Abstract
Microcos paniculata is a shrub used traditionally as folk medicine and to make herbal teas. Previous research into this species has mainly focused on its chemical composition and medicinal value. However, the lack of a reference genome limits the study of the molecular mechanisms of active compounds in this species. Here, we assembled a haplotype-resolved chromosome-level genome of M. paniculata based on PacBio HiFi and Hi-C data. The assembly contains two haploid genomes with sizes 399.43 Mb and 393.10 Mb, with contig N50 lengths of 43.44 Mb and 30.17 Mb, respectively. About 99.93% of the assembled sequences could be anchored to 18 pseudo-chromosomes. Additionally, a total of 482 Mb repeat sequences were identified, accounting for 60.76% of the genome. A total of 49,439 protein-coding genes were identified, of which 48,979 (99%) were functionally annotated. This haplotype-resolved chromosome-level assembly and annotation of M. paniculata will serve as a valuable resource for investigating the biosynthesis and genetic basis of active compounds in this species, as well as advancing evolutionary phylogenomic studies in Malvales.
Similar content being viewed by others
Background & Summary
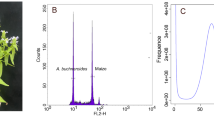
Microcos paniculata Linnaeus (Fig. 1a), known in Chinese as Buzhaye, is a shrub commonly used in traditional Chinese medicine and herbal cooling teas1, including Wanglaoji, Huoqizheng2 and Jiaduobao, with an annual demand of about 250 tons (http://bk.cnpharm.com/zgyyb/2008/04/28/246974.html). The leaves of M. paniculata are also commonly used in ethnomedicinal treatments for food stagnation, damp-heat jaundice and fever3. Up to now, numerous studies have extensively investigated the phytochemical composition and pharmacological properties of this species, revealing the existence of bioactive secondary metabolites such as flavonoids, alkaloids, triterpenoids and organic acids1,4 from M. paniculata extracts. However, due to the lack of a high-quality reference genome, the molecular basis and evolution of the secondary metabolite biosynthesis in M. paniculata are rarely reported5.
Morphological characters (a) and the landscape of genome assembly and annotation of M. paniculata (b). The tracks from outside to inside are: pseudo-chromosomes, density of class I TEs, density of class II TEs, density of protein-coding genes, proportion of tandem repeats, GC content and collinear blocks.
In the present study, we assembled the genome of M. paniculata using 106 × short reads (42 Gb), 35 × HiFi reads (14 Gb), 75 × Hi-C reads (30 Gb) and 50 × iso-seq reads (20 Gb). The final assembly (~792 Mb) consisted of two complete haplotypes, haplotype A (399.43 Mb) and haplotype B (393.10 Mb), with contig N50 lengths of 43.44 Mb and 30.17 Mb, respectively (Table 1). About 99.93% of the assembled sequences were anchored onto 18 (2n) pseudo-chromosomes (Fig. 1b). The chloroplast and mitochondrial genomes were 159,456 bp and 380,905 bp, respectively. A total of 1,080,648 repeat sequences, with an approximate length of 482 Mb were identified, accounting for 60.76% of the assembled genome. Of the identified repeats, long terminal repeats (LTRs) constituted the largest proportion, with a number of 394,112 and a cumulative length of 321,160,287 bp, accounting for 40.52% of the M. paniculata genome assembly (Table 2). The genome contained 65,874 genes, including 49,439 protein-coding genes and 16,435 non-coding genes (Table 3). A total of 48,979 genes were functionally annotated, accounting for 99% of the identified protein-coding genes (Table 4). Of these, 44,971 genes were annotated by all three methods together (Fig. 2). In particular, 639 genes have been annotated as being related to the biosynthesis or metabolism of flavonoids, alkaloids and triterpenoids (Table S1). The resulting high-quality reference genome and annotation of M. paniculata will be a valuable resource for improving our understanding of the evolutionary relationships within the Malvales, for studying the molecular basis and biosynthetic mechanisms of phytochemical compounds, and for further study and exploitation of M. paniculata.
Methods
Sample collection and genome sequencing
Samples of M. paniculata were collected at Xishuangbanna Tropical Botanical Garden (XTBG), Chinese Academy of Sciences, Mengla, Yunnan Province, China. Genomic DNA was extracted using a modified CTAB method6. DNA quality was assessed using a NanoDrop One spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA) and a Qubit 3.0 Fluorometer (Life Technologies, Carlsbad, CA, USA). Whole genome sequencing, Pacbio sequencing, Hi-C (high-through chromosome conformation capture) sequencing and full-length isoform sequencing (iso-seq) were performed at Wuhan Benagen Technology Co. Ltd. (Wuhan, China).
For whole genome sequencing, 1 μg of genomic DNA was sonicated to an approximate size range of 200–400 bp using a sonicator (Covaris, Brighton, UK). The short-read libraries were constructed following the manufacturer’s instructions and then sequenced on the DNBSEQ-T7 platform (BGI lnc., Shenzhen, China) using the PE (paired-end) 150 model.
For long-read sequencing, genomic DNA was sheared using the Megaruptor 3 shearing kit (Diagenode SA., Seraing, Belgium). The AMPure PB beads size selection kit (Pacbio, Menlo Park, CA, USA) was used to selectively deplete DNA fragments smaller than 5 kb. The libraries were prepared using the SMRTbell® prep kit 3.0 (Pacbio, Menlo Park, CA, USA) and then sequenced on a Revio system (Pacbio, Menlo Park, CA, USA). Raw sequencing data were converted to HiFi (high fidelity) reads using the CCS workflow 7.0.07 with parameters (--streamed --log-level INFO --stderr-json-log --kestrel-files-layout–min-rq 0.9 --non-hifi-prefix fail --knrt-ada --pbdc-model).
For Hi-C sequencing, leaf material from young shoots was fixed in 2% formaldehyde solution, and the Hi-C library was generated following a published protocol8. Briefly, the cross-linked materials were digested with 400 units of MboI, and marked with biotin-14-dCTP, and then subjected to blunt-end ligation of crosslinked fragments. After re-ligation, reverse crosslinking and purification, the chromatin DNA was sheared to a size of 200–600 bp using sonication. The biotin-labelled Hi-C fragments were then enriched using streptavidin magnetic beads. After the addition of A-tailing and an adapter, the Hi-C libraries were PCR-amplified (12–14 cycles) and then sequenced on the DNBSEQ-T7 platform (BGI lnc., Shenzhen, China) in PE150 mode.
Full-length isoform sequencing (iso-seq) was used to obtain high quality transcriptomic data. RNA was extracted from leaves, flowers and stems of M. paniculata using the R6827 Plant RNA Kit (Omega Bio-Tek, Norcross, GA, USA) following the manufacturer’s instructions. The cDNA-PCR Sequencing kit SQK-PCS109 by Oxford Nanopore (Oxford Nanopore Technologies, Oxford, UK) was used to prepare full-length cDNA libraries. The libraries were then sequenced on the PromethION sequencer (Oxford Nanopore Technologies, Oxford, UK).
Genome assembly
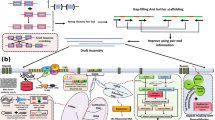
PacBio HiFi reads and Hi-C short reads were combined as input to Hifiasm v0.19.5-r5929 using the default parameters to generate haplotype-resolved contigs for subsequent analysis. Hi-C reads were mapped to the assembled haplotype contigs using Juicer v1.5.610, and a Hi-C-assisted initial chromosome assembly was then performed using the 3D-DNA v18092211 pipeline (with the parameters --early-exit -m haploid -r 0). Chromosome boundaries were then adjusted and the misjoins and switch errors were corrected manually using Juicebox v1.11.0812. This process generated chromosome-scale scaffolds and un-anchored contig sequences.
LR_Gapcloser v1.1.113 was used to fill gaps in the chromosome assembly based on HiFi reads (with the parameters -s p -r 2 -g 500 -v 500 -a 0.25). HiFi reads were then re-mapped to the chromosome scaffolds. The mapped reads located around the telomere repeat sequences (TTTAGGG)n14 were then extracted and assembled into contigs using Hifiasm v0.19.5-r592 with the default parameters. The resulting contigs were aligned back to the chromosome scaffold to extend the chromosome ends for telomere sequences, and totally 28 telomere sequences were obtained (Fig. 3a). In addition, GetOrganelle v1.7.515 was used to assemble the chloroplast and mitochondrial genomes.
Nextpolish2 v0.1.016 was used to polish the above assembly based on HiFi reads and short reads with default parameters. Redundant haplotigs and rDNA fragments were removed using the Redundans v0.13c17 pipeline (with the parameters -identity 0.98 -overlap 0.8) and manually curated. A high quality haplotype resolved genome assembly of M. paniculata was then obtained.
Repeat annotation
The EDTA (Extensive de novo TE Annotator) program v1.9.918 (with the parameters --sensitive 1 --anno 1) was used for the de novo identification of transposable elements (TE), generating a TE library. RepeatMasker v4.0.719 was utilized to identify repeat elements (with the parameters -no_is -xsmall).
Annotation of protein-coding genes and noncoding RNAs
A total of 314,962 publicly available non-redundant protein sequences from Theobroma cacao20, Durio zibethinus21, Corchorus capsularis22, Gossypium raimondii23, Heritiera littoralis24, Dipterocarpus turbinatus25, Aquilaria sinensis26, Arabidopsis thaliana27, Carica papaya28, Vitis vinifera29, and Bombax ceiba30 were used as homologous protein evidence for gene annotation. Iso-seq data were mapped to the genome using Minimap2 v2.2431 (with the parameters -a -x splice --end-seed-pen = 60 --G 200k), then assembled in StringTie v1.3.532 (with the parameters -L -t -f 0.05), and the resulting sequences were used as transcript evidence.
PASA (Program to Assemble Spliced Alignments) v2.4.133 was used to annotate the genomic structure based on transcript evidence with the default parameters. Then, full-length gene sequences were identified by aligning with homologous protein evidence using BLAT34 (-prot) and removing the hits with query or target coverage <95%. The gene model was trained and optimized for five rounds in AUGUSTUS v3.4.035 using the full-length gene set with the default parameters.
The MAKER2 v2.31.936 pipeline was used to perform annotation based on ab initio prediction, the transcript evidence and the homologous protein evidence. Briefly: (1) RepeatMasker v4.0.719 was used to mask repeat sequences in the genome; (2) AUGUSTUS v3.4.035 was used for ab initio prediction based on the genomic sequence; (3) BLASTN was used to align the transcript evidence to the repeat-masked genome, and BLASTX was employed to align the homologous protein evidence to the genome. Exonerate v2.2.037 was used to realign the BLAST hits to the genome; (4) Finally, the predicted gene models were integrated using MAKER2 based on the hints generated from the above alignments.
EvidenceModeler (EVM) v1.1.138 was further employed to merge the annotation results obtained from PASA v2.4.1 and MAKER2 v2.31.9, generating consensus annotations. TEsorter v1.4.139 was utilized to identify TE protein domains on the genome (with the parameters -genome -db rexdb -cov 30 -eval 1e-5 -prob 0.9), and these domains were masked in the EVM process. The results obtained from EVM were refined by incorporating UTR sequences and alternative splicing using PASA v2.4.1 with the default parameters. Annotations that were too short (<50 amino acids), lacked start or stop codons, contained an internal stop codon, or had ambiguous bases were excluded. All annotations were then merged, and redundant annotations were removed.
In addition, for non-coding RNA (ncRNA) annotations, tRNAScan-SE v1.3.140 was used to identify transfer RNA (tRNA), and Barrnap v0.9 (https://github.com/tseemann/barrnap) was used to identify ribosomal RNA (rRNA). To ensure accuracy, partial rRNA annotations were excluded. Furthermore, RfamScan v14.241 was used to identify other ncRNA.
We employed three strategies to predict the function of the protein-coding genes: (1) eggNOG-mapper v2.0.042 (--target_taxa Viridiplantae -m diamond) was utilized to search for homologous genes in the eggNOG database, enabling Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) annotation; (2) DIAMOND v0.9.2443 (--evalue 1e-5 --max-target-seqs 5) was employed to align protein-coding genes with the Swiss-Prot, TrEMBL, NR (non-redundant protein in NCBI), and the TAIR10 protein databases; (3) InterProScan v5.27-66.044 was used to annotate protein domains and motifs by searching multiple publicly available databases, such as PRINTS, Pfam, SMART, PANTHER, and CDD of the InterPro database. TBtools v1.13245 was then used to draw a Venn diagram to show unique and shared protein-coding genes annotated using the three described strategies.
Comparison between haplotype assemblies
SyRI (Synteny and Rearrangement Identifier) v1.646 was used to detect synteny and genomic structural variations (≥50 bp in size) between the two haplotypes, with the default parameters. In total, our analysis identified 3,011 syntenic regions (∼350 Mb), 768 translocations (∼45 Mb), 20 inversions (∼2 Mb), 2,175 duplications in haplotype A (~15 Mb) and 1,686 duplications in haplotype B (~8 Mb). Most duplications were found on chromosomes 4 and 8, and most inversions were found on chromosome 7 (Fig. 3b). SyRI v1.6 was also used to identify SNPs, small InDels (insertions and deletions, <50 bp in size) and tandem repeats. Finally, 1,264,264 SNPs (∼1 Mb), 105,563 insertions (∼2 Mb in haplotype B), 100,073 deletions (∼2 Mb in haplotype A) and 282 tandem repeats (∼1 Mb) were identified.
Data Records
The BGI short reads, PacBio HiFi long reads, Hi-C reads and Iso-Seq data have been deposited at the Sequence Read Archive database of NCBI (National Center for Bioinformation Information) under accession numbers SRR25456891-SRR2545689447,48,49,50. The final genome assembly has been deposited at the GenBank database under the accession numbers GCA_030664735.151 and GCA_030664755.152. The genome annotations are available from the Figshare repository53. The AUGUSTUS model trained and optimized for this genome, together with the configuration files for MAKER are available from the Figshare repository54.
Technical Validation
We first calculated the mapping rate as a measure of assembly accuracy. The short reads and the long reads were re-mapped to the assembly using BWA-MEM v0.7.17-r118855 and Minimap2 v2.2431, respectively, with the default parameters. The mapping rates were calculated after filtering out non-primary alignments. In total, 99.89% of HiFi reads, 97.75% of iso-seq reads and 99.81% of short reads were mapped (Table 5). Moreover, the read coverage depth of both short and long read data was evenly distributed along each phased chromosome, indicating high quality of our haplotype-resolved assembly (Figure S1).
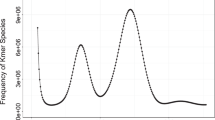
We evaluated the completeness of the genome assembly using BUSCO (Benchmarking Universal Single-Copy Orthologs) v5.3.256 based on the embryophyta_odb10 ortholog database. The BUSCO evaluation of the haplotype A identified 1,591 complete BUSCOs (including 1,561 single and 30 duplicated BUSCOs), accounting for 98.6% of the haplotype, while the missing BUSCOs represented merely 0.7% (Table 6). Similarly, the BUSCO assessment of the haplotype B identified 1,588 complete BUSCOs (including 1,560 single and 28 duplicated BUSCOs), accounting for 98.4% of the haplotype, while the missing BUSCOs were only 0.9% (Table 6). This indicates a relatively complete assembly. We used Merqury v1.357 to estimate the consensus and completeness of the genome assembly. Our results gave a consensus quality value (QV) of 73.38 for the genome assembly, and the completeness value was 99.19% (Table 6). We also used KAT (K-mer Analysis Toolkit) v2.4.058 to estimate the quality of the genome assembly by comparing k-mers in HiFi reads and in the assembly. Our results show high consistency between the reads and the genome assembly (Fig. 4a), with each haplotype representing approximately half of the heterozygous peak and nearly all of the homozygous peak (Fig. 4b,c).
In addition, we used BUSCO to evaluate the completeness of the genome annotation by retaining only the longest protein sequence for each gene, and found that the annotation of haplotype A was 97.6% complete, with only 17 (1.1%) genes missing, and the annotation of haplotype B was 97.1% complete, with only 19 (1.2%) genes missing (Table 7), indicating that the annotation was of high quality.
The Hi-C reads were aligned to the genome assembly using Juicer v1.5.610 with the default parameters. The Juicebox12 tools pre command (pre -n -q 0 or 1) was used to convert the raw file generated by Juicer into hic format, and dump command (dump observed BP 100000) was used to extract 100-kb contact matrix from the hic file. The hic file was visualized by Juicebox. Strong interactive signals were observed around the diagonal of the pseudo-chromosomes, and there was no obvious noise outside the diagonal (Fig. 5a), indicating the high quality of this chromosome assembly. In addition, no anomalies were observed across each homologous chromosome pair when duplicated reads were excluded (Fig. 5b), suggesting no switch errors between phased haplotypes.
Code availability
All commands and pipelines used were performed according to the manuals or protocols of the tools used in this study. The software and tools used are publicly accessible, with the version and parameters specified in the Methods section. If no detailed parameters were mentioned, default parameters were used. No custom code was used in this study.
References
Jiang, Y. Q. & Liu, E. H. Microcos paniculata: a review on its botany, traditional uses, phytochemistry and pharmacology. Chin. J. Nat. Med. 17, 561–574, https://doi.org/10.1016/S1875-5364(19)30058-5 (2019).
Wang, J. et al. Deciphering the formulation secret underlying Chinese Huo-Clearing herbal drink. Front. Pharmacol. 12, 654699, https://doi.org/10.3389/fphar.2021.654699 (2021).
National Medical Products Administration & National Health Commission of the People’s Republic of China. Pharmacopoeia of the People’s Republic of China Vol. 1 (China Medical Science Press, 2020).
Yang, Y., Li, S. G., Ye, W. C. & Jiang, R. W. Studies on chemical constituents of Microcos paniculata. Lishizhen Medicine and Materia Medica Research 21, 2790–2792 (2010).
Lin, S. Transcriptomic sequencing of Microcos paniculata and the excavation of key genes in ACGs biosynthesis. Master thesis, Guangdong Pharmaceutical University (2017).
Doyle, J. J. & Doyle, J. L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin 19, 11–15 (1987).
Wenger, A. M. et al. Accurate circular consensus long-read sequencing improves variant detection and assembly of a human genome. Nat. Biotechnol. 37, 1155–1162, https://doi.org/10.1038/s41587-019-0217-9 (2019).
van Berkum, N. L. et al. Hi-C: A method to study the three-dimensional architecture of genomes. J. Vis. Exp. 39, e1869 https://doi.org/10.3791/1869 (2010).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 18, 170–175, https://doi.org/10.1038/s41592-020-01056-5 (2021).
Durand, N. C. et al. Juicer provides a One-Click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98, https://doi.org/10.1016/j.cels.2016.07.002 (2016).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95, https://doi.org/10.1126/science.aal3327 (2017).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3, 99–101, https://doi.org/10.1016/j.cels.2015.07.012 (2016).
Xu, G. C. et al. LR_Gapcloser: a tiling path-based gap closer that uses long reads to complete genome assembly. GigaScience 8, giy157, https://doi.org/10.1093/gigascience/giy157 (2018).
Gao, D. et al. TAR30, a homolog of the canonical plant TTTAGGG telomeric repeat, is enriched in the proximal chromosome regions of peanut (Arachis hypogaea L.). Chromosome Res. 30, 77–90, https://doi.org/10.1007/s10577-022-09684-7 (2022).
Jin, J. J. et al. GetOrganelle: a fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biol. 21, 241, https://doi.org/10.1186/s13059-020-02154-5 (2020).
Hu, J. et al. NextPolish2: a repeat-aware polishing tool for genomes assembled using HiFi long reads. bioRxiv, 2023.2004.2026.538352 https://doi.org/10.1101/2023.04.26.538352 (2023).
Pryszcz, L. P. & Gabaldón, T. Redundans: an assembly pipeline for highly heterozygous genomes. Nucleic Acids Res. 44, e113, https://doi.org/10.1093/nar/gkw294 (2016).
Ou, S. et al. Benchmarking transposable element annotation methods for creation of a streamlined, comprehensive pipeline. Genome Biol. 20, 275, https://doi.org/10.1186/s13059-019-1905-y (2019).
Tarailo-Graovac, M. & Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr. Protoc. Bioinformatics 25, 4.10.11–4.10.14, https://doi.org/10.1002/0471250953.bi0410s25 (2009).
Motamayor, J. C. et al. The genome sequence of the most widely cultivated cacao type and its use to identify candidate genes regulating pod color. Genome Biol. 14, r53, https://doi.org/10.1186/gb-2013-14-6-r53 (2013).
Teh, B. T. et al. The draft genome of tropical fruit durian (Durio zibethinus). Nat. Genet. 49, 1633–1641, https://doi.org/10.1038/ng.3972 (2017).
Zhang, L. et al. Reference genomes of the two cultivated jute species. Plant Biotechnol. J. 19, 2235–2248, https://doi.org/10.1111/pbi.13652 (2021).
Wang, M. et al. Genomic innovation and regulatory rewiring during evolution of the cotton genus Gossypium. Nat. Genet. 54, 1959–1971, https://doi.org/10.1038/s41588-022-01237-2 (2022).
He, Z. et al. Evolution of coastal forests based on a full set of mangrove genomes. Nat. Ecol. Evol. 6, 738–749, https://doi.org/10.1038/s41559-022-01744-9 (2022).
Wang, S. et al. The chromosome-scale genomes of Dipterocarpus turbinatus and Hopea hainanensis (Dipterocarpaceae) provide insights into fragrant oleoresin biosynthesis and hardwood formation. Plant Biotechnol. J. 20, 538–553, https://doi.org/10.1111/pbi.13735 (2022).
Ding, X. et al. Genome sequence of the agarwood tree Aquilaria sinensis (Lour.) Spreng: the first chromosome-level draft genome in the Thymelaeceae family. GigaScience 9, giaa013, https://doi.org/10.1093/gigascience/giaa013 (2020).
Cheng, C.-Y. et al. Araport11: a complete reannotation of the Arabidopsis thaliana reference genome. Plant. J. 89, 789–804, https://doi.org/10.1111/tpj.13415 (2017).
Yue, J. et al. SunUp and Sunset genomes revealed impact of particle bombardment mediated transformation and domestication history in papaya. Nat. Genet. 54, 715–724, https://doi.org/10.1038/s41588-022-01068-1 (2022).
Jaillon, O. et al. The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449, 463–467, https://doi.org/10.1038/nature06148 (2007).
Gao, Y. et al. De novo genome assembly of the red silk cotton tree (Bombax ceiba). GigaScience 7, giy051, https://doi.org/10.1093/gigascience/giy051 (2018).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100, https://doi.org/10.1093/bioinformatics/bty191 (2018).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295, https://doi.org/10.1038/nbt.3122 (2015).
Haas, B. J. et al. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic Acids Res. 31, 5654–5666, https://doi.org/10.1093/nar/gkg770 (2003).
Kent, W. J. BLAT—The BLAST-Like Alignment Tool. Genome Res. 12, 656–664, https://doi.org/10.1111/10.1101/gr.229202 (2002).
Stanke, M., Diekhans, M., Baertsch, R. & Haussler, D. Using native and syntenically mapped cDNA alignments to improve de novo gene finding. Bioinformatics 24, 637–644, https://doi.org/10.1093/bioinformatics/btn013 (2008).
Holt, C. & Yandell, M. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinformatics 12, 491, https://doi.org/10.1186/1471-2105-12-491 (2011).
Slater, G. S. C. & Birney, E. Automated generation of heuristics for biological sequence comparison. BMC Bioinformatics 6, 31, https://doi.org/10.1186/1471-2105-6-31 (2005).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biol. 9, R7, https://doi.org/10.1186/gb-2008-9-1-r7 (2008).
Zhang, R. G. et al. TEsorter: An accurate and fast method to classify LTR-retrotransposons in plant genomes. Hortic. Res. 9, uhac017, https://doi.org/10.1093/hr/uhac017 (2022).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964, https://doi.org/10.1093/nar/25.5.955 (1997).
Nawrocki, E. P. et al. Rfam 12.0: updates to the RNA families database. Nucleic Acids Res. 43, D130–D137, https://doi.org/10.1093/nar/gku1063 (2014).
Huerta-Cepas, J. et al. Fast genome-wide functional annotation through orthology assignment by eggNOG-Mapper. Mol. Biol. Evol. 34, 2115–2122, https://doi.org/10.1093/molbev/msx148 (2017).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60, https://doi.org/10.1038/nmeth.3176 (2015).
Jones, P. et al. InterProScan5: genome-scale protein function classification. Bioinformatics 30, 1236–1240, https://doi.org/10.1093/bioinformatics/btu031 (2014).
Chen, C. et al. TBtools: An integrative toolkit developed for interactive analyses of big diological data. Mol. Plant 13, 1194–1202, https://doi.org/10.1016/j.molp.2020.06.009 (2020).
Goel, M., Sun, H., Jiao, W. B. & Schneeberger, K. SyRI: finding genomic rearrangements and local sequence differences from whole-genome assemblies. Genome Biol. 20, 277, https://doi.org/10.1186/s13059-019-1911-0 (2019).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR25456891 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR25456892 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR25456893 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR25456894 (2023).
NCBI Assembly https://identifiers.org/ncbi/insdc.gca:GCA_030664735.1 (2023).
NCBI Assembly https://identifiers.org/ncbi/insdc.gca:GCA_030664755.1 (2023).
Liu, D. T., Tian, X. L., Shao, S. C., Ma, Y. P. & Zhang, R. G. Haplotype-resolved chromosomal-level assembly of Buzhaye (Microcos paniculata), a medicinal plant in China. Figshare https://doi.org/10.6084/m9.figshare.22892834.v1 (2023).
Zhang, R. G. AUGUSTUS model for genome annotation of Microcos paniculata. Figshare https://doi.org/10.6084/m9.figshare.24493639 (2023).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. ArXiv, 1303.3997 https://doi.org/10.48550/arXiv.1303.3997 (2013).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212, https://doi.org/10.1093/bioinformatics/btv351 (2015).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol. 21, 245, https://doi.org/10.1186/s13059-020-02134-9 (2020).
Mapleson, D., Garcia Accinelli, G., Kettleborough, G., Wright, J. & Clavijo, B. J. KAT: a K-mer analysis toolkit to quality control NGS datasets and genome assemblies. Bioinformatics 33, 574–576, https://doi.org/10.1093/bioinformatics/btw663 (2016).
Acknowledgements
This work was supported by West Light Talent Programs of the Chinese Academy of Sciences (Grant Nos. E1XB011B01 & E2227151W1), Key Basic Research Programs of Yunnan Province (Grant Nos. 202302AE090018 & 202101BC070003) and the Ten Thousand Talent Program of Yunnan Province (Grant No. YNWR-QNBJ-2018-174).
Author information
Authors and Affiliations
Contributions
S.C.S., R.G.Z. and Y.P.M. designed and funded the study, S.C.S. collected and prepared the samples, R.G.Z. performed the bioinformatic analyses. D.T.L. and X.L.T. drafted the manuscript. R.G.Z., Y.P.M. and D.T.L. revised the manuscript. All authors approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
41597_2023_2821_MOESM2_ESM.xlsx
The annotated genes putatively involved in the biosynthesis or metabolism of flavonoids, alkaloids and triterpenoids in Buzhaye
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, D., Tian, X., Shao, S. et al. Haplotype-resolved chromosomal-level genome assembly of Buzhaye (Microcos paniculata). Sci Data 10, 901 (2023). https://doi.org/10.1038/s41597-023-02821-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02821-9