiation class of LMM. Plant PCD is different from animal PCD. Basing on rupture of the tonoplast followed by rapid clearance of the whole cytoplasm and sometimes most of the cell walls or not, plant PCD can be grouped into two types: autolytic and non-autolytic50. Ultrastructure observation indicated that many cells generated autophagosomes in lmd, but few autophagosomes was found in NT and oe21 cells. Dead cells clustered in lmd, leaving only the cell wall. That’s different from NT and oe21.
Transcriptome analysis explored the gene interactions and the change of transcriptional level in lmd leaves. WRKY superfamily transcription factors (TFs) play an important role in plant growth, development and responses to biotic and abiotic stress39, 51. NPR1 is a key regulator of the SAR pathway downstream of SA. In Arabidopsis, WRKY TFs act upstream or downstream of NPR1 to mediate defence responses to the pathogen52. Many WRKY TFs such as WRKY9, 19, 28, 32, 33, 37, 70, 71, 72, and 75 were all up-regulated which indicated defence responses and the regulation of plant growth changed in lmd. Some other TFs or proteins associated with defence response, senescence, hormone signals, and oxidation-reduction are also up-regulated. This indicated a complex signal network formation in lmd.
Some environmental factors such as light53, 54, temperature, and humidity55, 56 can influence the phenotype of LMM57. Both light and temperature influence the chloroplast number and morphology in the FZL mutant58. Certain mutants show total or partial degradation of chloroplast membranes and irregular degradation of thylakoids. In this study, we cultured the lmd mutant under natural conditions (45°43.2′N, 126°37.4′E). During the whole growing season (from May to October), plants experience different sunshine condition (long-day and short-day) and different temperatures (5 °C to 34 °C). The lmd mutant including one-year-old to five-year-old plants, showed necrotic spots on mature leaves steadily. We also simulated long-day (16 h light) and short-day (8 h light) conditions, and the necrotic spots on lmd leaves showed no difference. Temperature treatments of 30 °C and 16 °C did not result in changes in the formation of lesions. That meant the necrotic spots formation of lmd was independent of light and tempreture.
Many studies showed that chloroplasts play an important role in the formation of lesions7, 58. We tested the photosynthesis and chlorophyll fluorescence of lmd, oe21 and NT. The net photosynthetic rate (Pn) of lmd was lower than that of NT and oe21. The maximum PSII efficiency (Fv/Fm), photochemical efficiency of PSII (ΦPSII), photochemical quenching (qp), non-photochemical quenching (NPQ), stomatal conductance, intercellular CO2 concentration and transpiration rate in lmd showed no differences with oe21 and NT (Supplemental Table S1). The lower Pn may be due to the lesions on leaves. Ultrastructure observation showed that the number and morphology of chloroplasts in lmd was not different from those of oe21 and NT. These results indicated that the formation of lesions was not related to chloroplast in lmd.
Ethylene is considered as an important phytohormone related to plant senescence and fruit ripening59. Some studies also showed ethylene can promote leaf senescence at a defined age, but it is not the most significant factor60. We examined the ethylene release of lmd by gas chromatography. Our results (measuring at 1 ppm) showed that there was no difference among lmd, NT and oe21 on ethylene release which indicated that the early drop of leaf in lmd was independent of ethylene. Ethylene is not the most important factor for leaf dropping in lmd. In addition to ethylene, abscisic acid (ABA) is another important plant hormone for leaf abscission. HPLC-MS showed that ABA of lmd was higher than that of oe21 and NT, which may account for the early drop of leaves in lmd (Supplemental Fig. S4).
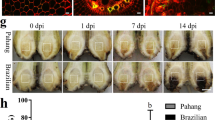
BpGH3.5 is an early auxin-response factor which regulates root elongation in birch61. Since the other 20 BpGH3.5 overexpression lines do not show a phenotype similar to lmd, the phenotype of lmd was not a result of the overexpression of BpGH3.5 but was related to the T-DNA insertion. Through the southern blotting assay, we found two insertion sites in lmd genome (Fig. 7A). Thus, we performed assays to confirm the T-DNA insertion sites. Genome re-sequencing is mainly used to analyse differences between individuals in a species and also a powerful tool to study plant or animal evolution62,63,64. In this study, we used it to confirm the T-DNA insertion sites and achieved a fine result in line with the TAIL-PCR result (Supplemental Fig. S5). Using the genomewide re-sequencing approach, a total of 56,361,128 clean reads were generated, covering 91.97% of the reference Betula genome. We used the L-border and R-border sequences of T-DNA to blast the re-sequencing data and then used the output reads to blast the Betula genome data. According to the blast results (Table 1), we achieved two insertion positions: one was a non-gene-coding region and the other was 598 bp before the first exon of BpEIL1 (Fig. 7B). qRT-PCR showed that the BpEIL1 expression level of lmd was significantly lower than that of NT and oe21 which indicated that the insert inhibited the expression of BpEIL1 (Fig. 7C). EIL1 is a member of the EIN3 family, which is an important transcription factor in ethylene signal transduction and junction of ET and other phytohormones65. In Arabidopsis, EIN3 and EIL1 are functional redundant. There are only two members of EIN3 family in birch which indicated that BpEIN3 and BpEIL1 may have their own special functions respectively. In Arabidopsis, through the negative regulation of SID2, EIN3/EIL1 have an inhibiting effect on SAR34. In rice spotted leaf3 (SPL3) mutant, EIN2 and EIN3 are both down-regulated66. But there are no reports to show that EIN3/EIL1 can directly induce lesion mimic phenotype in other species. However, gene functions may differ in different genetic backgrounds58, 67. In this study, the lower BpEIL1 expression level in lmd may result in a lesion mimic phenotype through H2O2 accumulation and SA increasing. However, how BpEIL1 regulates the lesion mimic phenotype formation still needs to be discussed.
The T-DNA insertion sites analysis. (A) Southern blotting result. The arrows indicate that there are two T-DNA insertion sites in lmd genome. (B) Structure of BpEIL1 gene. Boxes indicate exons, lines indicate introns, the arrow indicates the insertion site of T-DNA which is 598 bp before the first exon of BpEIL1. (C) BpEIL1 relative expression level of NT, oe21 and lmd determined by qRT-PCR. The expression level of BpEIL1 was significantly lower than that of NT and oe21.
Material and Methods
Plant material
The lmd mutant was isolated from the BpGH3.5 transgenic lines. In all experiments, we used both the BpGH3.5 overexpression lines lmd and oe21 and the non-transgenic line NT. All the plants used in this study were 3 years old unless otherwise mentioned. Plants were grown under the natural conditions of Harbin in Heilongjiang province of China. Some plants were grown in the culture room with 16 h/8 h light/dark at a temperature of 26 °C.
Plant height and ground diameter measurement
Plant height was measured using a metre stick. The ground diameter was measured using Vernier callipers. Thirty plants from each line were measured.
Callose deposition
The prepared paraffin sections were dewaxed with 100% xylol twice and then dehydrated in an ethanol series of different concentrations. After being rinsed by distilled water, the samples were dipped into 0.01% aniline blue for 1 h. Afte rinsing with glycerine, the sections were examined under a ZEISS AXIO Imager. A1 fluorescence microscope.
Chlorophyll Fluorescence and Photosynthesis Measurement
The functional leaves from the upper branch of triennial plants were used as materials. Chlorophyll fluorescence was measured using PAM-2500. Photosynthesis was measured using LI-6400XT. Ten plants from each line were measured.
Detection of cell death
The functional leaves were harvested and placed in 0.25% Evans’ blue solution for 30 min. The chlorophyll was removed by soaking in 95% ethanol until the tissues were free of chlorophyll. For measuring the dead cells, we eluted Evans’ blue by boiling 100 mg of stained leaves in 2 mL of SDS-methanol (methanol 50%: SDS 1%) solution at 50 °C for 1 h. After filtered, the eluent was measured using a 722 s spectrophotometer at 595 nm. The assay was performed in triplicate.
Cellular structure observation
Functional leaves from lmd, oe21 and NT were used as experimental materials. For microscopic observation, the leaves were cut into tablets (5 mm × 10 mm) and dipped into FAA for 24 h and then embedded in paraffin. The paraffin sections were stained by safranin-fast green. The leaves were collected and pre-fixed with a mixed solution of 2.5% glutaraldehyde for TEM and SEM observation. For TEM, the samples were post-fixed in 1% osmium tetroxide, dehydrated in an acetone concentration series, infiltrated in epoxy for 3 h and then embedded. The prepared tissue was cut with a diamond knife, stained with uranyl acetate and lead citrate, and then observed under a JEM-100X transmission electron microscope68. For SEM, we took the 1st to the 4th leaves as metarials. The samples were dehydrated in an ethanol concentration series and tertiary butanol and dried in an ES-2030 (HITACHI) freeze-drier for 4 h. The prepared tissue was fixed on a sample platform, coated with 100–150 Å gold, and observed under an S-3400 scanning electron microscope.
DAB staining
For the detection of H2O2 accumulation, fresh leaves from lmd, oe21 and NT were stained with 1 mg/ml (pH 3.8) DAB overnight at room temperature, followed by soaking in 95% ethanol until the chlorophyll was eluted.
DCFH-DA staining
Dimethyl sulfoxide (DMSO) was used to dissolve DCFH-DA to 10 μM. The functional leaves were torn from the hypodermis and immersed into the stomatal opening solution (30 mM KCl, 10 mM MES-KOH, pH 6.15) to induce stomatal opening under light. This was then placed into the DCFH-DA solution and stained for 10–15 min and observed under an LSM 700 laser scanning microscope.
RNA-seq libraries construction, sequencing, functional annotation and GO analysis
RNA-seq libraries were constructed from mRNA using NEB Next UltraTM RNA Library Prep Kit for Illumina (NEB, USA) according to the manufacturer’s instructions and sequenced on an Illumina Hiseq. 2500 platform. Gene function was annotated based on the following databases: Nr (NCBI non-redundant protein sequences), Nt (NCBI non-redundant nucleotide sequences), Pfam (Protein family), KOG/COG (Clusters of Orthologous Groups of proteins), Swiss-Prot (A manually annotated and reviewed protein sequence database), KO (KEGG Ortholog database), GO (Gene Ontology). GO enrichment analysis of the differentially expressed genes (DEGs) was implemented by the GOseq R packages based on Wallenius non-central hyper-geometric distribution69.
RNA extract and quantitative real-time PCR
Total RNA was extracted from the leaves of lmd, oe21 and NT using a Plant RNA Extract kit (Bioteke, Beijing, China). cDNA was synthesized using 1 μg of total RNA by ReverTra Ace qPCR RT Master Mix with gDNA Remover (TOYOBO, OSAKA, Japan) according to the manufacturer’s instructions. Quantitative real-time PCR (qRT–PCR) was performed on an ABI 7500 real-time PCR detection system using SYBR Green Real-time PCR Master Mix -Plus- (TOYOBO, OSAKA, Japan). Bp18S was chosen as a reference gene to normalize the relative expression. Gene-specific primers (Supplemental Table S2) were used to amplify each gene in triplicate.
Phytohormone and enzyme assays
Phytohormones were extracted from plant leaves with isopropanol/hydrochloric acid and dichloromethane and then dried using nitrogen. The residue was dissolved in methanol. After filtered by a 0.22-µm filter membrane, the phytohormones content was measured by HPLC-MS. The ethylene release was measured by gas chromatography as reported70. The total soluble protein content, MDA content, SOD activity, and POD activity of the leaves were determined using a total protein assay kit (A045-3), a thiobarbituric acid (TBA) assay kit (A003-1), an SOD assay kit (A001-1), and a POD assay kit (A084-3), respectively, all of which were acquired from the Nanjing Jiancheng Technology Company (Nanjing, China)68. All of the physiological and biochemical indexes above were measured in triplicate.
Pathogen injection
Alternaria alternata was cultured on potato dextrose agar (PDA) medium at 25 °C for 7 d. The spores were suspended in sterile water to an density of 2 × 105/mL and then sprayed onto 40-cm-high seedlings. The treated seedlings were cultured at 25 °C and 16 h/8 h light/dark with 1000–1500 lx intensity. 7 days later, we started to record the disease occurrence. Mobility was calculated by the rate of infected individuals. The severity of the infected plants was calculated by the proportion of infected leaves relative to all leaves on each plant. The assay was performed for 3 times.
Southern blotting and TAIL-PCR
DNA was extracted from the loose callus of lmd using the cetyl trimethylammonium bromide (CTAB) method71. Southern blotting was performed as reported72. Briefly, total DNA was digested by BamH I and separated by gel electrophoresis. Molecules were transferred from the gel to a nylon membrane by capillary action using absorbent paper. A specific sequence was detected on the membrane by molecular hybridization with DIG-labelled 35 S promotor fragment as probe (Supplemental Fig. S6). CDP-star (Roche, Mannheim, Germany) was used to detect the signal. TAIL-PCR was performed following TaKaRa Genome Walking Kit (TaKaRa, Daliang, China) instructions. Three gene-specific primers (Supplemental Table S2) were used in this assay.
DNA isolation and re-sequencing
Genomic DNA of lmd leaves were isolated using the CTAB method71. Illumina sequencing library was constructed according to manufacturer’s instructions and then sequenced on the Illumina Hiseq. 2500 platform. The Betula genome was download from http://birch.genomics.cn/ as reference genome. We searched for the reads containing L-border or R-border sequence of T-DNA in the clean reads data. The searching results were used to blast the reference genome to look for the insertion sites.
Data analysis
The treatments were analysed for significant differences using one-way ANOVA. Data are presented as the mean ± standard error.
References
Fomicheva, A. S. et al. Programmed cell death in plants. Biochemistry (Moscow). 77, 1452–1464 (2012).
Van Hautegem, T., Waters, A. J., Goodrich, J. & Nowack, M. K. Only in dying, life: programmed cell death during plant development. Trends Plant Sci. 20, 102–113 (2015).
Agarrwal, R., Padmakumari, A. P., Bentur, J. S. & Nair, S. Metabolic and transcriptomic changes induced in host during hypersensitive response mediated resistance in rice against the Asian rice gall midge. Rice. 9, 5, doi:https://doi.org/10.1186/s12284-016-0077-6 (2016).
Abad, M. S. et al. Characterization of Acquired Resistance in Lesion-Mimic Transgenic Potato Expressing Bacterio-Opsin. Mol Plant Microbe Interact. 10, 635–645 (1997).
Bouchez, O., Huard, C., Lorrain, S., Roby, D. & Balague, C. Ethylene is one of the key elements for cell death and defense response control in the Arabidopsis lesion mimic mutant vad1. Plant physiol. 145, 465–477 (2007).
Yin, Z. et al. Characterizing Rice Lesion Mimic Mutants and Identifying a Mutant with Broad-Spectrum Resistance to Rice Blast and Bacterial Blight. Mol Plant Microbe Interact. 13, 869–876 (2000).
Hu, G., Yalpani, N., Briggs, S. P. & Johal, G. S. A porphyrin pathway impairment is responsible for the phenotype of a dominant disease lesion mimic mutant of maize. Plant cell. 10, 1095–1105 (1998).
McGrann, G. R., Steed, A., Burt, C., Nicholson, P. & Brown, J. K. Differential effects of lesion mimic mutants in barley on disease development by facultative pathogens. J Exp Bot. 66, 3417–3428 (2015).
Anand, A., Schmelz, E. A. & Muthukrishnan, S. Development of a lesion-mimic phenotype in a transgenic wheat line overexpressing genes for pathogenesis-related (PR) proteins is dependent on salicylic acid concentration. Mol Plant Microbe Interact. 16, 916–925 (2003).
Kumar, V. et al. A novel, conditional, lesion mimic phenotype in cotton cotyledons due to the expression of an endochitinase gene from Trichoderma virens. Plant science. 183, 86–95 (2012).
Bruggeman, Q., Raynaud, C., Benhamed, M. & Delarue, M. To die or not to die? Lessons from lesion mimic mutants. Front plant sci. 6, 24, doi:https://doi.org/10.3389/fpls.2015.00024 (2015).
Fitzgerald, H. A., Chern, M.-S., Navarre, R. & Ronald, P. C. Overexpression of (At)NPR1 in rice leads to a BTH- and environment-induced lesion-mimic. Mol Plant Microbe Interact. 17, 140–151 (2004).
Jiang, H., Chen, Y., Li, M., Xu, X. & Wu, G. Overexpression of SGR results in oxidative stress and lesion-mimic cell death in rice seedlings. J Integr Plant Biol. 53, 375–387 (2011).
Dangl, J. L., Dietrich, R. A. & Richberg, M. H. Death don’t have no mercy Cell death programs in plant–microbe interactions. Plant cell. 8, 1793–1807 (1996).
Tsunezuka, H., Fujiwara, M., Kawasaki, T. & Shimamoto, K. Proteome analysis of programmed cell death and defense signaling using the rice lesion mimic mutant cdr2. Mol Plant Microbe Interact. 18, 52–59 (2005).
Liu, Z. et al. BIK1 interacts with PEPRs to mediate ethylene-induced immunity. P Natl Acad Sci USA 110, 6205–6210 (2013).
Clarke, J. D., Volko, S. M., Ledford, H., Ausubel, F. M. & Dong, X. Roles of Salicylic Acid, Jasmonic Acid, and Ethylene in cpr-Induced Resistance in Arabidopsis. Plant Cell. 12, 2175–2190 (2000).
Song, S. et al. Interaction between MYC2 and Ethylene Insensitive3 modulates antagonism between jasmonate and ethylene signaling in Arabidopsis. Plant cell. 26, 263–279 (2014).
Kaurilind, E., Xu, E. & Brosche, M. A genetic framework for H2O2 induced cell death in Arabidopsis thaliana. BMC genomics. 16, 837, doi:https://doi.org/10.1186/s12864-015-1964-8 (2015).
Kwon, S. I., Cho, H. J., Kim, S. R. & Park, O. K. The Rab GTPase RabG3b positively regulates autophagy and immunity-associated hypersensitive cell death in Arabidopsis. Plant physiol. 161, 1722–1736 (2013).
Laluk, K. et al. Biochemical and genetic requirements for function of the immune response regulator Botrytis-Induced Kinase1 in plant growth, ethylene signaling, and PAMP-triggered immunity in Arabidopsis. Plant cell. 23, 2831–2849 (2011).
Tran, P. T. et al. A simple method for screening of plant NBS-LRR genes that confer a hypersensitive response to plant viruses and its application for screening candidate pepper genes against Pepper mottle virus. J Virol Methods. 201, 57–64 (2014).
Zhang, L. et al. Cotton GhMKK5 affects disease resistance, induces HR-like cell death, and reduces the tolerance to salt and drought stress in transgenic Nicotiana benthamiana. J Exp Bot. 63, 3935–3952 (2012).
Malamy, J., Hennig, J. & Klessig, D. F. Temperature-Dependent lnduction of Salicylic Acid and Its Conjugates during the Resistance Response to Tobacco Mosaic Virus lnfection. Plant cell. 4, 359–366 (1992).
L. G., I., R. I., K., G. F., B. & A. R, A. The effect of salicylic and jasmonic acids on the activity and range of protective proteins during the infection of wheat by the septoriosis pathogen. Izv Akad Nauk Ser Biol. 1, 34–41 (2015).
Prithiviraj, B., Bais, H. P., Jha, A. K. & Vivanco, J. M. Staphylococcus aureus pathogenicity on Arabidopsis thaliana is mediated either by a direct effect of salicylic acid on the pathogen or by SA-dependent, NPR1-independent host responses. Plant J. 42, 417–432 (2005).
Angulo, C. et al. Role of dioxygenase alpha-DOX2 and SA in basal response and in hexanoic acid-induced resistance of tomato (Solanum lycopersicum) plants against Botrytis cinerea. J Plant Physiol. 175, 163–173 (2015).
Gaffney, T. et al. Requirement of Salicylic Acid for the Induction of Systemic Acquired Resistance. Science. 261, 754–766 (1993).
Zhang, W. et al. PeaT1-induced systemic acquired resistance in tobacco follows salicylic acid-dependent pathway. Mol Biol Rep. 38, 2549–2556 (2011).
Hibi, T. et al. Involvement of EIN3 homologues in basic PR gene expression and flower development in tobacco plants. J Exp Bot. 58, 3671–3678 (2007).
Chen, Y. F., Etheridge, N. & Schaller, G. E. Ethylene signal transduction. Ann Bot. 95, 901–915 (2005).
Solano, R., Stepanova, A., Chao, Q. & Ecker, J. R. Nuclear events in ethylene signaling_ a transcriptional cascade mediated by Ethylene-Insensitive3 and Ethylene-Response-Factor1. Genes Dev. 12, 3703–3714 (1998).
Li, Z., Peng, J., Wen, X. & Guo, H. Ethylene-insensitive3 is a senescence-associated gene that accelerates age-dependent leaf senescence by directly repressing miR164 transcription in Arabidopsis. Plant cell. 25, 3311–3328 (2013).
Chen, H. et al. Ethylene Insensitive3 and Ethylene Insensitive3-Like1 Repress Salicylic Acid Induction Deficient2 expression to negatively regulate plant innate immunity in Arabidopsis. Plant cell. 21, 2527–2540 (2009).
Zhang, L. et al. An AP2 domain-containing gene, ESE1, targeted by the ethylene signaling component EIN3 is important for the salt response in Arabidopsis. Plant Physiol. 157, 854–865 (2011).
Duan, X. et al. TaEIL1, a wheat homologue of AtEIN3, acts as a negative regulator in the wheat-stripe rust fungus interaction. Mol plant pathol. 14, 728–739 (2013).
Yang, Y. et al. The Arabidopsis Mediator subunit MED16 regulates iron homeostasis by associating with EIN3/EIL1 through subunit MED25. Plant J. 77, 838–851 (2014).
Bauer, P. & Blondet, E. Transcriptome analysis of ein3 eil1 mutants in response to iron deficiency. Plant Signal Behav. 6, 1669–1671 (2011).
Shimono, M. et al. Rice WRKY45 Plays a Crucial Role in Benzothiadiazole-Inducible Blast Resistance. Plant Cell. 19, 2064–2076 (2007).
Meng, X. et al. Phosphorylation of an ERF Transcription Factor by Arabidopsis MPK3/MPK6 Regulates Plant Defense Gene Induction and Fungal Resistance. Plant cell. 25, 1126–1142 (2013).
Groen, S. C. et al. Pathogen-triggered ethylene signaling mediates systemic-induced susceptibility to herbivory in Arabidopsis. Plant cell. 25, 4755–4766 (2013).
L. C., vL., M., R. & C. M., P. Significance of inducible defense-related proteins in infected plants. Annu Rev Phytopathol. 44, 135–162 (2006).
Wu, C. et al. Rice lesion mimic mutants with enhanced resistance to diseases. Mol Genet Genomics. 279, 605–619 (2008).
Chen, X. et al. Proteomic analysis of a disease-resistance-enhanced lesion mimic mutant spotted leaf 5 in rice. Rice. 6, doi:https://doi.org/10.1186/1939-8433-6-1 (2013).
Penning, B. W., Johal, G. S. & McMullen, M. D. A major suppressor of cell death,slm1, modifies the expression of the maize (Zea maysL.) lesion mimic mutationles23. Genome. 47, 961–969 (2004).
Zeng, L., Yin, Z., Chen, J., Leung, H. & Wang, G. L. Fine genetic mapping and physical delimitation of the lesion mimic gene Spl11 to a 160-kb DNA segment of the rice genome. Mol Genet Genomics. 268, 253–261 (2002).
Luna, E. et al. Callose deposition: a multifaceted plant defense response. Mol Plant Microbe Interact. 24, 183–193 (2011).
Wan, L. et al. A rice β-1,3-glucanase gene Osg1 is required for callose degradation in pollen development. Planta. 233, 309–323 (2010).
Moeder, W. & Yoshioka, K. Lesion mimic mutants A classical, yet still fundamental approach to study programmed cell death. Plant Signal Behav. 3, 764–767 (2008).
van Doorn, W. G. Classes of programmed cell death in plants, compared to those in animals. J Exp Bot. 62, 4749–4761 (2011).
Chen, C. & Chen, Z. Potentiation of developmentally regulated plant defense response by AtWRKY18, a pathogen- induced Arabidopsis transcription factor. Plant physiol. 129, 706–716 (2002).
Yu, D., Chen, C. & Chen, Z. Evidence for an Important Role of WRKY DNA Binding Proteins in the Regulation of NPR1 Gene Expression. Plant cell. 13, 1527–1539 (2001).
Mason, G. A., Lemus, T. & Queitsch, C. The Mechanistic Underpinnings of anago1-Mediated, Environmentally Dependent, and Stochastic Phenotype. Plant physiol. 170, 2420–2431 (2016).
Wang, J. et al. Characterization and fine mapping of a light-dependent leaf lesion mimic mutant 1 in rice. Plant Physiol Biochem. 97, 44–51 (2015).
Liu, J., Jambunathan, N. & McNellis, T. W. Transgenic expression of the von Willebrand A domain of the BONZAI 1/COPINE 1 protein triggers a lesion-mimic phenotype in Arabidopsis. Planta. 221, 85–94 (2005).
Mosher, S. et al. The lesion-mimic mutant cpr22 shows alterations in abscisic acid signaling and abscisic acid insensitivity in a salicylic acid-dependent manner. Plant physiol. 152, 1901–1913 (2010).
Jung, Y. H. et al. The rice (Oryza sativa) blast lesion mimic mutant, blm, may confer resistance to blast pathogens by triggering multiple defense-associated signaling pathways. Plant Physiol Biochem. 43, 397–406 (2005).
Landoni, M. et al. A mutation in the FZL gene of Arabidopsis causing alteration in chloroplast morphology results in a lesion mimic phenotype. J Exp bot. 64, 4313–4328 (2013).
Chang, K. N. et al. Temporal transcriptional response to ethylene gas drives growth hormone cross-regulation in Arabidopsis. eLIFE 2, doi:https://doi.org/10.7554/eLife.00675.001,10.7554/eLife.00675.002 (2013).
Jing, H. C., Schippers, J. H., Hille, J. & Dijkwel, P. P. Ethylene-induced leaf senescence depends on age-related changes and OLD genes in Arabidopsis. J Exp bot. 56, 2915–2923 (2005).
Yang, G. et al. BpGH3.5, an early auxin-response gene, regulates root elongation in Betula platyphylla × Betula pendula. Plant Cell PCTOC. 120, 239–250 (2014).
Natarajan, S. et al. Whole Genome Re-Sequencing and Characterization of Powdery Mildew Disease-Associated Allelic Variation in Melon. PloS one 11, e0157524, doi:https://doi.org/10.1371/journal.pone.0157524 (2016).
Thudi, M. et al. Whole genome re-sequencing reveals genome-wide variations among parental lines of 16 mapping populations in chickpea (Cicer arietinum L.). BMC plant biol. 16(Suppl 1), 10, doi:https://doi.org/10.1186/s12870-015-0690-3 (2016).
Li, Y.-h et al. Molecular footprints of domestication and improvement in soybean revealed by whole genome re-sequencing. BMC genomics. 14, 579–591 (2013).
Chao, Q., Rothenberg, M., Solano, R., Roman, G. & Terzaghi, W. E. J. R. Activation of the ethylene gas response pathway in Arabidopsis by the nuclear protein Ethylene-Insensitive3 and related proteins. Cell. 89, 1133–1144 (1997).
Wang, S. H. et al. Mutation of Spotted Leaf3 (SPL3) impairs abscisic acid-responsive signalling and delays leaf senescence in rice. J Exp bot. 66, 7045–7059 (2015).
Zhang, Y. F., Hou, M. M. & Tan, B. C. The requirement of WHIRLY1 for embryogenesis is dependent on genetic background in maize. PloS one 8, e67369, doi:https://doi.org/10.1371/journal.pone.0067369 (2013).
Li, Z. et al. Fine mapping of the lesion mimic and early senescence 1 (lmes1) in rice (Oryza sativa). Plant Physiol Biochem. 80, 300–307 (2014).
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq_ accounting for selection bias. Genome Biol. 11, R14 (2010).
Li, C.-z & Wang, G.-x Interactions between reactive oxygen species, ethylene and polyamines in leaves of Glycyrrhiza inflata seedlings under root osmotic stress. Plant Growth Regul. 42, 55–60 (2004).
Cheng, Y.-J., Guo, W.-W., Yi, H.-L., Pang, X.-M. & Deng, X. An efficient protocol for genomic DNA extraction from Citrus species. Plant Mol Biol Rep. 21, 177–178 (2003).
Southern, E. Southern blotting. Nat Protoc. 1, 518–525 (2006).
Acknowledgements
This study was supported by the following foundations: 1. National Natural Science Foundation of China (NSFC) (grant no. 31670673); 2.The 111 Project (B16010).
Author information
Authors and Affiliations
Contributions
Jing Jiang and Guifeng Liu provided experimental design and plant material. Ranhong Li performed experiments and statistical analyses. Su Chen analyzed genomic re-sequencing data. Rui Han performed qRT-PCR assay. Ranhong Li wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, R., Chen, S., Liu, G. et al. Characterization and Identification of a woody lesion mimic mutant lmd, showing defence response and resistance to Alternaria alternate in birch. Sci Rep 7, 11308 (2017). https://doi.org/10.1038/s41598-017-11748-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-11748-2
This article is cited by
-
Selection of elite lines of BpGH3.5-transgenic Betula platyphylla using growth adaptability analysis
Journal of Forestry Research (2022)
-
Growth adaptability and foreign gene stability of TaLEA transgenic Populus simonii × nigra
Annals of Forest Science (2021)
-
Inhibition of BpEIN3 causes plaques in leaves of Betula platyphylla × B. pendula
Trees (2020)
-
Transcriptome analysis provides new insights into leaf shape variation in birch
Trees (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.