Abstract
Pre-mRNA splicing by the spliceosome requires the biogenesis and recycling of its small nuclear ribonucleoprotein (snRNP) complexes, which are consumed in each round of splicing. The human U5 snRNP is the ~1 MDa ‘heart’ of the spliceosome and is recycled through an unknown mechanism involving major architectural rearrangements and the dedicated chaperones CD2BP2 and TSSC4. Late steps in U5 snRNP biogenesis similarly involve these chaperones. Here we report cryo-electron microscopy structures of four human U5 snRNP–CD2BP2–TSSC4 complexes, revealing how a series of molecular events primes the U5 snRNP to generate the ~2 MDa U4/U6.U5 tri-snRNP, the largest building block of the spliceosome.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The 3D cryo-EM density maps 1, 2, 3, 4, 5 and 6 of the U5 snRNP have been deposited into the Electron Microscopy Data Bank under the accession numbers EMD-18234, EMD-18229, EMD-18235, EMD-18237, EMD-18239 and EMD-18238. The coordinate file of the U5 snRNP in states I, II, III and IV have been deposited into Protein Data Bank (PDB) under the accession numbers 8Q7V, 8Q7Q, 8Q7W and 8Q7X. Source data are provided with this paper.
References
Kastner, B., Will, C. L., Stark, H. & Lührmann, R. Structural insights into nuclear pre-mRNA splicing in higher eukaryotes. Cold Spring Harb. Perspect. Biol. 11, a032417 (2019).
Wilkinson, M. E., Charenton, C. & Nagai, K. RNA splicing by the spliceosome. Annu. Rev. Biochem. 89, 359–388 (2020).
Wan, R., Bai, R., Zhan, X. & Shi, Y. How is precursor messenger RNA spliced by the spliceosome? Annu. Rev. Biochem. 89, 333–358 (2020).
Will, C. L. & Lührmann, R. Spliceosome structure and function. Cold Spring Harb. Perspect. Biol. 3, a003707 (2011).
Vorländer, M. K., Pacheco-Fiallos, B. & Plaschka, C. Structural basis of mRNA maturation: time to put it together. Curr. Opin. Struct. Biol. 75, 102431 (2022).
Grainger, R. J. & Beggs, J. D. Prp8 protein: at the heart of the spliceosome. RNA 11, 533–557 (2005).
Golas, M. M. et al. 3D cryo-EM structure of an active step I spliceosome and localization of its catalytic core. Mol. Cell 40, 927–938 (2010).
Bach, M. & Lührmann, R. Protein–RNA interactions in 20S U5 snRNPs. Biochim. Biophys. Acta 1088, 139–143 (1991).
Laggerbauer, B. et al. The human U5 snRNP 52K protein (CD2BP2) interacts with U5-102K (hPrp6), a U4/U6.U5 tri-snRNP bridging protein, but dissociates upon tri-snRNP formation. RNA 11, 598–608 (2005).
Liu, Y.-C. & Cheng, S.-C. Functional roles of DExD/H-box RNA helicases in pre-mRNA splicing. J. Biomed. Sci. 22, 54 (2015).
Plaschka, C., Newman, A. J. & Nagai, K. Structural basis of nuclear pre-mRNA splicing: lessons from yeast. Cold Spring Harb. Perspect. Biol. 11, a032391 (2019).
Bialkowska, A. & Kurlandzka, A. Proteins interacting with Lin 1p, a putative link between chromosome segregation, mRNA splicing and DNA replication in Saccharomyces cerevisiae. Yeast 19, 1323–1333 (2002).
Klimešová, K. et al. TSSC4 is a component of U5 snRNP that promotes tri-snRNP formation. Nat. Commun. 12, 3646 (2021).
Bergfort, A. et al. The intrinsically disordered TSSC4 protein acts as a helicase inhibitor, placeholder and multi-interaction coordinator during snRNP assembly and recycling. Nucleic Acids Res. 50, 2938–2958 (2022).
Boon, K. L. et al. prp8 mutations that cause human retinitis pigmentosa lead to a U5 snRNP maturation defect in yeast. Nat. Struct. Mol. Biol. 14, 1077–1083 (2007).
Weber, G. et al. Structural basis for dual roles of Aar2p in U5 snRNP assembly. Genes Dev. 27, 525–540 (2013).
Laggerbauer, B., Achsel, T. & Lührmann, R. The human U5-200kD DEXH-box protein unwinds U4/U6 RNA duplices in vitro. Proc. Natl Acad. Sci. USA 95, 4188–4192 (1998).
Plaschka, C., Lin, P.-C., Charenton, C. & Nagai, K. Prespliceosome structure provides insights into spliceosome assembly and regulation. Nature 559, 419–422 (2018).
Charenton, C., Wilkinson, M. E. & Nagai, K. Mechanism of 5′ splice site transfer for human spliceosome activation. Science 364, 362–367 (2019).
Zhan, X., Yan, C., Zhang, X., Lei, J. & Shi, Y. Structures of the human pre-catalytic spliceosome and its precursor spliceosome. Cell Res. 28, 1129–1140 (2018).
Agafonov, D. E. et al. Molecular architecture of the human U4/U6.U5 tri-snRNP. Science 351, 1416–1420 (2016).
Bell, M., Schreiner, S., Damianov, A., Reddy, R. & Bindereif, A. p110, a novel human U6 snRNP protein and U4/U6 snRNP recycling factor. EMBO J. 21, 2724–2735 (2002).
Nielsen, T. K., Liu, S., Lührmann, R. & Ficner, R. Structural basis for the bifunctionality of the U5 snRNP 52K protein (CD2BP2). J. Mol. Biol. 369, 902–908 (2007).
Galej, W. P., Oubridge, C., Newman, A. J. & Nagai, K. Crystal structure of Prp8 reveals active site cavity of the spliceosome. Nature 493, 638–643 (2013).
Jehle, S. et al. A human kinase yeast array for the identification of kinases modulating phosphorylation-dependent protein–protein interactions. Mol. Syst. Biol. 18, e10820 (2022).
Weber, G. et al. Mechanism for Aar2p function as a U5 snRNP assembly factor. Genes Dev. 25, 1601–1612 (2011).
Liu, S., Rauhut, R., Vornlocher, H. P. & Lührmann, R. The network of protein-protein interactions within the human U4/U6.U5 tri-snRNP. Rna 12, 1418–1430 (2006).
Schreib, C. C. et al. Functional and biochemical characterization of Dib1’s role in pre-messenger RNA splicing. J. Mol. Biol. 430, 1640–1651 (2018).
Mayeda, A. & Krainer, A. R. Preparation of HeLa cell nuclear and cytosolic S100 extracts for in vitro splicing. Methods Mol. Biol. 118, 309–314 (1999).
Kastner, B. et al. GraFix: sample preparation for single-particle electron cryomicroscopy. Nat. Methods 5, 53–55 (2008).
Tegunov, D. & Cramer, P. Real-time cryo-electron microscopy data preprocessing with Warp. Nat. Methods 16, 1146–1152 (2019).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Jamali, K., Kimanius, D. & Scheres, S. ModelAngelo: automated model building in cryo-EM maps. Preprint at bioRxiv https://doi.org/10.1101/2023.05.16.541002 (2023).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Evans, R. et al. Protein complex prediction with AlphaFold-Multimer. Preprint at bioRxiv https://doi.org/10.1101/2021.10.04.463034 (2022).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Kidmose, R. T. et al. Namdinator—automatic molecular dynamics flexible fitting of structural models into cryo-EM and crystallography experimental maps. IUCrJ 6, 526–531 (2019).
Croll, T. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. D 74, 519–530 (2018).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Behrens, S.-E. & Lührmann, R. Immunoaffinity purification of a [U4/U6.U5] tri-snRNP from human cells. Genes Dev. 5, 1439–1452 (1991).
Acknowledgements
We thank members of the Plaschka Group for their help and discussions; staff at the Protein Technologies Facility at the Vienna BioCenter Core Facilities (VBCF), a member of the Vienna BioCenter (VBC), for assistance with protein production; staff at the VBCF Electron Microscopy Facility, in particular T. Heuser and H. Kotisch, for support, data collection and maintaining facilities; staff at the IMP‐IMBA‐GMI BioOptics facility for performing FACS; R. Zimmermann and his team for computational support; K. Mechtler and his team for mass spectrometry; staff at the in-house Molecular Biology Service for reagents; C. Bernecky for discussions; C. Bernecky, E. Bassat and members of the Plaschka group for critical reading of the manuscript. D.R.B. was supported by a Marie Sklodowska-Curie fellowship (101028744). M.K.V. was supported by an EMBO Postdoctoral Fellowship. Research in the laboratory of C.P. is supported by Boehringer Ingelheim, the European Research Council under the Horizon 2020 research and innovation programme (ERC-2020-STG 949081 RNApaxport) and by the Austrian Science Fund (FWF) doc.funds program DOC177-B (RNA@core: Molecular mechanisms in RNA biology). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. For the purpose of open access, the author has applied for a CC BY public copyright license to any author accepted manuscript version arising from this submission.
Author information
Authors and Affiliations
Contributions
D.R.B. generated the GFP-3C-CD2BP2 K562 cell line, and with A.W.P. grew cells and prepared nuclear extract. S.V., D.R.B. and L.F. purified endogenous complexes and performed biochemical experiments. S.V. and M.K.V. collected cryo-EM data. D.R.B., S.V., M.K.V. and C.P. analyzed the cryo-EM data and modeled the U5 snRNP structure. D.R.B. and C.P. analyzed the data and prepared the manuscript with input from all authors. C.P. designed and supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Sebastian Klinge and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Dimitris Typas was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Purification and cryo-EM imaging of the human U5 snRNP complexes.
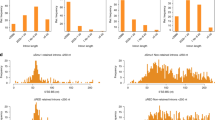
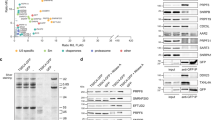
a. Endogenous CD2BP2 is detected by western blot in nuclear and cytoplasmic extracts (NE and CE) from K562 wild-type or K562 GFP-CD2BP2 overexpressing cells (representative image, n = 4). Note that the western blot band for CD2BP2 migrates at a higher molecular weight in CE compared to NE fractions, indicating that post-translational modifications could regulate CD2BP2 activity. Consistent with this, CD2BP2 has been reported to be phosphorylated, which may influence its interactions with the U5 snRNP41. b. GFP-CD2BP2 purifications from nuclear or cytoplasmic extracts (NE and CE) yield U5 snRNPs of similar protein composition (n = 2). c. Purification scheme (top) and SDS-PAGE analysis (bottom) of the human U5 snRNP complexes from K562 cells. The most abundant proteins in the samples are annotated according to mass spectrometry analysis of SDS-PAGE gel slices. The asterisk annotates contaminants (n = 9). d. Denoised cryo-EM micrographs of U5 snRNP complexes. The dataset contained 21,572 micrographs. Scale bar, 300 Å (n = 1). e. Cryo-EM 2D class averages of U5 snRNP complexes in State I, II, III, IV. The particles subsets for the shown States I-IV 2D class averages were obtained from U5 snRNP image processing shown in Extended Data Fig. 2. Scale bar, 100 Å.
Extended Data Fig. 2 Cryo-EM image processing of the human U5 snRNP complexes.
a. Three-dimensional image classification tree of the U5 snRNP cryo-EM data. The dataset contained 21,574 micrographs from which 788,114 particles were picked and extracted. An initial volume was generated from 57,138 particles in cryoSPARC33 using the ab-initio reconstruction algorithm, which served as a reference volume to classify the entire dataset using two rounds of heterogenous classification (see Methods). The cleaned U5 snRNP particle stack contained 256,427 particles and was refined to an overall resolution of 2.6 Å. Focused three-dimensional classifications and subsequent refinements, yielded six cryo-EM densities, labeled 1–6, in resolutions varying from 2.6 to 4.6 Å. b. Gold-standard Fourier Shell Correlation (FSC = 0.143) of U5 snRNP cryo-EM maps 1–6. c. Orientation distribution plots for all particles contributing to the respective U5 snRNP cryo-EM maps 1–6. d. Composite U5 snRNP cryo-EM maps 1–6 shown from the front view, colored by local resolution as determined by cryoSPARC33. e. As panel d. but the maps are colored by U5 snRNP subunits as in Fig. 1. f. Gallery of cryo-EM densities from the U5 snRNP subunits CD2BP2 (map 5), TSSC4 (map 5), SNU114 (map 6) and U5 snRNA (map 6), superimposed on the final coordinate models.
Extended Data Fig. 3 Structures of the human U5 snRNP State II and IV.
a, b. Back views of the final U5 snRNP coordinate models of States II (panel a) and IV (panel b), highlighting the differences PRP8–DDX23 interactions in State IV (and III) but absent from States II (and I). Colors as in Fig. 1. Transparent models indicate regions not observed in our densities.
Extended Data Fig. 4 Structural comparisons of U5 snRNP to U4/U6.U5 tri-snRNP and CD2BP2–DIM1 structures.
a. Comparison of U5 snRNP State II (this study) and U4/U6.U5 tri-snRNP structures (PDB ID 6QW6). For the U5 snRNP (left), PRP8N and PRP8L, U5 snRNA, TSSC4 and PRP6 are shown alone for clarity. For the U4/U6.U5 tri-snRNP (center) the same regions are shown, in addition to the PRP8-interaction segments of U4/U6 snRNAs, of PRP31, and complete DIM1. The structures were aligned on their PRP8L domains (right), with the U4/U6.U5 tri-snRNP rendered transparent. Black arrows indicate movements within U5 snRNP compared to the equivalent U4/U6.U5 tri-snRNP regions. b. As in panel a, the structures were aligned on the PRP8L domain (right), to visualize the clash of the CD2BP2 ‘Hook-N’ (U5 snRNP, left) with PRP8α-finger–PRP31 interfaces and with DIM1 in the U4/U6.U5 tri-snRNP (center). c. The CD2BP2 ‘GYF’ domain from the CD2BP2 GYF–DIM1 crystal structure (PDB ID 1SYX, center) clashes with PRP8L–PRP31 interfaces in the U4/U6.U5 tri-snRNP structure (PDB ID 6QW6, right). The CD2BP2 GYF–DIM1 crystal structure was aligned with the U4/U6.U5 tri-snRNP on DIM1 (right). d. As in panel a, the structures were aligned on the PRP8L domain (right), to visualize the clash between the CD2BP2 ‘Hook’ (U5 snRNP, left) and the extended PRP8 α-finger (U4/U6.U5 tri-snRNP, center). e. As in panel a, but the structures were aligned on the PRP8N domain (right) to visualize the clash between the CD2BP2 ‘Hook-C’ (U5 snRNP, left) and the PRP6 N-terminus (residues 72–99; U4/U6.U5 tri-snRNP, center). f. As in panel a, but the structures were aligned on the PRP8RT domain (right) to visualize the clash between TSSC4 (U5 snRNP, left) and the PRP6 N-terminus (residues 236–281; U4/U6.U5 tri-snRNP, center).
Extended Data Fig. 5 Structural comparison of the human U5 snRNP to the yeast Prp8-Aar2 structure.
a. Comparison of PRP8L domains in the human U5 snRNP (State I, left) and the yeast Saccharomyces cerevisiae Prp8-Aar2 crystal structures24 (PDB ID 4I43, right). The PRP8 α-finger is highlighted with a black outline. Colors as in Fig. 1; Aar2 (light pink). b. Comparison of PRP8 α-finger conformations in the human U5 snRNP (State I, left) and the yeast Prp8-Aar2 crystal structures24 (PDB ID 4I43, right).
Extended Data Fig. 6 Fourier shell correlations between U5 snRNP cryo-EM maps and respective coordinate models.
Fourier shell correlations between the cryo-EM maps 1, 2, 3, 4 with the respective refined coordinate models of U5 snRNP State I, II, III, and IV using phenix.mtriage.
Supplementary information
Supplementary Table 1
Mass spectrometry of human U5 snRNP complexes.
Source data
Source Data Fig. 1
Uncropped blots and gels as shown in Extended Data Fig. 1.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Riabov Bassat, D., Visanpattanasin, S., Vorländer, M.K. et al. Structural basis of human U5 snRNP late biogenesis and recycling. Nat Struct Mol Biol (2024). https://doi.org/10.1038/s41594-024-01243-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41594-024-01243-4