Abstract
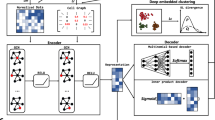
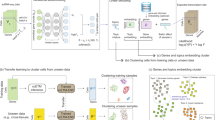
Single-cell proteomics sequencing technology sheds light on protein–protein interactions, posttranslational modifications and proteoform dynamics in the cell. However, the uncertainty estimation for peptide quantification, data missingness, batch effects and high noise hinder the analysis of single-cell proteomic data. It is important to solve this set of tangled problems together, but the existing methods tailored for single-cell transcriptomes cannot fully address this task. Here we propose a versatile framework designed for single-cell proteomics data analysis called scPROTEIN, which consists of peptide uncertainty estimation based on a multitask heteroscedastic regression model and cell embedding generation based on graph contrastive learning. scPROTEIN can estimate the uncertainty of peptide quantification, denoise protein data, remove batch effects and encode single-cell proteomic-specific embeddings in a unified framework. We demonstrate that scPROTEIN is efficient for cell clustering, batch correction, cell type annotation, clinical analysis and spatially resolved proteomic data exploration.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data used in this study are publicly available, and their usages are fully illustrated in Methods. The SCoPE2_Specht25 dataset was downloaded from ref. 48. The nanoPOTS dataset11 was downloaded at MassIVE data repository with ID MSV000084110. The N2 dataset12 was downloaded from MassIVE data repository with ID MSV000086809. The SCoPE2_Leduc dataset3 was downloaded from ref. 49. The plexDIA dataset5 was downloaded from ref. 50. The pSCoPE_Huffman dataset15 was downloaded from ref. 51 (derived from their original ‘Benchmarking experiments: Fig. 1b,e data’). The pSCoPE_Leduc dataset3 was downloaded from ref. 52. The ECCITE-seq dataset29 was downloaded from Gene Expression Omnibus with accession number GSE126310. The BaselTMA dataset30 was downloaded from Zenodo53. The T-SCP dataset24 was downloaded from the PRIDE partner repository (accession no. PXD024043). Source data are provided with this paper.
Code availability
The codes were implemented in Python and are released at GitHub (https://github.com/TencentAILabHealthcare/scPROTEIN) and Zenodo (https://doi.org/10.5281/zenodo.10547614)68 with detailed instructions.
References
Svensson, V., Vento-Tormo, R. & Teichmann, S. A. Exponential scaling of single-cell RNA-seq in the past decade. Nat. Protoc. 13, 599–604 (2018).
Slavov, N. Unpicking the proteome in single cells. Science 367, 512–513 (2020).
Leduc, A., Huffman, R. G., Cantlon, J., Khan, S. & Slavov, N. Exploring functional protein covariation across single cells using nPOP. Genome Biol. 23, 261 (2022).
Petelski, A. A. et al. Multiplexed single-cell proteomics using SCoPE2. Nat. Protoc. 16, 5398–5425 (2021).
Derks, J. et al. Increasing the throughput of sensitive proteomics by plexDIA. Nat. Biotechnol. 41, 50–59 (2023).
Doerr, A. Single-cell proteomics. Nat. Methods 16, 20 (2019).
Marx, V. A dream of single-cell proteomics. Nat. Methods 16, 809–812 (2019).
Perkel, J. M. Single-cell proteomics takes centre stage. Nature 597, 580–582 (2021).
Schoof, E. M. et al. Quantitative single-cell proteomics as a tool to characterize cellular hierarchies. Nat. Commun. 12, 3341 (2021).
Furtwängler, B. et al. Real-time search-assisted acquisition on a tribrid mass spectrometer improves coverage in multiplexed single-cell proteomics. Mol. Cell. Proteomics 21, 100219 (2022).
Dou, M. et al. High-throughput single cell proteomics enabled by multiplex isobaric labeling in a nanodroplet sample preparation platform. Anal. Chem. 91, 13119–13127 (2019).
Woo, J. et al. High-throughput and high-efficiency sample preparation for single-cell proteomics using a nested nanowell chip. Nat. Commun. 12, 6246 (2021).
Gatto, L. et al. Initial recommendations for performing, benchmarking and reporting single-cell proteomics experiments. Nat. Methods 20, 375–386 (2023).
Bennett, H. M., Stephenson, W., Rose, C. M. & Darmanis, S. Single-cell proteomics enabled by next-generation sequencing or mass spectrometry. Nat. Methods 20, 363–374 (2023).
Huffman, R. G. et al. Prioritized mass spectrometry increases the depth, sensitivity and data completeness of single-cell proteomics. Nat. Methods 20, 714–722 (2023).
Khan, Z. et al. Primate transcript and protein expression levels evolve under compensatory selection pressures. Science 342, 1100–1104 (2013).
Vogel, C. & Marcotte, E. M. Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nat. Rev. Genet. 13, 227–232 (2012).
Gygi, S. P., Rochon, Y., Franza, B. R. & Aebersold, R. Correlation between protein and mRNA abundance in yeast. Mol. Cell Biol. 19, 1720–1730 (1999).
Marguerat, S. et al. Quantitative analysis of fission yeast transcriptomes and proteomes in proliferating and quiescent cells. Cell 151, 671–683 (2012).
Irish, J. M., Kotecha, N. & Nolan, G. P. Mapping normal and cancer cell signalling networks: towards single-cell proteomics. Nat. Rev. Cancer 6, 146–155 (2006).
Vanderaa, C. & Gatto, L. Replication of single-cell proteomics data reveals important computational challenges. Expert Rev. Proteomics 18, 835–843 (2021).
Cheung, T. K. et al. Defining the carrier proteome limit for single-cell proteomics. Nat. Methods 18, 76–83 (2020).
Mund, A. et al. Deep Visual Proteomics defines single-cell identity and heterogeneity. Nat. Biotechnol. 40, 1231–1240 (2022).
Brunner, A.-D. et al. Ultra-high sensitivity mass spectrometry quantifies single-cell proteome changes upon perturbation. Mol. Syst. Biol. 18, e10798 (2022).
Specht, H. et al. Single-cell proteomic and transcriptomic analysis of macrophage heterogeneity using SCoPE2. Genome Biol. 22, 50 (2021).
Sticker, A., Goeminne, L., Martens, L. & Clement, L. Robust summarization and inference in proteome-wide label-free quantification. Mol. Cell. Proteomics 19, 1209–1219 (2020).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell. Proteomics 13, 2513–2526 (2014).
Kendall, A. & Gal, Y. What uncertainties do we need in Bayesian deep learning for computer vision? Adv. Neural Inf. Process. Syst. 30, 5580–5590 (2017).
Mimitou, E. P. et al. Multiplexed detection of proteins, transcriptomes, clonotypes and CRISPR perturbations in single cells. Nat. Methods 16, 409–412 (2019).
Jackson, H. W. et al. The single-cell pathology landscape of breast cancer. Nature 578, 615–620 (2020).
van Dijk, D. et al. Recovering gene interactions from single-cell data using data diffusion. Cell 174, 716–729 (2018).
Li, H., Brouwer, C. R. & Luo, W. A universal deep neural network for in-depth cleaning of single-cell RNA-seq data. Nat. Commun. 13, 1901 (2022).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Hie, B., Bryson, B. & Berger, B. Efficient integration of heterogeneous single-cell transcriptomes using Scanorama. Nat. Biotechnol. 37, 685–691 (2019).
Welch, J. D. et al. Single-cell multi-omic integration compares and contrasts features of brain cell identity. Cell 177, 1873–1887 (2019).
Boekweg, H. et al. Features of peptide fragmentation spectra in single-cell proteomics. J. Proteome Res. 21, 182–188 (2022).
Samimi, S. et al. Increased programmed death-1 expression on CD4+ T cells in cutaneous T-cell lymphoma: implications for immune suppression. Arch. Dermatol. 146, 1382–1388 (2010).
Keren, L. et al. A structured tumor-immune microenvironment in triple negative breast cancer revealed by multiplexed ion beam imaging. Cell 174, 1373–1387 (2018).
Zhu, Y. et al. Deep graph contrastive representation learning. in ICML Workshop on Graph Representation Learning and Beyond (2020).
Rong, Y., Huang, W., Xu, T. & Huang, J. DropEdge: towards deep graph convolutional networks on node classification. in International Conference on Learning Representations (2020).
Kipf, T. N. & Welling, M. Semi-supervised classification with graph convolutional networks. in International Conference on Learning Representations (2017).
Tschannen, M., Djolonga, J., Rubenstein, P. K., Gelly, S. & Lucic, M. On mutual information maximization for representation learning. in International Conference on Learning Representations (2019).
Chen, T., Kornblith, S., Norouzi, M. & Hinton, G. A simple framework for contrastive learning of visual representations. PMLR https://proceedings.mlr.press/v119/chen20j.html (2020).
van den Oord DeepMind, A., Li DeepMind, Y. & Vinyals DeepMind, O. Representation learning with contrastive predictive coding. Preprint at arXiv https://doi.org/10.48550/arxiv.1807.03748 (2018).
Wang, Y. & Yang, Y. Bayesian robust graph contrastive learning. Preprint at arXiv https://doi.org/10.48550/arxiv.2205.14109 (2022).
Ahmed, M., Seraj, R. & Islam, S. M. S. The k-means algorithm: a comprehensive survey and performance evaluation. Electronics 9, 1295 (2020).
Kingma, D. & Ba, J. Adam: A method for stochastic optimization. in International Conference on Learning Representations (2015).
SCoPE2 data processed to ASCII text matrices. slavovlab https://scp.slavovlab.net/Specht_et_al_2019 (2019).
Raw data from experiments benchmarking nPOP. slavovlab https://scp.slavovlab.net/Leduc_et_al_2021 (2021).
plexDIA data organized by experiments. slavovlab https://scp.slavovlab.net/Derks_et_al_2022 (2022).
pSCoPE data processed to ASCII text matrices. slavovlab https://scp.slavovlab.net/Huffman_et_al_2022_v1 (2022).
Model systems: cell lines of monocytes (U937 cells) and melanoma cells (WM989-A6-G3). slavovlab https://scp.slavovlab.net/Leduc_et_al_2022 (2022).
The single-cell pathology landscape of breast cancer. Zenodo https://doi.org/10.5281/zenodo.3518284 (2019).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291 (2019).
scrublet. GitHub https://github.com/swolock/scrublet (2019).
scikit-learn. scikit-learn https://scikit-learn.org/stable/ (2011).
scanpy. pypi https://pypi.org/project/scanpy/ (2018).
MAGIC. GitHub https://github.com/KrishnaswamyLab/MAGIC (2018).
harmony-pytorch. pypi https://pypi.org/project/harmony-pytorch/ (2019).
scanorama. pypi https://pypi.org/project/scanorama/ (2019).
AutoClass. GitHub https://github.com/datapplab/AutoClass (2022).
Reimand, J. et al. g:Profiler—a web server for functional interpretation of gene lists. Nucleic Acids Res. 44, W83–W89 (2016).
g:Profiler. Bioinformatics, Algorithmics and Data Mining Group https://biit.cs.ut.ee/gprofiler/gost (2016).
Hubert, L. & Arabie, P. Comparing partitions. J. Classif. 2, 193–218 (1985).
Rousseeuw, P. J. Silhouettes: a graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 20, 53–65 (1987).
Estévez, P. A., Tesmer, M., Perez, C. A. & Zurada, J. M. Normalized mutual information feature selection. IEEE Trans. Neural Netw. 20, 189–201 (2009).
Mogotsi, I. C. & Christopher, D. in Introduction to Information Retrieval (eds Manning C. D. et al.) 192–195 (Cambridge Univ. Press, 2009).
Li, W. A versatile deep graph contrastive learning framework for single-cell proteomics embedding. Zenodo https://doi.org/10.5281/zenodo.10547614 (2024).
Acknowledgements
The authors thank R. Aebersold for his valuable suggestion regarding this work, P. Zhao for model development advice and S. Zhu for providing valuable knowledge in the field of MS. This work was supported by the National Natural Science Foundation of China (61973174 to H.Z. and 62373200 to H.Z.), the Key-Area Research and Development Program of Guangdong Province (2021B0101420005 to F.Y.) and the Young Elite Scientists Sponsorship Program by CAST (2023QNRC001 to F.Y.). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
F.Y. and J.Y. conceived and designed the project. W.L. and H.Z. developed the method. W.L. performed the research and conducted the experiments under the supervision of F.Y., H.Z. and J.Y. W.L. and F.Y. analyzed the results. W.L. and F.Y. wrote the manuscript. W.L. finished the figures under the guidance of F.Y. and J.Y. F.W. helped polish the figures and manuscript. H.Z. and J.Y. revised the manuscript. Y.R. gave suggestions for building the graph model and improving the manuscript. L.L. helped with the revision and data analysis tasks. B.W. provided suggestions for utilizing trustworthy AI and improved the manuscript. All authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Arunima Singh, in collaboration with the Nature Methods team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Systematic analysis concerning the sensitivity of the hyperparameter d.
a, Influence of the hyperparameter d on the ARI and NMI scores achieved in the clustering task using the SCoPE2_Specht dataset. d is the dimensionality of the learned latent embeddings. b, Influence of hyperparameter d on the ARI_cell and NMI_cell results obtained with the cell type labels as the ground truth in the data integration task using the N2 and nanoPOTS datasets. c, Influence of hyperparameter d on the accuracy and macro-f1 score values attained in the label transfer task (transferring from N2 to nanoPOTS). d, Influence of hyperparameter d on the 1-ARI_batch and 1-NMI_batch results obtained with the batch labels as the ground truth in the data integration task using the SCoPE2_Leduc and plexDIA datasets.
Extended Data Fig. 2 Systematic analysis concerning the sensitivity of the hyperparameter K.
a, Influence of the hyperparameter K on the ARI and NMI scores achieved in the clustering task using the SCoPE2_Specht dataset. K is the number of prototypes in the attribute denoising process of stage 2. b, Influence of hyperparameter K on the ARI_cell and NMI_cell results obtained with the cell type labels as the ground truth in the data integration task using the N2 and nanoPOTS datasets. c, Influence of hyperparameter K on the accuracy and macro-f1 score values achieved in the label transfer task (transferring from N2 to nanoPOTS). d, Influence of hyperparameter K on the 1-ARI_batch and 1-NMI_batch results obtained with the batch labels as the ground truth in the data integration task using the SCoPE2_Leduc and plexDIA datasets.
Extended Data Fig. 3 Embedding visualizations produced for the SCoPE2_Specht, N2 and nanoPOTS datasets.
a, The embedding learning process for the SCoPE2_Specht dataset. From left to right, we depict the learning process from the raw peptide level, the learned peptide uncertainty, the aggregated protein levels after executing uncertainty adjustments and the final learned cell embeddings. b, Visualization of parts of the raw protein profiles and the embeddings learned by scPROTEIN on the N2 and nanoPOTS datasets. In the left panel, we can observe that the batch effect is exhibited in the same cell type across the two datasets. In the right panel, scPROTEIN can greatly mitigate the batch effect, and the same cell type tends to show similar patterns. c, Diagram showing the label transfer process based on the learned embeddings. In the left panel, the gray dots represent cells with unknown labels from the query set, and the dots with other colors represent cells with known labels from the reference set. When the batch effect is effectively removed (middle panel), the gray cells can then be annotated accurately by KNN (right panel).
Extended Data Fig. 4 Data integration results obtained on the pSCoPE_Huffman and plexDIA datasets.
a, t-SNE plots showing the cells of the pSCoPE_Huffman and plexDIA datasets, colored by their data acquisitions and cell lines. HPAFII is the shared cell line between the two datasets. b, ARI_cell, ASW_cell, NMI_cell, and PS_cell results produced by scPROTEIN and the comparison methods with the cell type labels as the ground truth (x-axis) and the 1-metrics with batch labels as the ground truth (y-axis) on the pSCoPE_Huffman and plexDIA datasets. c, Heatmap showing the estimated uncertainties of each peptide signal across cells, colored by the estimated uncertainty calculated on the pSCoPE_Huffman dataset. The batch information and protein information are shown below the heatmap and on the right-hand side of the heatmap, respectively.
Extended Data Fig. 5 Data integration results obtained on the pSCoPE_Leduc and plexDIA datasets.
a, t-SNE plots showing the cells of the pSCoPE_Leduc and plexDIA datasets, colored by their data acquisitions and cell lines. Melanoma and U-937 are the shared cell types between in the two datasets. b, ARI_cell, ASW_cell, NMI_cell, and PS_cell results produced by scPROTEIN and the comparison methods with the cell type labels as the ground truth (x-axis) and the 1-metrics with batch labels as ground truth (y-axis) on the pSCoPE_Leduc and plexDIA datasets.
Extended Data Fig. 6 Data integration results obtained on the pSCoPE_Leduc and SCoPE2_Leduc datasets.
a, t-SNE plots showing the cells of the pSCoPE_Leduc and SCoPE2_Leduc datasets, colored by their data acquisitions and cell lines. U-937 is the shared cell type between the two datasets. b, ARI_cell, ASW_cell, NMI_cell, and PS_cell results obtained by scPROTEIN and the comparison methods with the cell type labels as the ground truth (x-axis) and the 1-metrics with batch labels as ground truth (y-axis) on the pSCoPE_Leduc and SCoPE2_Leduc datasets.
Extended Data Fig. 7 Application of scPROTEIN to clinical proteomic dataset.
a, UMAP of the scPROTEIN embeddings, which shows the cells colored by their clustering results. b, Detailed ratio of the cluster 1 cells for the control donor and CTCL donor. c, Volcano plot showing the differentially expressed proteins found by contrasting the healthy cells and CTCL cells in cluster 1. d, Top GO terms in the BP for the identified upregulated proteins of the CTCL cells in cluster 1. The p-values are computed using Fisher’s one-tailed test and adjusted by the multiple-hypotheses testing method (g:SCS) of gProfiler. e, Detailed ratio of the cluster 8 cells for the control donor and CTCL donor. f, Volcano plot showing the differentially expressed proteins found by contrasting the healthy cells and CTCL cells in cluster 8. g, Top GO terms in the BP for the identified upregulated proteins of the CTCL cells in cluster 8. The p-values are computed using Fisher’s one-tailed test and adjusted by the multiple-hypotheses testing method (g:SCS) of gProfiler.
Extended Data Fig. 8 Application of scPROTEIN to spatial proteomic data.
a, Visualizations of the learned spatial informative embeddings and the spatial heterogeneity degrees within tumor samples. b, Visualizations of the learned spatial informative embeddings and the spatial heterogeneity degrees within nontumor samples.
Supplementary information
Supplementary Information
A combined supplementary file that includes Supplementary Figs. 1–10 and Tables 1 and 2.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, W., Yang, F., Wang, F. et al. scPROTEIN: a versatile deep graph contrastive learning framework for single-cell proteomics embedding. Nat Methods 21, 623–634 (2024). https://doi.org/10.1038/s41592-024-02214-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-024-02214-9