Abstract
We present an extensive and diverse dataset of bond separation energies associated with the homolytic cleavage of covalently bonded molecules (A-B) into their corresponding radical fragments (A. and B.). Our dataset contains two different classifications of model structures referred to as “Existing” (molecules with associated experimental data) and “Hypothetical” (molecules with no associated experimental data). In total, the dataset consists of 4502 datapoints (1969 datapoints from the Existing and 2533 datapoints from the Hypothetical classes). The dataset covers 49 unique X-Y type single bonds (except H-H, H-F, and H-Cl), where X and Y are H, B, C, N, O, F, Si, P, S, and Cl atoms. All the reference data was calculated at the (RO)CBS-QB3 level of theory. The reference bond separation energies are non-relativistic ground-state energy differences and contain no zero-point energy corrections. This new dataset of bond separation energies (BSE49) is presented as a high-quality reference dataset for assessing and developing computational chemistry methods.
Measurement(s) | bond separation energies |
Technology Type(s) | ab initio quantum chemistry computational method |
Factor Type(s) | existing or hypothetical model structure |
Machine-accessible metadata file describing the reported data: https://doi.org/10.6084/m9.figshare.16680787
Similar content being viewed by others
Background & Summary
Bond dissociation enthalpies (BDEs) are a central property in chemistry that have been studied for decades experimentally and computationally1,2,3,4. BDEs can be used to estimate the selectivity and reactivity of various molecules with free radicals (like ·OH, ·OOH, ·OR, ·OOR, ·NO, ·NO2, etc.) that are generated and transformed during chemical reactions relevant in chemistry and biology5,6,7,8,9,10. In this context, the calculation of BDEs for C-H, O-H, N-H, S-H, O-O, and S-S bonds in biologically relevant systems can help develop an understanding of the efficiency of antioxidants11,12,13. Furthermore, the calculation of BDEs is fundamental to develop a deeper understanding of various enzyme catalytic processes14,15,16 and surface functionalization chemistry17,18,19.
In 2012, Drew and Reynisson employed BDE calculations to predict the major metabolic sites of fifty known drug molecules20. Similarly, Andersson and co-workers applied BDE calculations to estimate the sensitivity of various drug candidates toward autoxidation21. The application of computed BDEs in these works shows how computational techniques can be incorporated into the risk assessment of drug products and guide further experimentation. Computationally obtained BDEs were also reported in different studies22,23,24, where the C-O and C-C BDEs were calculated for several substituted analogues of lignin, an abundant polymeric organic material and a potential renewable source of biofuels and chemicals22,23,24. The calculated BDEs were used to predict the homolytic dissociation of C-C and C-O bonds under thermal decomposition using model compounds representing the dominant linkages of lignin.
Given the importance of BDEs in many areas of chemistry and, consequently, the need to accurately predict bond energies computationally, a dataset of accurately predicted bond separation energies (BSEs) is developed here using an accurate computational chemistry method. Bond separation energies are a molecular property that can be computed in a straightforward manner in vacuum and provides direct information about the strength of a chemical bond. The BSEs presented in this work are differences between non-relativistic ground-state energies and contain no vibrational energy contributions, no zero-point energies, and no attempt has been made at thermally averaging over molecular conformations. As such, the reported BSEs are not comparable to experimental BDEs, but they serve as an ideal resource for developing and evaluating lower-cost computational chemistry methods used for a wide range of applications in chemistry and biology. Similar datasets to the one proposed in this work are available in the literature, but they tend to be small in terms of the total number of datapoints25, lack bond-type diversity26,27 or are calculated using less accurate computational chemistry methods compared to the one used in this work28,29,30. To the best of our knowledge, an accurate and extensive dataset of computationally predicted BSEs is not available in the literature. The main reason for this absence is that BSE calculations with high accuracy require computationally expensive methods that tend to scale poorly with system size.
This work addresses the aforementioned gap in the literature by constructing a large dataset (4502 datapoints) of computationally predicted BSEs of 49 unique bond types, all of which are determined with a high-level composite theoretical procedure denoted as (RO)CBS-QB331,32,33. This approach ensures uniform, high-quality reference data and eliminates the need to collect and verify data gathered from various sources, which may differ substantially in their accuracy. The (RO)CBS-QB3 method is known to produce BDEs of high accuracy8,33,34,35,36,37. Therefore, it is suitable for developing a database of BSEs that can be used to test and parametrize low-cost computational methods. One particular target application of our dataset is for the training of cost-effective computational approaches like atom-centered potentials38,39,40 (ACPs) or machine learning potentials28,29,30.
Methods
Dataset composition
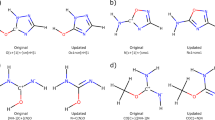
We present the BSE49 dataset, which comprises a broad range of bond separation energies for 49 unique bond types. The model systems present in the dataset are neutral molecules with X-H, X-F, X-Cl, X-X, and X-Y single bonds, where X and Y are B, C, N, O, Si, P, and S. The number of datapoints and the ranges of bond separation energies associated with each bond type are provided in Table 1. The structures of model systems on which the calculations were performed are divided into “Existing” and “Hypothetical” classes. The Existing type structures were built by selecting molecules with experimental data reported in the Comprehensive Handbook of Chemical Bond Dissociation Energies41. In contrast, the Hypothetical type structures were constructed by functional group substitutions of X-Y single bonds in order to include bond types that were not present in the handbook and to increase the diversity and number of datapoints for each bond type in the dataset. The candidate molecules for both Existing and Hypothetical subsets were generated using a partially automated computational workflow as described below.
Dataset generation
The calculated bond separation energies are defined as the negative of the difference in the ground-state electronic energies for the reaction
where A. and B. represent the two radical fragments formed by homolytically breaking the A-B covalent bond in vacuum. Based on this reaction, the equilibrium geometries of the parent molecules and their respective radical fragments are required to calculate the bond separation energies. The geometries of the parent molecule and the associated radicals were constructed manually for both Existing and Hypothetical subsets using the Avogadro42 program. The constructed geometries were then used as starting points for a conformer search. The CSD conformer generator43 and FullMonte44 codes were used to generate multiple conformers. The geometry of each conformer was relaxed to the corresponding local minimum using the Gaussian45 software package. This relaxation was carried out first by using a low-level method, combining the B3LYP46,47,48,49,50,51 density functional and 6-31G*52,53 basis set along with the D354,55,56 dispersion correction scheme using the Becke-Johnson57 damping (B3LYP-D3(BJ)/6-31G*). The optimized conformers were ranked using the B3LYP-D3(BJ)/6-31G* relative energies at the local minima. The ten lowest-energy conformers were then re-optimized at the higher-level CAM-B3LYP-D3(BJ)/def2-TZVP level of theory54,55,56,57,58,59. Range-separated functionals like CAM-B3LYP minimize the delocalization error, which could be important in the description of radical species60. The lowest-energy conformer obtained in this procedure was used for calculating the bond separation energies using the composite method described below. All calculations employed a default self-consistent field (SCF) convergence criterion of 10−8 Hartrees, ultrafine integration grid, and a tight optimization convergence criteria (maximum force = 1.5 × 10−5 Hartrees/Bohr, RMS force = 1 × 10−5 Hartrees/Bohr, maximum displacement = 6 × 10−5 Bohr, RMS displacement = 4 × 10−5 Bohr).
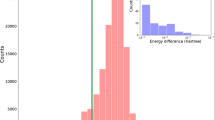
This partially automated workflow produced structures that are not necessarily the global minima. A visual inspection of the structures revealed that about 20% of the conformers generated do not correspond to the global minima, which reflects the difficulty of solving a global optimization problem (finding the most stable conformer) for such a large number of systems reliably. In addition, due to computational constraints, no attempt was made at evaluating the conformational energy landscape and statistically weighting the low-energy conformers associated with each molecule. Therefore, the dataset is not appropriate for direct comparison to bond separation energies obtained by back-correcting experimental BDEs, but it is suitable for testing and training computationally less expensive methods regarding their ability to accurately calculate the energy difference between the chosen conformers of products (A. and B.) and reactant (A-B).
The structures obtained from the workflow described above were then used for the final step of reference data calculation, using the composite (RO)CBS-QB331,32,33 method. The restricted-open-shell61 CBS-QB3 or ROCBS-QB3 was employed for the open-shell radical fragments, while restricted closed-shell calculations were performed for the closed-shell parent molecules with CBS-QB3. The composite (RO)CBS-QB3 method approximates energies at the complete-basis-set CCSD(T) level, using a series of computationally lower-cost methods including: (i) geometry optimization followed by vibration frequency calculation using the unrestricted-open-shell62 B3LYP/6-311G(2d,d,p) method46,47,48,49,50,51,63, (ii) ROMP2/6-311+G(3d2f,2df,2p) level63,64,65 energy extrapolated to the complete-basis-set limit, (iii) energy calculation at ROMP4(SDQ)/6-31+G(d(f),p) level63,64,66, and (iv) energy calculation at ROCCSD(T)/6-31+G† level63,64,67 (where 6-31+G† is a modified 6-31+G(d) basis set). Note that the final (RO)CBS-QB3 energy includes additional empirical correction terms described in Reference33. Structures were screened to remove any system for which the imaginary frequencies were obtained. The (RO)CBS-QB3 energies for the structures associated with a particular bond breaking reaction were used to obtain the bond separation energies for the dataset.
Data Records
The reference bond separation energies (in kcal/mol) and coordinates (in Å) of the structures presented in the BSE49 dataset are publicly available free-of-charge from the Figshare68 and GitHub (https://github.com/aoterodelaroza/bse49) repositories in the plain-text database file format (DB format) described in Table 2. The atomic coordinates of the model structures are stored in a plain-text XYZ format in the Geometries directory. The BSE49 dataset contains one DB format file and three XYZ format files for each bond separation energy. In total, deposited files include 4502 DB format files stored in the db-BSE49 directory and 13506 XYZ format files stored in their respective Existing or Hypothetical classification directories. Additional files labelled as BSE49_Existing.org and BSE49_Hypothetical.org are also provided. These files contain the necessary information about the reference data for all the model systems.
File format
For each molecule, the reference bond separation energy and the atomic coordinates are stored in a file named MoleculeName.db. The Cartesian coordinates of the atoms are stored in files called MoleculeName_AB.xyz, MoleculeName_A.xyz, and MoleculeName_B.xyz, where AB represents the parent molecule, A represents the first radical fragment, and B represents the second radical fragment.
The DB format file contains a header line specifying the reference energy value (in kcal/mol) followed by three ‘molc’ (short for molecule) blocks containing a unique integer identifier, charge, multiplicity, and the atomic coordinates (in Å) of the parent molecule and its corresponding radical fragments. The XYZ format file contains a header line defining the number of atoms N, a comment line containing the charge and multiplicity, and N lines with each containing element type and X, Y, Z coordinates (in Å). The BSE49_Existing.org and BSE49_Hypothetical.org files are special-character separated plain-text files (where the special character is ‘|’) containing multiple lines and eight columns. The columns are: (i) dataset name of the model system, (ii) unique integer identifier 1 indicating the A. fragment, (iii) geometry filename of the A. fragment, (iv) unique integer identifier 1 indicating the B. fragment, (v) geometry filename of the B. fragment, (vi) unique integer identifier -1 indicating the A-B model system, (vii) geometry filename of the A-B model system, and (viii) computational reference bond separation energy (in kcal/mol).
Technical Validation
For the generation of reference data, the reliable (RO)CBS-QB3 method was chosen for all the model systems considered in the BSE49 dataset. The (RO)CBS-QB3 method has been widely used in literature in recent years69,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90. The developers of the (RO)CBS-QB3 method reported that it predicts heats of formation at 298 K with a mean absolute deviation (MAD) from the experiment of 0.91 kcal/mol33. For bond dissociation enthalpies of eleven molecules with chemical structures typically found in amino acid sidechains, peptide termini, and peptide backbones, Moore et al. reported an MAD of 1.72 kcal/mol from the experimental values8. For small lignin model molecules, the CBS-QB3 approach was shown to yield bond dissociation enthalpies within 2.99 kcal/mol from experimental values34. (RO)CBS-QB3 has been used as a reference method for benchmarking various density functional theory methods to estimate bond dissociation enthalpies in a different study on small lignin model systems23. Hudzik and co-workers utilized the CBS-QB3 composite method to study the C-H bond separation energies of a few alkane molecules and reported a good agreement with literature values35. The (RO)CBS-QB3 has also been used for the prediction of bond dissociation enthalpies in a previous work by Menon et al.36 The MAD of (RO)CBS-QB3 was reported to be only 0.60 kcal/mol from the experiment and was suggested as being a reliable and efficient procedure for calculating bond separation energies in comparison to the other composite methods tested. In another work, bond dissociation enthalpies of 200 molecules were calculated using an earlier version of this work’s composite method, CBS-Q37. It was shown that the results of the CBS-Q composite procedure predicted bond dissociation enthalpies to within 2.39 kcal/mol of the reported experimental values. Collectively, these results support the selection of (RO)CBS-QB3 as a practical and accurate method for the generation of reference data in this work. Note that the reference bond separation energies reported in this work are non-relativistic (RO)CBS-QB3 energies without zero-point energy corrections. This makes the reference data suitable to support the development of low-cost computational chemistry methods like those described in references28,29,30,38,39,40.
Code availability
Throughout this work, the Gaussian software package was used for geometry optimizations, frequency calculations, and composite (RO)CBS-QB3 calculations. The Gaussian software package can be purchased from Gaussian Inc. (http://gaussian.com/) under a commercial license. CSD conformer generator was used for conformer generation. The CSD conformer generator can be purchased under a commercial license from https://www.ccdc.cam.ac.uk/solutions/csd-enterprise/applications/conformer-generator/. Fullmonte software package was also used along with MOPAC16 (PM6-DH2 method). Fullmonte software package can be downloaded free-of-cost from https://github.com/bobbypaton/FullMonte. Whereas MOPAC16 software package can be installed after acquiring a free license from http://openmopac.net/. The Avogadro molecular editor and visualizer is an open-source program available at https://avogadro.cc/.
References
Chan, B., Collins, E. & Raghavachari, K. Applications of isodesmic‐type reactions for computational thermochemistry. WIREs Comput. Mol. Sci. 11, e1501 (2020).
Johnson, E. R., Clarkin, O. J. & DiLabio, G. A. Density functional theory based model calculations for accurate bond dissociation enthalpies. 3. A single approach for X-H, X-X, and X-Y (X, Y = C, N, O, S, Halogen) bonds. J. Phys. Chem. A 107, 9953–9963 (2003).
DiLabio, G. A. & Pratt, D. A. Density functional theory based model calculations for accurate bond dissociation enthalpies. 2. Studies of X-X and X-Y (X, Y = C, N, O, S, Halogen) bonds. J. Phys. Chem. A 104, 1938–1943 (2000).
DiLabio, G. A., Pratt, D. A., LoFaro, A. D. & Wright, J. S. Theoretical study of X-H bond energetics (X = C, N, O, S): Application to substituent effects, gas phase acidities, and redox potentials. J. Phys. Chem. A 103, 1653–1661 (1999).
Hioe, J. & Zipse, H. Radical stability–Thermochemical aspects. In Encyclopedia of Radicals in Chemistry, Biology and Materials, eds. Chatgilialoglu, C. & Studer, A. (John Wiley & Sons, Ltd., 2012).
Zavitsas, A. A. Thermochemistry and hydrogen transfer kinetics. In Encyclopedia of Radicals in Chemistry, Biology and Materials, eds. Chatgilialoglu, C. & Studer, A. (John Wiley & Sons, Ltd., 2012).
Hioe, J., Mosch, M., Smith, D. M. & Zipse, H. Dissociation energies of Cα–H bonds in amino acids –a re-examination. RSC Adv. 3, 12403–12408 (2013).
Moore, B. N. & Julian, R. R. Dissociation energies of X–H bonds in amino acids. Phys. Chem. Chem. Phys. 14, 3148–3154 (2012).
Coote, M. L. & Zavitsas, A. A. Using inherent radical stabilization energies to predict unknown enthalpies of formation and associated bond dissociation energies of complex molecules. Tetrahedron 72, 7749–7756 (2016).
Adhikary, A., Kumar, A., Becker, D. & Sevilla, M. D. Understanding DNA radicals employing theory and electron spin resonance spectroscopy. In Encyclopedia of Radicals in Chemistry, Biology and Materials, eds. Chatgilialoglu, C. & Studer, A. (John Wiley & Sons, Ltd., 2012).
Wright, J. S., Johnson, E. R. & DiLabio, G. A. Predicting the activity of phenolic antioxidants: Theoretical method, analysis of substituent effects, and application to major families of antioxidants. J. Am. Chem. Soc. 123, 1173–1183 (2001).
Kancheva, V. D. et al. Antiradical and antioxidant activities of new bio-antioxidants. Biochimie 94, 403–415 (2012).
Bond dissociation energies and thermodynamic functions of antioxidants. In Handbook of Antioxidants: Bond Dissociation Energies, Rate Constants, Activation Energies, and Enthalpies of Reactions, Second Edition, eds. Denisov, E. T. & Denisova, T. (CRC Press LLC, 1999).
Jensen, K. P. & Ryde, U. How the Co-C bond is cleaved in coenzyme B12 enzymes: A theoretical study. J. Am. Chem. Soc. 127, 9117–9128 (2005).
Qu, Z. W., Hansen, A. & Grimme, S. Co-C bond dissociation energies in cobalamin derivatives and dispersion effects: Anomaly or just challenging? J. Chem. Theory Comput. 11, 1037–1045 (2015).
Kozlowski, P. M. et al. The cobalt-methyl bond dissociation in methylcobalamin: New benchmark analysis based on density functional theory and completely renormalized coupled-cluster calculations. J. Chem. Theory Comput. 8, 1870–1894 (2012).
Kruse, P., Johnson, E. R., DiLabio, G. A. & Wolkow, R. A. Patterning of vinylferrocene on H-Si(100) via self-directed growth of molecular lines and STM-induced decomposition. Nano Lett. 2, 807–810 (2002).
Tong, X., DiLabio, G. A. & Wolkow, R. A. A self-directed growth process for creating covalently bonded molecular assemblies on the H-Si(100)-3x1 surface. Nano Lett. 4, 979–983 (2004).
Piva, P. G. et al. Field regulation of single-molecule conductivity by a charged surface atom. Nature 435, 658–661 (2005).
Drew, K. L. M. & Reynisson, J. The impact of carbon–hydrogen bond dissociation energies on the prediction of the cytochrome P450 mediated major metabolic site of drug-like compounds. Eur. J. Med. Chem. 56, 48–55 (2012).
Andersson, T., Broo, A. & Evertsson, E. Prediction of drug candidates’ sensitivity toward autoxidation: Computational estimation of C-H dissociation energies of carbon-centered radicals. J. Pharm. Sci. 103, 1949–1955 (2014).
Parthasarathi, R., Romero, R. A., Redondo, A. & Gnanakaran, S. Theoretical study of the remarkably diverse linkages in lignin. J. Phys. Chem. Lett. 2, 2660–2666 (2011).
Kim, S. et al. Computational study of bond dissociation enthalpies for a large range of native and modified lignins. J. Phys. Chem. Lett. 2, 2846–2852 (2011).
Beste, A. & Buchanan, A. C. III Computational study of bond dissociation enthalpies for lignin model compounds. Substituent effects in phenethyl phenyl ethers. J. Org. Chem. 74, 2837–2841 (2009).
Chan, B. & Radom, L. BDE261: A comprehensive set of high-level theoretical bond dissociation enthalpies. J. Phys. Chem. A 116, 4975–4986 (2012).
Saito, T., Kambara, H. & Takano, Y. Quantitative assessment of reparameterized PM6 (rPM6) for hydrogen abstraction reactions. Mol. Phys. 118, e1700313 (2020).
Zhao, Y., Ng, H. T., Peverati, R. & Truhlar, D. G. Benchmark database for ylidic bond dissociation energies and its use for assessments of electronic structure methods. J. Chem. Theory Comput. 8, 2824–2834 (2012).
Wen, M., Blau, S. M., Spotte-Smith, E. W. C., Dwaraknath, S. & Persson, K. A. BonDNet: a graph neural network for the prediction of bond dissociation energies for charged molecules. Chem. Sci. 12, 1858–1868 (2021).
Qu, X., Latino, D. A. R. S. & Aires-De-sousa, J. A big data approach to the ultra-fast prediction of DFT-calculated bond energies. J. Cheminform. 5, 34 (2013).
St. John, P. C., Guan, Y., Kim, Y., Kim, S. & Paton, R. S. Prediction of organic homolytic bond dissociation enthalpies at near chemical accuracy with sub-second computational cost. Nat. Commun. 11, 2328 (2020).
Montgomery, J. A. Jr., Frisch, M. J., Ochterski, J. W. & Petersson, G. A. A complete basis set model chemistry. VI. Use of density functional geometries and frequencies. J. Chem. Phys. 110, 2822–2827 (1999).
Montgomery, J. A. Jr., Frisch, M. J., Ochterski, J. W. & Petersson, G. A. A complete basis set model chemistry. VII. Use of the minimum population localization method. J. Chem. Phys. 112, 6532–6542 (2000).
Wood, G. P. F. et al. A restricted-open-shell complete-basis-set model chemistry. J. Chem. Phys. 125, 094106 (2006).
Jarvis, M. W. et al. Direct detection of products from the pyrolysis of 2-phenethyl phenyl ether. J. Phys. Chem. A 115, 428–438 (2011).
Hudzik, J. M., Bozzelli, J. W. & Simmie, J. M. Thermochemistry of C7H16 to C10H22 alkane isomers: Primary, secondary, and tertiary C-H bond dissociation energies and effects of branching. J. Phys. Chem. A 118, 9364–9379 (2014).
Menon, A. S., Wood, G. P. F., Moran, D. & Radom, L. Bond dissociation energies and radical stabilization energies: An assessment of contemporary theoretical procedures. J. Phys. Chem. A 111, 13638–13644 (2007).
Feng, Y., Liu, L., Wang, J.-T., Huang, H. & Guo, Q.-X. Assessment of experimental bond dissociation energies using composite ab initio methods and evaluation of the performances of density functional methods in the calculation of bond dissociation energies. J. Chem. Inf. Comput. Sci. 43, 2005–2013 (2003).
Otero-de-la-Roza, A. & DiLabio, G. A. Improved basis-set incompleteness potentials for accurate density-functional theory calculations in large systems. J. Chem. Theory Comput. 16, 4176–4191 (2020).
Prasad, V. K., Otero-de-la-Roza, A. & DiLabio, G. A. Atom-centered potentials with dispersion-corrected minimal-basis-set Hartree–Fock: An efficient and accurate computational approach for large molecular systems. J. Chem. Theory Comput. 14, 726–738 (2018).
Otero-de-la-Roza, A. & DiLabio, G. A. Transferable atom-centered potentials for the correction of basis set incompleteness errors in density-functional theory. J. Chem. Theory Comput. 13, 3505–3524 (2017).
Comprehensive Handbook of Chemical Bond Dissociation Energies, ed. Luo, Y.-R. (Taylor & Francis Group, LLC, 2007).
Hanwell, M. D. et al. Avogadro: an advanced semantic chemical editor, visualization, and analysis platform. J. Cheminform. 4, 17 (2012).
https://www.ccdc.cam.ac.uk/solutions/csd-enterprise/applications/conformer-generator/.
Frisch, M. J. et al. Gaussian 16, Revision B.01 (Gaussian, Inc., Wallingford CT, 2016).
Becke, A. D. Density‐functional thermochemistry. III. The role of exact exchange. J. Chem. Phys. 98, 5648–5652 (1993).
Lee, C., Yang, W. & Parr, R. G. Development of the Colle-Salvetti correlation-energy formula into a functional of the electron density. Phys. Rev. B 37, 785–789 (1988).
Vosko, S. H., Wilk, L. & Nusair, M. Accurate spin-dependent electron liquid correlation energies for local spin density calculations: a critical analysis. Can. J. Phys. 58, 1200–1211 (1980).
Stephens, P. J., Devlin, F. J., Chabalowski, C. F. & Frisch, M. J. Ab Initio calculation of vibrational absorption and circular dichroism spectra using density functional force fields. J. Phys. Chem. 98, 11623–11627 (1994).
Becke, A. D. A new mixing of Hartree-Fock and local density-functional theories. J. Chem. Phys. 98, 1372–1377 (1993).
Becke, A. D. Density-functional exchange-energy approximation with correct asymptotic behavior. Phys. Rev. A 38, 3098–3100 (1988).
Hariharan, P. C. & Pople, J. A. The influence of polarization functions on molecular orbital hydrogenation energies. Theor. Chim. Acta 28, 213–222 (1973).
Francl, M. M. et al. Self‐consistent molecular orbital methods. XXIII. A polarization‐type basis set for second‐row elements. J. Chem. Phys. 77, 3654–3665 (1982).
Goerigk, L. A comprehensive overview of the DFT-D3 London-dispersion correction. In Non-Covalent Interactions in Quantum Chemistry and Physics: Theory and Applications, eds. Otero-de-la-Roza & DiLabio G. A. (Elsevier Inc., 2017).
Grimme, S., Antony, J., Ehrlich, S. & Krieg, H. A consistent and accurate ab initio parametrization of density functional dispersion correction (DFT-D) for the 94 elements H-Pu. J. Chem. Phys. 132, 154104 (2010).
Grimme, S., Ehrlich, S. & Goerigk, L. Effect of the damping function in dispersion corrected density functional theory. J. Comput. Chem. 32, 1456–1465 (2011).
Johnson, E. R. & Becke, A. D. A post-Hartree-Fock model of intermolecular interactions: Inclusion of higher-order corrections. J. Chem. Phys. 124, 174104 (2006).
Yanai, T., Tew, D. P. & Handy, N. C. A new hybrid exchange–correlation functional using the Coulomb-attenuating method (CAM-B3LYP). Chem. Phys. Lett. 393, 51–57 (2004).
Schäfer, A., Huber, C. & Ahlrichs, R. Fully optimized contracted Gaussian basis sets of triple zeta valence quality for atoms Li to Kr. J. Chem. Phys. 100, 5829–5835 (1994).
Johnson, E. R., Otero-de-la-Roza, A. & Dale, S. G. Extreme density-driven delocalization error for a model solvated-electron system. J. Chem. Phys. 139, 184116 (2013).
Roothaan, C. C. J. New developments in molecular orbital theory. Rev. Mod. Phys. 23, 69–89 (1951).
Pople, J. A. & Nesbet, R. K. Self-consistent orbitals for radicals. J. Chem. Phys. 22, 571–572 (1954).
AB INITIO Molecular Orbital Theory, eds. Hehre, W. J., Radom, L., Schleyer, P. V. R. & Pople, J. (Wiley, 1986)
Lauderdale, W. J., Stanton, J. F., Gauss, J., Watts, J. D. & Bartlett, R. J. Many-body perturbation theory with a restricted open-shell Hartree-Fock reference. Chem. Phys. Lett. 187, 21–28 (1991).
Knowles, P. J., Andrews, J. S., Amos, R. D., Handy, N. C. & Pople, J. A. Restricted Møller-Plesset theory for open-shell molecules. Chem. Phys. Lett. 186, 130–136 (1991).
Lauderdale, W. J., Stanton, J. F., Gauss, J., Watts, J. D. & Bartlett, R. J. Restricted open-shell Hartree-Fock-based many-body perturbation theory: Theory and application of energy and gradient calculations. J. Chem. Phys. 97, 6606–6620 (1992).
Bartlett, R. J. Coupled-cluster theory: An overview of recent developments. In Mordern Electronic Structure Theory Part II, ed. Yarkony, D. R. (World Scientific, 1995).
Prasad, V. K., Khalilian, H., Otero-de-la-Roza, A. & DiLabio, G. A. BSE49, a diverse, high-quality benchmark dataset of separation energies of chemical bonds. figshare https://doi.org/10.6084/m9.figshare.14544060.v1 (2021).
Abdel-Rahman, M. A. et al. Mechanistic insights of the degradation of an O-anisidine carcinogenic pollutant initiated by OH radical attack: theoretical investigations. New J. Chem. 45, 5907–5924 (2021).
Nguyen, T. D.-T., Pham, N., Mai, T. V.-T., Nguyen, H. M. & Huynh, L. K. Detailed kinetic mechanism of thermal decomposition of furyl radicals: Theoretical insights. Fuel 288, 119699 (2021).
Chen, Y. F. et al. Insights into evolution mechanism of PAHs in coal thermal conversion: A combined experimental and DFT study. Energy 222, 119970 (2021).
Wang, Q.-D., Sun, M.-M. & Liang, J. Theoretical study of the hydrogen abstraction reactions from substituted phenolic species. Comput. Theor. Chem. 1196, 113120 (2021).
Liao, Z., Zeng, M. & Wang, L. Atmospheric oxidation mechansim of polychlorinated biphenyls (PCBs) initiated by OH radicals. Chemosphere 240, 124756 (2020).
Wang, L. & Wang, L. Atmospheric oxidation mechanism of acenaphthene initiated by OH radicals. Atmos. Environ. 243, 117870 (2020).
Sun, W., Hamadi, A., Abid, S., Chaumeix, N. & Comandini, A. An experimental and kinetic modeling study of phenylacetylene decomposition and the reactions with acetylene/ethylene under shock tube pyrolysis conditions. Combust. Flame 220, 257–271 (2020).
Zeng, M., Liao, Z. & Wang, L. Atmospheric oxidation of gaseous anthracene and phenanthrene initiated by OH radicals. Atmos. Environ. 234, 117587 (2020).
Wang, S. et al. Aromatic photo-oxidation, a new source of atmospheric acidity. Environ. Sci. Technol. 54, 7798–7806 (2020).
Abdel-Rahman, M. A., Shibl, M. F., El-Demerdash, S. H. & El-Nahas, A. M. Simulated kinetics of the atmospheric removal of aniline during daytime. Chemosphere 255, 127031 (2020).
Khojandi, M., Seif, A., Zahedi, E., Domingo, L. R. & Karimkhani, M. Unravelling the kinetics and molecular mechanism of the degenerate Cope rearrangement of bullvalene. New J. Chem. 44, 6543–6552 (2020).
Padash, R. Mechanism and kinetic investigations of 5-fluorouracil tautomeric conversions in the gas phase: DFT and CBS-QB3 methods using multichannel Rice–Ramsperger–Kassel–Marcus steady-state approximation theory. Theor. Chem. Acc. 139, 1–9 (2020).
Doroshenko, I., Vaskivskyi, Y. & Chernolevska, Y. Structural transformations in solid and liquid n-butanol from FTIR spectroscopy. Mol. Cryst. Liq. Cryst. 697, 11–19 (2020).
Ning, H., Wu, J., Ma, L. & Ren, W. Exploring the pyrolysis chemistry of prototype aromatic ester phenyl formate: Reaction pathways, thermodynamics and kinetics. Combust. Flame 211, 337–346 (2020).
Poskrebyshev, G. A. The CBS values of ΔfH°298.15 and S°298.15 of the phenoxy radicals, formed by abstraction of H atom from the components of surrogate bio-oil. Comput. Theor. Chem. 1169, 112625 (2019).
Abdel-Rahman, M. A., Shibl, M. F., El-Demerdash, S. H. & El-Nahas, A. M. First-principle studies on the gas phase OH-initiated oxidation of O-toluidine. Comput. Theor. Chem. 1170, 112634 (2019).
Wagner, J. P. Gauging stability and reactivity of carbonyl: O-oxide Criegee intermediates. Phys. Chem. Chem. Phys. 21, 21530–21540 (2019).
Bietti, M. et al. Evaluation of polar effects in hydrogen atom transfer reactions from activated phenols. J. Org. Chem. 84, 1778–1786 (2019).
Vaskivskyi, Y. et al. 1-Hexanol conformers in a nitrogen matrix: FTIR study and high-level ab initio calculations. J. Mol. Liq. 278, 356–362 (2019).
Bain, M., Hansen, C. S., Karsili, T. N. V. & Ashfold, M. N. R. Quantifying rival bond fission probabilities following photoexcitation: C-S bond fission in t-butylmethylsulfide. Chem. Sci. 10, 5290–5298 (2019).
Poskrebyshev, G. A. The values of ΔfH°298.18(AlnOmHp), calculated using the CBS correction dependencies, as well as the thermochemistry of the isodesmic/homodesmotic reactions. Comput. Theor. Chem. 1164, 112540 (2019).
DiLabio, G. A. et al. Hydrogen atom transfer (HAT) processes promoted by the Quinolinimide-N-oxyl radical. A kinetic and theoretical study. J. Org. Chem. 82, 6133–6141 (2017).
Acknowledgements
GAD would like to thank the Natural Sciences and Engineering Research Council of Canada, the Canadian Foundation for Innovation, and the British Columbia Knowledge Development Fund for financial support. We are grateful to Compute Canada, Westgrid, and the University of British Columbia’s advanced research computing platforms for the generous allocation of computing resources. AOR thanks the Spanish Ministerio de Ciencia e Innovación and the Agencia Estatal de Investigación (AEI) for financial support (projects PGC2018-097520-A-100 and RED2018-102612-T), and the Spanish MINECO for a Ramón y Cajal fellowship (RyC-2016-20301).
Author information
Authors and Affiliations
Contributions
V.K.P. performed the selection, calculation, curation, and validation associated with the generation of the BSE49 dataset. M.H.K. assisted in the curation and cross-validation of results. A.O.R. and G.A.D. designed the idea of the study and co-supervised this work. V.K.P., M.H.K., A.O.R. and G.A.D. co-wrote the data descriptor.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files associated with this article.
About this article
Cite this article
Prasad, V.K., Khalilian, M.H., Otero-de-la-Roza, A. et al. BSE49, a diverse, high-quality benchmark dataset of separation energies of chemical bonds. Sci Data 8, 300 (2021). https://doi.org/10.1038/s41597-021-01088-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-021-01088-2