Abstract
Artemisinins are the current mainstay of malaria chemotherapy. Their exact mode of action is an ongoing matter of debate, and several factors have recently been reported to affect an early stage of artemisinin resistance of the most important human malaria parasite Plasmodium falciparum. Here, we identified a locus on chromosome 7 that affects the artemisinin susceptibility of P. falciparum in a quantitative trait locus analysis of a genetic cross between strains 7G8 and GB4. This locus includes the peroxiredoxin gene PFAOP. However, steady-state kinetic data with recombinant PfAOP do not support a direct interaction between this peroxidase and the endoperoxide artemisinin. Furthermore, neither the overexpression nor the deletion of the encoding gene affected the IC50 values for artemisinin or the oxidants diamide and tert-butyl hydroperoxide. Thus, PfAOP is dispensable for blood stage parasite survival, and the correlation between the artemisinin susceptibility and chromosome 7 is probably based on another gene within the identified locus.
Similar content being viewed by others
Introduction
Artemisinin from the herb Artemisia annua is a traditional, highly efficient antimalarial drug1, 2. Artemisinin-based combination therapies have been recommended by the World Health Organization for the treatment of Plasmodium falciparum malaria since 2001 and have become a key factor of global malaria control programs. For example, artemisinin and its derivatives (Fig. 1a) are estimated to reduce mortality in young children with uncomplicated malaria by more than 95%3, 4. However, early stage artemisinin resistance has emerged in form of a delayed parasite clearance phenotype5. The altered artemisinin susceptibility has been associated with discrete mutational changes within a gene that encodes the P. falciparum kelch protein K136,7,8. Mutations in the K13 gene alone cannot account for all resistance cases, suggesting that delayed parasite clearance is the result of convergent evolutionary events8, 9. Additional factors that might affect the artemisinin susceptibility of P. falciparum or facilitate K13 mutations were identified by whole genome sequencing of blood samples as well as reverse genetics and chemogenomic profiling in vitro 8, 10, 11. A quantitative trait locus (QTL) analysis also revealed an effect of the parasite’s multidrug resistance transporter PfMDR1 and of two additional loci on chromosomes 12 and 13 on artemisinin susceptibility12.
Chemical structures of artemisinin derivatives and QTL analysis. (a) Chemical structures of compounds indicated. (b) In vitro ART and ATM IC50 values of 32 F1 progeny from the 7G8 × GB4 cross and the two parental clones (indicated by arrows). Means ± SEM of at least five independent determinations. (c) QTL analysis. The logarithm of odds (LOD) scores are shown for ART (black lines) and ATM (grey lines) responses as a function of genomic location. Loci associated with decreased ART and ATM susceptibility are indicated. The dotted line represents the confidence line with p < 0.01.
The exact mode(s) of action of artemisinin as well as the mechanisms that contribute to artemisinin resistance are still a matter of debate and might be even interconnected. Numerous potential targets – including lipids and proteins such as the phosphatidylinositol-3-kinase PfPI3K – have been recently identified and localise to a variety of subcellular compartments13,14,15,16,17,18. Transcriptome analyses suggest that artemisinin resistance is linked to the lipid and protein metabolism of P. falciparum resulting in an altered cell cycle progression in accordance with experiments in cell culture11, 19,20,21,22. The activity of artemisinin depends on its endoperoxide moiety (Fig. 1a), which was suggested to be reduced and activated by heme-bound Fe2+ within the digestive vacuole, where the proteolytic degradation of hemoglobin takes place16, 18, 23, 24. Furthermore, artemisinin was hypothesized to promote the formation of reactive oxygen species resulting in so-called oxidative stress18, 24,25,26,27, although such an unspecific mode of action has also been questioned13, 23. In this regard it is interesting to note that the toggling of redox switches in molecular sensors and transducers is a specific process28, and that K13 shares a high similarity with Keap1, which is the primary redox sensor for Nrf2-dependent redox signalling in mammals7, 29. Whether K13 is involved in similar redox signaling cascades in P. falciparum is unknown.
Other crucial sensors in redox signalling belong to the ubiquitous protein family of peroxiredoxins (Prx). These hydroperoxidases catalyse the reduction of H2O2 and other hydroperoxides yielding water and the corresponding alcohol as products. Some Prx also exert functions as chaperones30,31,32. The P. falciparum genome encodes five different Prx-isoforms33,34,35,36. Furthermore, the parasite was shown to import functional human Prx-2 from erythrocytes37. According to current theories, Plasmodium Prx detoxify hydroperoxides in different subcellular compartments34, 35, 38,39,40,41. However, experimental data supporting the relevance of the Plasmodium Prx-isoforms for parasite survival is rather limited. While the cytosolic isoforms TPx-1 and 1-Cys-Prx were shown to be dispensable for the development of P. falciparum and P. berghei blood stage parasites42,43,44, attempts to delete the gene that encodes the nuclear isoform nPrx were unsuccessful and point towards an essential function35. The relevance of the other two Plasmodium Prx-isoforms for parasite survival remains unclear. One of these isoforms is the so-called antioxidant protein PfAOP which belongs to the Prx5 subfamily and is found in the plastid (apicoplast) and cytosol of P. falciparum 41. The dual localization of PfAOP presumably arose from a gene fusion event following a horizontal prokaryote-to-eukaryote gene transfer41. Although studies on recombinant PfAOP confirmed its peroxidase activity in vitro 45,46,47, its physiological relevance for hydroperoxide removal or redox homeostasis has not been addressed before.
Here we identified a locus on chromosome 7 that affects the artemisinin susceptibility in the genetic cross between the P. falciparum clones 7G8 and GB4. One of the 49 genes within the locus encodes PfAOP. Taking into account the established anti-oxidative activity of PfAOP and the hypothesized pro-oxidative mode of action of artemisinins, we investigated a potential contribution of PfAOP to artemisinin susceptibility and parasite survival.
Results
QTL mapping reveals PFAOP as a potential artemisinin susceptibility factor
In a previous study we have interrogated the contribution of the genetic background on artemisinin susceptibility by investigating artemisinin responses in the progeny of a genetic cross between the P. falciparum clones HB3 (Latin America) and Dd2 (Southeast Asia). We found that loci on chromosomes 5 (including Pfmdr1), 12 and 13 contribute to altered responsiveness12. Here we repeated the study, but this time using the genetic cross between strains 7G8 (Latin America) and GB4 (Africa)48. Responses to artemisinin and artemether were determined, using a standard cell proliferation assay where parasites were exposed to the drugs for 72 h49. The IC50 values of strains 7G8 and GB4 were for artemisinin 5.0 ± 0.6 nM and 5.7 ± 1.0 nM and for artemether 2.7 ± 0.4 nM and 3.6 ± 0.8 nM, respectively. Although the parental clones had comparable IC50 values for artemisinin and artemether, the responsiveness of the 32 progeny investigated covered a range from 2.1 ± 0.3 nM to 7.9 ± 0.6 nM in the case of artemisinin and from 0.3 ± 0.1 nM to 3.6 ± 0.8 nM in the case of artemether (Fig. 1b).
To identify determinants of altered artemisinin and artemether responsiveness, we conducted a QTL analysis by correlating the IC50 values with the genetic maps of the progeny and the parental clones48. One major QTL with a LOD score of 3.1 (p < 0.001) was identified for both drugs, mapping to the genetic marker C7MK11 on chromosome 7. According to the genome of the reference strain 3D7 this locus encodes 49 genes including PFAOP (PF3D7_0729200) (Table S1). The locus is distinct from the artemisinin susceptibility loci identified in the HB3 × Dd2 cross and it is also distinct from the pfcrt locus on chromosome 7 that is associated with chloroquine resistance and altered susceptibility to quinine and quinine-like antimalarial drugs50,51,52. We found no link with the K13 gene on chromosome 137 or the PfPI3K gene on chromosome 515. This finding was anticipated since the two parental clones carry identical wild type loci of these two genes.
Artemisinins are neither substrates nor inhibitors of PfAOP
In order to test a potential direct interaction between artemisinins and PfAOP, we first analysed whether recombinant PfAOP converts endoperoxides in an established coupled enzymatic assay (Fig. 2a). The detected consumption of NADPH in the presence of artemisinins or di-tert-butyl peroxide (DTBP) was comparable to the background activity of negative controls without peroxide substrate, whereas the conversion of tert-butyl hydroperoxide (tBOOH) served as a positive control as described previously45, 47. Thus, PfAOP does not convert endoperoxides and its enzymatic activity is restricted to hydroperoxide substrates.
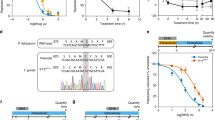
Peroxidase activity of recombinant PfAOP in the presence of artemisinins. (a) Coupled enzymatic assays with 2.5 µM PfAOP and artemisinins as potential substrates. Activities with 100 µM tBOOH or DTBP served as controls. (b) Reversible inhibition assays in the presence of variable concentrations of artemisinins or DTBP. Activities were determined with 2.5 µM PfAOP and 50 µM tBOOH and normalized to the activities of controls without endoperoxide. (c) Time-dependent irreversible inhibition assays. PfAOP (50 µM) was preincubated with 100 µM artemisinins for up to 10 min on ice before the enzyme was diluted 1:20 in a standard coupled enzymatic assay containing 100 µM tBOOH. Activities were normalized to controls that were preincubated without peroxide. A time-dependent inactivation of PfAOP by tBOOH served as a positive control (open circles) and was compared to a potential inactivation by artemisinins (closed circles). Assays containing DMSO are labelled with an asterisk. All data are the mean ± standard deviation of at least two independent duplicate assays.
Next, we analysed whether PfAOP is inhibited by artemisinins. A reversible competitive inhibition was tested for the conversion of tBOOH in the presence of variable concentrations of artemisinins (Fig. 2b). Furthermore, a time-dependent irreversible inhibition was addressed by preincubating recombinant PfAOP with artemisinins, DTBP or tBOOH before transferring the enzyme to standard assays with tBOOH (Fig. 2c). No inhibitory effects were detected except for the positive control with tBOOH, which is not only a substrate but also an irreversible inhibitor that ‘overoxidizes’ PfAOP45, 47. In summary, artemisinins are neither reversible nor irreversible inhibitors of recombinant PfAOP and enzymatic assays do not support a direct effect of PfAOP on artemisinins and vice versa. However, our enzymatic assays neither exclude a potential interaction between PfAOP and Fe2+-activated artemisinin nor indirect effects of PfAOP on the artemisinin susceptibility of P. falciparum. In order to address these aspects, we genetically manipulated the reference strain 3D7 and determined the artemisinin susceptibility of overexpressing and knockout strains.
Overexpression of PfAOP does not alter the artemisinin susceptibility
Artemisinins have been suggested to promote the formation of reactive oxygen species including H2O2 18, 24,25,26,27. The removal of H2O2 or other hydroperoxides by PfAOP might, therefore, affect the artemisinin susceptibility of P. falciparum. PfAOP has a bipartite topogenic signal (BTS), which is required for correct apicoplast targeting of the dual localized protein41. In order to test a potential role of the cytosolic and/or apicoplast form of PfAOP, we sequenced exon 1 (which encodes the BTS) of the parental strains and selected progenies. No single nucleotide polymorphisms or other mutations were found. Next, we overexpressed cytosolic as well as apicoplast-targeted truncated or full length GFP-fusion constructs of PfAOP in 3D7 wild type cells as described previously41. In addition, we generated and analysed enzymatically inactive serine mutants45 of these constructs as controls (Fig. 3a). All GFP-fusion constructs were successfully expressed in P. falciparum resulting, on average, in an approximately 2- to 6-fold increase of the total cellular PfAOP content (Fig. 3b). None of the overexpressed constructs led to an increase of the IC50 value for artemisinin or had a dominant-negative effect (Fig. 3c–f). In summary, irrespective of the protein localization or the presence or absence of the catalytic cysteine residue, overexpression of PFAOP does not alter the artemisinin susceptibility of P. falciparum.
IC50 values for artemisinin of parasites that overexpress PfAOP. (a) The modular gene architecture of PfAOP is shown on top. Exon 1 encodes the bipartite topogenic signal (BTS) and exon 2 the Prx5 domain. Constructs encoding cytosolic truncated as well as apicoplast-targeted full length GFP-fusion proteins of PfAOP are depicted below and were overexpressed in strain 3D7 as described previously41. Residue Cys117 at the active site of PfAOP was shown to be essential for the hydroperoxidase activity45 and is labelled with an asterisk. The residue was mutated to serine in controls. (b) Western blot analyses confirming the overexpression of the GFP-fusion constructs listed in panel (a). Detection of GFP in the upper blot revealed no significant degradation of the fusion proteins. Hence, detection of PfAOP at approximately 22 kDa reflects only endogenous PfAOP (endo) whereas the upper bands in the same blot reflect the GFP-fusion constructs (fusion). Representative images were obtained by standard western blotting. The estimated ratio between both protein species is summarized on top and is based on semi-quantitative western blotting using a C-DiGit Blot Scanner in parallel experiments. (c–e) Artemisinin dose-response curves of strain 3D7 blood stage cultures that express the indicated GFP-fusion constructs from panel a. All data are the mean ± standard deviation of at least three independent triplicate measurements. (f) IC50 values obtained from panels c-e. None of the differences between the IC50 values was found to be significant (p > 0.05) based on statistical analyses in SigmaPlot 12.5 using the one way ANOVA method.
PfAOP is not essential and does not protect against artemisinin or external oxidants
To test whether the complete loss of PfAOP affects the artemisinin susceptibility of P. falciparum, we cloned the plasmid pL7-PFAOP and deleted the encoding gene in strain 3D7 by double crossover using the CRISPR/Cas9 system53 (Fig. 4a). The genetic manipulation was monitored for three different clones following positive selection and limited dilution. All three clones had lost the endogenous gene and had integrated the selection marker after successful 5′- and 3′-crossovers as revealed by analytical PCR (Fig. 4b). Furthermore, the loss of PfAOP was confirmed by western blotting using a specific antibody41 (Fig. 4c). Giemsa-stained blood smears of the three different knockout strains were subsequently analysed by light microscopy but revealed no suspicious morphologies. The growth rates were also identical to the wild type strain (Fig. 5). Next, we determined the IC50 values of the three knockout strains for artemisinin as well as the oxidants tBOOH and diamide as described previously12, 54. Loss of PfAOP affected none of the IC50 values as compared to the wild type strain (Fig. 6). Please note that two different batches of artemisinin were used for the two independent sets of experiments in Figs 3 and 6, which presumably explains the systematic deviation of the IC50 value for artemisinin. The IC50 values for tBOOH in Fig. 6b (between 90 ± 8 µM and 99 ± 7 µM) and diamide in Fig. 6c (between 75 ± 4 µM and 84 ± 11 µM) were in accordance with previous measurements54.
Generation and validation of Δpfaop knockout parasites. (a) Schematic summary of the knockout strategy by double crossover using the plasmid pL7-PFAOP. Expected product sizes from analytical PCR reactions 1–3 are highlighted. The two external primers should anneal to chromosome 7, whereas the two internal primers should anneal to the gene that encodes the selection marker human dihydrofolate reductase (hDHFR). (b) Following the genetic manipulation of the wild type strain 3D7 (3D7 WT) and isolation of three clonal cell lines (Δaop#1-3), products from PCR reactions 1-3 were analysed by agarose gel electrophoresis. Marker (M) and expected product sizes are indicated on the left and right side, respectively. (c) Western blot analysis of parasite cell lines Δaop#1-3. Protein extracts from 107 parasites were loaded per lane. Hsp70 was decorated as a control.
Growth curve analysis. The parasitemia in standard blood stage cultures of strains Δaop#1-3 from Fig. 4 was determined by counting parasites from Giesma-stained blood smears. Wild type strain 3D7 served as a control. All data are the mean ± standard deviation of triplicate measurements for each strain. None of the differences between the parasitemias was found to be significant (p > 0.05) based on statistical analyses in SigmaPlot 12.5 using the one way ANOVA method.
IC50 values for artemisinin and oxidants of Δpfaop knockout parasites. (a) Artemisinin dose-response curves (upper panel) and IC50 values (lower panel) of blood stage cultures of strains Δaop#1-3 from Fig. 4. (b,c) tBOOH and diamide (DIA) dose-response curves (upper panel) and IC50 values (lower panel). All data are the mean ± standard deviation of at least three independent triplicate measurements. Wild type strain 3D7 served as a control. None of the differences between the IC50 values was found to be significant (p > 0.05) based on statistical analyses in SigmaPlot 12.5 using the one way ANOVA method.
Recent studies revealed that ring-stage survival assays can be advantageous to address a potential artemisinin resistance, for example, because of short artemisinin half-lives and drug-induced parasite dormancy55,56,57,58. We therefore analyzed the ring-stage survival percentage of our knockout and overexpressing strains after treatment with 700 nM artemisinin. Wild type strain 3D7 and a mutant NF54 strain that encodes the resistance factor K13C580Y were analyzed in parallel as negative and positive controls53, respectively (Fig. 7). Neither the deletion nor the overexpression of PFAOP had an effect on the ring-stage survival percentage, whereas the survival percentage of the positive control around 12% was twofold increased as compared to strain 3D7. (The difference between the controls was, in our hands, less pronounced as previously reported53, 56,57,58, most likely because we used 700 nM artemisinin instead of dihydroartemisininin in accordance with our QTL analysis, which was also performed with artemisinin). Taken together, the assays in Fig. 7 indicate that PfAOP has no effect on the survival percentage of P. falciparum ring stage parasites under artemisinin pressure.
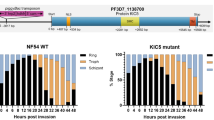
Ring-stage survival assays. Highly synchronous ring stage parasites were treated with or without 700 nM artemisinin for 6 h, washed and further incubated for 66 h. The parasitemia was subsequently determined from Giesma-stained blood smears. The parasite survival percentage was calculated for wild type strain 3D7 (3D7 WT), a strain overexpressing GFP-tagged full length PfAOP from Fig. 3 (PfAOP), a knockout strain from Fig. 4 (Δaop#1) and a positive control that carries the mutation for K13C580Y (NF54K13C580Y). All data are the mean ± standard deviation of four independent experiments. Statistical analyses were performed in SigmaPlot 12.5 using the one way ANOVA method. Results are summarized on top of the bar chart (n.s., not significant; **p < 0.01; ***p < 0.001).
In summary, PfAOP is not essential in asexual blood stage parasites and is dispensable for the removal of external oxidants such as tBOOH and diamide. Furthermore, the loss of PfAOP taken alone does not affect the artemisinin susceptibility of P. falciparum. Thus, the correlation between the artemisinin susceptibility and chromosome 7 is either based on another gene within the identified locus or might require the interplay of several genetic factors.
Discussion
Immunofluorescence microscopy and transcriptome analyses revealed that dual localized PfAOP is constitutively expressed in blood stage parasites33, 41. What could be the function of PfAOP in the apicoplast? Specific iron-sulfur clusters are, at least in prokaryotes, the major targets of oxidative challenges by superoxide anions and H2O2 59. The apicoplast is essential for parasite survival because of the biosynthesis of the isoprenoid precursor isopentenyl pyrophosphate60, 61. Two enzymes of this biosynthetic pathway – the (E)-4-hydroxy-3-methylbut-2-enyl diphosphate synthase and reductase – bind iron-sulfur clusters62, 63. Interestingly, two of the five artemisinin susceptibility factors that were identified from Southeast Asian blood samples in the study by Miotto et al. localise to the apicoplast10. One of the apicoplast artemisinin susceptibility factors is ferredoxin, which also carries an iron-sulfur cluster64, 65. The iron-sulfur clusters of ferredoxin and the (E)-4-hydroxy-3-methylbut-2-enyl diphosphate synthase and reductase (as well as other apicoplast proteins) are probably synthesized by an essential machinery within the stroma of the organelle50. Considering the presence and relevance of redox-sensitive iron-sulfur cluster-containing apicoplast enzymes, it is quite surprising that the loss of the hydroperoxidase PfAOP has no effect on parasite survival. One possible explanation could be the potential absence of major sources of superoxide anions or hydroperoxides in the apicoplast. However, the presence of an apicoplast superoxide dismutase66 seems to contradict this null hypothesis. Another explanation for the absent phenotype of our Δpfaop knockout strains is that the cytosol and apicoplast harbor additional hydroperoxidases with redundant functions. Likely candidates in the cytosol are the dithiol Prx-isoform TPx-1, the Prx6-isoform 1-Cys-Prx and imported human Prx-237, 39, 40, 67. The only alternative hydroperoxidase in the apicoplast appears to be the glutathione peroxidase-like thioredoxin peroxidase TPxGl 68, 69. At this point it is interesting to note that the expression of Plasmodium peroxidases might be interdependent and differ significantly depending on the growth conditions as reported for P. berghei ookinetes70. A third explanation for the absent knockout phenotype is that PfAOP exerts another non-essential function, e.g., as a redox sensor that interacts with transducer molecules29. However, attempts to trap mixed disulfide bonds between PfAOP and potential interaction partners so far only revealed false positive candidates (data not shown). In summary, the physiological function of dual localized PfAOP remains unknown and the encoding gene is fully dispensable for the growth of asexual blood stage parasites in cell culture.
Neither traditional growth inhibition assays nor ring-stage survival assays revealed an altered artemisinin susceptibilty of PFAOP overexpressing and knockout strains. Since non-essential PFAOP on its own does not affect the artemisinin susceptibility, PFAOP might either contribute to a polygenic trait, or another gene within the identified locus on chromosome 7 might be the cause for the detected effects in Fig. 1. Which of the other genes from Table S1 might be relevant within the identified locus? Among the seven genes that were recently discovered as (potential) artemisinin susceptibility factors by chemogenomic profiling11, only one gene (PF3D7_0727100) is located on chromosome 7. PF3D7_0727100 encodes a conserved Plasmodium protein of unknown function, which was also identified in our QTL analysis (Table S1). It is therefore possible that PF3D7_0727100 is the crucial factor within the artemisinin susceptibility locus. Two other candidates from Table S1 – namely PF3D7_0727400, which encodes a putative proteasome subunit alpha type-5, and PF3D7_0729900, which encodes a putative dynein heavy chain – were also identified by mass spectrometry screens using modified artemisinin probes and click chemistry16, 17. PF3D7_0727400 might be of particular interest because of recent models that link artemisinin susceptibility to proteasome-dependent protein turnover15, 18, 20, 71. Future analyses are necessary to address the relevance of the candidates PF3D7_0727100, PF3D7_0727400 and PF3D7_0729900 for artemisinin susceptibility.
Methods
Materials
ART, DHA, ATS and DTBP were purchased from Sigma. ATM was from Sciphar. Recombinant PfAOP with an N-terminal MRGSH6GS-tag replacing the first 59 amino acid residues of the full length protein as well as P. falciparum glutathione reductase (PfGR) and P. falciparum glutaredoxin (PfGrx) were produced in Escherichia coli and purified by affinity chromatography as described previously45, 72. Plasmid pUF1-Cas9 for the expression of Cas9 and plasmid pL6 for guide RNA expression and homologous recombination in P. falciparum 53 were kind gifts from J.J. Lopez-Rubio. Primers were from Metabion. WR99210 was a gift from Jacobus Pharmaceutical Company.
Steady-state kinetics
Coupled enzymatic assays containing recombinant PfAOP, PfGR and PfGrx were carried out at 25 °C using a thermostatted Jasco V-650 UV-visual spectrophotometer as described previously45, 47. All assays were performed in assay buffer containing 0.1 M Tris-HCl, 1 mM EDTA, pH 8.0. Stock solutions of 4 or 6 mM NADPH, 2 or 4 mM tBOOH, 40 mM GSH and 10 mM ATS were freshly prepared in assay buffer before each experiment. Stock solutions of 10 mM ART, ATM, DHA and DTBP were dissolved in DMSO. Whether PfAOP converts endoperoxide substrates was tested in standard assays containing 150 µM NADPH, 1 U/mL PfGR, 1 mM GSH and 5 µM PfGrx. The consumption of NADPH was monitored at 340 nm (ε = 6.22 mM−1cm−1). A baseline was recorded for 60 sec before the addition of 100 µM ART, ATM, DHA, ATS or DTBP. All assays were subsequently started by adding 2.5 µM recombinant PfAOP. Assays containing 100 µM tBOOH with or without the same volume of DMSO served as positive controls. Initial activities were corrected by subtracting the final slope (ΔAbs/min) of the baseline using the Spectra Analysis program (Spectra Manager version 2, Jasco).
To test whether endoperoxides are reversible inhibitors of PfAOP, standard assays containing 50 µM tBOOH were supplemented with 0–100 µM ART, ATM, DHA, ATS or DTBP before starting the assay with 2.5 µM recombinant PfAOP. The activity was determined and normalized to the negative controls without endoperoxide. A potential irreversible inhibition of PfAOP was analysed in a time-course experiment. Samples containing 50 µM recombinant PfAOP with or without 100 µM ART, ATM, DHA, ATS, DTBP or tBOOH were preincubated on ice for up to 10 min before transferring 50 µL of the mixture to a standard 1 mL assay with 100 µM tBOOH as a substrate. The residual activity was determined in the standard assay and normalized to the activity of negative controls that were preincubated in parallel without peroxide.
Cloning of expression- and knockout constructs
The generation of pARL plasmids for the expression of full length, truncated and mutated fusion constructs between PfAOP and GFP has been described previously41. Plasmid pL7-PFAOP was generated as follows: 1) A suitable guide sequence was identified in PfAOP with a confidence value of 0.010 using the protospacer software (http://www.protospacer.com). 2) The 3′-homology region of PfAOP was PCR-amplified using primers PfAOP/3′MCS/pL6/KO/s (5′-GATCGAATTCCAAACGATTTTACTTCAATAGATAC-3′) and PfAOP/3′MCS/pL6/KO/as (5′-GATCCCATGGCTGATTATTTTTTAAAAACTCTTTTAC-3′) and plasmid pQE30/PFAOP C117S as a template45. The PCR product was first subcloned according to the manufacturer’s protocol using the TopoTA cloning kit (Invitrogen) and subsequently cloned into pL6 using the restriction sites EcoRI and NcoI. 3) The 5′-homology region of PfAOP was PCR-amplified from genomic DNA of strain 3D7 using primers PfAOP/5′MCS/KO/s (5′-GATCCCGCGGCTGTTCTTTTTATTATATGAATGAAGAG-3′) and PfAOP/5′MCS/KO/as (5′-GATCTCTAGAGGATTCATAAACTTTTTGGGAAAACC-3′). The PCR product was first subcloned into the Topo cloning vector and subsequently cloned into pL6 using SacII and the compatible restriction sites generated by XbaI (insert) and SpeI (pL6). 4) The guide sequence was cloned into pL6 yielding pL7-PFAOP as described previously53. Plasmid pL6 containing the 5′- and 3′-homology regions was digested with AvrII (2 h, 37 °C) and BtgZI (2 h, 60 °C). The insert was generated by annealing oligonucleotide pL6/gRNA/PfAOP/s (5′-TAAGTATATAATATTACAACATATCTGATACCGATGTTTTAGAGCTAGAA-3′) and the complementary reverse oligonucleotide pL6/gRNA/PfAOP/as. (The 15 bp stretches that are homologous to pL6 and that flank the 20 bp guide sequence are highlighted in the sequence). The insert was mixed with digested pL6 and fused using the In-Fusion HD Cloning Kit (Clontech) according to the manufacturer’s protocol. Correct sequences of the intermediate and final constructs were confirmed by analytic restriction digests and commercial sequencing (GATC Biotech).
Parasite culture, QTL analysis and genetic manipulation
P. falciparum strain 3D7 was cultured at 37 °C according to standard protocols73 using fresh human A+ erythrocytes at a hematocrit of 3% in RPMI 1640 medium that was supplemented with 0.45% (w/v) albumax II, 0.2 mM hypoxanthine and 2.7 μg/mL gentamicin. Progeny of the 7G8 × GB4 cross and the two parental clones were maintained at a hematocrit of 5% in RPMI 1640 medium supplemented with 10% inactivated human A+ serum instead of albumax II. Parasites were cultured at 80% humidity, 3 or 5% CO2, 5% O2 and 90% or 92% N2. Synchronization of ring stage parasites was carried out with 5% (w/v) sorbitol74. The F1 progeny from the GB4 × 7G8 cross were obtained from MR448. The identity of all progeny was verified using eight polymorphic microsatellite markers51. QTL analysis was performed according to the method by Haley and Knott75 in line with previous independent studies12, 51. The genetic maps have been published48.
Transfections were performed as described76 using 100 μg of plasmid DNA per construct. Transfectants were usually detected in Giemsa-stained blood smears between two and four weeks post transfection. Parasites that were transfected with pARL plasmids were selected with 5 nM WR99210 and the expression of GFP-fusion constructs was confirmed by live cell imaging as described previously41. Parasites that were transfected with plasmid pUF1-Cas9 were selected with 100 nM atovaquone. After removal of atovaquone the strain was transfected with plasmid pL7-PFAOP and selected with 5 nM WR99210. Clonal parasite lines were obtained after limiting dilution and analysed by PCR and western blotting. Genomic DNA was isolated with phenol/chloroform as described previously77. Primers for analytical PCR reactions 1–3 in Fig. 4 were 3D7/P1/PfAOP/s (5′-ATATGATATATCTTATCGGTCCC-3′), probe/hDHFR/as (5′-CCTTTCTCCTCCTGGACATC-3′), probe/hDHFR/s (5′-CATGGTTCGCTAAACTGCATC-3′) and 3D7/P4/PfAOP/as (5′-TGGTATTACAAATAAGGGAAGAC-3′). Western blots against PfAOP were performed using the purified peptide antibody αPfAOP-5 as described previously41 with slight modifications. Parasites were isolated without magnetic cell separation by treatment with 0.05% (w/v) saponin78, 79 in ice-cold buffer containing 1.84 mM KH2PO4, 10 mM Na2HPO4, 137 mM NaCl, 2.7 mM KCl, pH 7.4. Isolated parasites were resuspended in Laemmli buffer containing 30% (v/v) 2-mercaptoethanol, boiled for 10 min and analysed by SDS-PAGE and western blotting. Growth curves were determined for three asynchronous clonal knockout lines by counting approximately 1000 erythrocytes per Giemsa-stained blood smear.
In vitro growth inhibition (IC50) assays
Cell proliferation assays in the presence of ART, ATM, tBOOH and diamide were performed using the SYBR Green method49 as described previously12, 51, 54. Synchronized ring stage parasites of overexpressing and knockout strains were incubated in 96-well plates with 1:3 serial drug dilutions. Drug concentrations ranged from 300 nM to 46 pM for ART and from 3.33 mM to 1 µM for tBOOH and diamide. Wells containing uninfected erythrocytes and infected erythrocytes without drug served as controls. Following incubation at 37 °C for 72 h, plates were frozen and stored at −80 °C. The plates were subsequently thawed at room temperature and the parasites were lysed with 2x lysis buffer containing 20 mM Tris-HCl, pH 7.5, 5 mM EDTA, 0.08% Triton X-100, 0.008% saponin and 0.12 µL/mL SYBR Green 1. The fluorescence was measured using a FLUOstar plate reader (BMG Labtech) with a gain set at 60 and an excitation and emission wavelength of 485 nm and 535 nm, respectively. Dose-response curves and IC50 values were computed using the four parameter Hill function in SigmaPlot 12.5. As a control, the PfAOP content of overexpressing parasites was analyzed by standard western blotting and quantified using SuperSignal West Femto chemiluminiscent substrate (Thermo Scientific), a C-DiGit Blot Scanner and the software ImageStudioLite (LI-COR).
Ring-stage survival assays
The in vitro ring-stage survival assay was adapted from Witkowski et al. 56, 57. Tightly sorbitol-synchronized early ring stage parasites (parasitemia 0.5–1%, hematocrit 2%, culture volume 2 mL) were treated in six well plates with 700 nM artemisinin for 6 h, washed five times with 5 mL RPMI, resuspended in complete medium and cultured for another 66 h. Blood smears were prepared, stained with Giemsa and approximately 10.000 erythrocytes were counted by light microscopy. Four independent measurements were conducted for each strain. Ring-stage survival percentages were calculated by comparing the number of viable parasites between drug-treated and mock-treated controls.
References
Klayman, D. L. Qinghaosu (artemisinin): an antimalarial drug from China. Science 228, 1049–55 (1985).
Tu, Y. The discovery of artemisinin (qinghaosu) and gifts from Chinese medicine. Nat Med 17, 1217–20 (2011).
W.H.O. Global report on antimalarial efficacy and drug resistance: 2000-2010. http://www.who.int/malaria/publications/atoz/9789241500470/en/ (2010).
W.H.O. World Malaria Report 2015. http://www.who.int/malaria/publications/world-malaria-report-2015/report/en/ (2016).
Dondorp, A. M. et al. Artemisinin resistance in Plasmodium falciparum malaria. N Engl J Med 361, 455–67 (2009).
Ashley, E. A. et al. Spread of artemisinin resistance in Plasmodium falciparum malaria. N Engl J Med 371, 411–23 (2014).
Ariey, F. et al. A molecular marker of artemisinin-resistant Plasmodium falciparum malaria. Nature 505, 50–5 (2014).
Straimer, J. et al. Drug resistance. K13-propeller mutations confer artemisinin resistance in Plasmodium falciparum clinical isolates. Science 347, 428–31 (2015).
Wilairat, P., Kumpornsin, K. & Chookajorn, T. Plasmodium falciparum malaria: Convergent evolutionary trajectories towards delayed clearance following artemisinin treatment. Med Hypotheses 90, 19–22 (2016).
Miotto, O. et al. Genetic architecture of artemisinin-resistant Plasmodium falciparum. Nat Genet 47, 226–34 (2015).
Pradhan, A. et al. Chemogenomic profiling of Plasmodium falciparum as a tool to aid antimalarial drug discovery. Sci Rep 5, 15930 (2015).
Beez, D., Sanchez, C. P., Stein, W. D. & Lanzer, M. Genetic predisposition favors the acquisition of stable artemisinin resistance in malaria parasites. Antimicrob Agents Chemother 55, 50–5 (2011).
Cui, L. & Su, X. Z. Discovery, mechanisms of action and combination therapy of artemisinin. Expert Rev Anti Infect Ther 7, 999–1013 (2009).
O’Neill, P. M., Barton, V. E. & Ward, S. A. The molecular mechanism of action of artemisinin–the debate continues. Molecules 15, 1705–21 (2010).
Mbengue, A. et al. A molecular mechanism of artemisinin resistance in Plasmodium falciparum malaria. Nature 520, 683–7 (2015).
Wang, J. et al. Haem-activated promiscuous targeting of artemisinin in Plasmodium falciparum. Nat Commun 6, 10111 (2015).
Ismail, H. M. et al. Artemisinin activity-based probes identify multiple molecular targets within the asexual stage of the malaria parasites Plasmodium falciparum 3D7. Proc Natl Acad Sci USA 113, 2080–5 (2016).
Tilley, L., Straimer, J., Gnadig, N. F., Ralph, S. A. & Fidock, D. A. Artemisinin Action and Resistance in Plasmodium falciparum. Trends Parasitol 32, 682–96 (2016).
Siwo, G. H. et al. An integrative analysis of small molecule transcriptional responses in the human malaria parasite Plasmodium falciparum. BMC Genomics 16, 1030 (2015).
Mok, S. et al. Drug resistance. Population transcriptomics of human malaria parasites reveals the mechanism of artemisinin resistance. Science 347, 431–5 (2015).
Shaw, P. J. et al. Plasmodium parasites mount an arrest response to dihydroartemisinin, as revealed by whole transcriptome shotgun sequencing (RNA-seq) and microarray study. BMC Genomics 16, 830 (2015).
Hott, A. et al. Artemisinin-resistant Plasmodium falciparum parasites exhibit altered patterns of development in infected erythrocytes. Antimicrob Agents Chemother 59, 3156–67 (2015).
Meshnick, S. R., Taylor, T. E. & Kamchonwongpaisan, S. Artemisinin and the antimalarial endoperoxides: from herbal remedy to targeted chemotherapy. Microbiol Rev 60, 301–15 (1996).
Klonis, N. et al. Artemisinin activity against Plasmodium falciparum requires hemoglobin uptake and digestion. Proc Natl Acad Sci USA 108, 11405–10 (2011).
Antoine, T. et al. Rapid kill of malaria parasites by artemisinin and semi-synthetic endoperoxides involves ROS-dependent depolarization of the membrane potential. J Antimicrob Chemother 69, 1005–16 (2014).
Berman, P. A. & Adams, P. A. Artemisinin enhances heme-catalysed oxidation of lipid membranes. Free Radic Biol Med 22, 1283–8 (1997).
Krungkrai, S. R. & Yuthavong, Y. The antimalarial action on Plasmodium falciparum of qinghaosu and artesunate in combination with agents which modulate oxidant stress. Trans R Soc Trop Med Hyg 81, 710–4 (1987).
Deponte, M. & Lillig, C. H. Enzymatic control of cysteinyl thiol switches in proteins. Biol Chem 396, 401–13 (2015).
Brigelius-Flohe, R. & Flohe, L. Basic principles and emerging concepts in the redox control of transcription factors. Antioxid Redox Signal 15, 2335–81 (2011).
Rhee, S. G. & Woo, H. A. Multiple functions of peroxiredoxins: peroxidases, sensors and regulators of the intracellular messenger H(2)O(2), and protein chaperones. Antioxid Redox Signal 15, 781–94 (2011).
Deponte, M. Glutathione catalysis and the reaction mechanisms of glutathione-dependent enzymes. Biochim Biophys Acta 1830, 3217–66 (2013).
Perkins, A., Nelson, K. J., Parsonage, D., Poole, L. B. & Karplus, P. A. Peroxiredoxins: guardians against oxidative stress and modulators of peroxide signaling. Trends Biochem Sci 40, 435–45 (2015).
Deponte, M., Rahlfs, S. & Becker, K. Peroxiredoxin systems of protozoal parasites. Subcell Biochem 44, 219–29 (2007).
Kawazu, S., Komaki-Yasuda, K., Oku, H. & Kano, S. Peroxiredoxins in malaria parasites: parasitologic aspects. Parasitol Int 57, 1–7 (2008).
Richard, D. et al. A genome-wide chromatin-associated nuclear peroxiredoxin from the malaria parasite Plasmodium falciparum. J Biol Chem 286, 11746–55 (2011).
Gretes, M. C., Poole, L. B. & Karplus, P. A. Peroxiredoxins in parasites. Antioxid Redox Signal 17, 608–33 (2012).
Koncarevic, S. et al. The malarial parasite Plasmodium falciparum imports the human protein peroxiredoxin 2 for peroxide detoxification. Proc Natl Acad Sci USA 106, 13323–8 (2009).
Akerman, S. E. & Muller, S. 2-Cys peroxiredoxin PfTrx-Px1 is involved in the antioxidant defence of Plasmodium falciparum. Mol Biochem Parasitol 130, 75–81 (2003).
Kawazu, S. et al. Expression profiles of peroxiredoxin proteins of the rodent malaria parasite Plasmodium yoelii. Int J Parasitol 33, 1455–61 (2003).
Yano, K. et al. Expression of mRNAs and proteins for peroxiredoxins in Plasmodium falciparum erythrocytic stage. Parasitol Int 54, 35–41 (2005).
Djuika, C. F. et al. Prokaryotic ancestry and gene fusion of a dual localized peroxiredoxin in malaria parasites. Microbial Cell 2, 5–13 (2015).
Yano, K. et al. 2-Cys Peroxiredoxin TPx-1 is involved in gametocyte development in Plasmodium berghei. Mol Biochem Parasitol 148, 44–51 (2006).
Komaki-Yasuda, K., Kawazu, S. & Kano, S. Disruption of the Plasmodium falciparum 2-Cys peroxiredoxin gene renders parasites hypersensitive to reactive oxygen and nitrogen species. FEBS Lett 547, 140–4 (2003).
Kimura, R., Komaki-Yasuda, K., Kawazu, S. & Kano, S. 2-Cys peroxiredoxin of Plasmodium falciparum is involved in resistance to heat stress of the parasite. Parasitol Int 62, 137–43 (2013).
Djuika, C. F. et al. Plasmodium falciparum antioxidant protein as a model enzyme for a special class of glutaredoxin/glutathione-dependent peroxiredoxins. Biochim Biophys Acta 1830, 4073–90 (2013).
Nickel, C., Rahlfs, S., Deponte, M., Koncarevic, S. & Becker, K. Thioredoxin networks in the malarial parasite Plasmodium falciparum. Antioxid Redox Signal 8, 1227–39 (2006).
Staudacher, V. et al. Plasmodium falciparum antioxidant protein reveals a novel mechanism for balancing turnover and inactivation of peroxiredoxins. Free Radic Biol Med 85, 228–36 (2015).
Sa, J. M. et al. Geographic patterns of Plasmodium falciparum drug resistance distinguished by differential responses to amodiaquine and chloroquine. Proc Natl Acad Sci USA 106, 18883–9 (2009).
Smilkstein, M., Sriwilaijaroen, N., Kelly, J. X., Wilairat, P. & Riscoe, M. Simple and inexpensive fluorescence-based technique for high-throughput antimalarial drug screening. Antimicrob Agents Chemother 48, 1803–6 (2004).
Gisselberg, J. E., Dellibovi-Ragheb, T. A., Matthews, K. A., Bosch, G. & Prigge, S. T. The suf iron-sulfur cluster synthesis pathway is required for apicoplast maintenance in malaria parasites. PLoS Pathog 9, e1003655 (2013).
Sanchez, C. P., Mayer, S., Nurhasanah, A., Stein, W. D. & Lanzer, M. Genetic linkage analyses redefine the roles of PfCRT and PfMDR1 in drug accumulation and susceptibility in Plasmodium falciparum. Mol Microbiol 82, 865–78 (2011).
Fidock, D. A. et al. Mutations in the P. falciparum digestive vacuole transmembrane protein PfCRT and evidence for their role in chloroquine resistance. Mol Cell 6, 861–71 (2000).
Ghorbal, M. et al. Genome editing in the human malaria parasite Plasmodium falciparum using the CRISPR-Cas9 system. Nat Biotechnol 32, 819–21 (2014).
Wezena, C.A., Krafczyk, J., Staudacher, V. & Deponte, M. Growth inhibitory effects of standard pro- and antioxidants on the human malaria parasite Plasmodium falciparum. Exp Parasitol (2017).
Tucker, M. S., Mutka, T., Sparks, K., Patel, J. & Kyle, D. E. Phenotypic and genotypic analysis of in vitro-selected artemisinin-resistant progeny of Plasmodium falciparum. Antimicrob Agents Chemother 56, 302–14 (2012).
Witkowski, B. et al. Reduced artemisinin susceptibility of Plasmodium falciparum ring stages in western Cambodia. Antimicrob Agents Chemother 57, 914–23 (2013).
Witkowski, B. et al. Novel phenotypic assays for the detection of artemisinin-resistant Plasmodium falciparum malaria in Cambodia: in-vitro and ex-vivo drug-response studies. Lancet Infect Dis 13, 1043–9 (2013).
Kite, W. A., Melendez-Muniz, V. A., Moraes Barros, R. R., Wellems, T. E. & Sa, J. M. Alternative methods for the Plasmodium falciparum artemisinin ring-stage survival assay with increased simplicity and parasite stage-specificity. Malar J 15, 94 (2016).
Imlay, J. A. The molecular mechanisms and physiological consequences of oxidative stress: lessons from a model bacterium. Nat Rev Microbiol 11, 443–54 (2013).
Jomaa, H. et al. Inhibitors of the nonmevalonate pathway of isoprenoid biosynthesis as antimalarial drugs. Science 285, 1573–6 (1999).
Yeh, E. & DeRisi, J. L. Chemical rescue of malaria parasites lacking an apicoplast defines organelle function in blood-stage Plasmodium falciparum. PLoS Biol 9, e1001138 (2011).
Rekittke, I. et al. Structure of the (E)-4-hydroxy-3-methyl-but-2-enyl-diphosphate reductase from Plasmodium falciparum. FEBS Lett 587, 3968–72 (2013).
Ralph, S. A. et al. Tropical infectious diseases: metabolic maps and functions of the Plasmodium falciparum apicoplast. Nat Rev Microbiol 2, 203–16 (2004).
Kimata-Ariga, Y., Saitoh, T., Ikegami, T., Horii, T. & Hase, T. Molecular interaction of ferredoxin and ferredoxin-NADP + reductase from human malaria parasite. J Biochem 142, 715–20 (2007).
Vollmer, M., Thomsen, N., Wiek, S. & Seeber, F. Apicomplexan parasites possess distinct nuclear-encoded, but apicoplast-localized, plant-type ferredoxin-NADP + reductase and ferredoxin. J Biol Chem 276, 5483–90 (2001).
Pino, P. et al. Dual targeting of antioxidant and metabolic enzymes to the mitochondrion and the apicoplast of Toxoplasma gondii. PLoS Pathog 3, e115 (2007).
Kawazu, S., Ikenoue, N., Takemae, H., Komaki-Yasuda, K. & Kano, S. Roles of 1-Cys peroxiredoxin in haem detoxification in the human malaria parasite Plasmodium falciparum. FEBS J 272, 1784–91 (2005).
Kehr, S., Sturm, N., Rahlfs, S., Przyborski, J. M. & Becker, K. Compartmentation of redox metabolism in malaria parasites. PLoS Pathog 6, e1001242 (2010).
Chaudhari, R., Narayan, A. & Patankar, S. A novel trafficking pathway in Plasmodium falciparum for the organellar localization of glutathione peroxidase-like thioredoxin peroxidase. FEBS J 279, 3872–88 (2012).
Turturice, B. A. et al. Expression of cytosolic peroxiredoxins in Plasmodium berghei ookinetes is regulated by environmental factors in the mosquito bloodmeal. PLoS Pathog 9, e1003136 (2013).
Dogovski, C. et al. Targeting the cell stress response of Plasmodium falciparum to overcome artemisinin resistance. PLoS Biol 13, e1002132 (2015).
Urscher, M., More, S. S., Alisch, R., Vince, R. & Deponte, M. Tight-binding inhibitors efficiently inactivate both reaction centers of monomeric Plasmodium falciparum glyoxalase 1. FEBS J 279, 2568–78 (2012).
Trager, W. & Jensen, J. B. Human malaria parasites in continuous culture. Science 193, 673–5 (1976).
Lambros, C. & Vanderberg, J. P. Synchronization of Plasmodium falciparum erythrocytic stages in culture. J Parasitol 65, 418–20 (1979).
Haley, C. S. & Knott, S. A. A simple regression method for mapping quantitative trait loci in line crosses using flanking markers. Heredity (Edinb) 69, 315–24 (1992).
Deitsch, K., Driskill, C. & Wellems, T. Transformation of malaria parasites by the spontaneous uptake and expression of DNA from human erythrocytes. Nucleic Acids Res 29, 850–3 (2001).
Beck, H. P. Extraction and purification of Plasmodium parasite DNA. Methods Mol Med 72, 159–63 (2002).
Zuckerman, A., Spira, D. & Hamburger, J. A procedure for the harvesting of mammalian plasmodia. Bull World Health Organ 37, 431–6 (1967).
Benting, J., Mattei, D. & Lingelbach, K. Brefeldin A inhibits transport of the glycophorin-binding protein from Plasmodium falciparum into the host erythrocyte. Biochem J 300(Pt 3), 821–6 (1994).
Acknowledgements
This work was supported by the Deutsche Forschungsgemeinschaft (Grant DE 1431/8–1 of the priority program SPP 1710 to M.D.). The position of M.D. was funded by the Deutsche Forschungsgemeinschaft in the frame of the Heisenberg programme (Grant DE 1431/9–1). We thank Stefan Prior and Marina Müller for technical assistance with the QTL analysis. We thank J.J. Lopez-Rubio for the Cas9 and pL6 plasmids and Jacobus Pharmaceutical Company for WR99210.
Author information
Authors and Affiliations
Contributions
C.F.D. performed the kinetic measurements & generated and analysed the overexpressing strains. V.S. generated and analysed the knockout strain. C.P.S. performed the QTL analysis. M.L. conceived and supervised the QTL experiments. M.D. conceived and supervised all other experiments and wrote the manuscript. All authors discussed the results and gave approval to the final version of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Djuika, C.F., Staudacher, V., Sanchez, C.P. et al. Knockout of the peroxiredoxin 5 homologue PFAOP does not affect the artemisinin susceptibility of Plasmodium falciparum . Sci Rep 7, 4410 (2017). https://doi.org/10.1038/s41598-017-04277-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-04277-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.